A High-Resolution, Single-Grain, In Vivo Pollen Hydration Bioassay for Arabidopsis thaliana
In This Article
Summary
An improved method to measure pollen hydration profiles in Arabidopsis thaliana is described here. The new method offers higher resolution, is noninvasive, and is highly reproducible. The protocol represents a new tool for a finer dissection of the processes that regulate the early stages of pollination.
Abstract
Sexual reproduction in flowering plants requires initial interaction between the pollen grain and the stigmatic surface, where a molecular dialog is established between the interacting partners. Studies across a range of species have revealed that a series of molecular checkpoints regulate the pollen-stigma interaction to ensure that only compatible, generally intraspecific pollen is successful in effecting fertilization. In species that possess a 'dry stigma', such as the model plant Arabidopsis thaliana, the first post-pollination, prezygotic compatibility checkpoint is the establishment of pollen hydration.
This phase of pollination is tightly regulated, whereby signals from the pollen grain elicit the release of water from the stigma, thus permitting pollen hydration. The ability to accurately measure and track pollen hydration over time is key to the design of experiments directed at understanding the regulation of this critical step in reproduction. Published protocols frequently utilize flowers that have been excised from the parent plant, maintained on liquid or solid media, and bulk pollinated.
This paper describes a noninvasive, in vivo pollination bioassay that permits minute-by-minute hydration tracking of individual A. thaliana pollen grains at high resolution. The assay is highly reproducible, able to detect very subtle variations of pollen hydration profiles, and thus is suitable for the analysis of mutants that affect pathways regulating pollination. Although the protocol is lengthier than those described for bulk pollinations, the precision and reproducibility it provides, along with its in vivo nature, make it ideal for the detailed dissection of pollination phenotypes.
Introduction
Successful sexual reproduction in angiosperms typically relies on the transfer of intraspecific pollen grains from the anther to the stigma, either within or between individuals (i.e. pollination). This transfer of pollen grains to a receptive flower is usually mediated by pollinators or abiotic factors; as such, this also frequently results in the deposition of heterospecific pollen under natural conditions. With a few exceptions, the progression of pollination by heterospecific pollen is evolutionarily disadvantageous, reducing reproductive fitness through lost mating opportunities, with most resulting hybrid progeny either failing to develop appropriately or being sterile1. Thus, mechanisms have evolved to block pollination by 'incompatible' heterospecific pollen2. Rapid recognition of compatible pollen is therefore arguably the most important process in the early stages of sexual reproduction in many flowering plants.
In the Brassicaceae family, where stigmas are of the 'dry' type, a series of molecular checkpoints act at multiple stages in the reproductive process regulating pollination, such that only compatible pollen is successful. Pollen hydration is one of the most important checkpoints (Figure 1), as pollen that fails to hydrate cannot progress to produce a pollen tube and subsequently deliver sperm to the female gametophyte. Frequently, incompatible grains fail to pass this first pollination checkpoint, and thus do not gain access to stigmatic water3. Amongst members of the Brassicaceae family, the recognition of pollen occurs rapidly, with compatibility being established within minutes of pollen grain attachment to the pistil4,5. In recent years, much progress has been made, and we are now beginning to understand the molecular mechanisms that regulate key pollination checkpoints.
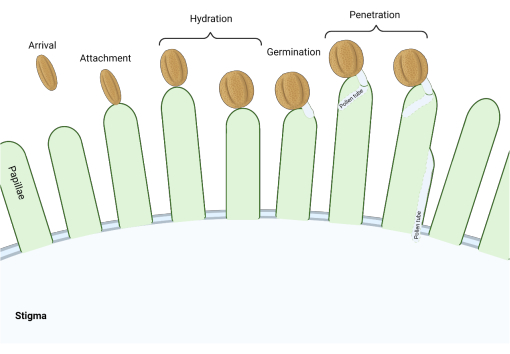
Figure 1: Overview of key events during compatible pollination. These stages, such as pollen hydration and pollen tube germination, are also pollination 'checkpoints' that must be successfully navigated to effect compatible pollination. The diagram represents a 'dry' type stigma, which is typical to species from the Brassicaceae family2,20. Please click here to view a larger version of this figure.
