Deep Learning-Based Segmentation of Cryo-Electron Tomograms
In This Article
Summary
This is a method for training a multi-slice U-Net for multi-class segmentation of cryo-electron tomograms using a portion of one tomogram as a training input. We describe how to infer this network to other tomograms and how to extract segmentations for further analyses, such as subtomogram averaging and filament tracing.
Abstract
Cryo-electron tomography (cryo-ET) allows researchers to image cells in their native, hydrated state at the highest resolution currently possible. The technique has several limitations, however, that make analyzing the data it generates time-intensive and difficult. Hand segmenting a single tomogram can take from hours to days, but a microscope can easily generate 50 or more tomograms a day. Current deep learning segmentation programs for cryo-ET do exist, but are limited to segmenting one structure at a time. Here, multi-slice U-Net convolutional neural networks are trained and applied to automatically segment multiple structures simultaneously within cryo-tomograms. With proper preprocessing, these networks can be robustly inferred to many tomograms without the need for training individual networks for each tomogram. This workflow dramatically improves the speed with which cryo-electron tomograms can be analyzed by cutting segmentation time down to under 30 min in most cases. Further, segmentations can be used to improve the accuracy of filament tracing within a cellular context and to rapidly extract coordinates for subtomogram averaging.
Introduction
Hardware and software developments in the past decade have resulted in a "resolution revolution" for cryo-electron microscopy (cryo-EM)1,2. With better and faster detectors3, software to automate data collection4,5, and signal boosting advances such as phase plates6, collecting large amounts of high-resolution cryo-EM data is relatively straightforward.
Cryo-ET delivers unprecedented insight into cellular ultrastructure in a native, hydrated state7,8,9,10. The primary limitation is sample thickness, but with the adoption of methods such as focused ion beam (FIB) milling, where thick cellular and tissue samples are thinned for tomography11, the horizon for what can be imaged with cryo-ET is constantly expanding. The newest microscopes are capable of producing well over 50 tomograms a day, and this rate is only projected to increase due to the development of rapid data collection schemes12,13. Analyzing the vast amounts of data produced by cryo-ET remains a bottleneck for this imaging modality.
Quantitative analysis of tomographic information requires that it first be annotated. Traditionally, this requires hand segmentation by an expert, which is time-consuming; depending on the molecular complexity contained within the cryo-tomogram, it can take hours to days of dedicated attention. Artificial neural networks are an appealing solution to this problem since they can be trained to do the bulk of the segmentation work in a fraction of the time. Convolutional neural networks (CNNs) are especially suited to computer vision tasks14 and have recently been adapted for the analysis of cryo-electron tomograms15,16,17.
Traditional CNNs require many thousands of annotated training samples, which is not often possible for biological image analysis tasks. Hence, the U-Net architecture has excelled in this space18 because it relies on data augmentation to successfully train the network, minimizing the dependency on large training sets. For instance, a U-Net architecture can be trained with only a few slices of a single tomogram (four or five slices) and robustly inferred to other tomograms without retraining. This protocol provides a step-by-step guide for training U-Net neural network architectures to segment electron cryo-tomograms within Dragonfly 2022.119.
Dragonfly is commercially developed software used for 3D image segmentation and analysis by deep learning models, and it is freely available for academic use (some geographical restrictions apply). It has an advanced graphical interface that allows a non-expert to take full advantage of the powers of deep learning for both semantic segmentation and image denoising. This protocol demonstrates how to preprocess and annotate cryo-electron tomograms within Dragonfly for training artificial neural networks, which can then be inferred to rapidly segment large datasets. It further discusses and briefly demonstrates how to use segmented data for further analysis such as filament tracing and coordinate extraction for sub-tomogram averaging.
Protocol
NOTE: Dragonfly 2022.1 requires a high-performance workstation. System recommendations are included in the Table of Materials along with the hardware of the workstation used for this protocol. All tomograms used in this protocol are binned 4x from a pixel size of 3.3 to 13.2 ang/pix. Samples used in the representative results were obtained from a company (see the Table of Materials) that follows animal care guidelines that align to this institution's ethical standards. The tomogram used in this protocol and the multi-ROI that was generated as training input have been included as a bundled dataset in Supplemental File 1 (which can be found at https://datadryad.org/stash/dataset/doi:10.5061/dryad.rxwdbrvct) so the user can follow along with the same data should they wish to. Dragonfly also hosts an open access database called the Infinite Toolbox where users can share trained networks.
