Exploring the Effects of Spaceflight on Mouse Physiology using the Open Access NASA GeneLab Platform
In This Article
Summary
The NASA GeneLab platform provides unfettered access to precious omics data from biological spaceflight experiments. We describe how a typical mouse experiment is conducted in space and how data from such experiments can be accessed and analyzed.
Abstract
Performing biological experiments in space requires special accommodations and procedures to ensure that these investigations are performed effectively and efficiently. Moreover, given the infrequency of these experiments it is imperative that their impacts be maximized. The rapid advancement of omics technologies offers an opportunity to dramatically increase the volume of data produced from precious spaceflight specimens. To capitalize on this, NASA has developed the GeneLab platform to provide unrestricted access to spaceflight omics data and encourage its widespread analysis. Rodents (both rats and mice) are common model organisms used by scientists to investigate space-related biological impacts. The enclosure that house rodents during spaceflight are called Rodent Habitats (formerly Animal Enclosure Modules), and are substantially different from standard vivarium cages in their dimensions, air flow, and access to water and food. In addition, due to environmental and atmospheric conditions on the International Space Station (ISS), animals are exposed to a higher CO2 concentration. We recently reported that mice in the Rodent Habitats experience large changes in their transcriptome irrespective of whether animals were on the ground or in space. Furthermore, these changes were consistent with a hypoxic response, potentially driven by higher CO2 concentrations. Here we describe how a typical rodent experiment is performed in space, how omics data from these experiments can be accessed through the GeneLab platform, and how to identify key factors in this data. Using this process, any individual can make critical discoveries that could change the design of future space missions and activities.
Introduction
The overall goal of this manuscript is to provide a clear methodology of how to use NASA's GeneLab platform1 and how rodent experiments done in space are translated to omics data for analysis. Spacefaring humans are exposed to numerous health risks from altered gravity fields, space radiation, isolation from Earth, and other hostile environmental factors2,3,4,5,6. Biological experiments performed in space and on the ground have helped to define and quantify these risks7,8,9,10,11,12,13,14. In space, these experiments have been conducted on the International Space Station (ISS), the Space Shuttle, and other orbital platforms. Conducting these experiments requires specialized hardware and methodology given the unique concerns of performing experiments in space including limited crew time and the microgravity environment. Various platforms now exist for performing sophisticated experiments in space using plant, animal, and microbial models15.
Rodent models have been particularly important to advancing our understanding of how mammals, including humans, respond to spaceflight. These include the impact of spaceflight on the muscle structure16,17,18 and immune functions19,20,21. The standard vivarium cages used for housing rodents on Earth are not suitable for spaceflight experiments22,23. Therefore, over the years mice and rats have been flown and housed in various cages including the Japanese Aerospace Exploration Agency (JAXA) Habitat Cage24, animal carrying space capsules used on the BION-M1 unmanned Russian satellite25,26,27, the Mice Drawer System (MDS) designed by the Italian Space Agency28,29,30, the NASA Animal Enclosure Module (AEM), and now the NASA Rodent Transporter and Habitats23. Rodent experiments first started on board the Space Shuttle using cages referred to as the Animal Enclosure Module (AEM). This hardware was used in 27 rodent experiments on the Space Shuttle23. The AEM was originally developed for relatively short experiments on-board the shuttle (< 20 days). Since the development of the ISS, the AEMs have been modified for longer duration experiments and are now referred to as Rodent Habitats22,23. The new Rodent Habitats are designed to support long-duration missions in the ISS using the EXpedite the PRocessing of Experiments for Space Station (EXPRESS) Rack interface. Rodent Habitats are substantially different from standard vivarium cages in their dimensions, air flow, filter and exhaust system, and access to food and water (Figure 1). Nevertheless, this hardware has proven to be an effective research platform, enabling key insights into the spaceflight-induced changes to mammalian physiology19,31,32,33,34,35,36.
Large volumes of omics data can now be generated from biological spaceflight experiments including those performed with rodents. Recently, data from these omics experiments have been made publicly available through the NASA GeneLab platform1 which is a comprehensive data repository and analysis platform that allows anyone to develop hypotheses from spaceflight experiments. GeneLab provides tools for the discovery, access, sharing and analysis of data. We utilized GeneLab datasets to show that differences between the standard vivarium cages and specialized Rodent Habitats used in space cause massive differences in the transcriptome of mice36. We analyzed four different publicly available datasets, comparing different tissues from rodents housed in either the Rodent Habitats or standard vivarium cages. Using an unbiased systems biology analysis, we determined that the main drivers and pathways that were changed were consistent with a hypoxic response due to the high CO2 levels caused by higher CO2 concentrations on ISS, which leads to higher CO2 concentrations in the Rodent Habitat given that they are passive systems that take in the ambient air. This demonstrates how scientists can use open source tools and data to generate novel findings with implication on how the environment of the ISS impacts astronaut health.
Here we describe how rodent experiments are performed in space and how data from these experiments can be accessed through an open-source, omic platform related to space biology. We discuss the configuration of the Rodent Habitats used for space missions, and how spaceflight tissues are processed. We also describe how spaceflight omics data can be discovered and accessed on GeneLab and how key factors driving the overall response to spaceflight can be identified36. The specific example we will present on how this protocol is implemented will be comparing the biological differences occurring in rodents housed in Rodent Habitat and the vivarium controls that were published by Beheshti et al.36. It is important to note that ground controls are essential for spaceflight rodent experiments. As described in this protocol, these controls are done with both identical conditions (i.e., CO2 conditions, humidity, temperature, cage dimensions, etc.) in the Rodent Habitats on the ISS and in standard vivarium cages that have the standard environmental (i.e., CO2 conditions, humidity, and temperature) conditions on Earth. The rodents housed in the Rodent Habitat ground controls allow for the direct comparison to rodents in space. While rodents housed in vivarium cages allow for the biological comparison between the different housing (e.g., vivarium cages vs. Rodent hardware). The Rodent Habitat is different than vivarium cages in that it has constant air flow (0.1–0.3 m/s), a long duration, and a secondary exhaust filter that captures and absorbs the animal waste guided to the exhaust filter by continuous air flow in microgravity. In addition, Rodent Habitats have passive systems and intake ambient air; therefore, they also have higher CO2 concentrations due to elevated levels in the ISS cabin (~5,000 ppm).
