Quantitative Polymerase Chain Reaction-based Analyses of Murine Intestinal Microbiota After Oral Antibiotic Treatment
In This Article
Summary
Here we provide detailed protocols for the oral administration of antibiotics to mice, collection of fecal samples, DNA extraction and quantification of fecal bacteria by qPCR.
Abstract
The gut microbiota has a central influence on human health. Microbial dysbiosis is associated with many common immunopathologies such as inflammatory bowel disease, asthma and arthritis. Thus, understanding the mechanisms underlying microbiota-immune system crosstalk is of crucial importance. Antibiotic administration, while aiding pathogen clearance, also induces drastic changes in the size and composition of intestinal bacterial communities which can have an impact on human health. Antibiotic treatment in mice recapitulates the impact and long-term changes in human microbiota from antibiotic treated patients, and enables investigation of the mechanistic links between changes in microbial communities and immune cell function. While several methods for antibiotic treatment of mice have been described, some of them induce severe dehydration and weight-loss complicating the interpretation of the data. Here, we provide two protocols for oral antibiotic administration which can be used for long-term treatment of mice without inducing major weight-loss. These protocols make use of a combination of antibiotics that target both Gram-positive and Gram-negative bacteria and can be provided either ad libitum in the drinking water or by oral gavage. Moreover, we describe a method for the quantification of microbial density in fecal samples by qPCR which can be used to validate the efficacy of the antibiotic treatment. The combination of these approaches provides a reliable methodology for the manipulation of the intestinal microbiota and the study of the effects of antibiotic treatment in mice.
Introduction
The mammalian gastrointestinal mucosa is a unique environment colonized by a highly complex mixture of microorganisms that establish a mutualistic relationship with the host. The defense system of the intestinal mucosa comprises an epithelial layer and a plethora of immune cells that restrict commensals within the intestine while preserving their number and diversity. Conversely, commensal organisms are required for the development of a fully functional immune system. While interactions between host and commensal bacteria are normally beneficial, it is becoming increasingly clear that dysregulated immune system-microbiota crosstalk can favor the development of chronic inflammatory diseases, such asinflammatory bowel disease, arthritis or asthma1,2.
The gut microbiota can be altered by various factors, but perhaps the most drastic changes are induced by antibiotic treatment that severely alters both the size and composition of bacterial communities3,4. While the benefits of antibiotics to treat infections are unquestionable, the microbiota changes induced by antibiotic exposure in humans can also modify immune defenses which can lead to detrimental effects on health. For instance, antibiotic treatment in humans has been linked to an increased risk of Clostridium difficile-induced diarrhea, asthma and certain types of cancer3. Antibiotic treatment in mice recapitulates the impact and long-term alterations found in gut communities of antibiotic-treated patients, and has enabled investigation of the mechanistic links between changes in microbial communities and immune cell function. However, several reports have shown that administration of antibiotics on the drinking water ad libitum results in very noticeable weight loss as mice refrain from drinking water, presumably due to its foul taste5,6. Thus, in these models the severe dehydration concomitant to oral antibiotic administration may complicate the interpretation of experiments aiming to identify the effect of antibiotic treatment in immune cell function.
Several approaches can be used to explore the size and composition of microbial communities in the intestinal compartment7. Next generation sequencing technologies have provided invaluable data on this matter8, however these methods are relatively expensive and require expert bioinformatic analyses for interpretation of the data. On the other hand, traditional microbiological culture methods allow detection of the bacterial species, but they have low sensitivity and a large fraction of commensal bacteria (particularly anaerobes) are very difficult or impossible to cultivate with currently available methods8. Quantitative polymerase chain reaction (qPCR) techniques are being used increasingly for quantification and identification of fecal bacterial species, as they provide a fast and reliable culture-independent measure of total microbial load. Accordingly, qPCR methods have proved useful to study changes in the microbiota associated with age or with progression of several diseases including inflammatory bowel disease9,10. In line with this, qPCR methods provide a fast and cost-effective approach to validate the effect of various treatments (including antibiotics) in fecal bacterial loads and microbiota composition10,11,12.
