Method Article
クローン性増殖のためのCD133 +肝幹細胞の単離
要約
Here we describe the isolation of CD133 expressing liver stem cells and cancer stem cells from whole murine liver, a process that requires tissue digestion, cell enrichment, and flow cytometry isolation. We include methods for advanced single cell isolation and clonal expansion.
要約
Liver stem cell, or oval cells, proliferate during chronic liver injury, and are proposed to differentiate into both hepatocytes and cholangiocytes. In addition, liver stem cells are hypothesized to be the precursors for a subset of liver cancer, Hepatocellular carcinoma. One of the primary challenges to stem cell work in any solid organ like the liver is the isolation of a rare population of cells for detailed analysis. For example, the vast majority of cells in the liver are hepatocytes (parenchymal fraction), which are significantly larger than non-parenchymal cells. By enriching the specific cellular compartments of the liver (i.e. parenchymal and non-parenchymal fractions), and selecting for CD45 negative cells, we are able to enrich the starting population of stem cells by over 600-fold.The proceduresdetailed in this report allow for a relatively rare population of cells from a solid organ to be sorted efficiently. This process can be utilized to isolateliver stem cells from normal murine liver as well as chronic liver injury models, which demonstrate increased liver stem cell proliferation. This method has clear advantages over standard immunohistochemistry of frozen or formalin fixed liver as functional studies using live cells can be performed after initial co-localization experiments. To accomplish the procedure outlined in this report, a working relationship with a research based flow-cytometry core is strongly encouraged as the details of FACS isolation are highly dependent on specialized instrumentation and a strong working knowledge of basic flow-cytometry procedures. The specific goal of this process is to isolate a population of liver stem cells that can be clonally expanded in vitro.
プロトコル
すべてのソリューション、メディア、楽器、フィルタ、およびチューブは滅菌および汚染のリスクを減らすために無菌操作で処理する必要があります。 4で、事前とストア内のすべてのバッファとメディア24時間を準備℃に
1。全体の肝臓からの実質及び非実質分離
- 5 mgのコラゲナーゼ、5 mgのプロナーゼ、および1 mg DNaseを含む10 mlの滅菌PBSでネジのトップで15 ccのチューブを使用して消化するための酵素溶液を調製します。
- 機関認定品(動物実験用委員会)CO 2窒息などのメソッドを使用して、マウスを安楽死させる。 70%エタノール溶液で安楽死させた動物の腹部領域を拭いてください。滅菌器、オープン腹腔と外植片の肝臓エンブロックを使用する。外植肝臓から胆嚢、膀胱を削除します。閉じた滅菌シャーレに層流フードへの全肝を転送する。
- この手順は、層流フードで完了します。滅菌カミソリの刃を使用して、滅菌シャーレに1分間に複数の水平方向および垂直方向のカットの組み合わせで肝臓を飾らない。場所¼上記のステップ1.1から10ccのコラゲナーゼを含むPBS、プロナーゼ、およびDNアーゼで15 ccのチューブのみじん切り肝臓のパルプの。各¼みじん切り肝臓で繰り返します。
- 37℃の水浴に¼みじん切り肝臓のパルプと酵素℃で45分間、振とう器で1〜サイクル/秒でのある場所のチューブ。層流フードへ転送する前に、水浴から取り出した後に70%エタノールでチューブを拭きます。
- この手順は、層流フードで完了します。滅菌シャーレに収集するために70ミクロンメッシュフィルターを通して消化された肝臓のパルプをひずみ。滅菌DMEM 2mlのアリコートを使用して:10%熱不活化ウシ胎児血清を含むF12培地を、フィルターをすすぎ、フィルターを介してマッシュに消化されたパルプをシリンジのプランジャーのゴムの端を使用してください。ろ液約20mLの総量を作るために5回を繰り返します。ろ液は、2つの等しい15 mLの試験管に分ける。