Protocol
The animal protocols for housing and tissue processing follow standard guidelines for laboratory animal care and have been approved by NASA's flight and ground Institutional Animal Care and Use Committees (IACUC).
1. Configuration of Rodent Habitats
NOTE: The NASA Rodent Habitats (previously AEMs) have different features from the vivarium cages to accommodate for operations in space (Figure 1).
- House 10 mice in each Rodent Habitat (up to 30 g per mouse). House 5 mice per compartment when the habitat is configured into two compartments or 10 mice if there is a single compartment.
NOTE: NASA Rodent Habitats have a larger accessible surface area per rodent than the standard vivarium cages. - For ground controls animals, house mice in Rodent Habitat inside the ISS Environmental Simulator (ISSES) under identical environmental conditions as the flight animals including CO2 concentrations, temperature, and relative humidity.
- Provide animals with ad libitum access to custom made NASA Nutrient Upgraded Rodent Foodbars (NuRFB) in accordance with National Research Council (NRC) nutritional requirements for mice37, and to water through pressure activated lixits.
- Monitor the animals' health and behavior which will be enabled in the Rodent Habitats with the 12:12 h light cycle similar to vivarium cages in standard facilities with LED lighting during the day and infrared lighting during video health checks that take place during the dark cycle.
- Place four cameras in the Rodent Habitat cages for the daily monitoring of animals' health and behavior and collect videos during the night with infrared lighting.
- Deliver the rodents to the ISS in a Transporter (Figure 2B) aboard the Dragon Capsule or similar launch vehicle.
- Ensure that the rodents are observed and examined by the NASA flight veterinarian before being loaded into the Transporter for launch, and by the trained crew members upon arrival to ISS and before transfer to Rodent Habitats.
- For this transition period, house up to 20 mice (10 on each side) or 12 rats in the transporter.
NOTE: Similar to the Rodent Habitat, the Transporter is a passive unit for environmental conditions. During this short transition period, this single-unit can house up to 20 mice.
2. Rodent Handling for Spaceflight Experiments
- Procure rodents from standard vendors.
NOTE: Following delivery, group rodents within standard vivarium cages and have the animals acclimate to NASA NuRFB, lixits, and raised wire floors until the animals are loaded into the Transporter. Leaving the rodents in the cages will allow the animals to adapt naturally. The handling of mice in and out of both the Rodent Habitats and vivarium cages follows protocols commonly used for all rodent experiments12,27,28. The Rodent Habitat system (Figure 1A) will be utilized for both spaceflight mission on the STS and ISS, respectively, and for ground controls simulating ISS or STS environmental conditions. - For some missions use standard vivarium cages (Figure 1B) for the vivarium control. Use 5 or 10 mice per standard vivarium cage.
- For Rodent Habitats, place 10 mice in two different compartments with 5 mice per compartment. Remove the cage divider to house 10 mice per Habitat in a single compartment.
- Utilize three components of the Rodent hardware during spaceflight missions as described below (Figure 2).
- Place rodents in a Transporter (Figure 2B) for the travel between the Earth and the ISS or vice versa at double density (10 mice per side, 20 mice per Transporter).
- Once on ISS, attach the Animal Access Unit (AAU) (Figure 2C) to the Transporter. Transfer rodents from the Transporter to the Habitats using Mouse Transfer Boxes (MTB) (5 mice per MTB) (Figure 2D).
NOTE: The AAU is used to contain any animal products (e.g., feces, urine, fur) from getting to the ISS cabin. - Detach the AAU from the Transporter and attach to the Rodent Habitat. Then transfer the animals from the MTB to the Rodent Habitat (Figure 2A) where they reside for the duration of the mission.
NOTE: The CO2 concentration due to elevated levels in the ISS cabin for all Rodent Habitats is at 5,000 ppm.
- Monitor the temperature and humidity of the Rodent Habitats, but there are no active thermal controls. Ensure that the Rodent Research team works with ISS to maintain and control the cabin temperature, which determines the temperature in the Rodent Habitat.
NOTE: The light and dark cycle in the Rodent Habitats occurs every 12 h (e.g., 5:00–17:00 GMT, lights on) and the ISS crew performs regular and frequent change out of the food (weekly or biweekly) and refills the water (every ~28 days).
3. Euthanasia of Rodents and Processing Tissue
- For euthanasia, give rodent an overdose of a general anesthetic (Ketamine/Xylazine up to 150/45 mg/kg body mass diluted in phosphate-buffered saline for a total volume of 0.3 mL) via intraperitoneal injection (IP) paired with a secondary method of euthanasia (cervical dislocation or thoracotomy).
- For experiments conducted on the ISS:
- Return rodents either live, or
- Euthanize on ISS.
- Freeze rodent carcasses at -95 ± 2 °C in the freezers on the ISS and return to Earth on the available return vehicle (currently SpaceX Dragon capsule).
- Once the rodents are returned to Earth, dissect all organs and tissues (i.e., liver, kidney, skin, muscles, heart, spleen, eyes, adrenal glands, lungs, and brain) and store at -80 °C or in RNA stabilizing solution.
- Follow the same procedures and timings for all control ground experiments as the flight experiment with a 3–5 day offset to match the ISS telemetry data.
- From the preserved tissues isolate RNA, protein, and DNA isolation using standard protocols that are described in detail associated with each dataset on the GeneLab platform (genelab.nasa.gov).
NOTE: Rodent tissues not utilized by the primary investigator(s) become part of NASA's Institutional Scientific Collection. These samples are stored in Ames Research Center's (ARC) Non-Human Biobank where they are cataloged and made available for request by the science community. Available tissues can be found on the Life Sciences Data Archive Public Website at: https://Lsda.jsc.nasa.gov/Biospecimen.