Here, we present a detailed step-by-step account of two distinct protocols for oral antibiotic administration to mice, fecal sample collection, DNA extraction, preparation of standards and quantification of bacteria in fecal samples by qPCR. These protocols provide a reliable method to manipulate the intestinal microbiota in mice and to study of the effects of antibiotic treatment in intestinal homeostasis and disease.
Protocol
Experiments described here were performed using 6-8 weeks old wild-type (C57BL6/J) mice maintained in a specific pathogen free (SPF) facility. All animal experiments were approved by the King’s College London and the Francis Crick Institute Animal Welfare and Ethical Review Body and the United Kingdom Home Office. Prior to beginning any animal procedure, ensure that the appropriate permissions are obtained through the local institution/organization.
1. Administration of Antibiotics
NOTE: Two alternative methods for antibiotic treatment are provided: oral gavage (step 1.1) and administration of antibiotics via drinking water (step 1.2).
- Oral gavage
- Prepare stocks of individual antibiotics by dissolving them in autoclaved water at the following concentrations: Ampicillin at 100 mg/mL, Gentamicin at 100 mg/mL, Neomycin at 100 mg/mL, Metronidazole at 10 mg/mL and Vancomycin at 100 mg/mL. Filter sterilize using a 0.45 µm filter. Aliquot and store at -20 °C.
- Prepare a cocktail of antibiotics by mixing the stocks prepared above. For a volume of 1 mL mix 50 µL of Ampicillin (5 mg/mL final), 50 µL of Gentamicin (5 mg/mL final), 50 µL of Neomycin (5 mg/mL final), 500 µL of Metronidazole (5 mg/mL final), 25 µL of Vancomycin (2.5 mg/mL final) and 325 µL of water. Prepare the cocktail fresh before use.
- Load the antibiotic mix into a sterile 1 mL syringe and fix an appropriate gauge gavage needle (20 G for 15–20 g mice) while eliminating any bubbles.
- Gavage mice with 200 µL of antibiotic mix.
- Grab the skin over the mouse shoulder firmly, stretch the head and neck to straighten the esophagus. Direct the ball-tip of the feeding needle along the roof of the mouth and toward the back of the pharynx. Then, gently pass it down into the esophagus and inject the 200 μL solution.
- Administer the antibiotic cocktail once daily for the duration of the experiment.
- Antibiotics in the Drinking Water
CAUTION: Carefully monitor mouse weight daily for the first two weeks of antibiotic administration in the drinking water.- Prepare a cocktail of antibiotics by dissolving the following in 1 L of autoclaved water: 1 g of Ampicillin (1 g/L final), 1 g of Neomycin (1 g/L final), 1 g of Metronidazole (1 g/L final), 0.5 g of Vancomycin (0.5 g/L final) and 8 sachets (0.75 g each) of artificial sweetener (60 g/L final). Shake until dissolved. Store at 4 °C.
NOTE: Sweetener is added to hide the flavor of antibiotics and prevent mice dehydration. While several sweetener brands may work, the final concentration required may be different for each specific brand. - Fill the antibiotic cocktail into a water bottle (~100 mL/bottle) and place the bottle on a mouse cage.
NOTE: Use brown bottles or cover the bottles with foil to protect antibiotics from light. - Replace the antibiotic cocktail with fresh stock twice a week for the duration of the experiment.
- Prepare a cocktail of antibiotics by dissolving the following in 1 L of autoclaved water: 1 g of Ampicillin (1 g/L final), 1 g of Neomycin (1 g/L final), 1 g of Metronidazole (1 g/L final), 0.5 g of Vancomycin (0.5 g/L final) and 8 sachets (0.75 g each) of artificial sweetener (60 g/L final). Shake until dissolved. Store at 4 °C.
2. Collection of Fecal Samples from Stool, Ileum Content, and Ileum Wall
- Weigh and label 2 mL autoclaved tubes for sample collection.
- For collection of fresh stool samples, place each mouse in a restrainer and collect fecal pellets directly from the anus in a collection tube.
