A Tuberculosis Molecular Bacterial Load Assay (TB-MBLA)
In This Article
Summary
We describe a tuberculosis molecular bacterial load assay test performed after heat inactivation of sputum. Heat inactivation renders sputum samples noninfectious and obviates the need for containment level 3 laboratories for tuberculosis molecular tests.
Abstract
Tuberculosis is caused by Mycobacterium tuberculosis (Mtb), a pathogen classified by the United Nations (UN) as a dangerous category B biological substance. For the sake of the workers’ safety, handling of all samples presumed to carry Mtb must be conducted in a containment level (CL) 3 laboratory. The TB molecular bacterial load assay (TB-MBLA) test is a reverse transcriptase quantitative polymerase chain reaction (RT-qPCR) test that quantifies Mtb bacillary load using primers and dual-labelled probes for 16S rRNA. We describe the use of heat inactivation to render TB samples noninfectious while preserving RNA for the TB-MBLA. A 1 mL aliquot of the sputum sample in tightly closed 15 mL centrifuge tubes is boiled for 20 min at either 80 °C, 85 °C, or 95 °C to inactivate Mtb bacilli. Cultivation of the heat inactivated and control (live) samples for 42 days confirmed the death of TB. The inactivated sample is then spiked with 100 µL of the extraction control and RNA is extracted following the standard RNA isolation procedure. No growth was observed in the cultures of heat treated samples. The isolated RNA is subjected to real-time RT-qPCR, which amplifies a specific target in the Mtb 16S rRNA gene, yielding results in the form of quantification cycles (Cq). A standard curve is used to translate Cq into bacterial load, or estimated colony forming units per mL (eCFU/mL). There is an inverse relationship between Cq and the bacterial load of a sample. The limitation is that heat inactivation lyses some cells, exposing the RNA to RNases that cause a loss of <1 log10eCFU/mL (i.e., <10 CFU/mL). Further studies will determine the proportion of very low burden patients that cause false negative results due to heat inactivation.
Introduction
Caused by Mycobacterium tuberculosis (Mtb), over 7 x 106 new cases of tuberculosis (TB) are reported globally of which over 1 x 106 die per year1,2. To reverse the trend, the World Health Organization (WHO) launched a three-pillar approach including developing effective diagnostic and treatment tools3. Classified as a dangerous biological substance B by the UN, working with samples presumed positive for Mtb requires a containment level (CL) of 3. CL3 laboratories are expensive to build and maintain. Consequently, most countries have centralized TB culture services at regional or national levels. This means that smear microscopy is the most available diagnostic tool in the peripheral healthcare facilities.
There is WHO approval to implement rapid molecular tests like Xpert MTB/RIF at level 4 healthcare facilities, mostly situated at district levels4,5. Some districts are quite large and less accessible to some people. While beneficial, Xpert MTB/RIF works by detecting the Mtb DNA. DNA is a stable molecule that survives long after cells have died and thus is not a good standard for measuring viable cells critical for monitoring treatment response5,6. RNA-based assays offer an alternative for accurate measurement of viable cells7,8,9,10,11,12,13. RNA exists in different species of varying stability: ribosomal RNA (rRNA), transfer RNA (tRNA), and messenger RNA (mRNA). Messenger RNA is associated with gene expression and thus the most closely associated with cell activity and viability14. It is important to note that absence of gene expression is not equivalent to cell death because pathogens like Mtb are known to exist in inactive (dormant) but viable states15,16. Stable RNA species such as rRNA are therefore better markers of both active and inactive states of viable cells.
Using Escherichia coli, Sheridan showed that 16S rRNA proportionally increased with bacterial growth measured by colony forming units (CFU) counts17. There was a concurrent decline in CFU counts and 16S rRNA when E. coli bacteria were exposed to antibiotics. The fall in rRNA following cell death was an indicator that it could be used as a marker for cell viability13,17. Drawing from this principle, the TB molecular bacterial load assay (TB-MBLA) was developed for targeting M. tuberculosis 16S rRNA to measure viable TB bacillary load as a marker of treatment response for patients on anti-TB therapy11,18,19. We have further developed and optimized the TB-MBLA to incorporate a cellular extraction control that reflects lysis of M. tuberculosis bacilli and is robust in different environmental settings20. The TB-MBLA procedure requires the first steps of RNA isolation from Mtb to be conducted in a CL3 laboratory until Mtb cells are completely lysed to ensure the safety of the workers. It also includes sample preservation for retrospective batched analysis to be maintained at -80 °C in guanidine thiocyanate, a level 4 toxic substance. To this end, we have used heat to inactivate Mtb and render samples safe for TB-MBLA to be performed in smear microscopy level laboratories.
Use of heat in laboratory and clinical applications has been around for centuries21,22. However, some microorganisms like Mtb are tough to kill, and shorter exposure to heat is insufficient to kill all the cells23,24. A study revealed that 20 min heating of TB cultures at 80 °C killed all Mtb bacilli without destroying the DNA needed for PCR25. Subsequently, a number of laboratory DNA extraction techniques currently heat to 95 °C. We have applied the same principle to show that boiling TB samples at either 80 °C, 85 °C, or 95 °C inactivates Mtb while preserving sufficient RNA for TB-MBLA to be performed. Inactivated culture or sputum can be maintained in tightly closed containers at room temperature or refrigerated for 7 days without reducing the amount of quantifiable rRNA.
The TB-MBLA, currently used as a research use only (RUO) test is adaptable and has been applied to different sample types including sputum, lung tissue, and cerebral spinal fluid. It is yet to be applied on bronchial alveolar fluid, blood, and other sample types. Using sputum as the sample, results from multisite evaluation in Africa (unpublished data) and previous publications18,26 show that the sensitivity of MBLA is consistent with mycobacterium growth indicator tube (MGIT) liquid culture. However, TB-MBLA is faster, giving results in hours as opposed to days or weeks of culture, specific, not affected by non-TB microorganisms in the sample, and gives a quantitative measure of disease severity. The WHO has recently recognized TB-MBLA as a candidate to replace smear microscopy and culture for monitoring TB treatment2.
In this article we describe in detail the heat inactivation and TB-MBLA protocol published in Sabiiti et al.27. This detailed protocol will provide a one-stop visual resource for the TB-MBLA users across the globe.
Protocol
1. Sample Preparation
- Culture
- Working on a clean bench or class 1 cabin, harvest 1 mL aliquots of exponential phase Bacillus Calmette-Guérin (BCG) culture into 15 mL plastic centrifuge tubes. Tightly close the tubes.