- すべての転送は、層流フードで完了し、℃で使用可能な場合4で冷却遠心機を使用する必要があります。 1分間に50 × gで遠心する。 #1の上清を保存し、実質ペレットを捨てる。 1分間に50 × gで上清第1位を遠心分離します。 #2の上清を保存し、ペレットを捨てる。 1分間に50 × gで上清第2位を遠心分離します。 #3を上清保存し、ペレットを捨てる。非実質画分を得るために8分、180 × gでの最終的な清第3位を遠心分離します。
2。赤血球溶解
層流フードで働く、細胞は冷たいまま、および使用のソリューションは、4℃に冷却した。
- 手順の前の夜、10X濃度株式BD薬学は滅菌蒸留水で10倍希釈でバッファを溶解希釈することにより赤血球溶解バッファーを準備。 1Xは、4℃、30日℃で保存することがsolutionshould
- 手続き前の晩には、滅菌PBS、0.5%ウシ血清アルブミン 、および2mM EDTAを用いて国内でのみ有効のバッファを準備。 0.45ミクロンのフィルターで吸引フィルターユニットを使用してフィルタソリューション。オリジナルのプラスチック製の蓋でルターユニットの上部をカバーし、プラスチック製のラップで固定します。 4でStoreentireフィルタユニット°脱EDTAに12時間。フィルタユニットは、12時間後に標準的な滅菌キャップで置き換えることができます。
- この手順は、層流フードで完了します。 5ミリリットル滅菌チューブを用いて、上記2.1から1X希釈赤血球細胞溶解緩衝液1 mLに、上記のステップ1.6から非実質ペレットを再懸濁する。層流フードのアウト転送用チューブにフタを。
- 4℃15分間5秒間インキュベートするための穏やかにボルテックス℃で光から保護。
- 5分間、200 xgで遠心します。
- この手順は、層流フードで完了します。 1ミリリットルの氷冷し、滅菌国内でのみ有効バッファに上清に溶解した赤血球を破棄し、そして再度サスペンドペレット。
- 5分間、200 xgで遠心します。
- この手順は、層流フードで完了します。上清を捨て、1 mlの氷冷したと滅菌国内でのみ有効のバッファで再びサスペンドペレット。
- PBSの細胞懸濁液10μlを削除し、10μlのtryanブルーを追加。血球計算板を用いてremainingnon -実質細胞を数える。
3。非実質画分からCD45造血細胞枯渇
層流フードで働く、細胞は冷たいまま、および使用のソリューションは、4℃に冷却した。
- 最大108の合計のセルに107個の細胞あたりの国内でのみ有効緩衝液100μL中に細胞を懸濁する。
- 各107個の細胞を20μL国内でのみ有効CD45マイクロビーズ抗体を適用し、4℃でインキュベート℃で15分間。
- 5分間、200 xgで追加の2ミリリットル国内でのみ有効のバッファと遠心を追加。上清を取り除く。細胞ペレットは、(最大〜108総細胞)を1ml国内でのみ有効の緩衝液に再懸濁する。
- 層流フードでは、フィルタセルは国内でのみ有効LD磁気カラムを使用して。マグネットホルダー(ミルテニーMidiMACSまたはQuadroMACS)にカラムを配置することによって開始します。ろ液をキャッチするフィルター下に滅菌5 mlのチューブを置きます。 2ミリリットルMilteをロードすることによって、カラムを準備NYIバッファ。
- プレフィルターの洗浄が完了すると、LDの列にセルをロードします。細胞懸濁液がカラム内になると、1ミリリットルミルテニーのバッファを追加し、1mlミルテニーバッファは2つの追加回を洗う繰り返す。ろ過の速度を上げるために列に付属してプランジャーを使用しないでください。 ONLYのフィルタは、磁気ホルダーに入れ、ろ液を集める。
- 5分間、200 xgでCD45 -枯渇の非実質細胞の約5mlのコレクトろ液を(列が残り1 mlを保持する)心する。保持されたCD45陽性細胞を含む列を捨てる。
4。 CD133陽性細胞のフローサイトメトリー分離
- 卵形細胞のメディアを準備します。 F12ベースとして10%の熱不活性化ウシ胎児血清を含む培地、及び(1μg/ ml)のインスリンを追加し、HEPES(5モル/ L)、およびペニシリン/ストレプトマイシン(1%体積/体積):1:1 DMEMを使用してください。 0.45ミクロンのフィルターで吸引フィルターユニットを使用してフィルタソリューション。
- 10 7個の細胞あたり100μLで国内でのみ有効バッファーで細胞を再懸濁する。 CD133 - PE標識抗体の2μLを加える。対照としてのIgG - PE標識抗体とインキュベートする細胞の第二のグループを、使用する。 FACS用染色の対照として染色することなく細胞の第三のグループを保持する。
- 4℃でインキュベートする暗所で15分間° C。 2 mlの染色バッファーで再懸濁する。 5分間、200 xgで遠心します。上清を廃棄し、1ml国内でのみ有効のバッファで再びサスペンドペレット。
- このステップは、機関の特定の可能性がある標準のフローサイトメトリー細胞選別の手順を、使用して行われている。未染色の細胞およびIgG PE染色された細胞を用いて、CD133 +細胞集団の最適化されたゲーティングのためのパラメータをソート調整する。 PEは、(R - Phytoerythrin)一般的に488nmで発光するレーザーを持つ任意のフローサイトメーターで使用することができます。 PEの発光ピークは575 nmであり、FL - 2チャネルで検出される。
注:データ収集のためのセルのクエストのプログラムで、BD FACSソウルキャリバーまたはBD FACSヴァンテージのマシンを使用して、我々は250に設定されて側方散乱光で、前方の細胞集団を識別するために、ログスケールの散布図と側方散乱のビューを使用してください。 FL1とFL2は、ログスケールの両方で、550に設定します。これらのパラメータは、肝臓の非実質細胞を表示するには、最初の出発点を提供し、正と負の個体群の強度を染色に基づいて、必要に応じて調整されます。 - CD133 +ゲートを使用してCD133 +細胞集団を分離し、滅菌濾過した細胞培地で細胞を集める。
5。細胞培養法
- 遠心FACSは、5分間200 xgで細胞を集めた。細胞ペレットは、1mlあたり約5000細胞と卵細胞の培地に再懸濁する。初期起動実験のための50,000細胞/ mlに、より高い濃度で始まり、そしてテクニックの向上と全体的な細胞の生存率と歩留まりの向上と低下する場合があります。