1. Setup
- Changing default workspace:
- To change the workspace to mirror the one used in this protocol, on the left side in the Main panel, scroll down to the Scene's View Properties section and deselect Show Legends. Scroll down to the Layout section and select the Single scene and the Four Equal Views view.
- To update the default unit, go to File | Preferences. In the window that opens, change the Default unit from millimeters to nanometers.
- Helpful default keybinds:
- Press Esc to display the crosshairs in the 2D views and allow 3D volume rotation in 3D view. Press X to hide the crosshairs in the 2D views and allow 2D translation and 3D volume translation in 3D view.
- Hover over the crosshairs to see small arrows that can be clicked and dragged to change the angle of the viewing plane in the other 2D views.
- Press Z to enter the zoom state in both views, allowing users to click and drag anywhere to zoom in and out.
- Double-click on a view in the Four View scene to focus only that view; double-click again to return to all four views.
- Periodically save progress by exporting everything in the Properties tab as an ORS object for easy import. Select all the objects in the list and right click Export | As ORS object. Name the file and save. Alternatively, go to File | Save Session. To use the autosave feature in the software, enable it through File | Preferences | Autosave.
2. Image import
- For image import, go to File | Import Image Files. Click Add, navigate to the image file, and click Open | Next | Finish.
NOTE: The software does not recognize .rec files. All tomograms must have the .mrc suffix. If using the provided data, instead go to File | Import Object(s). Navigate to the Training.ORSObject file and click Open, then click OK.
3. Preprocessing (Figure 1.1)
- Create a custom intensity scale (used to calibrate image intensities across datasets). Go to Utilities | Dimension Unit Manager. In the lower left, click + to create a new Dimension Unit.
- Choose a high intensity (bright) and low intensity (dark) feature that is in all the tomograms of interest. Give the unit a name and an abbreviation (e.g., for this scale, set fiducial beads to 0.0 Standard intensity and the background to 100.0). Save the custom dimension unit.
NOTE: A custom intensity scale is an arbitrary scale that is created and applied to the data to ensure that all data is on the same intensity scale despite being collected at different times or on different equipment. Choose light and dark features that best represent the range that the signal falls within. If there are no fiducials in the data, simply choose the darkest feature that is going to be segmented (the darkest region of protein, for instance). - To calibrate images to the custom intensity scale, right-click the dataset in the Properties column on the right-hand side of the screen and select Calibrate Intensity Scale. On the Main tab on the left side of the screen, scroll down to the Probe section. Using the circular probe tool with an appropriate diameter, click a few places in the background region of the tomogram and record the average number in the Raw Intensity column; repeat for fiducial markers, then click Calibrate. If necessary, adjust the contrast to make structures visible again with the Area tool in the Window Leveling section of the Main tab.
- Image filtering:
NOTE: Image filtering can reduce noise and boost the signal. This protocol uses three filters that are built into the software as they work the best for this data, but there are many filters available. Once settled on an image filtering protocol for the data of interest, it will be necessary to apply exactly the same protocol to all tomograms prior to segmentation.- In the main tab on the left side, scroll down to the Image Processing Panel. Click Advanced and wait for a new window to open. From the Properties panel, select the dataset to be filtered and make it visible by clicking the eye icon to the left of the dataset.
- From the Operations panel, use the dropdown menu to select Histogram Equalization (under the Contrast section) for the first operation. Select Add Operation | Gaussian (under the Smoothing section). Change the kernel dimension to 3D.
- Add a third operation; then, select Unsharp (under the Sharpening section). Leave the output for this one. Apply to all slices and let the filtering run, then close the Image Processing window to return to the main interface.
4. Create training data (Figure 1.2)
- Identify the training area by first hiding the unfiltered dataset by clicking the eye icon to the left of it in the Data Properties panel. Then, show the newly filtered dataset (which will be automatically named DataSet-HistEq-Gauss-Unsharp). Using the filtered dataset, identify a subregion of the tomogram that contains all the features of interest.