NOTE: To process a whole 5 mL sample, five 15 mL centrifuge tubes are required.
CAUTION: A biosafety cabinet is required when working with any TB culture.
- Working on a clean bench or class 1 cabin, harvest 1 mL aliquots of exponential phase Bacillus Calmette-Guérin (BCG) culture into 15 mL plastic centrifuge tubes. Tightly close the tubes.
- Patient sputum specimen
- Working in a well-ventilated space and wearing a nasal mask, carefully open the specimen cup, pipette 1 mL aliquots into 15 mL plastic centrifuge tubes and tightly close the tubes.
NOTE: A wide mouth tip is recommended to pipette sputum. Using a pair of scissors, clip off the fine part of the 1 mL tip mouth to create a wider mouth.
- Working in a well-ventilated space and wearing a nasal mask, carefully open the specimen cup, pipette 1 mL aliquots into 15 mL plastic centrifuge tubes and tightly close the tubes.
2. Heat Inactivation
- Prior to sample preparation, set the water bath to 95 °C.
NOTE: The 95 °C temperature increases the chance of reducing RNase activity and thus preserve more RNA for downstream TB-MBLA. - Transfer the sample tubes to a holding rack immersed in the water bath. Ensure that three-fourths of each sample tube is immersed in the water.
- Boil at 95 °C for 20 min, then transfer the tubes to the bench to cool at room temperature for 5 min before starting the RNA extraction.
NOTE: Complete heat inactivation of M. tuberculosis bacilli and BCG was verified by incubating heat inactivated samples and controls at 37 °C for 42 days to verify growth. An optical density measurement at 600 nm (OD600) was taken at baseline and then weekly for the 42 day incubation period.
3. RNA Extraction
NOTE: The RNA extraction process described here is for the RNA kit listed in the Table of Materials. Other suitable RNA extraction kits by different manufacturers can be used.
- Extraction control (EC) addition
- Transfer 1 mL aliquots of the heat inactivated samples to 1.5 mL tubes. Spike 100 µL of the EC into each sample, close the tube, and mix by inverting the tube upside down 3x.
NOTE: The EC is supplied with the TB-MBLA vital bacteria kit (Table of Materials).
- Transfer 1 mL aliquots of the heat inactivated samples to 1.5 mL tubes. Spike 100 µL of the EC into each sample, close the tube, and mix by inverting the tube upside down 3x.
- Cell sedimentation
- Using a benchtop microcentrifuge, centrifuge the tubes at 20,000 x g for 10 min at room temperature. Pipette off the supernatant, leaving 50 µL of sediment.
- Suspend the sediment in 950 µL of lysis buffer by pipetting up and down, and transfer the whole suspension into the lysing matrix tube supplied with the RNA extraction kit (Table of Materials). Ensure the tubes are tightly closed and label both the lid and the side of the tube.
- For cell lysis, transfer the tubes from step 3.2.2 to a homogenizer. Homogenize the samples for 40 s at 6,000 rpm.
- Nucleic acid purification
- Centrifuge the lysate from step 3.3 at 12,000 x g for 5 min at room temperature.
- Prepare fresh 1 mL tubes and add 300 µL of chloroform into each tube.
- Using a 1 mL tip carefully pipette off the supernatant without touching the lysing matrix.
- Transfer the supernatant to the chloroform containing tubes and vortex for 5 s. Leave the tube to settle for 5 min or longer until three phases (upper, middle, and bottom) are clearly visible.
- Centrifuge at 12,000 x g for 5 min at room temperature. Carefully pipette the upper phase and transfer into fresh 1.5 mL tubes.
- To the tubes in step 3.4.5, add 500 µL of ice-cold 100% ethanol, close the tubes, and mix by gently inverting upside down 3x. Incubate the tubes at -80 °C for 15 min or -20 °C for 30 min and continue the extraction, or leave at -20 °C overnight to complete the extraction the following day.
- Set the microcentrifuge to 4 °C and leave to chill to at least 12 °C before commencing centrifugation. Load the tubes into the microcentrifuge and centrifuge for 20 min at 13,000 x g. Discard the supernatant, replace with 70% ice-cold ethanol, and centrifuge for another 10 min at 13,000 x g.
NOTE: The 70% ethanol should be made with molecular grade nuclease free water. - Discard all the supernatant from step 3.4.7 and transfer the tubes to an incubator set at 50 °C. Incubate for 20 min to dry the RNA/DNA pellet. Keep the tubes partially open to enable evaporation of all ethanol.
- Add 100 µL of nuclease free water to the dry pellet and incubate for 5 min at room temperature. Vortex for 3 s to mix the contents.
NOTE: At this stage the extract may be stored 2−3 days in the fridge or longer at -80 °C until section 3.5 is performed.
- DNA removal
NOTE: This step is crucial because the presence of DNA in the extract invalidates the MBLA result. This section is based on a DNA removal kit (Table of Materials).- Prepare a mix of the enzyme DNase I 10x buffer and DNase I enzyme for the number of samples (10 µL of buffer and 1 µL of DNase per sample) plus 10% extra to cover any loss from pipetting. Mix by vortexing and then pipette 11 µL into each tube containing the RNA extract.
- Mix by vortexing 3 s and then spin briefly (10 s at 13,000 x g) to remove any droplets on the walls. Incubate at 37 °C for 30 min in the hot block or incubator. Add an additional 1 μL of DNase I enzyme directly into each tube, mix well by vortexing, and incubate for a further 30 min at 37 °C.
- Thaw the DNase inactivation reagent 10 min prior to the end of the DNase incubation. Vortex 20 s to ensure a homogenous, milky suspension and then add 10 μL of DNase inactivation reagent into each RNA extract from step 3.5.2.
- Incubate the mixture at room temperature for 5 min. Vortex 3x during the 5 min incubation step.
- Centrifuge the mixture at 13,000 x g for 2 min. Carefully transfer the supernatant to 1.5 mL RNase free tubes without touching any of the inactivation matrix.
- Store the RNA extract in the fridge if running the RT-qPCR on the same day or at -80 °C for long-term storage.
4. Reverse Transcriptase qPCR
- For unknown samples, dilute all RNA extracts to be used in a 1:10 ratio in RNase free water. Mix well by vortexing for 5 s and briefly spin down to remove any droplets or air bubbles.
- For standard samples for a standard curve, take the Mtb and EC RNA standards from the -80 °C freezer and thaw at room temperature. Make seven and six 10-fold dilutions of Mtb and EC standard samples respectively. Change the tips before transferring the mixture from one tube to another.