NOTE: Samples can also be obtained by placing mice in a clean autoclaved cage, and collecting stool samples with clean sterilized forceps. - Euthanize the mice with CO2 asphyxiation followed by cervical dislocation.
- Lay a mouse carcass with the abdomen fully exposed and spray the abdominal area with 70% ethanol.
- Using sterilized forceps and scissors, make a transverse incision in the abdomen to expose the peritoneum without damaging any internal tissues. Lift the peritoneum and make an incision to expose the intestines.
- Remove the intestines (from colon to stomach) with the forceps and scissors and place it in a sterile Petri dish.
- Carefully use forceps to tease the small intestine (SI) away from the mesenteric arteries and fat. Extend the intestine and place it on a clean lab wipe.
- With a ruler, measure and cut 4 cm of the distal ileum of the SI (the closest part to the cecum). Cut and discard the 1 cm of intestine proximal to the cecum. There will be a 3 cm portion of ileum left which will be used to collect intestinal bacteria.
- Hold the ileum portion (3 cm) over a 2 mL sterile tube. Collect the intestinal content by directly extruding the intestine and colleting the sample in the tube. This sample will have bacteria from the ileum content.
- Prepare a 20 mL syringe with cold phosphate buffered saline (PBS) and flush the ileum portion (discard the flow-through).
- Place the ileum portion on a clean paper towel and open it longitudinally with scissors.
- Scrape the inside of the ileum wall with a scalpel.
- Collect any bacteria on the scalpel by washing the scalpel with 1 mL of PBS over a clean centrifuge tube. Spin at 8,000 × g for 5 min to pellet the bacteria and discard the supernatant. This sample will contain bacteria from the ileum wall.
NOTE: Fecal samples from stool, ileum content and wall can be frozen and stored at -80 °C until use. - Weigh the tubes containing the samples from stool and ileum content, and subtract the weight from the empty tubes (from step 2.1) to obtain the fecal weight in each sample.
- Extract bacterial DNA from stool, ileum content and ileum wall samples using commercially available kits. Store DNA samples at -20 °C until use.
3. Quantification of Intestinal Microbiota by qPCR
NOTE: This procedure includes the generation of a standard (step 3.1) and the method for qPCR set-up for standard and fecal samples (step 3.2)
- Generation of a standard for qPCR
- Use polymerase chain reaction (PCR) to amplify the 16S rRNA gene from the genomic DNA extracted from a bacterial culture using the reagents13 and PCR conditions detailed in Table 1.
- Run the PCR product on a 1.5% agarose gel and purify the DNA band using a commercially available DNA extraction kit.
- Ligate the purified DNA fragment using a cloning kit (see the Table of Materials, use a vector containing antibiotic resistance markers and β-galactosidase (LacZ) gene fusion for blue/white colony selection) and transform DH10 competent E. coli following the manufacturer’s instructions. Plate transformation onto Ampicillin (100 µg/mL), X-gal (20 µg/mL) Luria Bertani (LB) agar plates (10 g/L Tryptone, 5 g/L Yeast Extract, 10 g/L NaCl, 15 g/L Agar). Incubate overnight (O/N) at 37 °C.
- Select a single positive (white) colony from the plate. Inoculate it in 5 mL LB broth containing 100 µg/mL Ampicillin and incubate O/N at 37 °C, shaking at 250 rpm.
- From the O/N culture, isolate and purify the plasmid using a commercial miniprep kit according to manufacturer's instructions.
NOTE: It is important to sequence the plasmid insert at this stage, to ensure that the plasmid contains only one copy of the 16S rRNA gene and to determine its length in base-pairs (bp). - Linearize the plasmid with a restriction enzyme which cuts the plasmid only once.
- Purify the linearized plasmid using a commercially available kit.
- Determine the concentration of the plasmid by measuring absorbance at 260 nm using a spectrophotometer.
- Calculate the number of plasmid copies/µL of sample using the following formula14:
number of copies = (amount * 6.022 × 1023) / (length * 1 × 109 * 650)
where amount is the DNA concentration obtained in step 3.1.8 (in ng/µL); and length is the total length of the plasmid (with the insert) in bp. The number of copies will be obtained as copies/µL.