- To create a box around the region of interest, on the left side, in the main tab, scroll down to the Shapes category and select Create a Box. While in the Four View panel, use the different 2D planes to help guide/drag the edges of the box to enclose only the region of interest in all dimensions. In the data list, select the Box region and change the color of the border for easier viewing by clicking the gray square next to the eye symbol.
NOTE: The smallest patch size for a 2D U-Net is 32 x 32 pixels; 400 x 400 x 50 pixels is a reasonable box size to start. - To create a multi-ROI, in the left side, select the Segmentation tab | New and check Create as Multi-ROI. Ensure that the number of classes corresponds to the number of features of interest + a background class. Name the multi-ROI Training Data and ensure that the geometry corresponds to the dataset before clicking OK.
- Segmenting the training data
- Scroll through the data until within the bounds of the boxed region. Select the Multi-ROI in the properties menu on the right. Double-click the first blank class name in the multi-ROI to name it.
- Paint with the 2D brush. In the segmentation tab on the left, scroll down to 2D tools and select a circular brush. Then, select Adaptive Gaussian or Local OTSU from the dropdown menu. To paint, hold left ctrl and click. To erase, hold left shift and click.
NOTE: The brush will reflect the color of the currently selected class. - Repeat the previous step for each object class in the multi-ROI. Ensure that all structures within the boxed region are fully segmented or they will be considered background by the network.
- When all structures have been labeled, right-click the Background class in the Multi-ROI and select Add All Unlabeled Voxels to Class.
- Create a new single-class ROI named Mask. Ensure that the geometry is set to the filtered dataset, then click apply. In the properties tab on the right, right-click the Box and select Add to ROI. Add it to the Mask ROI.
- To trim the training data using the Mask, in the Properties tab, select both the Training Data multi-ROI and the Mask ROI by holding Ctrl and clicking on each. Next, click Intersect beneath the data properties list in the section labeled Boolean operations. Name the new dataset Trimmed Training Input, and ensure the geometry corresponds to the filtered dataset before clicking OK.
5. Using the segmentation wizard for iterative training (Figure 1.3)
- Import the training data to the segmentation wizard by first right-clicking the filtered dataset in the Properties tab, and then selecting the Segmentation Wizard option. When a new window opens, look for the input tab on the right side. Click Import Frames from a Multi-ROI and select the Trimmed Training Input.
- (Optional) Create a Visual Feedback Frame to monitor training progress in real time.
- Select a frame from the data that is not segmented and click + to add it as a new frame. Double-click the mixed label to the right of the frame and change it to Monitoring.
- To generate a New Neural Network Model, on the right side in the Models tab, click the + button to generate a new model. Select U-Net from the list, and then for input dimension, select 2.5D and 5 slices, then click Generate.
- To train the Network, click Train on the bottom right of the SegWiz window.
NOTE: Training can be stopped early without losing progress. - To use the trained network to segment new frames, when the U-Net training is complete, create a new frame and click Predict (bottom right). Then, click the Up arrow in the upper right of the predicted frame to transfer the segmentation to the real frame.
- To correct the prediction, Ctrl-Click two classes to change the segmented pixels of one to the other. Select both classes and paint with the brush to paint only pixels belonging to either class. Correct the segmentation in at least five new frames.
NOTE: Painting with the brush tool while both classes are selected means that instead of shift-click erasing, as it normally does, it will convert pixels of the first class to the second. Ctrl-click will accomplish the reverse. - For iterative training, click the Train button again and allow the network to train further for another 30-40 more epochs, at which point stop the training and repeat steps 4.5 and 4.6 for another round of training.
NOTE: In this way, a model can be iteratively trained and improved using a single dataset. - To publish the network, when satisfied with its performance, exit the Segmentation Wizard. In the dialog box that automatically pops up asking which models to publish (save), select the successful network, name it, then publish it to make the network available for use outside of the segmentation wizard.
6. Apply the network (Figure 1.4)
- To apply to the training tomogram first, select the filtered dataset in the Properties panel. In the Segmentation panel on the left, scroll down to the Segment with AI section. Make sure that the correct dataset is selected, choose the model that was just published in the dropdown menu, then click Segment | All Slices. Alternatively, select Preview to view a one-slice preview of the segmentation.