NOTE: Standard samples are supplied with the TB-MBLA kit. - Master mix preparation
NOTE: Master mix (MM) is a solution of PCR reagents sufficient to amplify all samples, standards, and water for a no template control (NTC). The water used as NTC should be the same water used in the extraction and for preparing the MM. Ensure that the standards, each RNA sample, and its decimal dilution are amplified 2x for the reverse transcriptase positive (RT+) reaction and 1x for the reverse transcriptase negative (RT-) reaction. The RT- reaction is a control to determine the efficiency of DNA removal (Table 1).- Transfer 16 µL of MM into each PCR reaction tube.
- Add 4 µL of RNA extract into each RT+ and RT- reaction tube and water into the NTC reaction tubes.
- Load the reaction tubes into a real time PCR machine and set the PCR conditions as follows: 50 °C for 30 min, 95 °C for 15 min, 40x cycles at 94 °C for 45 s, and 60 °C for 1 min with acquisition with fluorophores that absorb in green and yellow channels.
NOTE: The green channel is the Mtb detection fluorophore and the yellow is the extraction control detection fluorophore.
- Result interpretation
NOTE: Ensure that the duplicate reactions of the same sample do not differ by more than 1 standard deviation. Mtb and EC Cq values higher than 30 are considered negative. See further interpretation details in Table 2.- To interpret the treatment response, convert the Cq values into bacterial load (eCFU/mL) using the standard curve. Read the treatment response as the change in bacterial load over the treatment follow-up period.
NOTE: The fall in bacterial load following treatment signifies a positive response (i.e., anti-TB drugs killing the TB bacteria) while no change or rise in bacterial load implies a negative response, which may mean resistance of TB bacteria to anti-TB drugs or the patient not appropriately adhering to their treatment dose. The fall in bacterial load measured by TB-MBLA correlates with the increase in MGIT time to culture positivity (TTP).
- To interpret the treatment response, convert the Cq values into bacterial load (eCFU/mL) using the standard curve. Read the treatment response as the change in bacterial load over the treatment follow-up period.
5. Transmission Electron Microscopy
- Transfer 2 mL aliquots of heat inactivated cultures and controls into 2 mL microcentrifuge tubes. Centrifuge for 10 min at 20,000 x g.
- Discard the supernatant and suspend the pellet in 700 µL of cell fixation buffer and incubate at room temperature for 5 min to fix the cells.
- Centrifuge the suspension for 30 min at 16,000 x g to obtain a hard pellet. Discard the supernatant and replace with 1% sucrose in phosphate-buffered saline (PBS). Store the pellets in sucrose at 4 °C until sectioning for electron microscopy.
- Sectioning and TEM
NOTE: The protocol below is adapted from Griffiths et al.28.- Cryoprotect the cell pellet by embedding it in 2.1 M sucrose in PBS overnight at 4 °C. Wash 3x with ice-cold water.
- Cool the cryoprotected cell pellets in liquid nitrogen and mount them cryomicrotomy stubs. Using the cryomicrotome, cut ultrathin sections at 90 nm.
- Retrieve the cut sections using the tungsten wire loops of 1:1 mixture of 2% methyl cellulose and 2.1 M sucrose in PBS.
- Transfer the sections to pioloform coated 150 mesh hexagonal copper TEM supports (grids) and store at 4 °C or proceed to step 5.4.5.
- Contrast the grids in uranyl acetate and air-dry in a film of methyl acetate. Wash the grids with ice-cold water followed by two drops of PBS over 5 min and dry for 15−30 min. Examine the sections under TEM following the manufacturer’s guidelines.
NOTE: All labelling steps were performed at ice temperature (liquid temperature) except for the washing steps, which were performed at ambient temperature.
Representative Results
Heat inactivates all M. tuberculosis bacilli
The optical density (OD) of controls (live cells) increased over time, (0.04OD–0.85OD) and no OD change was observed in the heat inactivated samples, signifying growth and no growth respectively (Figure 1)27. Similarly, control clinical sputum grew positive by day 3 in MGIT while heat inactivated clinical samples did not flag positive until the end of incubation. Growth of Mtb in MGIT was confirmed by Ziehl-Neelsen smear microscopy and the antigen MPT6429.
RNA in inactivated samples is stable at 37 °C for 4 days
Heat inactivated samples were incubated at 37 °C to determine whether RNA degrades following heat inactivation of cells. No difference was found between the RNA harvested at Day (D) 0 immediately after heat inactivation and the RNA isolated at D1, 2, 3, and 4 in both BCG cultures (Figure 2A-C) and TB positive sputum (Figure 2D).
Exogenous RNase increases the rate of RNA degradation in the heat inactivated fractions
To determine why RNA was not degrading, RNase A enzyme was exogenously added at 1,000 U/mL before and/or after heat inactivation. This caused RNA loss equivalent to bacterial load 1.5 ± 0.3 -, 1.8 ± 0.2 -, and 1.3 ± 0.1 – log10 eCFU/mL at 80 °C, 85 °C, and 95 °C respectively across 4 days of incubation. There was a difference in the RNA degraded in samples where RNase was added before and/or after heat inactivation (Figure 3A-C).
Sufficient RNA is preserved for TB-MBLA using 16S rRNA as reference marker
The effect of heat inactivation on rRNA was measured in BCG cultures and sputum from TB positive patients. The measured bacterial load of control BCG culture, 5.3 ± 0.2 log10 eCFU/mL, was 0.2 ± 0.1 log10 eCFU/mL higher than the combined 5.1 ± 0.3 log10 eCFU/mL of heat inactivated culture (ANOVA p < 0.0001) at 80 °C, 85 °C, and 95 °C (Figure 4A)27. Similarly, the bacterial load of control patient sputum, 7.1 log10 eCFU/mL, was 0.8 ± 0.1 log10 eCFU/mL higher than the combined 6.3 ± 0.41 log10eCFU/mL at 80 °C, 85 °C, and 95 °C (Figure 4B)27. The bacterial load reduction was <1log in the two types of samples tested.
Sidak’s multiple comparisons test revealed a significant difference between the bacterial load at 95 °C versus that of 80 °C and 85 °C (p = 0.001). No difference was found in TB samples at all temperatures, p = 0.8.
Cell wall integrity is not destroyed by heat in the majority of Mtb cells
Using transmission electron microscopy (TEM) we investigated whether cells were lysed by heat inactivation. Thin sections of paraformaldehyde fixed pellets of cells were made and embedded in an electron rich medium prior to examination by TEM. Inspection of the cells at lower and higher magnification revealed intact cell wall and visible intracellular lipid bodies. Cells morphologically appeared elongated but not lysed. Figure 5 illustrates the morphology of mycobacterium cells at different magnifications. The top two panels reveal intracellular lipid bodies and the unhampered rope-like cording, a morphological characteristic typical of mycobacterium species. The lower two panels are higher magnification expanding the view of lipid bodies and revealing some micro-intracellular structures.