Pioneering research on the Brassica self-incompatibility (SI) system, where 'self' pollen is recognized and rejected, established the paradigm for pollen-stigma recognition in Brassicaceae6,7,8,9,10. SI in Brassica and its relatives is mediated by 'recognition' proteins that reside on the surface of pollen and at the stigmatic plasma membrane that, upon interaction, lead to pollen rejection. SI pollen rejection operates by disruption of the basal pollen-stigma compatibility system which, when fully activated by the perception of compatible pollen, leads to targeted secretion by the stigma, thus driving pollen hydration (for reviews of the pollen compatibility mechanism, see11,12). In the example of SI, the pollen-borne ligand is a small cysteine-rich protein, S-locus cysteine rich (SCR/SP11), and the stigmatic receptor is the S-locus receptor kinase (SRK).
Recently, in Arabidopsis thaliana, another group of small cysteine-rich pollen-borne proteins, pollen coat protein class Bs (AtPCP-Bs), have been found to be important regulators of pollen acceptance through the activation of pollen hydration13. Stigmatic receptors of the AtPCP-Bs and aspects of the downstream regulatory pathway have also recently been described14,15. Interestingly, mutational studies of genes encoding potential pollen-borne and stigmatic signaling mediators of pollen hydration (including AtPCP-Bs) have failed to generate plants that have a complete block to the pollen hydration checkpoint. This strongly suggests that multiple other, yet undiscovered, factors play a role in the regulation of pollen hydration. Building upon the method first described by Wang et al.13, here we describe an improved high-resolution in vivo bioassay suitable for the identification of subtle pollen hydration defects in candidate mutant A. thaliana lines.
Protocol
1. Plant growth and preparation of flowers
- Stratify A. thaliana seeds in 0.1% agarose or sterile water for 3 days at 4 °C, or as dry seeds for 16-24 h at -20 °C (uNASC, personal communication).
- Transfer the stratified seeds to pots of compost and place in an environmentally controlled growth chamber. Propagate plants with a 16:8 h, light:dark photoperiod provided by fluorescent tubes (130 µmol m-2 s-1). Maintain the temperature at 21 ± 2 °C with approximately 40% relative humidity.
- Ensure that pollen donor and recipient plants, along with any other appropriate 'control' plant lines, are sown together to ensure synchronous flowering. Propagate the plants for approximately 6 weeks until the inflorescences are well established.
- Select stage 12 flower buds on the pollen recipient plant 1 day prior to carrying out the bioassay for emasculation16,17-these are unopened flower buds that will complete flower opening and anther dehiscence the following day18.
NOTE: Avoid the first three flowers produced on the main inflorescence, as these typically exhibit unusual reproductive behavior. If available and appropriate for the study, use a male sterile plant line, such as the A. thaliana (accession Col-0) pA9-barnase line, where anthers fail to mature19. - To emasculate flowers of the pollen recipient, place the plant, in its pot, on its side. Tape the plant stem, in the region close to the flowers that will be emasculated, to a glass slide that is in position under a stereo dissecting microscope.
- Using a pair of fine-tipped forceps, carefully tease open the flower bud and remove all the flower petals and anthers. Ensure that the pistil is undamaged and that the stigma is free of contaminating pollen.
NOTE: Male sterile plant lines do not require emasculation. - Return the plants to the growth chamber and ensure the emasculated flowers do not come into contact with other plants or foreign objects.
2. Pollen hydration assay-raw data acquisition
- The following morning, remove the plants from the growth chamber. Lay the pollen recipient plant on its side and position the flower on the stage of an inverted microscope (Figure 2) so that the stigma can be clearly imaged.
- Fix the position of the flower to be imaged by immobilizing the stem to a glass slide using strips of masking tape. Maintain the temperature between 18 °C and 25 °C and relative humidity below 60%.
NOTE: For the pA9-barnase male sterile plant line, it is optimal to perform the assay in the morning when the flowers open and the petals do not hinder the field of view. - Next, remove a healthy and freshly opened flower from the pollen-donor plant. Place under a dissecting microscope and gather a few pollen grains on the tip of clean fine-tipped forceps by gently touching the anthers (Figure 3A). An eyelash taped to a short rod is also an effective tool for collecting and transferring pollen (Figure 3C).
- Remove excess pollen from the forceps by lightly touching them against the petals of the flowers from where the pollen grains were harvested, until a monolayer of pollen grains is formed at the tip of the forceps.