- To apply to an inference dataset, import the new tomogram. Preprocess according to Step 3 (Figure 1.1). In the Segmentation panel, go to the Segment with AI section. Making sure that the newly filtered tomogram is the dataset selected, choose the previously trained model, and click Segment | All Slices.
7. Segmentation manipulation and cleanup
- Quickly clean up noise by first choosing one of the classes that has segmented noise and the feature of interest. Right-click | Process Islands | Remove by Voxel Count | Select a voxel size. Start small (~200) and gradually increase the count to remove most of the noise.
- For segmentation correction, Ctrl-click two classes to paint only pixels belonging to those classes. Ctrl-click + drag with the segmentation tools to change pixels of the second class to the first and Shift-click + drag to accomplish the opposite. Continue doing this to quickly correct incorrectly labeled pixels.
- Separate connected components.
- Choose a class. Right-click a class in Multi-ROI | Separate Connected Components to create a new class for each component that is not connected to another component of the same class. Use the buttons below the Multi-ROI to easily merge the classes.
- Export the ROI as Binary/TIFF.
- Choose a class in the Multi-ROI, then right-click and Extract Class as a ROI. In the properties panel above, select the new ROI, right-click | Export | ROI as Binary (ensure that the option to export all images into one file is selected).
NOTE: Users can easily convert from tiff to mrc format using the IMOD program tif2mrc20. This is useful for filament tracing.
- Choose a class in the Multi-ROI, then right-click and Extract Class as a ROI. In the properties panel above, select the new ROI, right-click | Export | ROI as Binary (ensure that the option to export all images into one file is selected).
8. Generating coordinates for sub-tomogram averaging from the ROI
- Extract a class.
- Right-click Class to be used for averaging | Extract Class as ROI. Right-click class ROI | Connected Components | New Multi-ROI (26 connected).
- Generate coordinates.
- Right-click the new Multi-ROI | Scalar Generator. Expand Basic Measurements with Dataset | check Weighted Center of Mass X, Y and Z. Select the dataset and compute. Right-click Multi-ROI | Export Scalar Values. Check Select all Scalar slots, then OK to generate centroid world coordinates for each class in the multi-ROI as a CSV file.
NOTE: If particles are close together and the segmentations are touching, it may be necessary to perform a watershed transform to separate the components into a multi-ROI.
- Right-click the new Multi-ROI | Scalar Generator. Expand Basic Measurements with Dataset | check Weighted Center of Mass X, Y and Z. Select the dataset and compute. Right-click Multi-ROI | Export Scalar Values. Check Select all Scalar slots, then OK to generate centroid world coordinates for each class in the multi-ROI as a CSV file.
9. Watershed transform
- Extract the class by right-clicking class in Multi-ROI to be used for averaging | Extract Class as ROI. Name this ROI Watershed Mask.
- (Optional) Close holes.
- If the segmented particles have holes or openings, close these for the watershed. Click the ROI in Data Properties. In the Segmentation tab (on the left), go to Morphological Operations and use whatever combination of Dilate, Erode, and Close necessary to achieve solid segmentations with no holes.
- Invert the ROI by clicking the ROI | Copy Selected Object (below Data Properties). Select the copied ROI, and on the left side in the Segmentation tab click Invert.
- Create a distance map by right-clicking the inverted ROI | Create Mapping Of | Distance Map. For later use, make a copy of the distance map and invert it (right-click | Modify And Transform | Invert Values | Apply). Name this inverted map Landscape.
- Create seed points.
- Hide the ROI and display the Distance Map. In the Segmentation tab, click Define Range and decrease the range until only a few pixels in the center of each point are highlighted and none are connected to another point. At the bottom of the Range section, click Add to New. Name this new ROI Seedpoints.
- Perform watershed transform.
- Right-click Seedpoints ROI | Connected Components | New Multi-ROI (26 connected). Right-click the newly generated Multi-ROI | Watershed Transform. Select the distance map named Landscape and click OK; select the ROI named Watershed Mask and click OK to calculate a watershed transform from each seed point and separate individual particles into separate classes in the multi-ROI. Generate coordinates as in step 8.2.