- BDアンジオジェネシスラミニン上に細胞をプレートに1000細胞/ cm 2を用いて96ウェルプレートをコーティング。 37℃加湿インキュベーター内で場所° C、5%CO 2で。 24時間後、肝細胞増殖因子(50 ngの/ NL)および上皮成長因子(20 ng / ml)を加える。
- 単一セルの場合、96ウェルラミニンコートプレートの各ウェルに楕円形の細胞培地の50μlに直接細胞を分離する。唯一の陽性細胞の厳密な選択のための単一細胞のFACSの設定を使用します。 24時間後、ステップ5.2で上記のようにHGFとEGFとの楕円形の細胞培地50μlを加える。 5〜7日後に完全にメディアを変更します。
- 拡大するコロニーは、通常のセルの合計数メッキと細胞生存率に応じて、2週間後に発生する50%コンフルエント、より大きいと、細胞は、以下のように午前一時03分割されることがあります。
- トリプシン0.05パーセント- EDTAを用いて細胞を分割する。ちょうど足りるだけのウェル底面カバーするために、50〜100μL/ウェル、96ウェルプレート上で適用されます。 37℃インキュベーターで場所° 3-5分のためのC。
- 各ウェルに培地100μLを加え、5 mlのチューブにすべての液体を転送する。 5分間、200 xgで各チューブと遠心分離機に1 mLのメディアを追加します。
- よく、1〜3の新しい井戸(1:3比)にプレートに細胞を用いて、ラミニンコートしたディッシュにメディアのプレートに細胞を再懸濁する。
6。 RT - PCRを用いて双方向性のステータスの確認
この手順をRNeasyキットに同梱さをRNeasyプロトコルハンドブック、に詳述されている。
- 96ウェルプレートのコロニーのためのしたRNeasyミクロカラムを使用してください。各ウェルから培地を吸引除去する。直接各ウェルに75μlのBuffer RLTを(RNeasyキットから)を追加します。滅菌ラバーポリスマンでプレートの底を掻き取る。マイクロ遠心チューブと1分間ボルテックス混合物にPipettelysate。溶解液に70%エタノールを追加し、ピペッティングにより混和する。
- 10,000 rpmで15秒間のマイクロ遠心機で2 mlのコレクションチューブ(としてRNeasyキットに付属)と遠心分離機に入れRNeasyカラムへの解決策を転送する。溶出液を捨てる。
- コレクションチューブを再使用してください。カラムを洗浄するために、10,000 rpmで15秒間RNeasyカラムや遠心に350μlのBuffer RW1(RNeasyキット)を追加膜。溶出洗浄を捨てる。
- 70μlのBuffer RDD(両方はRNeasyキットに付属)に10μLのDNase Iストック溶液を加え、室温で15分間RNeasyカラムのメンブレンとインキュベートして80μLのDNase IバッファーRDDのインキュベーションミックスを追加する。
- メンブレンを洗浄するために、10,000 rpmで15秒間RNeasyカラムや遠心に350μlのBuffer RW1(RNeasyキット)を追加します。溶出洗浄とコレクションチューブを捨てる。
- 新しい2 mlコレクションチューブ(RNeasyキットに付属)でRNeasyカラムを置きます。メンブレンを洗浄するために、10,000 rpmで15秒間RNeasyカラムや遠心に500μlのBuffer RPE(RNeasyキット)を追加します。溶出洗浄を捨てる。
- RNaseフリー水(RNeasyキット)を使用して80%エタノールを準備します。 10,000 rpmで2分間をRNeasy columnandの遠心に80%エタノール500μLを加える。溶出洗浄を捨てる。
- 5分間、フルスピードで新しい2 mlコレクションチューブ(RNeasyキット)と遠心でRNeasyカラムを置きます。任意の溶出洗浄とコレクションチューブを捨てる。
- 新しい1.5 mlコレクションチューブ(RNeasyキット)でRNeasyカラムを置きます。フルスピードで1分間カラムのメンブレンと遠心分離機の中心に14μLのRNaseフリー水(RNeasyキット)を追加します。精製したRNAとのろ液を収集し、マイナスで、すぐにまたはストアcDNAを作成する場合は氷に転送する80 ° C今後の使用のために。
- 最後の10単位の濃度/氷冷した1 ×バッファーRTでμLにOmniscript.DiluteのRNase阻害剤を用いて逆転写。 5秒、5秒間のパルス遠心のための渦。 Omniscriptプロトコルの13ページにしたがって、氷上で新鮮なマスターミックスを調製する。 5秒、5秒間のパルス遠心のための渦。すべての反応に必要な合計のために必要なものより10%増しにマスターミックスのボリュームを準備することをお勧めします。
- マスターミックスを含む個々のチューブにテンプレートRNAを追加。 5秒、5秒間のパルス遠心のための渦。 37℃で60分間インキュベート℃を
- のHotStarTaq DNAポリメラーゼを用いて肝細胞( アルブミン )とcholangiocyte特定の遺伝子のPCR増幅。のHotStarTaqプロトコルの本の15ページあたりの反応ミックスを調製する。 20時/μlの濃度に希釈株式のプライマーをお勧めします、そしてフォワードプライマーとリバースプライマーに用い、1μL/反応チューブで。最初はRNA抽出のために使用される細胞の数に応じて、我々は、起動に3反応チューブ(コントロールをロードするためのβ-アクチン、 アルブミン 、およびKRT19)の間で最終的なcDNAを割ることをお勧めします。
- PCRプライマーの設計を表1に記載されています。
- サーモサイクラーに反応チューブを置きます。最初の実験のために次のプログラムをお勧めします。95 ° CFOR 15分30秒、95℃、55℃で30秒、72℃30秒C 72に続い× 35サイクル、℃で10分間、続い× 1サイクル。
- PCR産物はアガロースゲルを含浸エチジウムブロマイドを用いて分析している。
7。 CD133 +幹細胞の腫瘍性の確認
- この手順は、手順4または手順5から培養された細胞を使用してから新たに分離した細胞で行うことができます。新たに分離した細胞は生存率と縮小歩留まりが低下しているので我々は、培養細胞を用いた初期の実験を行うことをお勧めします。