Bacterial load measured by TB-MBLA is inversely correlated to MGIT culture time to positivity
For patients responding to therapy, the bacterial load falls at an average 1 log10 eCFU/mL per week over the course of treatment. Fast responders clear faster, converting to negative (zero bacterial load) by 2 weeks of treatment. The fall in bacterial load measured by TB-MBLA corresponded to the rise in MGIT culture time to positivity (TTP). Figure 6 demonstrates the inverse correlation found between bacterial load and MGIT TTP. The difference, however, is that the TB-MBLA results are available in 4 h and time-to-result is independent of the level of bacterial load. This is in contrast with 5–25 days for MGIT culture tests. Contamination with non-TB bacteria further compromise the results from culture tests.
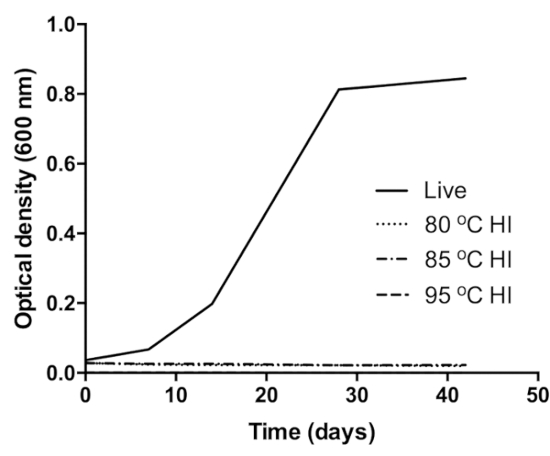
Figure 1: Verification of BCG inactivation at 80 °C (dot pattern curve), 85 °C (dot-dash pattern curve), and 95 °C (dash pattern curve). The control (black curve) was live (unheated) BCG culture inoculated into same growth medium. Growth in the control was confirmed by the increase in the OD of the culture over the incubation period. This figure was modified from Sabiiti et al.27. Please click here to view a larger version of this figure.
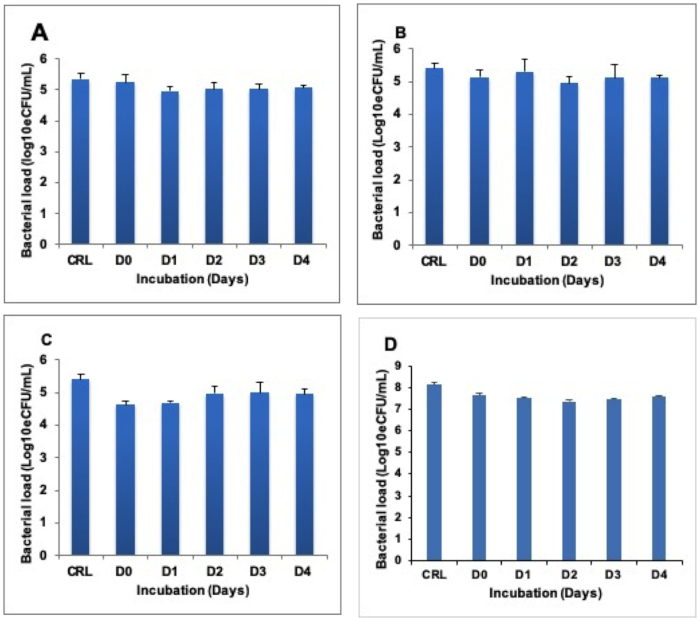
Figure 2: Stability of quantifiable RNA in heat inactivated sample incubated at 37 °C for 4 days (D0–D4). (A), (B), and (C) show BCG cultures inactivated at 80 °C, 85 °C, and 95 °C respectively. (D) shows the TB sputum inactivated at 80 °C. The control is the untreated (live) fraction of the same sample. Error bars = standard deviation. Three repeats, nine replicates per run. Please click here to view a larger version of this figure.
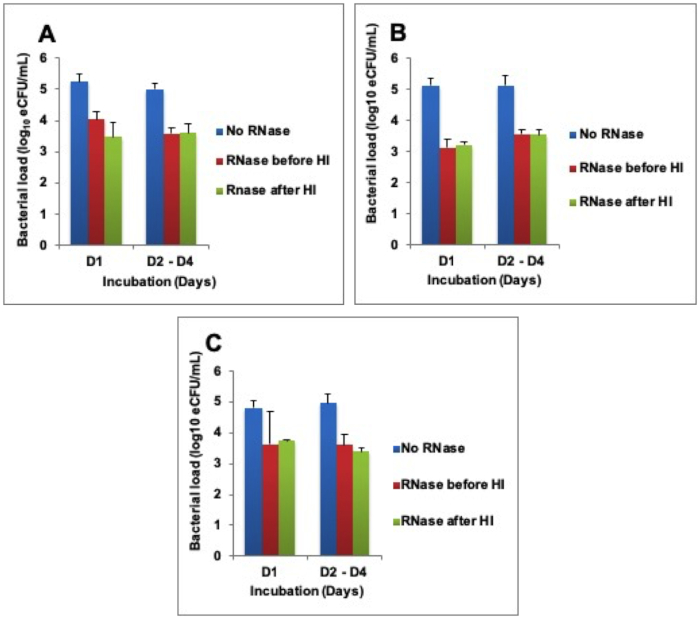
Figure 3: Higher RNA degradation after exogenous addition of RNase A enzyme. (A) Heat inactivation (HI) at 80 °C, (B) HI at 85 °C, and (C) HI 95 °C. Error bars = standard error of the mean. Please click here to view a larger version of this figure.
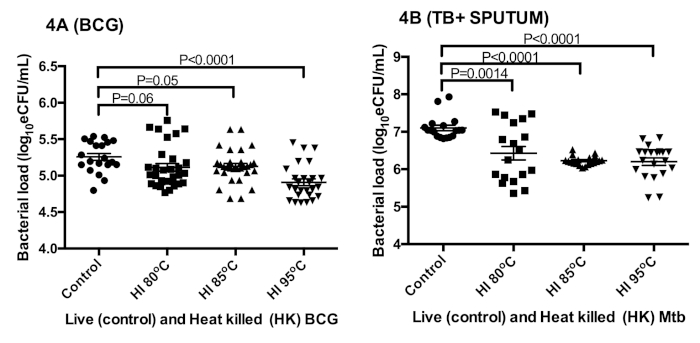
Figure 4: The preservation of sufficient RNA for TB-MBLA bacterial load measurement using 16S rRNA as a marker. (A) Bacterial load estimated from in vitro BCG cultures. (B) Bacterial load estimated from tuberculosis positive sputum. Error bars = standard error of the mean (n = 18 and 20 replicates for A and B respectively). This figure was modified from Sabiiti et al.27. Please click here to view a larger version of this figure.