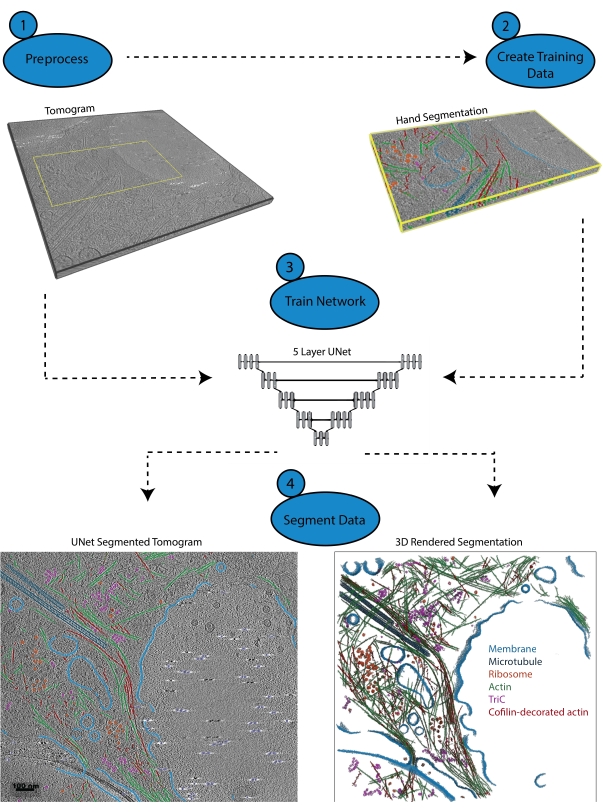
Figure 1: Workflow. 1) Preprocess the training tomogram by calibrating the intensity scale and filtering the dataset. 2) Create the training data by hand-segmenting a small portion of a tomogram with all appropriate labels the user wishes to identify. 3) Using the filtered tomogram as the input and the hand segmentation as the training output, a five-layer, multi-slice U-Net is trained in the segmentation wizard. 4) The trained network can be applied to the full tomogram to annotate it and a 3D rendering can be generated from each segmented class. Please click here to view a larger version of this figure.
Representative Results
Following the protocol, a five-slice U-Net was trained on a single tomogram (Figure 2A) to identify five classes: Membrane, Microtubules, Actin, Fiducial markers, and Background. The network was iteratively trained a total of three times, and then applied to the tomogram to fully segment and annotate it (Figure 2B,C). Minimal cleanup was performed using steps 7.1 and 7.2. The next three tomograms of interest (Figure 2D,G,J) were loaded into the software for preprocessing. Prior to image import, one of the tomograms (Figure 2J) required pixel size adjustment from 17.22 Å/px to 13.3 Å/px as it was collected on a different microscope at a slightly different magnification. The IMOD program squeezevol was used for resizing with the following command:
'squeezevol -f 0.772 inputfile.mrc outputfile.mrc'
In this command, -f refers to the factor by which to alter the pixel size (in this case: 13.3/17.22). After import, all three inference targets were preprocessed according to steps 3.2 and 3.3, and then the five-slice U-Net was applied. Minimal cleanup was again performed. The final segmentations are displayed in Figure 2.
Microtubule segmentations from each tomogram were exported as binary (step 7.4) TIF files, converted to MRC (IMOD tif2mrc program), and then used for cylinder correlation and filament tracing. Binary segmentations of filaments result in much more robust filament tracing than tracing over tomograms. Coordinate maps from filament tracing (Figure 3) will be used for further analysis, such as nearest neighbor measurements (filament packing) and helical sub-tomogram averaging along single filaments to determine microtubule orientation.
Unsuccessful or inadequately trained networks are easy to determine. A failed network will be unable to segment any structures at all, whereas an inadequately trained network typically will segment some structures correctly and have a significant number of false positives and false negatives. These networks can be corrected and iteratively trained to improve their performance. The segmentation wizard automatically calculates a model's Dice similarity coefficient (called score in the SegWiz) after it is trained. This statistic gives an estimate of the similarity between the training data and the U-Net segmentation. Dragonfly 2022.1 also has a built-in tool to evaluate a model's performance that can be accessed in the Artificial Intelligence tab at the top of the interface (see documentation for usage).