4. Generating Omics Data from RNA, DNA, and Protein Extracts
- From the extracted macromolecules (RNA, DNA, protein) use standard protocols to generate omics data. These are described in detail in the respective study metadata on GeneLab.
5. GeneLab Repository and Submitting Data
NOTE: Space biology related omics data are submitted to the GeneLab Data Repository. GeneLab accepts and hosts space-related omics data funded by multiple space agencies around the world.
- Generate omics related data that can be hosted on the GeneLab repository.
- Submit generated data to GeneLab, either when analysis is complete or based on the discretion of the investigator.
NOTE: Data submitted to other public omics databases are imported and published into the GeneLab repository. GeneLab generated data are curated and published without an embargo period. GeneLab, specifically the Sample Processing Lab, generates data from various spaceflight experiments using optimized extraction protocols and techniques to increase the omics data from spaceflight experiments.
- Submit generated data to GeneLab, either when analysis is complete or based on the discretion of the investigator.
- When the data are ready to be submitted, format and transfer the metadata and data to GeneLab with the following method (Supplemental Figure 1):
- Use the ISAcreator tools to define an experimental study and store the metadata.
NOTE: The ISAcreator tool is available for download with a guided tutorial here38. - Refer the data listed here39 to understand accepted data types and formats for raw and processed data files.
- To optimize upload and storage, compress data files.
- Transfer the metadata and raw and/or processed data to GeneLab data curators through the workspace40.
- Create a username and password and upload the data.
- Use the ISAcreator tools to define an experimental study and store the metadata.
- Once the data have been uploaded to the workspace, share data to a GeneLab curator.
NOTE: Detailed steps on how to upload and share files can be found in the Data Submission Guide41. - Each submission is verified by a curator and published in the GeneLab repository42.
6. Finding Datasets for Analysis using Search Features on GeneLab
- Search for different datasets on GeneLab by going to the link (Supplemental Figure 2)38.
- Specifically related to a previous publication36, search for the following terms: GLDS-21, GLDS-111, GLDS-25, and GLDS-63.
- Access the GeneLab homepage by clicking on "GeneLab Data System" on the left-hand side of the screen.
- Enter the keywords in the "search data" box to search for specific areas of interest. In this case enter each of the following dataset identifiers separately: GLDS-21, GLDS-111, GLDS-25, and GLDS-63.
- In addition to searching the GeneLab repository, search across other databases including NIH GEO, EBI Pride, and ANL MG-RAST by selecting the desired check boxes under the search bar.
NOTE: Currently only for the GeneLab repository, a user can search by using the following filter categories: Organisms, Assay Type, Factors, and Project Type.
7. Storing and Transferring Files of Interest for Analysis
NOTE: The GeneLab Workspace is designed to store and transfer files directly from the GeneLab database (Supplemental Figure 3).
- Click on "Workspace" on top of the Data Systems menu.
- If new user, register for a new account.
NOTE: The GeneLab Workspace is powered by GenomeSpace43. - Access detailed instructions on how to use the workspace by selecting "Help" on the top menu and clicking on User Guide.
- For each user, access all the datasets in the GeneLab repository by selecting the "Public/genelab" folder on the left-hand menu.
- Copy the datasets of interest to a local directory workspace by going to the folder with the data of interest. Right click on the specific file, select "copy/move" in the menu that appears, select the folder to copy the file to, and then click on "copy".
- Find the following datasets related to a previous publication36 as instructed above and copy over to the local workspace: GLDS-21, GLDS-111, GLDS-25, and GLDS-63.
8. Accessing Metadata and Description of Each Study
NOTE: Metadata files for each dataset in the GeneLab repository are in the "Public/genelab" dataset subfolder on the left-side menu.
- Find the metadata information for the dataset of interest by accessing one or more metadata files contained in a "metadata" subfolder of each dataset. For example, for GLDS-100, there are 2 files in the "Public/genelab/GLDS-100/metadata" subfolder: "GLDS-100_metadata_RR1_BIOBANK-Eye-ISA.zip" and "GLDS-100_metadata_RR1ExpDesign.pdf".
- Ensure that every dataset has a single zipped file that provides metadata according to the ISATab specification (which subsumes the MIAME, MIAPE, and other MIBBI framework standards for minimum metadata requirements). Always end this type of file name in "ISA.zip". For example, for GLDS-100, this file is "GLDS-100_metadata_RR1_BIOBANK-Eye-ISA.zip".
- Use the ISACreator tool44 or a text editor to visualize and access the ISATab metadata, which contains the text description for the study and assay metadata for each dataset.
NOTE: Within the ISATab metadata, samples are described and associated with bioassays, and bioassays are described and associated with output data files. - Check for the presence of the output assay data files that are located within each dataset in subfolders by type of assay. For example, for GLDS-100, RNA-Seq output assay files are located in the "Public/genelab/GLDS-100/transcriptomics/" folder.
9. Analysis of GeneLab Data
NOTE: Various pipelines can be implemented for various omics data. Here, the specific example focuses on an unbiased systems biology transcriptomic pipeline which is used to determine the "key drivers" of the system being studied.
- Check previously published literatures36,45,46,47,48,49,50 to understand this pipeline.
- Once a specific dataset of interest is selected for analysis, download the data to a local machine with the following method:
- Click on the specific dataset.
- Click on the "Study Files" tab on the far left of the headers.
- Ensure that all datafiles and metadata are available in this menu.
- To download each file, click on the specific file names.
- For the microarray datasets that will be downloaded from GeneLab, use the following pre-processing steps.
- Process the raw data for each dataset separately using background subtraction and Quantile normalized using RMAExpress51 for the commercial microarrays.
- Create principle component analysis (PCA) plots using R to determine how closely the biological replicates grouped together.