- 上記のステップ5.5からトリプシン0.05パーセント- EDTAを用いて細胞をトリプシン処理。 37で3〜5分間インキュベーション° Cの後、各ウェルに培地100μLを追加し、個々の5 mlのチューブ(1ウェル= 1チューブ)を各ウェルから全ての液体を転送する。 5分間、200 xgで各チューブと遠心分離機に1 mlのメディアを追加します。
- 細胞を1 mlの氷冷PBSで再懸濁する。 PBSの細胞懸濁液10μlを削除し、10μlのトリパンブルーを追加。トリパンブルー排除を用いて、生きた細胞の数を決定する。 FACS分離した細胞を使用する場合、細胞数をFACS分離数によって決定されます。
- 5分間、200 xgで細胞を遠心分離し、1 × 10 6ライブcells/100μlの濃度でPBSに再懸濁した。 100μlのマトリゲルを追加。 6週齢の免疫不全ヌードマウスは、28 gagueの針を用いて皮下注射した。サイトあたり200μlの1 × 10 6個の細胞を注入する。
- マウスは、毎日の腫瘍の成長のために監視されています。一度腫瘍の形態は、一般的に3〜4週間のインキュベーション後、腫瘍体積をキャリパー(高さ×長さ×幅)を使用して測定されます。
8。代表的な結果:
正常な、健康なマウスの肝臓から、CD133 +肝臓の幹細胞の予想される細胞の収率は、肝臓あたり1,000〜5,000です。これらの細胞は静止肝臓では比較的まれであり、文化の中でうまく展開しません。我々は、正常な肝臓の単一細胞解析を使用しないことをお勧めして拡大する生細胞の収率は極めて低いです。
/ - -または肝臓特異的PTEN - / -マウス、2-4セル数の期待値などMAT1aなどの6週間1または慢性的な損傷につながる遺伝的改変のためのDDC 0.1%食として重要な慢性的なけが、、と肝臓のための最大100,000個の細胞を持つLS孤立増加、孤立/肝臓(図2)。遺伝的モデルを使用している場合は肝臓の幹細胞を分離するためのこれらの手順は腫瘍フリー動物を用いて実施されるべきであるとして、自発的な腫瘍の率の知識は、非常に重要です。 / - - MAT1aはたとえば、モデルが18ヶ月で自然発生腫瘍を形成する、我々は前に報告された自然発生腫瘍に、遅くとも15から16ヶ月以上の動物を使用しないことをお勧めします。
慢性的な傷害のモデルからの単一細胞の単離は、手続きがマスターされている、と細胞の生存率が確保されると、単一の細胞から拡大(3-9 colonies/96ウェルプレート)いくつかのコロニーが得られます。 7日後に拡大単一のCD133 +細胞由来のコロニーの3現在の代表的な位相コントラスト像を示しています。
双方向性のステータスの確認は、 アルブミンとKrt19 RT - PCRを使用して行われている。拡張された単一細胞からコロニーが肝細胞のマーカー( アルブミン )と胆管(Krt19)のための表現の両方を紹介します。図4は、7日間で約25セル/コロニーでは、3つの分離されたコロニーから双方向の潜在的な発現を示しています。
CD133 +正常な肝臓と化学的に誘発された肝臓損傷(6週間用などDDC 0.1%食)からの幹細胞はヌードマウスで腫瘍を形成しません。 CD133 +特定の遺伝的モデル(MAT1a - / -または肝臓特異的PTEN - / -マウス)由来の幹細胞は、後半にプレ腫瘍慢性傷害期後期に分離された場合、ヌードマウスで腫瘍を形成します。この腫瘍形成表現型は、現在、癌幹細胞として識別される2〜4を例えば、MAT1a - 。/ -マウスは生後18週で自然発生肝腫瘍を形成する。肝疾患の後期慢性傷害の段階で、15〜16週で分離されたCD133 +肝臓の幹細胞は、ヌードマウスで腫瘍を形成します。図5は、単一のCD133 +肝臓の幹細胞からin vitroで拡大1 × 10 6細胞の二国間の注入から代表的な腫瘍を示しています。
| 遺伝子 | フォワードプライマー | リバースプライマー |
| β-アクチン | 5' - TGTTACCAACTGGGACGACA - 3' | 5' - GGGGTGTTGAAGGTCTCAAA - 3' |
| アルブミン | 5' - CATGCCAAATTAGTGCAGGA - 3' | 5' - GCTGGGGTTGTCATCTTTGT - 3' |
| ケラチン19 | 5' - TGCTGGATGAGCTGACTCTG - 3' | 5' - AATCCACCTCCACACTGACC - 3' |
表1:RT - PCR用プライマーの設計
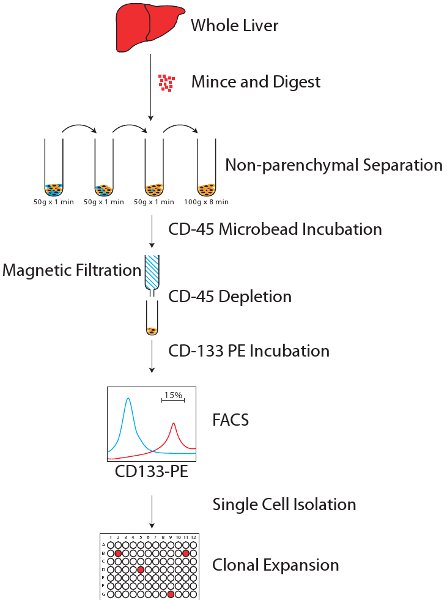
図1:メソッドの全体の概略図。
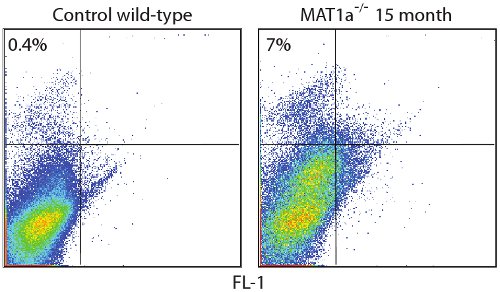
図2:野生型マウスから高度に濃縮されたCD45枯渇非実質割合で識別されたCD133 +肝臓の幹細胞制御無傷肝臓は高濃縮CD45枯渇非実質分数内で0.4パーセントCD133 +細胞を示しています。遺伝的にMAT1aノックアウト- / - 、慢性肝障害のモデルである、CD133の人口は、同じ高濃縮画分の10倍以上拡大する。
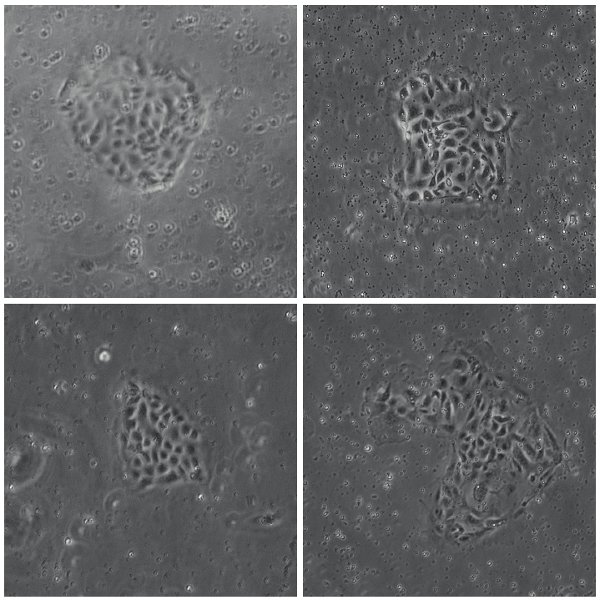
図3:単一のCD133 +クローンからのコロニーを拡大ラミニンコート96ウェルプレート上に展開単一のCD133 +細胞に由来するfourコロニーの位相コントラスト像が。。
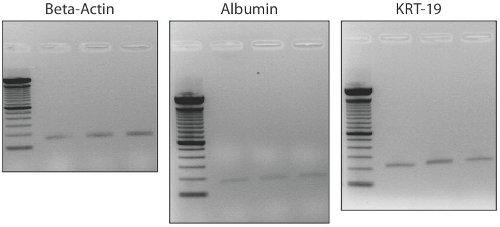
図4:。CD133 +肝臓の幹細胞クローン拡大コロニーからのBi -潜在的な遺伝子発現は、7日目では、コロニーは約25細胞である。遺伝子発現は、CD133 +細胞の双方向性の状態を確認し、肝細胞マーカーアルブミンとcholangiocyteマーカーKrt19の発現を示しています。
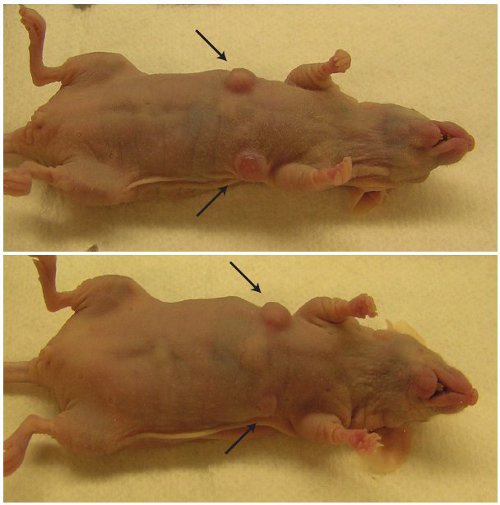
図5: - / -モデルCD133 +肝臓の幹細胞はヌードマウスで腫瘍を形成する矢印は、4週間MAT1aから注入された1 × 10 6 CD133 +細胞の後に増殖する腫瘍を示している。 CD133 +などのCCl 4(0.1%)DDC食事などの毒素誘発される慢性肝障害から細胞、腫瘍を形成しない。腫瘍形成は悪性の可能性、または幹細胞集団内のがん幹細胞の表現型を識別するために使用されます。
ディスカッション