NOTE: There are online tools based on the above formula that enable easy calculation of the number of plasmid copies. - Aliquot and store the standard as needed at -20 °C until use.
- qPCR Set-up for Standard and Fecal Samples
- Thaw the qPCR standard (from step 3.1.10), fecal samples DNA (from step 2.15) and qPCR reagents (from a commercially available kit) on ice.
NOTE: qPCR reagents used in this example include the SYBR Green qPCR reaction mix and forward and reverse primers (Table 2). - Dilute the standard in sterile DNA-free water in a range from 107 to 102 copies/µL (e.g., 1/5 serial dilutions from 107 to 6.4 x 102 copies/µL). Dilute fecal sample DNA to 1/2, 1/5, 1/10.
- Make a master mix reaction for the total number of reactions plus 1 (Table 2).
- Mix 30 µL of the master mix and 5 µL of template (standard, sample or water for negative control)
- Add 10 µL of this mix to each well in a 384-well optical qPCR plate. Perform each reaction in triplicate.
- Seal the qPCR place, centrifuge briefly and load the plate on the qPCR machine programmed with the following cycling conditions: 95 °C for 20 min followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min.
- Obtain CT values for standard and samples.
- Generate a standard curve by plotting the CT values for the standards versus the logarithm of the copy number of the plasmid (as calculated in step 3.1.9). Perform lineal regression of the standard curve.
- Calculate the 16S rDNA copy numbers for fecal samples by interpolating CT values (obtained in step 3.2.2) in the standard curve.
NOTE: 16S rDNA copy numbers for each sample should be corrected considering the dilution factor of the sample (as prepared in step 3.2.2), the final volume nucleic acids that were eluted (in step 2.15) and the quantity of fecal sample (calculated in step 2.14) to obtain gene copies per gram of feces.
- Thaw the qPCR standard (from step 3.1.10), fecal samples DNA (from step 2.15) and qPCR reagents (from a commercially available kit) on ice.
Representative Results
Here we provide two alternative protocols for oral antibiotic treatment of mice. Figure 1 shows percentage of body-weight (related to original base line weight for each animal) in mice treated with antibiotics either by oral gavage (red) or in the drinking water (blue) for 10 consecutive days. No noticeable weight-loss is found in mice that receive antibiotics by oral gavage. However, when mice are treated with antibiotics ad libitum in the drinking water, they lose weight (~10%) within the first few days of antibiotic administration, but recover normal weight gain thereafter (Figure 1). Nonetheless, around 5-10% of mice receiving antibiotics in the drinking water can reach >20% weight loss within the first week of treatment, in which case they are euthanized.
The quantification of bacteria in fecal samples requires the use of an adequate standard curve which is obtained by plotting the log of copy number for the standard (as calculated in step 3.2.2) versus the CT values obtained from qPCR (step 3.2.7). Figure 2A shows a representative example from a standard curve which meets the standard curve performance criteria with a R2 value of 0.99827, a slope of -3.09 and a efficiency ((-1 + 10^(-1/slope))*100) of 110%. R2 values of 0.99 and PCR efficiencies within the range of 90 to 110% are preferred. Within the linear range, the regression analysis equation enables quantification of the 16S rDNA abundance within the fecal samples. Figure 2B shows the number of 16S rDNA copies in fecal stool, SI content and SI wall. In Figure 2B data are shown as 16S rDNA copies/g of fecal sample for fecal stool and SI content. For SI wall, data are presented as total number of 16S rDNA copies obtained from bacteria recovered from the 3 cm of SI wall (as the quantity of starting material is too small to obtain a precise weight).
To evaluate the effect of antibiotics on the density of bacteria in fecal samples, mice were treated with antibiotics by oral gavage daily for 10 days (days 1 to 10) and stool samples were collected before (day 0), at different time-points during antibiotic treatment (days 5 and 10), and 7 days after stopping antibiotic administration (day 17; Figure 3A). As shown in Figure 3B, antibiotic treatment induces a strong decrease in the number of 16S rDNA copies/g of feces detected at days 5 and 10, while the density of bacteria in the feces recovered to normal levels (comparable to pre-treatment) 1 week after antibiotic administration was stopped (day 17).