- Import data into MultiExperiment Viewer52 and calculate significant genes first using the false discovery rate (FDR) statistics starting with FDR < 0.05. If no significant genes appeared with FDR statistics, then use standard t-tests starting with a p-value < 0.05 to determine the significant genes.
- Once the statistically significant regulated genes have been determined, implement a fold-change cut-off of ≥ 1.2 or ≤ -1.2 to compare the experimental samples with the controls.
- Use Gene Set Enrichment Analysis (GSEA)53 for pathway and functional predictions.
- Use GSEA either through GenePattern54,55, directly through GSEA, or using R programming environment.
- Determine the significantly regulated pathways using the following gene sets: C2, C5, and hallmarks.
- Perform leading-edge analysis on the significantly regulated gene sets and determine leading edge genes associated with each experimental comparison and Gene Set.
- Find the leading-edge genes that overlap between all the gene sets for each experimental condition.
- Use another platform to determine predicted functions and pathways that are being significantly regulated. In this case use ingenuity pathway analysis (IPA) to determine the significant upstream regulators, biofunctions, and canonical pathways.
- Upload the list of genes with fold-change values for the statistically significant genes determined in step 9.4.4.
- Follow IPA's instructions to generate upstream regulators, biofunctions, and canonical pathways for each experimental comparison.
- Determine the gene associated for upstream regulators, biofunctions, and canonical pathways which have an activation z-score ≥ 2 (indicated activation) or ≤ -2 (indicating inhibition).
- Find the overlapping genes related to all the predictions above.
- Determine common/overlapping genes between steps 9.4 and 9.5.
NOTE: These genes are considered as the key/driver genes controlling the majority of the predicted functions and activity with the experimental conditions being analyzed. Previous studies have shown that knocking out or promoting these genes will make the experimental condition or system being studied non-functional45,46,49.- Construct networks through IPA (or any network assembly software) to determine the connectivity of the genes.
- Consider the most connected gene as central hub driving the key genes.
- To determine the connectivity between the datasets, group all key genes in one network and repeat connectivity test to determine the central hub that is occurring among all key genes from all datasets being analyzed.
10. Using Galaxy56 Interface on GeneLab to Analyze Transcriptomic Data
NOTE: Here a protocol for using the GeneLab Galaxy interface (available Fall 2018) to analyze transcriptomic data from GeneLab is described. Galaxy tutorials abound. Example tutorials on how to use Galaxy in general are available elesewhere57,58.
- Users can sign in to GeneLab using Google or NASA credentials. GeneLab Galaxy tools are located under the "Analyze" menu.
- Follow these three ways to bring data into the GeneLab Galaxy platform.
- Upload data from the local file system using the "Upload data" function.
- Import data from GeneLab GenomeSpace using the GenomeSpace importer tool under "Get Data" section.
NOTE: All GeneLab data files are available in the "public" folder, organized by the dataset accession number (see above). - Import data appear in the "history" of analysis section on the right-hand side. Users can have multiple histories, which are managed using either "History Options" or "View All Histories" buttons at the top of the history pane.
- Tools for analysis are listed and searchable on the left side of the interface.
- Check for the appearance of datasets that have been imported on the current history.
NOTE: Many details regarding the data are available for inspection for each dataset. - Select a tool on the left-hand side to populate a form in the center panel, with options for analysis and specification of data inputs. Create jobs for executing the analysis by completing the form and pressing "Execute".
- Check for jobs submitted which are represented in the history and color-coded to indicate status of execution (queued, executing, completed with or without errors).
- Link the tools into complex workflows. Manage workflows through tools located in the "Workflows" menu. Figure 3 shows an example workflow created for processing RNA-seq data.
- Share datasets, workflows, and histories with others using the "Shared Data" menu.
Representative Results
Determining key drivers from spaceflight transcriptomic data will assist NASA with determining health risks and developing potential countermeasures to combat negative effects on astronaut health. In our recent publication, we have followed the steps above and utilized GeneLab datasets to successfully show a novel finding that CO2 concentrations on the ISS can impact health36. We have also used the technique above in other studies to successfully determine the key factors driving the system being studied45,46,47,48,49,50. Here we will show how the results from using this protocol can be successfully used to determine the key drivers.
In this study, we primarily focused on the biological differences that occur in rodents housed in the Rodent Habits ground controls and the vivarium controls. As described above, it is the key to better understanding these two habitats, which will provide us information on possible confounding factors that can impact health due to the environment on the ISS. For all rodent spaceflight experiments, these ground controls are also essential to determine which biological factors are associated directly with spaceflight or due to the environmental conditions on the ISS. As stated in the protocol, the environmental condition for the vivarium habitat is not exposed to the higher CO2 level that is present for the Rodent Habitat. The vivarium habitat has the normal CO2 level that is present on Earth (currently being 300 to 380 ppm). The temperature and humidity for both habitats are similar.
We used the following datasets from the GeneLab platform to determine the key genes between the rodents housed in the Rodent Habitat ground controls and vivarium ground controls that are responsible for driving the differences between the two habitats: GLDS-21, GLDS-111, GLDS-25, and GLDS-63. Analysis to determine the significant genes was carried out as described above between the Rodent Habitat (previously AEM) and vivarium controls independently for each dataset. PCA plots showed grouping of the biological replicates (Figure 4 shows the PCA plots for GLDS-21). From the pre-processed data, we determined the leading-edge genes from the different GSEA gene sets. Using the genes with 1.2-fold-change (log2), we were able to predict the genes involved with predictions for upstream regulators, canonical pathways, and biofunctions. For each dataset we then found the common/overlapping genes involved for all the genes (Figure 5). These genes are now believed to be driving the response between the rodents in the Rodent Habitats (or AEM) and vivarium controls. Network representation of how these key genes connect shows the central hubs for each dataset being analyzed (Figure 6). For example, MAPK1 is the central hub for STS-108 skeletal muscle tissues from mice (Figure 6A). This would be interpreted as the gene that is driving the key genes and most likely the central player for causing biological differences for mice housed in Rodent Habitats versus the vivarium cages. In our previous work, we discuss how these key genes are associated with CO2 response from the existing scientific literature and how these genes can be responsible for biological changes observed in the mice36.