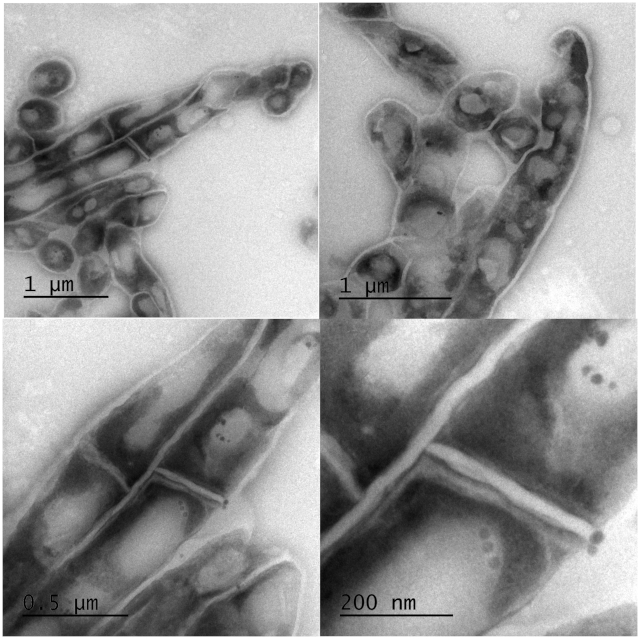
Figure 5: Electron micrographs of intact Mtb bacilli after inactivation at 95 °C for 20 min. Clear intact cell wall and enumerable lipid bodies were observed with TEM. Top: low magnification images of a group of cells revealing unhampered mycobacterial rope-like cording morphology and intracellular lipid bodies. Bottom: high magnification of the top panels to expand the view of lipid bodies and revealing some micro-intracellular structures. Please click here to view a larger version of this figure.
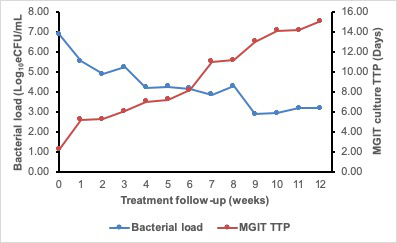
Figure 6: The 12 week treatment response curve of a patient on anti-TB therapy showing an inverse correlation of TB-MBLA measured bacterial load and MGIT TTP. The bacterial load (blue curve) falls while TTP rises (red curve) as the patient responds to treatment. Please click here to view a larger version of this figure.
| Master mix | RT positive reaction | RT negative reaction |
| Volume per reaction x no. of reactions + 5 | Volume per reaction x no. of reactions + 5 | |
| Quantitect mix | 10.0 μL | 10.0 μL |
| Mtb16S primer mix (F + R) | 0.4 μL | 0.4 μL |
| Mtb16S probe | 0.2 μL | 0.2 μL |
| EC primer mix (F + R) | 0.4 μL | 0.4 μL |
| EC probe | 0.2 μL | 0.2 μL |
| RT enzyme | 0.2 μL | ------- |
| RNase free water | 4.6 μL | 4.8 μL |
| Total volume | 16 µL | 16 μL |
Table 1: Master mix preparation guide for the TB-MBLA qPCR.
| Type | MTB channel | EC channel | Interpretation |
| Sample | Positive | Positive | Valid |
| Sample | Positive | Negative | Indeterminate |
| Sample | Negative | Positive | Valid |
| Sample | Negative | Negative | Invalid |
| MTB + Control | Positive | Negative | Valid |
| Extraction Control (EC) | Negative | Positive | Valid |
| DNA control | Negative | Negative | Valid |
| Negative Control (NTC) | Negative | Negative | Valid |
Table 2: TB-MBLA results interpretation guide.
Discussion
This article shows that heat treatment for 20 min at 80 °C, 85 °C, and 95 °C inactivates tuberculosis specimens effectively, making it possible for TB-MBLA to be performed in a non-CL3 facility without risk of infection to laboratory workers. The findings confirm observations made in previous studies while contrasting with some on the effectiveness of heat inactivation of Mtb25,30. For instance, some reports indicate that heating at 80 °C is not effective on high bacillary load samples25,31,32,33. The high-density inoculum effect was avoided in our study by ensuring that all sputum and pure cultures were heated at a 1 mL volume per 15 mL centrifuge tube providing adequate space to expose every part of the sample to boiling27.
RNA preservation following heat inactivation makes it possible for TB-MBLA to be performed. This finding concurs with two studies that demonstrated RNA preservation after heat inactivation12,34. We showed that the RNA in heat inactivated samples is stable at 37 °C for 4 days, implying that laboratories could batch tests by maintaining inactivated samples at room temperature for a week. By applying RNA extraction kits that require refrigeration or freezing, the ability to maintain heat inactivated samples at room temperature obviates the need for both cold chain and Category 3 laboratories to perform TB-MBLA in resource limited settings.
Less than 1log bacterial load was lost using the 16S rRNA as a marker. Although there was a difference between live and heat inactivated sample, the amount lost to heat inactivation is too small to compromise downstream results. Increasing temperature did not increase the amount of RNA lost, implying that the observed loss is independent of the heat treatment. Heat treatment at high temperatures most likely causes cell lysis, exposing RNA to degradation by RNases. Indeed, exogenous addition of RNase A to the heat inactivated fraction increased the rate of RNA degradation. Boiling did not reduce the activity of RNase, implying that it is a very resilient protein.
It is important to note that an average RNA degradation of 1.5 log10 eCFU/mL is not a large loss. To this end we hypothesize that heating lyses a small proportion of Mtb bacilli, thus exposing a small amount of RNA to RNase. Using TEM we showed that Mtb cell morphology and integrity of the cell walls is hardly affected by heating at 95 °C. This means that it may require various physiological factors and sufficient supply of RNases such as in the host to significantly degrade RNA35,36. Furthermore, being a structural ribosome, 16S rRNA is potentially less susceptible to RNase37,38. A single Mtb cell contains ~700 ribosomes/0.1 mm3 of cytoplasm37, implying that there are higher quantities of rRNA per cell. Thus, smaller quantities of RNase may have a smaller impact38. The existence of high numbers of ribosomes gives an advantage to TB-MBLA in terms of sensitivity and ability to detect low burden TB patients.