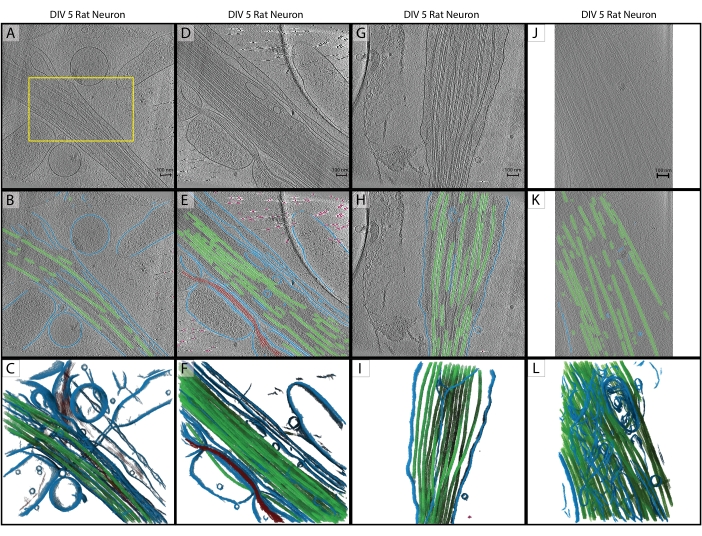
Figure 2: Inference. (A-C) Original training tomogram of a DIV 5 hippocampal rat neuron, collected in 2019 on a Titan Krios. This is a backprojected reconstruction with CTF correction in IMOD. (A) The yellow box represents the region where hand segmentation was performed for training input. (B) 2D segmentation from the U-Net after training is complete. (C) 3D rendering of the segmented regions showing membrane (blue), microtubules (green), and actin (red). (D-F) DIV 5 hippocampal rat neuron from the same session as the training tomogram. (E) 2D segmentation from the U-Net with no additional training and quick cleanup. Membrane (blue), microtubules (green), actin (red), fiducials (pink). (F) 3D rendering of the segmented regions. (G-I) DIV 5 hippocampal rat neuron from the 2019 session. (H) 2D segmentation from the U-Net with quick cleanup and (I) 3D rendering. (J-L) DIV 5 hippocampal rat neuron, collected in 2021 on a different Titan Krios at a different magnification. Pixel size has been changed with the IMOD program squeezevol to match the training tomogram. (K) 2D segmentation from the U-Net with quick cleanup, demonstrating robust inference across datasets with proper preprocessing and (L) 3D rendering of segmentation. Scale bars = 100 nm. Abbreviations: DIV = days in vitro; CTF = contrast transfer function. Please click here to view a larger version of this figure.
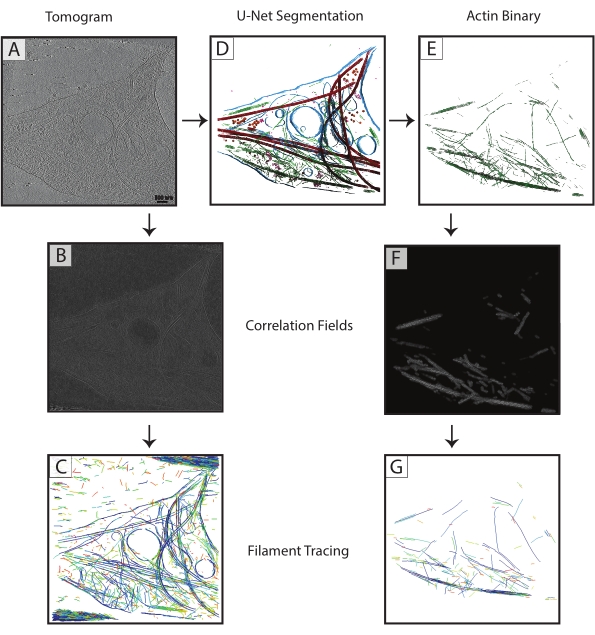
Figure 3: Filament tracing improvement. (A) Tomogram of a DIV 4 rat hippocampal neuron, collected on a Titan Krios. (B) Correlation map generated from cylinder correlation over actin filaments. (C) Filament tracing of actin using the intensities of the actin filaments in the correlation map to define parameters. Tracing captures the membrane and microtubules, as well as noise, while trying to trace just actin. (D) U-Net segmentation of tomogram. Membrane highlighted in blue, microtubules in red, ribosomes in orange, triC in purple, and actin in green. (E) Actin segmentation extracted as a binary mask for filament tracing. (F) Correlation map generated from cylinder correlation with the same parameters from (B). (G) Significantly improved filament tracing of just actin filaments from the tomogram. Abbreviation: DIV = days in vitro. Please click here to view a larger version of this figure.