Taking a systems biology approach, we next determined a "master regulator" that connects all the datasets/tissues and is potentially responsible for universal biological effects in rodents housed in AEMs compared to vivarium cages. This was done by determining the gene from all the datasets that is the most connected when constructing a network from all the key genes. We were able to show that MAPK1 is the most connected gene and central hub from all the key genes (Figure 7). To confirm if MAPK1 might be responsible for biological changes in mice from the higher CO2 levels in AEMs, we looked through the scientific literature for supporting evidence. We found several studies indicating the correlation of MAPK1 with CO259 and hypoxia19,60,61.
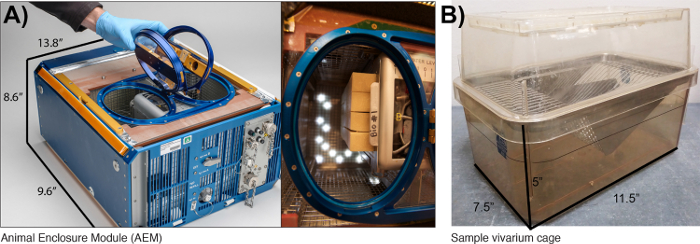

Figure 1: The Rodent Habitat (previously AEM) compared to the vivarium cages. (A) Image of the AEM cage provided by NASA (Credits: NASA/Dominic Hart). (B) The standard vivarium cage that is currently used (photo taken by our laboratory). This figure has been modified from Beheshti et al.36. Please click here to view a larger version of this figure.
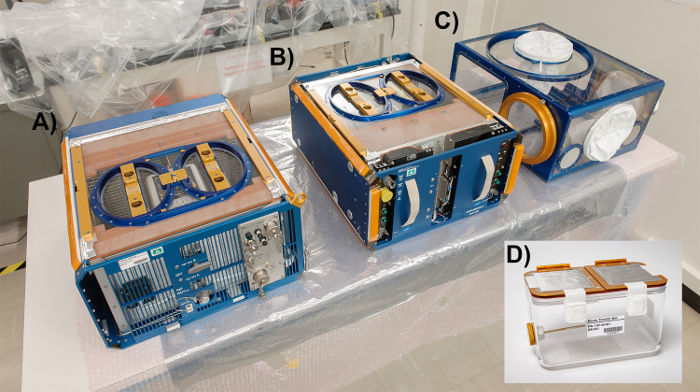

Figure 2: The Rodent Habitat Hardware System with the three different modules involved during transportation to and from the space missions. The left module (A) is the Rodent Habitat module (previously AEM), the center module (B) is the Transporter, and the right module (C) is the Animal Access Unit (AAU). (D)The Mouse Transfer Box (MTB). (Credits: NASA/Dominic Hart). Please click here to view a larger version of this figure.
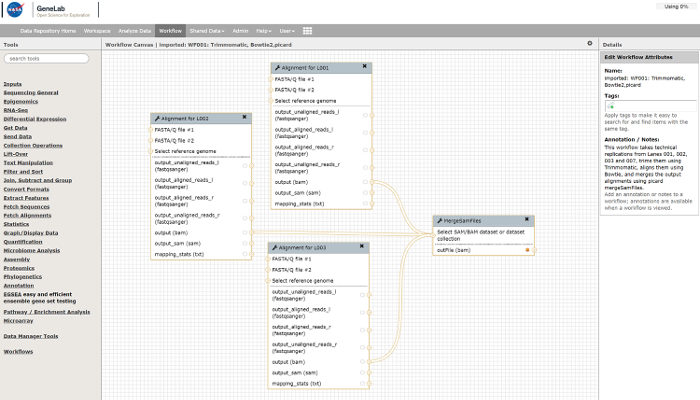
Figure 3: Example analysis workflow which can be used in the GeneLab Galaxy interface to process RNA-seq data. Please click here to view a larger version of this figure.
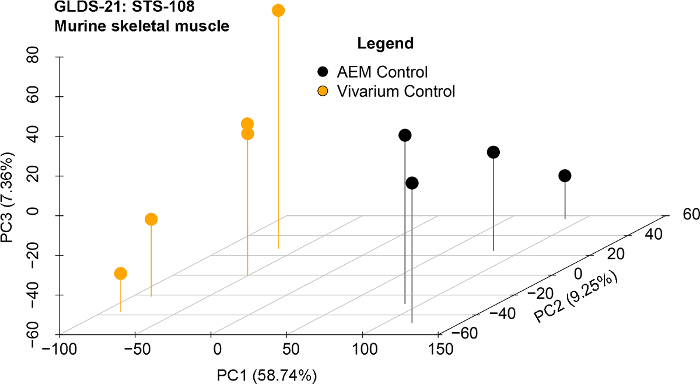
Figure 4: Principal component analysis (PCA) of representative dataset after pre-processing steps. GLDS-21 dataset for AEM vs. vivarium cage is shown for the murine skeletal muscle from the STS-118 mission. Please click here to view a larger version of this figure.
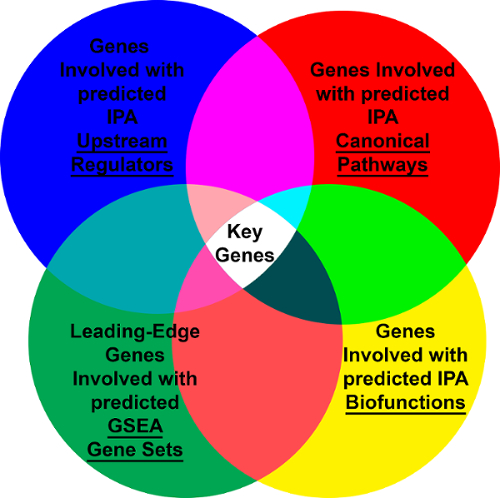
Figure 5: Venn diagram representing what key genes are determined using different pathway prediction tools. Please click here to view a larger version of this figure.