NOTE: A monolayer of pollen grains on the tip of fine-tipped forceps will significantly facilitate single grain transfer in subsequent steps. It is also possible to obtain one single pollen grain on the forceps by this technique (Supplementary Video S1). - Return to the pollen recipient plant and, using a low-power objective lens (e.g., 10x objective lens; Figure 3B), focus the inverted microscope on the stigma to be pollinated.
- Holding the forceps along the opening between the arms of the forceps (Figure 4), carefully approach an unpollinated ('virgin') stigmatic papilla cell.
NOTE: We have found this method of holding the forceps aids dexterity and reduces the impact of shaking hands. A micromanipulator can be used for users who are less experienced or have difficulty precisely applying a single pollen grain by hand. - Select a suitably positioned pollen grain on the forceps for transfer to the stigma. Continue to approach the unpollinated stigmatic papilla cell until the selected pollen grain makes light contact with its surface. Slowly withdraw the forceps and confirm pollen attachment (Figure 5).
NOTE: Supplementary Video S1 and Supplementary Video S2 demonstrate this step with both single and multiple pollen grains present on the forceps. - Ensure that the pollen grain is oriented such that its equatorial axis is clearly visible and in sharp focus. Immediately switch to a higher power objective lens (e.g., 20x) and capture an image of the pollen grain. This first image is T = 0. Continue to capture further images at 1 min intervals for a total of 10 min.
- Adjust the focus as necessary to accommodate small movements in the pollen grain or stigma. Record the ambient temperature and relative humidity of the room every 2 min to allow for future comparisons between experimental replicates.
- Once all the images have been captured, save them in a lossless format, such as the manufacturer's proprietary format or as TIFFs.
NOTE: There will be 11 images per pollen grain sampled (Supplementary Figure S1). The automated/manual timelapse acquisition settings in most proprietary image acquisition software are useful features to facilitate the organization of each time-series. - Repeat steps 2.4 to 2.9 for additional pollen grains. Acquire data for near-equivalent numbers of control (wild type [WT]) and experimental pollinations.
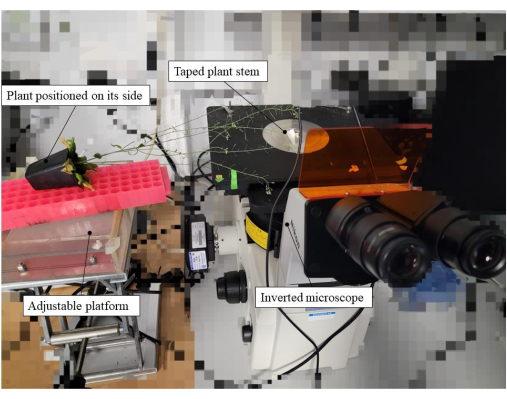

Figure 2: Equipment setup utilized for the pollen hydration bioassay. In this example, a pA9-barnase male sterile plant line was the pollen recipient. The plant, within its pot, was placed on its side, and the stem taped to a glass slide positioned on the stage of the microscope. To reduce mechanical stress and aid positioning of the plant, an adjustable platform was used for supporting the plant pot. Please click here to view a larger version of this figure.
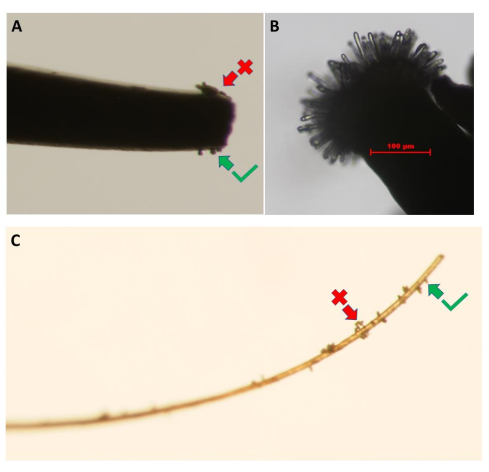
Figure 3: Collection of pollen grains from the pollen-donor flower. Images show the use of (A) fine-tipped forceps and (C) a piece of eyelash. Clumps of pollen (red arrow) should be removed by lightly touching them against the petals of the donor flowers until a monolayer of pollen grains is obtained (green arrow). (B) High-resolution image of an unpollinated A. thaliana (Col-0) pA9-barnase male sterile line stigma that has reached the appropriate developmental stage for the pollen hydration bioassay. Scale bar = 100 µm (B). Please click here to view a larger version of this figure.