Supplemental File 1: The tomogram used in this protocol and the multi-ROI that was generated as training input are included as a bundled dataset (Training.ORSObject). See https://datadryad.org/stash/dataset/doi:10.5061/dryad.rxwdbrvct.
Discussion
This protocol lays out a procedure for using Dragonfly 2022.1 software to train a multi-class U-Net from a single tomogram, and how to infer that network to other tomograms that do not need to be from the same dataset. Training is relatively quick (can be as fast as 3-5 min per epoch or as slow as a few hours, depending entirely on the network that is being trained and the hardware used), and retraining a network to improve its learning is intuitive. As long as the preprocessing steps are carried out for every tomogram, inference is typically robust.
Consistent preprocessing is the most critical step for deep learning inference. There are many imaging filters in the software and the user can experiment to determine which filters work best for particular datasets; note that whatever filtering is used on the training tomogram must be applied in the same way to the inference tomograms. Care must also be taken to provide the network with accurate and sufficient training information. It is vital that all features segmented within the training slices are segmented as carefully and precisely as possible.
Image segmentation is facilitated by a sophisticated commercial-grade user interface. It provides all the necessary tools for hand segmentation and allows for the simple reassignment of voxels from any one class into another prior to training and retraining. The user is allowed to hand-segment voxels within the whole context of the tomogram, and they are given multiple views and the ability to rotate the volume freely. Additionally, the software provides the ability to use multi-class networks, which tend to perform better16 and are faster than segmenting with multiple single-class networks.
There are, of course, limitations to a neural network's capabilities. Cryo-ET data are, by nature, very noisy and limited in angular sampling, which leads to orientation-specific distortions in identical objects21. Training relies on an expert to hand-segment structures accurately, and a successful network is only as good (or as bad) as the training data it is given. Image filtering to boost signal is helpful for the trainer, but there are still many cases where accurately identifying all pixels of a given structure is difficult. It is, therefore, important that great care is taken when creating the training segmentation so that the network has the best information possible to learn during training.
This workflow can be easily modified for each user's preference. While it is essential that all tomograms be preprocessed in exactly the same manner, it is not necessary to use the exact filters used in the protocol. The software has numerous image filtering options, and it is recommended to optimize these for the user's particular data before setting out on a large segmentation project spanning many tomograms. There are also quite a few network architectures available to use: a multi-slice U-Net has been found to work best for the data from this lab, but another user might find that another architecture (such as a 3D U-Net or a Sensor 3D) works better. The segmentation wizard provides a convenient interface for comparing the performance of multiple networks using the same training data.
Tools like the ones presented here will make hand segmentation of full tomograms a task of the past. With well-trained neural networks that are robustly inferable, it is entirely feasible to create a workflow where tomographic data is reconstructed, processed, and fully segmented as quickly as the microscope can collect it.
Acknowledgements
This study was supported by the Penn State College of Medicine and the Department of Biochemistry and Molecular Biology, as well as Tobacco Settlement Fund (TSF) grant 4100079742-EXT. The CryoEM and CryoET Core (RRID:SCR_021178) services and instruments used in this project were funded, in part, by the Pennsylvania State University College of Medicine via the Office of the Vice Dean of Research and Graduate Students and the Pennsylvania Department of Health using Tobacco Settlement Funds (CURE). The content is solely the responsibility of the authors and does not necessarily represent the official views of the University or College of Medicine. The Pennsylvania Department of Health specifically disclaims responsibility for any analyses, interpretations, or conclusions.