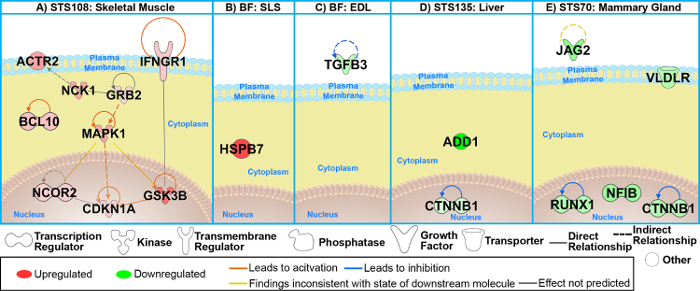
Figure 6: The key genes determined for all conditions and murine tissues between the AEM vs. vivarium cages. (A-E) Network representation of the key genes for each dataset/rodent tissue. Log2 fold-changes (with a cutoff of 1.2-fold-change) to the gene expression were used to obtain different shades of green for fold-change in downregulated genes, while different shades of red depict fold-change in upregulated genes. The darker the shade of green or red, the greater the fold-change. This figure has been modified from Beheshti et al.36. Please click here to view a larger version of this figure.
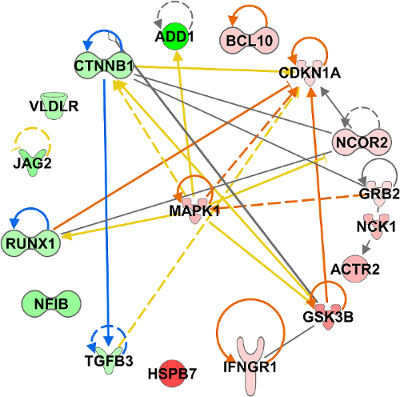
Figure 7: Determining the "master regulator" for rodents in Rodent Habitat housing compared to vivarium cages. Connections between all individual key genes (Figure 6) were determined and displayed as a network through IPA. Network is represented as a radial plot with the most connected key gene, MAPK1, in the center. Please click here to view a larger version of this figure.
Supplemental Figure 1: GeneLab-GenomeSpace Integration with ISACreator for Streamlining Data Processing Operations. Please click here to download this figure.
Supplemental Figure 2: Screenshot of GeneLab searches using federation/integration with heterogeneous bioinformatics external databases (GEO, PRIDE, MG-RAST). Please click here to download this figure
Supplemental Figure 3: Screenshot of the GeneLab collaborative workspace showing the user account management, and access controls (e.g., private, shared, public folders). Please click here to download this figure
Discussion
The NASA GeneLab platform is a comprehensive omics database and analysis platform that will allow the scientific community to generate novel hypotheses related to space biology. Here we have presented a comprehensive procedure for rodent experiments from the beginning of spaceflight to the generation of novel hypothesis from analyzing data utilizing a publicly available space biology platform. In addition, we have also provided an extensive protocol on an unbiased systems biology analysis for identifying key genes driving the system being studied. We have used our recent study36 as an example of how this protocol is effectively utilized to generate a novel hypothesis for space biology. We hope that this helps investigators better understand how spaceflight experiments are performed and how data from them lead to the data available on GeneLab, and ultimately allow for clearer interpretation of publicly available space biology omics data.
There are several critical steps within our protocol regarding both rodent spaceflight experiments and analysis of the data produced. Understanding the Rodent Habitat setup is critical to develop and design the optimal experiment for spaceflight. This would specifically entail the protocol and description we have provided in step 1 of our protocol. Once an investigator fully understands the different conditions existing in the Rodent Habitats compared to vivarium cages, the biological results being interpreted can be correlated properly to the environmental conditions in space. In additions, modifications to the Rodent Habitat cannot be done, since the Rodent Habitat has been optimally designed and approved by NASA for use of spaceflight.
To interpret the biological results, we have provided a thorough protocol on every step involved from uploading your data to GeneLab to analysis of the data to generate novel space biology hypothesis. Although all the steps are important in understanding how to generate data, the most critical steps for data analysis are steps 9 and 10. Step 9 provides a protocol to analyze transcriptomic data using an unbiased systems biology method to determine genes/pathways that are truly driving the experimental condition being analyzed. Step 10 is critical as it provides users with an easy methodology to analyze omics GeneLab datasets using the GeneLab platform. Modifications to the protocol provided can be done for some steps regarding analyzing data. Specifically, steps 9.4–9.6 can be done using R programming or any other favorite tools that the user prefers. Depending on the dataset, different statistics and fold-change cutoffs can be used to determine the significantly regulated genes. In addition, for determining the key genes in steps 9.5 and 9.6, the user can modify this protocol and use any tool that utilizes the significantly regulated genes to predict functions. The important concept is that using multiple predictive functional omics tools allows for determination of genes involved with the majority of functions being regulated in the system being studied.
The GeneLab platform continues to develop, and while the analyses described here were performed after data download, the next phase of GeneLab will allow for analysis of omics data directly on GeneLab platform, which will provide an easy workflow to generate processed data for higher-order analysis. Moreover, whereas we have focused on a protocol for interpreting transcriptomic data, GeneLab contains a wide variety of omics data including proteomic, genomic, metabolomic, and epigenomic data. The eventual platform will contain pipelines and guidelines for analysis of these different types of omics. The last phase of GeneLab will also implement a system level visualization interface to allow the basic user to easily generate space biology hypotheses.
Lastly, our systems biology analysis provides a unique and unbiased method to determine the key driving genes/pathways in any system being studied using omics datasets. We have used this methodology in several different independent studies with great success to determine the key drivers involved36,45,46,47,48,49,50. In a cancer related omics study, using this methodology we experimentally validated that our predicted key genes/pathways were actually driving the drug treatment response by knocking out the key genes in vitro45. We observed, as we had predicted through this protocol, that the treatment was not effective anymore due to the absence of the key genes. We believe that this unbiased systems biology protocol can be a useful tool to determine key pathways for any omics study.