Unlike the hematopoietic system, in which hematopoietic stem cells are responsible for maintaining a cellular differentiation system that replacesnormal physiologic turn-over of leukocytes, red blood cells, and platelets, liver stem cell, or adult liver progenitor cells, do not participate in normal liver homeostasis.5,6 After acute liver injury or partial hepatectomy, hepatocytes, as differentiated liver epithelium, undergo several rounds of proliferation to replace the lost liver mass.5,6Only during chronic injury are liver stem cells observed to proliferate.1,5-11 These adult, organ specific stem cells are proposed to differentiate into both hepatocytes and cholangiocytes.1,5-8Interestingly, the vast majority of liver cancer develops on the background of chronic injury, and thus liver stem cells are also hypothesized to be the precursors for a subset of liver cancer.2-4,12-17
One of the major challenges to stem cell work in the liver is isolation of rare populations of cells for functional analysis. For example, the vast majority of cells in the liver are hepatocytes, which are significantly larger than cholangiocytes and other smaller non-parenchymal cells. By breaking the whole liver into cell compartments (large hepatocytes - parenchymal fraction and smaller cells - non-parenchymal fraction), and further selecting for CD45 negative cells (non-hematopoietic cells), we are able to enrich the starting population of stem cells by over 600-fold.1,18-21Of note, we are by no means indicating that the CD133+ population is a 100% pure stem cell population, but clearly represents a heterogeneous population of cells, with different lineage and repopulation potential. One of the major limitations of the field is definitions of stem cells and progenitor cells. We use the term "stem cell" more broadly in this work, but in strict definition, CD133+ non-parenchymal cells represent a bi-lineage progenitor population. Given the state of current stem and progenitor research in the liver, this report does provide a starting place for investigators who are interested in this field. As new markers emerge, such as EpCAM,22,23 or transcription factors, such as Sox9,24 they can be incorporated. For example, we have found a fairly high rate of overlap between EpCAM+ cells and CD133+ cells.
In this report, we detail a process for stem cell isolation from normal murine liver as well as chronic liver injury models, which demonstrate increased liver stem cell proliferation. This method has clear advantages over standard immunohistochemistry of frozen or formalin fixed liver as functional studies using live cells can be performed after isolation.1,3,4 The specific goal of this process is to isolate a relatively pure population of liver stem cells that can be clonally expanded in vitro.