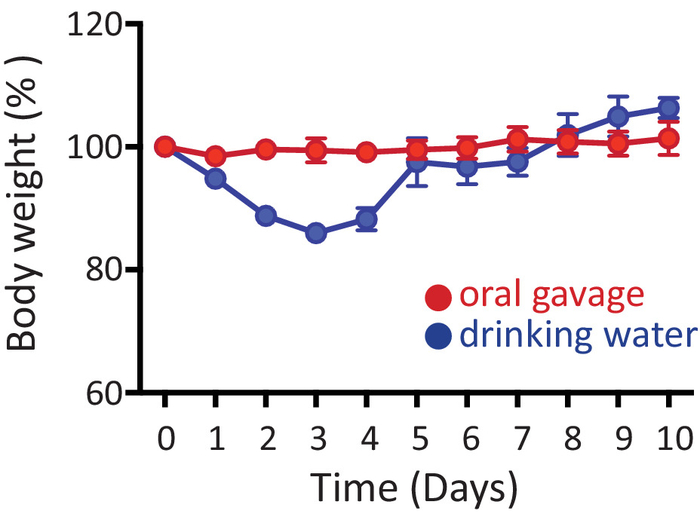
Figure 1: Administration of antibiotics. Mice received antibiotics either by oral gavage (red) or in the drinking water (blue) for 10 consecutive days. Plot shows weights of mice throughout the duration of the experiments relative to original weight before antibiotic administration (day 0). Data are shown as mean ± SEM. Please click here to view a larger version of this figure.
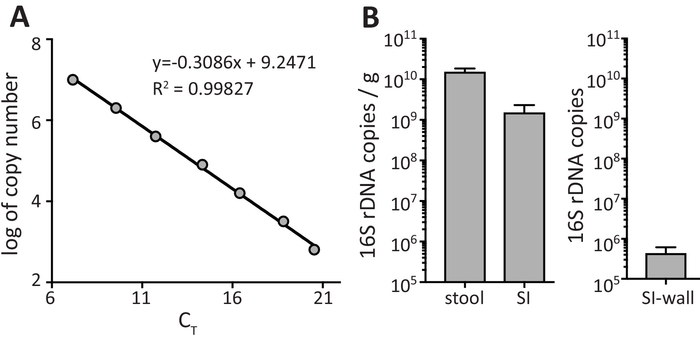
Figure 2: 16S rRNA gene qPCR amplification of standards and fecal samples. (A) Linear regression of standard curve with standard curve descriptors. (B) Calculation of gene abundances from fecal samples. Data are shown as mean ± SEM. Please click here to view a larger version of this figure.
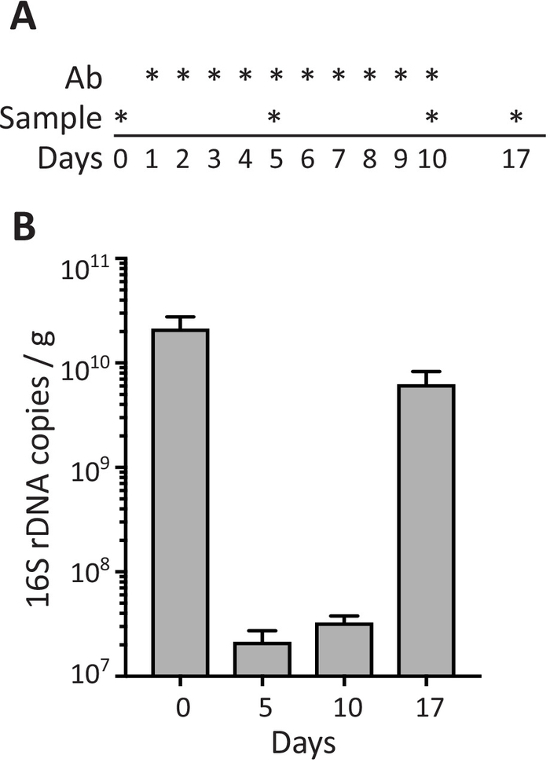
Figure 3: Fecal bacteria during antibiotic treatment. (A) Schematic of the schedule for antibiotic administration by oral gavage (Ab) and sample collection as depicted with *. (B) 16S rDNA copies per gram of feces in stool samples from mice collected at the indicated days. Data are shown as mean ± SEM. Please click here to view a larger version of this figure.