This protocol provides a quick and efficient method for the generation of novel space biology hypotheses. The data generated from GeneLab can be leveraged by investigators for future funding opportunities, experimental validation, and potential targets for development of countermeasures against microgravity and space radiation. The protocol provided here will permit for future space biology investigations to occur with optimal efficiency to allow for safe long-term space missions.
Acknowledgements
We would like to thank Alison French at NASA Ames Life Science Data Archive for her assistance with obtaining the video related to the Rodent Habitats and overall help with obtaining cage related information. We also like to thank Marla Smithwick at NASA Ames Research Center for her help with obtaining the proper information. Research funding was provided by the GeneLab Project at NASA Ames Research Center, through NASA's Space Biology Program in the Division of Space Life and Physical Sciences Research and Applications (SLPSRA). Any use of trade names is for descriptive purposes only and does not imply endorsement by the US Government.
Materials
| Name | Company | Catalog Number | Comments |
| C57BL/6 Mice | The Jackson Laboratoy | C57BL/6J | C57BL/6 mice were used for datasets related to Rodent Research-1 experiments |
| BALB/C Mice | Taconic | BALB | BALB/C mice were used for datasets related to Rodent Research-3 experiments |
| Vivarium Cages | Charles River Laboratory | Standard murine cages purchased from Charles River Laboratory | |
| Rodent Habitat | NASA | This cage and all components are built internally at NASA | |
| RNAlater | ThermoFisher Scientific | AM7020 | RNAlater is used to store the tissue for further RNA isolation |
References
- Cortese, F. Vive la radioresistance!: converging research in radiobiology and biogerontology to enhance human radioresistance for deep space exploration and colonization. Oncotarget. 9 (18), 14692-14722 (2018).
- Beheshti, A. NASA GeneLab Project: Bridging Space Radiation Omics with Ground Studies. Radiation Research. , (2018).
- Fernandez-Gonzalo, R., Baatout, S., Moreels, M. Impact of Particle Irradiation on the Immune System: From the Clinic to Mars. Frontiers in Immunology. 8, 177 (2017).
- Bloomfield, S. A., Martinez, D. A., Boudreaux, R. D., Mantri, A. V. Microgravity Stress: Bone and Connective Tissue. Comprehensive Physiology. 6, 645-686 (2016).
- Giuliani, A. High-Resolution X-Ray Tomography: A 3D Exploration Into the Skeletal Architecture in Mouse Models Submitted to Microgravity Constraints. Frontiers in Physiology. 9, 181 (2018).
- Boice, J. D. The Final Frontier-Research Relevant to Mars. Health Physics. 112 (4), 392-397 (2017).
- Chancellor, J. C. Limitations in predicting the space radiation health risk for exploration astronauts. NPJ Microgravity. 4, 8 (2018).
- Cucinotta, F. A. Space radiation risks for astronauts on multiple International Space Station missions. PLoS One. 9 (4), e96099 (2014).
- Cucinotta, F. A. Review of NASA approach to space radiation risk assessments for Mars exploration. Health Physics. 108 (2), 131-142 (2015).
- Frippiat, J. P. Towards human exploration of space: The THESEUS review series on immunology research priorities. NPJ Microgravity. 2, 16040 (2016).
- Goel, N. Effects of sex and gender on adaptation to space: behavioral health. Journal of Women's Health (Larchmt). 23 (11), 975-986 (2014).
- Mortazavi, S. M. J., Bevelacqua, J. J., Fornalski, K. W., Welsh, J., Doss, M. Comments on "Space: The Final Frontier-Research Relevant to Mars". Health Physics. 114 (3), 344-345 (2018).
- Blottner, D. Morphological, physiological and behavioural evaluation of a 'Mice in Space' housing system. Journal of Comparative Physiology B. 179 (4), 519-533 (2009).
- Karouia, F., Peyvan, K., Pohorille, A. Toward biotechnology in space: High-throughput instruments for in situ biological research beyond Earth. Biotechnology Advances. 35 (7), 905-932 (2017).
- Shen, H. Effects of spaceflight on the muscles of the murine shoulder. The FASEB Journal. 31 (12), 5466-5477 (2017).
- Spatz, J. M. Sclerostin antibody inhibits skeletal deterioration in mice exposed to partial weight-bearing. Life Sciences in Space Research (Amst). 12, 32-38 (2017).
- Tascher, G. Proteome-wide Adaptations of Mouse Skeletal Muscles during a Full Month in Space. Journal of Proteome Research. 16 (7), 2623-2638 (2017).
- Pecaut, M. J. Is spaceflight-induced immune dysfunction linked to systemic changes in metabolism?. PLoS One. 12 (5), e0174174 (2017).
- Ward, C. Effects of spaceflight on the immunoglobulin repertoire of unimmunized C57BL/6 mice. Life Sciences in Space Research (Amst). 16, 63-75 (2018).
- Rettig, T. A., Ward, C., Pecaut, M. J., Chapes, S. K. Validation of Methods to Assess the Immunoglobulin Gene Repertoire in Tissues Obtained from Mice on the International Space Station. Gravitational and Space Research. 5 (1), 2-23 (2017).
- Allen, D. L. Effects of spaceflight on murine skeletal muscle gene expression. Journal of Applied Physiology (1985). 106 (2), 582-595 (2009).
- Moyer, E. L. Evaluation of rodent spaceflight in the NASA animal enclosure module for an extended operational period (up to 35 days). NPJ Microgravity. 2, 16002 (2016).
- Shimbo, M. Ground-based assessment of JAXA mouse habitat cage unit by mouse phenotypic studies. Experimental Animals. 65 (2), 175-187 (2016).
- Aseyev, N. Adaptive Changes in the Vestibular System of Land Snail to a 30-Day Spaceflight and Readaptation on Return to Earth. Frontiers in Cellular Neuroscience. 11, 348 (2017).