The primary limitation to this procedure is that the majority of cells isolated will not be viable after flow cytometry (see Trouble Shooting section). This is the result of the hours required to prepare the cells, and the numerous procedures required to refine the population prior to isolation. If the liver is digested in single cell suspension for immediate FACS analysis, the size difference between hepatocytes and other non-parenchymal cells will make effective FACS gate creation impossible. If the liver non-parenchymal cells are utilized, without elimination of hematopoietic cells, there is a risk that a significant number of CD133+ cells may be of hematopoietic origin may contaminate the fraction. Furthermore, by processing the cells through the Miltenyi filter, only single cells and very small clusters of cells are collected. This ensures that the sample will not clog the FACS intake needle.
Alternatives for this procedure include modification for any alternative cell surface marker, such as CD49f, EpCAM, or combination of markers, such as CD133+EpCAM+A second published report utilizes a density gradient to isolate liver stem cells from a non-parenchymal fraction. This procedure requires an Ultra-centrifuge (8000 x g), adds significant time to the procedure, and in our experience, significantly reduced pre-FACS cellular yield and post-FACS viability.8
Future experiments include a broader range of gene expression analysis, including fetal liver genes, such as HNF3, HNF4α, and αfp.In addition, Western blot analysis and immunocytochemistry of expressed proteins can be utilized to confirm RT-PCR results of cells in culture.
One issue to note is recent work that identified CD133 expression on hepatic stellate cells.25 We now routinely screen our samples for markers of stellate cells (see Trouble Shooting section), and have not identified significant contamination in our fractions. This may be related to different techniques of liver digestion and cell isolation.2,3,26
Based on the fact that the majority of cells will not be viable immediately after FACS isolation, we recommend that the cells be plated, either as bulk CD133+ cells or single cells, prior to use in animals. 5-7 days in vitro will significantly improve results of tumor analysis. In addition, given the rigors of single cell isolation, this should only be conducted once colonies can be expanded from bulk CD133+ isolated cells. A more thorough discussion of the various culture conditions and scaffold proteins which may be utilized to culture liver stem and progenitor cells has been well characterized by Lola Reid. This work provides alternative conditions and modifications, which investigators may incorporate into their research program once basic isolation techniques are mastered.27,28 Dr. Reid's work also provides a more detailed analysis of the lineage biology and maturation between hepatic stem cells and committed progenitors.