| Reagent | Volume | |
| DNA | 8 µL | |
| Buffer 10X | 2 µL | |
| dNTPs (10mM) | 0.4 µL | |
| Eubacteria - F primer (10 mM) | 1 µL | |
| Eubacteria - R primer (10 mM) | 1 µL | |
| Taq Polimerase | 0.2 µL | |
| H2O | 7.4 µL | |
| Total volume | 20 µL | |
| Primer sequences | ||
| Eubacteria - F primer | 5' ACTCCTACGGGAGGCAGCAGT 3' | |
| Eubacteria - R primer | 5' ATTACCGCGGCTGCTGGC 3' | |
| Cycling conditions | ||
| Temperature | Time | Cycles |
| 94 °C | 5 min | 1x |
| 94 °C | 30 s | 30x |
| 60 °C | 30 s | 30x |
| 72 °C | 1 min | 30x |
| 72 °C | 5 min | 1x |
| 4 °C | ∞ | 1x |
Table 1: PCR reagents and conditions. This table provides the reagents and PCR cycling conditions to amplify 16S rRNA gene from bacterial culture for generation of a standard to use in the qPCR assays. Primer sequences were originally published by Kruglov et al.13.
| Reagent | Volume |
| SYBR Green master mix (2x) | 17.5 µL |
| Eubacteria - F primer (10 mM) | 0.7 µL |
| Eubacteria - R primer (10 mM) | 0.7 µL |
| H2O | 11.1 µL |
Table 2: qPCR master mix. The volumes shown (final volume 35 µL) are for a single sample to be run on triplicate (10 µL each) on a 384-well qPCR plate (accounting for 5 µL extra for pipetting error). The amount may be scaled up according to the number of samples to be analyzed.
Discussion
Here we provide experimental protocols for oral administration of antibiotics to mice and quantification of fecal bacteria by qPCR. The combination of antibiotics used in this protocol (containing ampicillin, gentamicin, neomycin, metronidazole and vancomycin) targets both Gram-positive and Gram-negative bacteria, offering bactericidal activity against a full spectrum of bacteria. Both oral gavage and administration of antibiotics in the drinking water greatly decrease fecal bacterial load5,6,12. Moreover, both treatments have a profound effect on the phenotype of the mice as they develop several characteristics typical of germ-free mice including reduced spleen size and enlarged cecum. The selection of a particular method for antibiotic administration may possibly depend on the length of the experiment as the oral gavage method requires daily administration of antibiotics, being more labor-intensive and possibly causing more discomfort to the animals in the long-run.
For administration of antibiotics in the drinking water, caution must be taken with the addition of the sweetener to the antibiotic mixture as this is a crucial factor to keep mice from dehydration. Several groups have shown how administration of antibiotics in drinking water (without addition of sweetener) leads to very severe and rapid weight-loss with all mice losing more than 20% of initial body weight within the first few days of the experiment5,6. In our protocol, the use of the saccharine-based sweetener seemed to be sufficient to mask the antibiotic taste in the water and mice lost weight in the first few days after antibiotic administration, but recovered their weights quickly after that (Figure 1). Nonetheless, in our experiments 5-10% of mice still reach the human end-point of >20% loss of baseline body weight and needed to be euthanized. We have also tested sucralose-based sweeteners which completely failed to prevent mice dehydration (100% of mice lost >20% of weight) while other authors have published similar failures for aspartame-based sweeteners5,6. Added to this, the age, genetic background and general health status of the mice used for the experiments should be considered, as they may influence weight-loss and animal well-being during antibiotic treatment. Thus, careful monitoring of mice weight and general health status should be performed daily during the first two weeks of oral antibiotic administration.