There was a strong correlation between bacterial load measured by TB-MBLA and MGIT culture TTP. This confirms bacterial load as the driver of culture positivity to some extent. However, the advantage of TB-MBLA is that it directly quantifies bacillary load present in the sample and does not require Mtb cell proliferation before detection. This contrasts with culture whose time to positivity depends on the level of bacterial load and rate of Mtb cell proliferation. Future studies will evaluate the TB-MBLA workflow, including heat inactivation, in routine clinical settings. The study will also explore samples with a range of bacterial loads to understand the number that might change from positive to negative (i.e., those with fewer bacteria following heat inactivation).
The TB-MBLA protocol for molecular quantification of bacterial load is the first of its kind in bacteriology. The method directly quantifies Mtb bacillary load from patient sputum and requires no culture to do so. This makes it faster and increases its potential to inform a clinical decision about patient progress. The heat inactivation step reduces the risk of infection and increases applicability of TB-MBLA in settings that do not have a category 3 laboratory. Following heat inactivation of the sample, there are three protocol steps to achieve TB-MBLA results: RNA extraction, reverse transcriptase (RT)-qPCR, and qPCR results analysis.
The higher the efficiency of isolating Mtb RNA from a patient sample, the higher the quality of the results. It is important to note that the quality of the sputum sample affects the amount of RNA isolated. For instance, salivary sputum is considered low quality and has been associated with low bacillary load. This means that training the patient for quality sputum expectoration is important. To assess the efficiency of the extraction process, an extraction control (i.e, the known number of non-Mtb cells) is spiked into the sample prior to RNA extraction. Retrieval of the extraction control confirms the efficiency of the RNA extraction process. The RNA isolation process cannot be valid unless the extraction control has been retrieved. Given the fact that Mtb is a resilient organism makes mechanical lysis a crucial part of the process. Homogenization of the sample at high speed (600 rpm) in the presence of beads (i.e., lysing matrix) effectively lyses the cells. Purification of the lysate yields an extract containing both RNA and DNA. Removal of DNA is a crucial last step of the RNA extraction. TB-MBLA aims to measure viable bacilli by quantifying RNA. Thus, failure to remove genomic DNA means that the results will have a signal from the DNA, which is not a good marker for cell viability6.
The RT-qPCR is a duplex running dual labelled probes for Mtb and extraction control. It involves three steps: 30 min reverse transcription by reverse transcriptase at 50 °C, 15 min denaturation at 95 °C, and 40 cycles of amplification at 94 °C and 60 °C. Acquisition of fluorescence from the probes occurs at 60 °C (i.e., the fragment elongation stage). It is important to note that the TB-MBLA has been optimized using a particular qPCR machine, so operators using other qPCR platforms should optimize the conditions for their equipment. The efficiency of DNA removal is controlled for by running a single reaction per sample in the absence of RT. A positive result from this reaction signifies incomplete removal of DNA. High burden samples that have high amounts of DNA may require double the amount of DNase enzyme to completely remove DNA. Fortunately, in high bacillary load samples, the presence of small amounts of DNA is less likely to affect the result from the RNA. Ribosomal RNA, the TB-MBLA target, naturally occurs in twice the amount of DNA37. In the PCR, a no template control (NTC), which is the water used to dissolve the PCR reagents, controls for cross contamination with exogenous DNA or RNA. A positive signal in the NTC implies cross contamination and the result considered invalid. This means all solutions constituted using this water have to be discarded and new ones made using a fresh vial of water. It is advisable to keep PCR water in separate aliquots to avoid contamination of all the water. A positive control (Mtb RNA) is used to control for the overall efficiency of PCR.
Result analysis involves the conversion of PCR Cqs into bacterial load (i.e., the estimated colony forming units per mL) using the standard curve. Setting and optimising the standard curve is crucial for this step. A standard curve efficiency of 0.95–1 is recommended. Standard curves for MTb and extraction control should be set up and optimized before patients or other test samples are run on the machine. Standard samples are provided with the TB-MBLA kit. A ten-fold dilution of the RNA extract is recommended for PCR. This implies that the bacterial load result has to be multiplied by a factor of 10 to obtain the final bacterial load result per mL. It is important to note that Cqs above 30 are considered negative for TB-MBLA. A minimum of two prospective bacterial load results measured at different time points is required to make an inference on treatment response. It is strongly recommended that one of the two results should be baseline, before initiation of treatment. However, if the patient initiated bacterial load assessment occurs midway through treatment, there must be a second time point bacterial load measurement to evaluate the treatment response. TB-MBLA can distinguish bacterial load in a space of 3 days on treatment but the ideal is two bacterial load measurements taken 7 days apart.
While the protocol generates informative quantitative results for treatment response, it is still largely manual and demands substantial hands on time for RNA extraction. Technicians in busy labs may not have this time. Arrangements are underway to automate the RNA extraction and PCR processes.
Acknowledgements
The study was made possible by funding from the European and Developing Countries Clinical Trials Partnership (EDCTP) – Pan African Biomarker Expansion program (PanBIOME) grant SP.2011.41304.008. Support was also obtained the University of St Andrews School of Medicine research grant. The publication of this manuscript and TB-MBLA protocol as a visual resource has been made possible by the funding from Scottish Funding Council and Global Challenges Research Fund to the University of St Andrews. Thanks to the Maputo Maternal hospital and Mavalane Health Centre which provided the clinical sputum specimens, and the Maputo Tuberculosis Treatment Unit team who helped with the heat inactivation experiments of clinical sputum specimens.