In terms of in vivo tumor analysis, we have had success using freshly isolated cells and cells from clonally expanded CD133+ cells in vitro, primarily using 1x106 cells. We have focused ontwo genetic strains of chronic liver injury, the MAT1a-/- and liver specific Pten-/- mice, and we have utilized both nude mice and wild-type mice as hosts for tumor growth. In our experience, CD133+ liver stem cells will only form tumors if they are isolated from significant liver injury models that are pre-malignant. Note that the tumors formed from CD133+ cells general have both hepatocellular carcinoma and cholangiocarcinoma features, suggesting a stem cell or progenitor cell origin to the tumors.2-4,29
Follow-up work after tumors are documented includes standard pathologic analysis of tumor tissue (H&E staining) and immunohistochemical staining. In addition, tumors can be minced and digested for FACS analysis or re-culture.2-4,30
In conclusion, we have detailed a procedure for the isolation, expansion, and basic characterization of CD133+ liver stem cells and CD133+ cancer stem cells.
Trouble Shooting:
CD45 contamination:
For Step 3, if there is contamination of CD45+ cells, which can be assessed by adding a CD45-FITC Ab prior to FACS analysis and isolation, check to ensure that the CD45 microbead antibody is not expired and that the filtrate was collected only while the filter was in the magnetic holder. Any filtrate collected while the filter is not in the magnetic holder will contain CD45+ cells.
Low cell number:
For the uninjured liver, the total number of CD133+ non-parenchymal isolated may be less than 10,000. These cells are rare in the quiescent liver. For a chronic injury model, such as the DDC 0.1% diet, the number will increase greatly to 100,000 cells. If the total cell number is significantly below these numbers, one issue to consider is the FACS Ab staining. Check to ensure that the CD133 Ab is not expired, as poor staining will result in a poor yield. Also, we recommend performing a FACS analysis of liver non-parenchymal cells to determine the relative population prior to attempting FACS isolation.
Low cell viability after FACS:
One of the issues of viability may be related to how the cells are processed and over what time. Ideally, the entire cell isolation procedure, steps 1-4, should be conducted without any delays between steps and completed in the same day. Any significant delay between steps 1-4 will greatly reduce the cell viability. A second issue related to cell viability may be related to the sheath pressure used for FACS isolation. We recommend using a lower sheath pressure. Lastly, once the cells are isolated, they should be immediately plated, as any significant storage on ice after sorting will also reduce viability.
CD133+ non-parenchymal heterogeneity:
Recent reports indicate that hepatic stellate cells may also have CD133 expression and have some plasticity.25 Therefore, in addition to verifying bi-potency genes with Albumin and Krt19, additional verification can include genes associated with stellate cells, such as glial fibrillary acidic protein in quiescent stellate cells and alpha-smooth muscle actin and desmin in activated stellate cells.3,4,26 In addition, the CD133+ population represents a broader progenitor population. The addition of a second marker, such as EpCAM, may help to refine the population further, and limit heterogeneity.
開示事項
Dr. Rountree receives research support from Bayer Pharmaceutics. Drs. Ding and Crooks and Ms. Dang and Ms. VanKirk have no disclosures to report.
謝辞
Dr. Rountree acknowledges current support from the Children's Miracle Network, National Institute of Health, K08DK080928 and R03DK088013, and the American Cancer Society Research Scholar Award, MGO-11651. Dr. Rountree acknowledges that this procedure was initially developed and refined while funded by the Pediatric Scientist Development Program (NICHD Grant Award K12-HD00850).
資料
| Name | Company | Catalog Number | Comments |
| DMEM:F12 | Invitrogen | 10565-018 | With phenol red |
| CD45 microbeads | Miltenyi Biotec | 130-052-301 | |
| Hepatocyte Growth Factor | Sigma-Aldrich | H1404 | |
| Epidermal Growth Factor | Sigma-Aldrich | E4127 | |
| DNase | Sigma-Aldrich | DN25 | 1 gram |
| Collagenase D | Roche Group | 1088874 | |
| Pronase | Roche Group | 0165921 | |
| 70 micron mesh strainer | Fisher Scientific | 352350 | |
| Omniscript RT | Qiagen | 205111 | 50 reactions |
| HotStarTaq | Qiagen | 203203 | |
| Miltenyi LD column | Miltenyi Biotec | 130-042-901 | |
| CD133-PE FACS Ab | eBioscience | 12-1331-82 | |
| Laminin coated plates | BD Biosciences | 354410 | 96 well |
| Trypsin 0.05% EDTA | Invitrogen | 25300-354 | 100 mL |
| RNeasy Micro Kit | Qiagen | 74004 | 50 columns for 5x105 cells or less |
| Pharm Lyse | BD Biosciences | 555899 | 10X concentration |
参考文献
- Rountree, C. B. A CD133-expressing murine liver oval cell population with bilineage potential. Stem Cells. 25, 2419-2429 (2007).
- Ding, W. CD133+ liver cancer stem cells from methionine adenosyl transferase 1A-deficient mice demonstrate resistance to transforming growth factor (TGF)-beta-induced apoptosis. Hepatology. 49, 1277-1286 (2009).