Materials
| Name | Company | Catalog Number | Comments |
| Dragonfly 2022.1 | Object Research Systems | https://www.theobjects.com/dragonfly/index.html | |
| E18 Rat Dissociated Hippocampus | Transnetyx Tissue | KTSDEDHP | https://tissue.transnetyx.com/faqs |
| IMOD | University of Colorado | https://bio3d.colorado.edu/imod/ | |
| Intel® Xeon® Gold 6124 CPU 3.2GHz | Intel | https://www.intel.com/content/www/us/en/products/sku/120493/intel-xeon-gold-6134-processor-24-75m-cache-3-20-ghz/specifications.html | |
| NVIDIA Quadro P4000 | NVIDIA | https://www.nvidia.com/content/dam/en-zz/Solutions/design-visualization/productspage/quadro/quadro-desktop/quadro-pascal-p4000-data-sheet-a4-nvidia-704358-r2-web.pdf | |
| Windows 10 Enterprise 2016 | Microsoft | https://www.microsoft.com/en-us/evalcenter/evaluate-windows-10-enterprise | |
| Workstation Minimum Requirements | https://theobjects.com/dragonfly/system-requirements.html |
References
- Bai, X. -. C., Mcmullan, G., Scheres, S. H. W. How cryo-EM is revolutionizing structural biology. Trends in Biochemical Sciences. 40 (1), 49-57 (2015).
- de Oliveira, T. M., van Beek, L., Shilliday, F., Debreczeni, J., Phillips, C. Cryo-EM: The resolution revolution and drug discovery. SLAS Discovery. 26 (1), 17-31 (2021).
- Danev, R., Yanagisawa, H., Kikkawa, M. Cryo-EM performance testing of hardware and data acquisition strategies. Microscopy. 70 (6), 487-497 (2021).
- Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. Journal of Structural Biology. 152 (1), 36-51 (2005).
- . Tomography 5 and Tomo Live Software User-friendly batch acquisition for and on-the-fly reconstruction for cryo-electron tomography Datasheet Available from: https://assets.thermofisher.com/TFS-Assets/MSD/Datasheets/tomography-5-software-ds0362.pdf (2022)
- Danev, R., Baumeister, W. Expanding the boundaries of cryo-EM with phase plates. Current Opinion in Structural Biology. 46, 87-94 (2017).
- Hylton, R. K., Swulius, M. T. Challenges and triumphs in cryo-electron tomography. iScience. 24 (9), (2021).
- Turk, M., Baumeister, W. The promise and the challenges of cryo-electron tomography. FEBS Letters. 594 (20), 3243-3261 (2020).
- Oikonomou, C. M., Jensen, G. J. Cellular electron cryotomography: Toward structural biology in situ. Annual Review of Biochemistry. 86, 873-896 (2017).
- Wagner, J., Schaffer, M., Fernández-Busnadiego, R. Cryo-electron tomography-the cell biology that came in from the cold. FEBS Letters. 591 (17), 2520-2533 (2017).
- Lam, V., Villa, E. Practical approaches for Cryo-FIB milling and applications for cellular cryo-electron tomography. Methods in Molecular Biology. 2215, 49-82 (2021).
- Chreifi, G., Chen, S., Metskas, L. A., Kaplan, M., Jensen, G. J. Rapid tilt-series acquisition for electron cryotomography. Journal of Structural Biology. 205 (2), 163-169 (2019).
- Eisenstein, F., Danev, R., Pilhofer, M. Improved applicability and robustness of fast cryo-electron tomography data acquisition. Journal of Structural Biology. 208 (2), 107-114 (2019).
- Esteva, A., et al. Deep learning-enabled medical computer vision. npj Digital Medicine. 4 (1), (2021).
- Liu, Y. -. T., et al. Isotropic reconstruction of electron tomograms with deep learning. bioRxiv. , (2021).
- Moebel, E., et al. Deep learning improves macromolecule identification in 3D cellular cryo-electron tomograms. Nature Methods. 18 (11), 1386-1394 (2021).
- Chen, M., et al. Convolutional neural networks for automated annotation of cellular cryo-electron tomograms. Nature Methods. 14 (10), 983-985 (2017).
- Ronneberger, O., Fischer, P., Brox, T. U-net: Convolutional networks for biomedical image segmentation. Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics). 9351, 234-241 (2015).
- Kremer, J. R., Mastronarde, D. N., McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. Journal of Structural Biology. 116 (1), 71-76 (1996).
- Iancu, C. V., et al. A "flip-flop" rotation stage for routine dual-axis electron cryotomography. Journal of Structural Biology. 151 (3), 288-297 (2005).
This article has been published
Video Coming Soon
ABOUT JoVE
Copyright © 2024 MyJoVE Corporation. All rights reserved