qPCR methods provide a fast and cost-effective approach for quantification of 16S rRNA in fecal samples. However, some limitations should be considered regarding this technique including: i) the requirement for a reliable high-quality standard; ii) the design and efficiency of the qPCR primers; iii) the fact that microorganisms may have different copy numbers of the 16S rRNA gene, thus gene copies may not directly equal cell counts15. Nonetheless, qPCR is a robust and sensitive method which enables rapid analysis of fecal samples. This method can be particularly useful to quickly validate the effect of various treatments (including antibiotics) in fecal bacterial loads as detailed here. Moreover, although we provide a protocol for quantification of total 16S rRNA, this method can be easily adapted (by designing specific primers16) to enable identification of individual bacterial taxa, thus providing both quantitative and qualitative information about microbiome size and composition.
In summary, we have provided two protocols for oral antibiotic treatment of mice and a qPCR-based method to quantify antibiotic-induced changes in fecal bacteria. Although these protocols can be further optimized and combined with other approaches according to individual experimental needs, they may serve as quick, cost-effective and reliable tools to manipulate the murine intestinal microbiota and to study of the effects of antibiotic treatment in intestinal homeostasis and disease.
Acknowledgements
This work was funded by the UK Medical Research Council (grant to P.B. MR/L008157/1); R.J. was supported by a Marie Curie Intra-European Fellowship (H2020-MSCA-IF-2015-703639); P.M.B. was supported by a studentship from the UK Medical Research Council and King's College London Doctoral Training Partnership in Biomedical Sciences (MR/N013700/1).
Materials
| Name | Company | Catalog Number | Comments |
| Ampicillin sodium salt | Sigma-Aldrich (Merck) | A9518 | |
| Neomycnin trisulfate salt hydrate | Sigma-Aldrich (Merck) | N1876 | |
| Metronidazole | Sigma-Aldrich (Merck) | M3761 | |
| Vancomycin hydrochloride | Sigma-Aldrich (Merck) | V2002 | |
| Gentamicin sulfate salt | Sigma-Aldrich (Merck) | G3632 | |
| Tryptone | Sigma-Aldrich (Merck) | T7293 | |
| Yeast Extract | Sigma-Aldrich (Merck) | Y1625 | |
| NaCL | Sigma-Aldrich (Merck) | S7653 | |
| Sweetener Sweet'n Low | Sweet'N Low | Available in the UK from Amazon.co.uk | |
| X-Gal (5-brom-4-chloro-3-indoyl B-D-galactopyranoside) | Fisher scientific | 10234923 | |
| Phosphate Buffered Saline | Thermo Fisher Scientific (Gibco) | 10010023 | |
| Ultrapure Agarose | Thermo Fisher Scientific (Invitrogen) | 16500500 | |
| RT-PCR grade water | Thermo Fisher Scientific (Invitrogen) | AM9935 | |
| Phusion High-Fidelity DNA Polymerase | New England BioLabs | M0530 | |
| Deoxynucleotide (dNTP) Solution Mix | New England BioLabs | N0447 | |
| iTaq Universal SYBR Green Supermix | Bio-Rad | 1725124 | with ROX |
| TOPO TA cloningTM for sequencing | Thermo Fisher Scientific (Invitrogen) | 450030 | |
| QIAamp fast DNA Stool mini kit | Qiagen | 51604 | |
| QIAprep spin Miniprep kit | Qiagen | 27106 | |
| QIAquick gel extraction kit | Qiagen | 28704 | |
| Syringe filter 0.45µm | Fisher scientific | 10460031 | |
| Swann-MortonTM Carbon steel sterile scalpel blades | Fisher scientific | 11792724 | |
| Syringe (1 ml) | BD Plastipak | 303172 | |
| Syringe (20 ml) | BD Plastipak | 300613 | |
| 1.5ml Crystal clear microcentriguge tube | StarLab | E1415-1500 | |
| 2ml Ultra high recovery microcentrifuge tube | StarLab | I1420-2600 | |
| Oral dosing needles 20Gx38 mm curved (pk/3) | Vet-Tech | DE008A | |
| Sterilin petri dish 50 mm | Scientific Laboratory Supplies | PET2020 | |
| Absolute qPCR plate seals | Thermo Fisher Scientific | AB1170 | |
| MicroAmpTM optical 384-well plate | Thermo Fisher Scientific (Applied Biosystems) | 4309849 | |
| ViiA7TM 7 real-time PCR system with 384-well block | Thermo Fisher Scientific (Applied Biosystems) | 4453536 | |
| Spectrophotometer (Nanodrop 1000) | Thermo Fisher Scientific | ND-1000 | |
| Labnet Prism microcentrifuge | Labnet | C2500 | |
| MultiGene Optimax Thermal cycler | Labnet | TC9610 |
References
- Belkaid, Y., Hand, T. W. Role of the microbiota in immunity and inflammation. Cell. 157 (1), 121-141 (2014).