Materials
| Name | Company | Catalog Number | Comments |
| Heat inactivation: | |||
| 15 mL Centrifuge tubes | Fisher-Scientific | 10136120 | 50 mL tubes may be used if larger sample volumes are involved |
| 15 mL Wire Tube rack | Fisher-Scientific | 11749128 | Other brands with similar make can be used |
| Water bath | Fisher-Scientific | 15700619 | Other brands with similar make can be used |
| RNA extraction: | |||
| Chloroform | Sigma | 372978-1L | Not necessary in other RNA extraction kit brands |
| Ethanol, absolute 99-100% | Sigma | 51976-500ML-F | Larger volume available |
| Extraction control | SOI group St Andrews | VitalbacteriaEC | Supplied with the TB-MBLA Vitalbacteria kit |
| FASTRNA Pro blue Kit | MP Biomedicals | 116025050 | Supplied with lysing matrix tubes |
| Molecular grade water (RNase free) | Sigma | W4502-1L | Qiagen, Fisherscientific can also supply same product |
| Precellys 24 homogeniser | VWR | 432-3750 | FastPrep MP Biomedicals can be used too |
| Refrigerated micro-centrifuge, Fresco 21, Heraeus | Fisher-Scientific | 15352117 | Other brands with similar capacity can be used |
| Thermomixer | Eppendorf | BLD-455-010C | Works with 1-2 mL tubes |
| TURBO DNA-free | Fisher-Scientific | AM1907M | Takes longer but more effective than shorter DNA removal procedures. |
| RT-qPCR: | |||
| Primers and Taqman probes | SOI group St Andrews | VitalbacteriaPP | Supplied with the TB-MBLA Vitalbacteria kit. Probe fluorophores are FAM (green) for Mtb and HEX (yellow) for Extraction control (EC). Applied Biosystems qPCR platforms use VIC instead of HEX. |
| QuantiTect Multiplex RT-PCR NR Kit | Qiagen | 204845 | PCR mix plus the RT enzyme |
| RotorGene Q 5plex machine | Qiagen | 9001580 | Other qPCR machines: Vii7, Quantistudio, Steponelus etc can be used |
| Strip Tubes and Caps, 0.1 ml | Qiagen | 981103 | Other tube sizes available depending on the rotor size. For other PCR platforms 96 well PCR plates are needed. |
| Non-heat inactivation Sample preservation materials | |||
| 1M Tris-HCl pH 7.5 | Sigma | 93313 | Supplied ready to use |
| 2-Mercaptoethanol | Sigma | 63689 | Many competent suppliers |
| Guanidine thiocyanate (GTC) | Promega | V2791 | Promega brand recommended for quality |
| Molecular grade water (RNase free) | Sigma | W4502-1L | Ensures that GTC solution is free of nucleases |
| General materials (used but not specific to TB-MBLA) | |||
| 500 mL plastic containers | 734-5087 | Nalgene brand recommeded | |
| Biological waste discard jars | Many competent suppliers | ||
| Chemical waste discard jars | Many competent suppliers | ||
| Disposable gloves, chemical resistant | Many competent suppliers | ||
| Freezer (-20 °C) | Many competent suppliers | ||
| Freezer (-80 °C) | Many competent suppliers | ||
| Fridge (0-8 °C) | Many competent suppliers | ||
| Fume hood | For safe handling of toxic reagents | ||
| Laboratory scales | For accurate measurement of reagents | ||
| Measuring cylinders, plastic | To ensure accurate measurement of reagents | ||
| PCR reaction tubes | Should be suitable to the PCR instrument used | ||
| Pipettes and matching sterile filtered pipette tips, | DNAse and RNAse-free, range: P1000, P200, P10, P2 recommended | ||
| Qiagility loading plates | Qiagen | 5-well master mix plate, 16-well reagent plate, 32-well sample plate, 72-well and 96-well reaction plates | |
| QIAgility Pipetting Robot | Qiagen | Important for high throughput pipetting | |
| Racks for 1.5 mL and 2 mL microtubes and for 15 mL and 50 mL Falcon tubes | Chemical-resistant and autoclavable recommended | ||
| RNase Away (Removes RNases enzymes from working space) | Fisher Scientific | 10666421 | Important to ensure services and devices are free from RNase enzyme |
| Safety goggles, chemical resistant | Fisher Scientific | Can also be got from Sigma. Protect eyes from toxic reagents. | |
| Sterile Pasteur pipettes, 1.5 mL - 3 mL | Many competent suppliers | ||
| Sterile RNAse-free microtubes | 1.5 mL tubes suitable for freezing at -80 °C recommended | ||
| TB disinfectant, e.g. Tristel Fuse | Ensure it is freshly prepared or prepared within a week | ||
| Vortex | Genie 2 brand recommended | ||
| Transmission electron microscopy | |||
| Glutaldehyde (8%) | Sigma | G7526-10ML | Part of the Cell fixation buffer (HEPES buffer) |
| HEPES (pH 7.4, 100 mM) | Sigma | H4034-25G | Part of the Cell fixation buffer (HEPES buffer) |
| Leica FCS ultracryo | Leica | Cryomicrotome for tissue sectioning | |
| Methyl cellulose | Agar Scientific | AGG6425 | Cold mounting resin |
| Paraformaldehyde (4% in PBS) | Sigma | P6148-500G | Part of the Cell fixation buffer (HEPES buffer) |
| Sucrose | Sigma | 84097-250G | Preservation and mounting medium |
| Uranyl acetate | Agar Scientific | AGR1260A | Universal Electron microscopy stain |
References
- World Health Organization. END TB Global Tuberculosis Report 2017. World Health Organization. , (2017).
- World Health Organization. Global tuberculosis report 2018. World Health Organization. , (2018).
- World Health Organization. TB Elimination in Low-Incidence Countries. World Health Organization. , (2014).
- Boehme, C. C., et al. diagnostic accuracy, and effectiveness of decentralised use of the Xpert MTB/RIF test for diagnosis of tuberculosis and multidrug resistance: a multicentre implementation study. The Lancet. 377, 1495-1505 (2011).
- World Health Organization. Xpert MTB/RIF assay for the diagnosis of pulmonary and extrapulmonary TB in adults and children. World Health Organization. , (2010).
- Friedrich, S. O., et al. Assessment of the sensitivity and specificity of Xpert MTB/RIF assay as an early sputum biomarker of response to tuberculosis treatment. The Lancet Respiratory Medicine. 1, 462-470 (2013).
- Desjardin, L. E., et al. Measurement of Sputum Mycobacterium tuberculosis Messenger RNA as a Surrogate for Response to Chemotherapy. American Journal of Respiratory Critical Care Medicine. 160, 203-210 (1999).
- Hellyer, T. J., et al. Detection of Viable Mycobacterium tuberculosis by Reverse Transcriptase-Strand Displacement Amplification of mRNA. Journal of Clinical Microbiology. 37, 518-523 (1999).
- Honeyborne, I., et al. Molecular Bacterial Load Assay , a Culture-Free Biomarker for Rapid and Accurate Quantification of Sputum Mycobacterium tuberculosis Bacillary Load during Treatment. Journal of Clinical Microbiology. 49, 3905-3911 (2011).
- Li, L., et al. Sputum Mycobacterium tuberculosis mRNA as a Marker of Bacteriologic Clearance in Response to Antituberculosis Therapy. Journal of Clinical Microbiology. 48, 46-51 (2010).