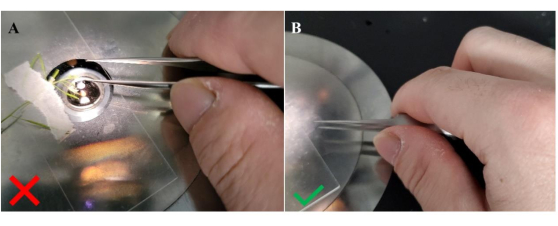
Figure 4: Method for holding forceps when transferring pollen to the recipient stigma. (A) Incorrect orientation for holding the forceps; (B) correct orientation for holding the forceps. Holding the forceps sideways in this configuration, as signified by the position of the thumb in between the arms of the forceps, provides greater stability to facilitate the transfer of pollen grains to unpollinated stigmatic papillae. Please click here to view a larger version of this figure.
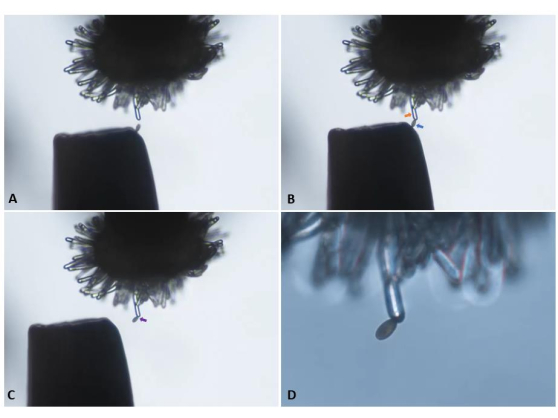
Figure 5: Transfer of a single pollen grain from the tip of a pair of forceps to an unpollinated ('virgin') stigmatic papilla cell of a pA9-barnase male sterile plant. (A) Careful approach toward the papilla cell. (B) Attachment of an appropriately positioned pollen grain (blue arrow) to the papilla cell (orange arrow). (C) Withdrawal of the forceps and visual confirmation of pollen attachment (purple arrow). Panels A-C were imaged with a 10x objective lens (10.5 mm working distance; 0.25 numerical aperture) and are snapshots derived from the video clip presented in Supplementary Video S1. (D) Transition to a 20x objective lens (2.1 mm working distance; 0.5 numerical aperture) for initiation of image capture in a time-series. Please click here to view a larger version of this figure.
3. Pollen hydration assay-measurements
- Define the rate of pollen hydration as the change in length of the semi-minor axis (Figure 6) of the pollen grain (i.e., the equatorial radius) over time and present it as percentage change (equation [1]):
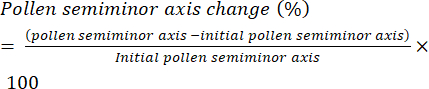 (1)
(1) - Using image analysis software, record semi-minor axis values for each pollen grain in the experimental series.
NOTE: The name of this measurement option is software-dependent, such as 'Rotated Ellipse' or '5-point ellipse'. - Repeat steps 3.1-3.2 for all other pollen grains to be measured. For consistency, apply the same degree of digital zoom and the same approach to defining the 'pollen boundary' for all measurements in the datasets.
- Once all the measurements are completed for a time-series, export the raw semi-minor axis values of each image stack to a spreadsheet and present the data in columns per image stack. Ensure the inclusion of data from at least 15 hydrated pollen grains in the analysis for each plant line (Supplementary Table S1).
- It is not unusual for a small number of pollen grains to fail to hydrate or hydrate significantly more slowly than expected. These 'dud' grains may be the result of poor contact between the grain and papilla cell or related to pollen viability. Look for and exclude these from the dataset unless these are required in their experimental design.
- Calculate the mean values for each timepoint per plant line. Use unpaired t-tests and one-way ANOVA for the statistical analysis of hydration data from WT and mutant lines at each timepoint. Use a multiple t-test for the simultaneous comparison of the means between WT and mutant lines across multiple timepoints.
NOTE: XY plots are also very useful for visualizing the overall trend of pollen hydration between the plant lines being compared.