- Hooper, L. V., Littman, D. R., Macpherson, A. J. Interactions between the microbiota and the immune system. Science. 336 (6086), 1268-1273 (2012).
- Ubeda, C., Pamer, E. G. Antibiotics, microbiota, and immune defense. Trends in Immunology. 33 (9), 459-466 (2012).
- Dethlefsen, L., Huse, S., Sogin, M. L., Relman, D. A. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biology. 6 (11), e280 (2008).
- Reikvam, D. H., et al. Depletion of murine intestinal microbiota: effects on gut mucosa and epithelial gene expression. PLoS One. 6 (3), e17996 (2011).
- Hill, D. A., et al. Metagenomic analyses reveal antibiotic-induced temporal and spatial changes in intestinal microbiota with associated alterations in immune cell homeostasis. Mucosal Immunology. 3 (2), 148-158 (2010).
- Fraher, M. H., O'Toole, P. W., Quigley, E. M. Techniques used to characterize the gut microbiota: a guide for the clinician. Nature Reviews Gastroenterology & Hepatology. 9 (6), 312-322 (2012).
- Eckburg, P. B., et al. Diversity of the human intestinal microbial flora. Science. 308 (5728), 1635-1638 (2005).
- Sokol, H., et al. Low counts of Fecalibacterium prausnitzii in colitis microbiota. Inflammatiry Bowel Diseases. 15 (8), 1183-1189 (2009).
- Bartosch, S., Fite, A., Macfarlane, G. T., McMurdo, M. E. Characterization of bacterial communities in feces from healthy elderly volunteers and hospitalized elderly patients by using real-time PCR and effects of antibiotic treatment on the fecal microbiota. Applied Environmental Microbiology. 70 (6), 3575-3581 (2004).
- Ubeda, C., et al. Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. Journal of Clinical Investigation. 120 (12), 4332-4341 (2010).
- Saez de Guinoa, J., et al. CD1d-mediated lipid presentation by CD11c(+) cells regulates intestinal homeostasis. EMBO Journal. 37 (5), (2018).
- Kruglov, A. A., et al. Nonredundant function of soluble LTalpha3 produced by innate lymphoid cells in intestinal homeostasis. Science. 342 (6163), 1243-1246 (2013).
- Wilhelm, J., Pingoud, A., Hahn, M. Real-time PCR-based method for the estimation of genome sizes. Nucleic Acids Research. 31 (10), e56 (2003).
- Kembel, S. W., Wu, M., Eisen, J. A., Green, J. L. Incorporating 16S gene copy number information improves estimates of microbial diversity and abundance. PLoS Computational Biology. 8 (10), e1002743 (2012).
- Yang, Y. W., et al. Use of 16S rRNA Gene-Targeted Group-Specific Primers for Real-Time PCR Analysis of Predominant Bacteria in Mouse Feces. Applied Environmental Microbiology. 81 (19), 6749-6756 (2015).
This article has been published
Video Coming Soon
ABOUT JoVE
Copyright © 2024 MyJoVE Corporation. All rights reserved