- Honeyborne, I., et al. The Molecular Bacterial Load Assay Replaces Solid Culture for Measuring Early Bactericidal Response to Antituberculosis Treatment. Journal of Clinical Microbiology. 52, 3064-3067 (2014).
- Hellyer, T. J., Jardin, L. E. D. E. S., Hehman, G. L., Cave, M. D. Quantitative Analysis of mRNA as a Marker for Viability of Mycobacterium tuberculosis. Journal of Clinical Microbiology. 37, 290-295 (1999).
- Aellen, S., Que, Y., Guignard, B., Haenni, M., Moreillon, P. Detection of Live and Antibiotic-Killed Bacteria by Quantitative Real-Time PCR of Specific Fragments of rRNA. Antimicrobial agents chemotherapy. 50, 1913-1920 (2006).
- Deutscher, M. P. Degradation of RNA in bacteria: Comparison of mRNA and stable RNA. Nucleic Acids Research. 34, 659-666 (2006).
- Gupta, R. K., Srivastava, R. Resuscitation Promoting Factors: A Family of Microbial Proteins in Survival and Resuscitation of Dormant Mycobacteria. Indian Journal of Microbiology. 52, 114-121 (2012).
- Mukamolova, G. V., Turapov, O., Malkin, J., Woltmann, G., Barer, M. R. Resuscitation-promoting factors reveal an occult population of tubercle bacilli in sputum. American Journal of Respiratory Critical Care Medicine. 181, 174-180 (2010).
- Sheridan, G. E. C., Masters, C. I., Shallcross, J. A., Mackey, B. M. Detection of mRNA by Reverse Transcription-PCR as an Indicator of Viability in Escherichia coliCells Detection of mRNA by Reverse Transcription-PCR as an Indicator of Viability in Escherichia coli Cells. Applied Environment Microbiology. 64, 1313-1318 (1998).
- Honeyborne, I., et al. Molecular bacterial load assay, a culture-free biomarker for rapid and accurate quantification of sputum Mycobacterium tuberculosis bacillary load during treatment. Journal of Clinical Microbiology. 49, 3905-3911 (2011).
- Sabiiti, W., et al. Molecular Bacterial Load Assay: a Fast and Accurate Means for Monitoring Tuberculosis Treatment Response. BMJ Global Health. 2, 1-8 (2017).
- Gillespie, H., Stephen, S. W., Oravcova, K., Bishop, A. K. Mybacterial Load Assay. Diagnostic Bacteriology. Methods and Protocols. , 89-105 (2017).
- Juffs, H., Deeth, H. Scientific Evaluation of Pasteurisation for Pathogen Reduction in Milk and Milk Products. Food Standards Australia New Zealand. , (2007).
- Holmes, C. J., Degremont, A., Kubey, W. Effectiveness of Various Chemical Disinfectants versus Cleaning Combined with Heat Disinfection on Pseudomonas Biofi lm. Blood Purification. 22, 461-468 (2004).
- Chedore, P., et al. Method for inactivating and fixing unstained smear preparations of Mycobacterium tuberculosis for improved laboratory safety. Journal of Clinical Microbiology. 40, 4077-4080 (2002).
- Cardoso, C. L., et al. Survival of Tubercle Bacilli in Heat-fixed and Stained Sputum Smears. Memórias do Instituto Oswaldo Cruz. 96, 277-280 (2001).
- Doig, C., Seagar, A. L., Watt, B., Forbes, K. J. The efficacy of the heat killing of Mycobacterium tuberculosis. Journal of Clinical Pathology. 55, 778-779 (2002).
- Honeyborne, I., et al. The molecular bacterial load assay replaces solid culture for measuring early bactericidal response to antituberculosis treatment. Journal of Clinical Microbiology. 52, 3064-3067 (2014).
- Sabiiti, W., et al. Heat Inactivation Renders Sputum Safe and Preserves Mycobacterium tuberculosis RNA for Downstream Molecular tests. Journal of Clinical Microbiology. , 1-8 (2019).
- Griffiths, G., McDowall, A., Back, R., Dubochet, J. On the preparation of cryosections for immunocytochemistry. Journal of Ultrasructure Research. 89, 65-78 (1984).
- Kumar, V. G., Urs, T. A., Ranganath, R. R. MPT 64 Antigen detection for Rapid confirmation of M. tuberculosis isolates. BMC Research Notes. 4, 79 (2011).
- Zwadyk, P., Down, J. A., Myers, N., Dey, M. S. Rendering of mycobacteria safe for molecular diagnostic studies and development of a lysis method for strand displacement amplification and PCR. Journal of Clinical Microbiology. 32, 2140-2146 (1994).
- Blackwood, K. S., et al. Viability testing of material derived from Mycobacterium tuberculosis prior to removal from a Containment Level-III Laboratory as part of a Laboratory Risk Assessment Program. BMC Infectious Diseases. 5, 3-9 (2005).
- Somerville, W., Thibert, L., Schwartzman, K., Behr, M. A. Extraction of Mycobacterium tuberculosis DNA: a Question of Contaiment. Journal of Clinical Microbiology. 43, 2996-2997 (2005).
- Bemer-Melchior, P., Drugeon, H. B. Inactivation of Mycobacterium tuberculosis for DNA typing analysis. Journal of Clinical Microbiology. 37, 2350-2351 (1999).
- McKillip, J. L., Jaykus, L. A., Drake, M. rRNA stability in heat killed and UV-irradiated enterotoxigenic Staphylococcus aureus and Escherichia coli O157:H7. Journal of Appied Environmental Microbiology. 64, 4264-4268 (1998).
- Blank, A., Dekker, C. A. Ribonucleases of Human Serum, Urine, Cerebrospinal Fluid, and Leukocytes. Activity Staining following Electrophoresis in Sodium Dodecyl Sulfate-Polyacrylamide Gels. Biochemistry. 20, 2261-2267 (1981).
- O'Leary, T. J. Reducing the impact of endogenous ribonucleases on reverse transcription-PCR assay systems. Clinical Chemistry. 45, 449-450 (1999).
- Yamada, H., Yamaguchi, M., Chikamatsu, K., Aono, A., Mitarai, S. Structome analysis of virulent Mycobacterium tuberculosis, which survives with only 700 ribosomes per 0.1 fl of cytoplasm. PLoS One. 10, 1-14 (2015).
- Yang, K., et al. Structural insights into species-specific features of the ribosome from the human pathogen Mycobacterium tuberculosis. Nucleic Acids Research. 45, 10884-10894 (2017).
Explore More Articles
This article has been published
Video Coming Soon
ABOUT JoVE
Copyright © 2024 MyJoVE Corporation. All rights reserved