- Markina, E., Andreeva, E., Andrianova, I., Sotnezova, E., Buravkova, L. Stromal and Hematopoietic Progenitors from C57/BI/6N Murine Bone Marrow After 30-Day "BION-M1" Spaceflight. Stem Cells and Development. , (2018).
- Radugina, E. A. Exposure to microgravity for 30 days onboard Bion M1 caused muscle atrophy and impaired regeneration in murine femoral Quadriceps. Life Sciences in Space Research (Amst). 16, 18-25 (2018).
- Albi, E. Reinterpretation of mouse thyroid changes under space conditions: the contribution of confinement to damage. Astrobiology. 14 (7), 563-567 (2014).
- Cancedda, R. The Mice Drawer System (MDS) experiment and the space endurance record-breaking mice. PLoS One. 7 (5), e32243 (2012).
- Neutelings, T. Skin physiology in microgravity: a 3-month stay aboard ISS induces dermal atrophy and affects cutaneous muscle and hair follicles cycling in mice. NPJ Microgravity. 1, 15002 (2015).
- Anselm, V., Novikova, S., Zgoda, V. Re-adaption on Earth after Spaceflights Affects the Mouse Liver Proteome. International Journal of Molecular Sciences. 18 (8), (2017).
- Baqai, F. P. Effects of spaceflight on innate immune function and antioxidant gene expression. Journal of Applied Physiology (1985). 106 (6), 1935-1942 (2009).
- Blaber, E. A., Pecaut, M. J., Jonscher, K. R. Spaceflight Activates Autophagy Programs and the Proteasome in Mouse Liver. International Journal of Molecular Sciences. 18 (10), (2017).
- Jonscher, K. R. Spaceflight Activates Lipotoxic Pathways in Mouse Liver. PLoS One. 11 (4), e0152877 (2016).
- Moskaleva, N. Spaceflight Effects on Cytochrome P450 Content in Mouse Liver. PLoS One. 10 (11), e0142374 (2015).
- Beheshti, A., Cekanaviciute, E., Smith, D. J., Costes, S. V. Global transcriptomic analysis suggests carbon dioxide as an environmental stressor in spaceflight: A systems biology GeneLab case study. Scientific Reports. 8 (1), 4191 (2018).
- National Resource Council. . Nutrient Requirements of Laboratory Animals, Fourth Revised Edition, 1995. , (1995).
- . NASA GeneLab Data System Available from: https://genelab-data.ndc.nasa.gov/genelab/ (2018)
- . GeneLab FAQ Available from: https://genelab.nasa.gov/faq/#6 (2018)
- . GeneLab Workspace Available from: https://genelab.nasa.gov/faq/#6 (2017)
- . GeneLab Data Submission Guide Available from: https://genelab-data.ndc.nasa.gov/genelab/help/GeneLab_Submission_Guide_2.0.pdf (2017)
- . GeneLab repository Available from: https://genelab-data.ndc.nasa.gov/genelab/projects (2017)
- Qu, K. Integrative genomic analysis by interoperation of bioinformatics tools in GenomeSpace. Nature Methods. 13 (3), 245-247 (2016).
- . ISACreator Available from: https://isa-tools.org/category/isacreator/index.html (2014)
- Ravi, D. Proteasomal Inhibition by Ixazomib Induces CHK1 and MYC-Dependent Cell Death in T-cell and Hodgkin Lymphoma. Cancer Research. 76 (11), 3319-3331 (2016).
- Wage, J. Proton irradiation impacts age-driven modulations of cancer progression influenced by immune system transcriptome modifications from splenic tissue. Journal of Radiation Research. 56 (5), 792-803 (2015).
- Beheshti, A. Tumor-host signaling interaction reveals a systemic, age-dependent splenic immune influence on tumor development. Oncotarget. 6 (34), 35419-35432 (2015).
- Beheshti, A., Neuberg, D., McDonald, J. T., Vanderburg, C. R., Evens, A. M. The Impact of Age and Sex in DLBCL: Systems Biology Analyses Identify Distinct Molecular Changes and Signaling Networks. Cancer Informatics. 14, 141-148 (2015).
- Beheshti, A. Host age is a systemic regulator of gene expression impacting cancer progression. Cancer Research. 75 (6), 1134-1143 (2015).
- Beheshti, A., Peluso, M., Lamont, C., Hahnfeldt, P., Hlatky, L. Proton irradiation augments the suppression of tumor progression observed with advanced age. Radiation Research. 181 (3), 272-283 (2014).
- Bolstad, B. M., Irizarry, R. A., Astrand, M., Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 19 (2), 185-193 (2003).
- Saeed, A. I. TM4 microarray software suite. Methods in Enzymology. 411, 134-193 (2006).
- Subramanian, A. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America. 102 (43), 15545-15550 (2005).
- Kuehn, H., Liberzon, A., Reich, M., Mesirov, J. P. Using GenePattern for gene expression analysis. Current Protocols in Bioinformatics. , (2008).
- Reich, M. GenePattern 2.0. Nature Genetics. 38 (5), 500-501 (2006).
- Afgan, E. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2016 update. Nucleic Acids Research. 44 (W1), W3-W10 (2016).
- Xu, Y. J., Elimban, V., Dhalla, N. S. Suppression of phosphorylated MAPK and caspase 3 by carbon dioxide. Molecular and Cellular Biochemistry. 436 (1-2), 23-28 (2017).
- Sang, N. MAPK signaling up-regulates the activity of hypoxia-inducible factors by its effects on p300. Journal of Biological Chemistry. 278 (16), 14013-14019 (2003).
- Seta, K. A., Kim, R., Kim, H. W., Millhorn, D. E., Beitner-Johnson, D. Hypoxia-induced regulation of MAPK phosphatase-1 as identified by subtractive suppression hybridization and cDNA microarray analysis. Journal of Biological Chemistry. 276 (48), 44405-44412 (2001).
This article has been published
Video Coming Soon
ABOUT JoVE
Copyright © 2024 MyJoVE Corporation. All rights reserved