- Rountree, C. B., Ding, W., He, L., Stiles, B. Expansion of CD133-expressing liver cancer stem cells in liver-specific phosphatase and tensin homolog deleted on chromosome 10-deleted mice. Stem Cells. 27, 290-299 (2009).
- Rountree, C. B., Senadheera, S., Mato, J. M., Crooks, G. M., Lu, S. C. Expansion of liver cancer stem cells during aging in methionine adenosyltransferase 1A-deficient mice. Hepatology. 47, 1288-1297 (2008).
- Michalopoulos, G. K. Liver regeneration. J. Cell. Physiol. 213, 286-300 (2007).
- Fausto, N., Campbell, J. S., Riehle, K. J. Liver regeneration. Hepatology. 43, 45-53 (2006).
- Theise, N. D. Gastrointestinal Stem Cells. III. Emergent themes of liver stem cell biology: niche, quiescence, self-renewal, and plasticity. Am. J. Physiol. Gastrointest. Liver Physiol. 290, 189-193 (2006).
- Wang, X. The origin and liver repopulating capacity of murine oval cells. Proc Natl Acad Sci U. S. A. 100, 11881-11888 (2003).
- Greenbaum, L. E., Wells, R. G. The role of stem cells in liver repair and fibrosis. Int. J. Biochem. Cell Biol. 43, 222-229 (2011).
- Choi, S. S., Omenetti, A., Syn, W. K., Diehl, A. M. The role of Hedgehog signaling in fibrogenic liver repair. Int. J. Biochem. Cell. Biol. 43, 238-244 (2011).
- Khurana, S. Scopolamine treatment and muscarinic receptor subtype-3 gene ablation augment azoxymethane-induced murine liver injury. J. Pharmacol. Exp. Ther. 333, 639-649 (2010).
- Sell, S., Leffert, H. L. Liver cancer stem cells. J. Clin. Oncol. 26, 2800-2805 (2008).
- Lee, J. S. A novel prognostic subtype of human hepatocellular carcinoma derived from hepatic progenitor cells. Nat. Med. 12, 410-416 (2006).
- Sell, S., Dunsford, H. A. Evidence for the stem cell origin of hepatocellular carcinoma and cholangiocarcinoma. Am. J. Pathol. 134, 1347-1363 (1989).
- Amin, R., Mishra, L. Liver stem cells and tgf-Beta in hepatic carcinogenesis. Gastrointest Cancer Res. 2, 27-30 (2008).
- Tang, Y. Progenitor/stem cells give rise to liver cancer due to aberrant TGF-beta and IL-6 signaling. Proc Natl Acad Sci. 105, 2445-2450 (2008).
- Kitisin, K., Pishvaian, M. J., Johnson, L. B., Mishra, L. Liver stem cells and molecular signaling pathways in hepatocellular carcinoma. Gastrointest Cancer Res. 1, 13-21 (2007).
- Rountree, C. B. Bone marrow fails to differentiate into liver epithelium during murine development and regeneration. Hepatology. 45, 1250-1260 (2007).
- Shimano, K. Hepatic oval cells have the side population phenotype defined by expression of ATP-binding cassette transporter ABCG2/BCRP1. Am J Pathol. 163, 3-9 (2003).
- Suzuki, A. Flow cytometric isolation and clonal identification of self-renewing bipotent hepatic progenitor cells in adult mouse liver. Hepatology. , (2008).
- Suzuki, A. Clonal identification and characterization of self-renewing pluripotent stem cells in the developing liver. J Cell Biol. 156, 173-184 (2002).
- Yamashita, T. EpCAM-positive hepatocellular carcinoma cells are tumor-initiating cells with stem/progenitor cell features. Gastroenterology. 136, 1012-1024 (2009).
- Yamashita, T., Budhu, A., Forgues, M., Wang, X. W. Activation of hepatic stem cell marker EpCAM by Wnt-beta-catenin signaling in hepatocellular carcinoma. Cancer Res. 67, 10831-10839 (2007).
- Furuyama, K. Continuous cell supply from a Sox9-expressing progenitor zone in adult liver, exocrine pancreas and intestine. Nat Genet. 43, 34-41 (2011).
- Kordes, C. CD133+ hepatic stellate cells are progenitor cells. Biochem Biophys Res Commun. 352, 410-417 (2007).
- Berg, T. Fibroblast Growth Factor 10 is critical for liver growth during embryogenesis and controls of hepatoblast survival via b-catenin activation. Hepatology. , (2007).
- Turner, R. Human hepatic stem cell and maturational liver lineage biology. Hepatology. 53, 1035-1045 (2011).
- Wang, Y. Lineage restriction of human hepatic stem cells to mature fates is made efficient by tissue-specific biomatrix scaffolds. Hepatology. 53, 293-305 (2011).
- Galicia, V. A. Expansion of hepatic tumor progenitor cells in Pten-null mice requires liver injury and is reversed by loss of AKT2. Gastroenterology. 139, 2170-2182 (2010).
- Ding, W. Epithelial-to-mesenchymal transition of murine liver tumor cells promotes invasion. Hepatology. 52, 945-953 (2010).
転載および許可
このJoVE論文のテキスト又は図を再利用するための許可を申請します
許可を申請さらに記事を探す
This article has been published
Video Coming Soon
Copyright © 2023 MyJoVE Corporation. All rights reserved