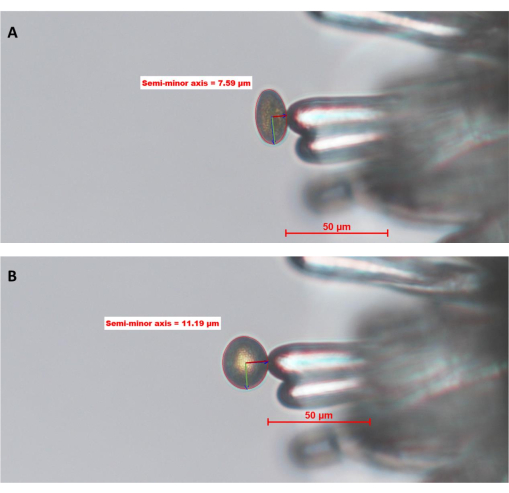
Figure 6: WT pollen grain hydrating on a stigmatic papilla cell of A. thaliana (Col-0; pA9-barnase male sterile line). (A) Time point zero, 0 (0 MAP) and (B) 10 MAP. The red circle around the pollen grain is the "pollen boundary" defined and drawn by the operator using image analysis software. The green and dark red lines inside the pollen represent the semi-major and semi-minor axes, respectively. The length of the semi-minor axis is used to calculate the degree of pollen hydration. A full time-series of this dataset can be found in Supplementary Figure S1. The image was taken with a 20x objective lens (2.1 mm working distance; 0.5 numerical aperture). Scale bars = 50 µm. Abbreviation: MAP = min-after-pollination. Please click here to view a larger version of this figure.
Results
This section presents two sets of example pollen hydration data, collected as described above, for A. thaliana. The first set of data consists of three replicates of a pollen hydration time-series for WT plants, with each replicate collected on a different day. Each replicate contains no less than 18 individual pollen grain values, totaling 55 pollen grains across all three replicates. Minimum and maximum values for the means between replicates, for all time points, were within 3% (Figure 7 and Supplementary Table S1). These representative data for WT pollinations clearly demonstrate the high degree of consistency that can be obtained utilizing the methodology detailed here for relatively low sample numbers and across different days.
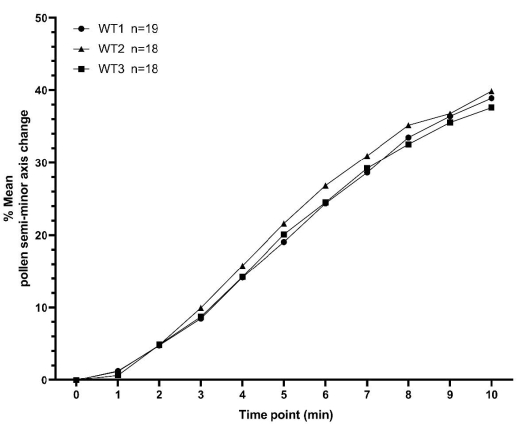
Figure 7: XY plot showing consistency of A. thaliana wild type pollen hydration profiles over a 10 min time period. The pollen parent was the Col-0 accession of A. thaliana and the pistil parent was the pA9-barnase male sterile A. thaliana (Col-0) line. The data represent three independent datasets collected on different days and demonstrate a high degree of consistency. A box and whisker diagram and statistical analysis of the means for these datasets is presented in Supplementary Figure S2. The number of measured pollen ('n') for each independent dataset is displayed next to the syntax (WT1/WT2/WT3) on the figure. Abbreviation: WT = wild type. Please click here to view a larger version of this figure.
The second set of data was obtained for a plant line harboring a T-DNA insertion in a pollen coat protein-encoding gene generating a 'knockdown' mutation, herein refer to as 'KD mutant'. Mutant pollen was deposited onto pA9-barnase male sterile stigmas for hydration profiling, as described in the protocol. As can be seen from the resulting data (Figure 8), mutant and WT pollen had indistinguishable hydration profiles over the first 5 min. However, 5-10 min after pollination (MAP), the mean semi-minor axis change for mutant pollen started to fall behind that of WT pollen, with the difference becoming statistically significant at 10 MAP. This result not only demonstrates that this pollen coat protein has a role in mediating pollen hydration, but also nicely illustrates the utility of this high-resolution, single-grain bioassay for tracking pollen hydration. In this particular example, its sensitivity was able to detect the subtle effect of a 'knockdown' of a pollen coat protein-encoding gene.
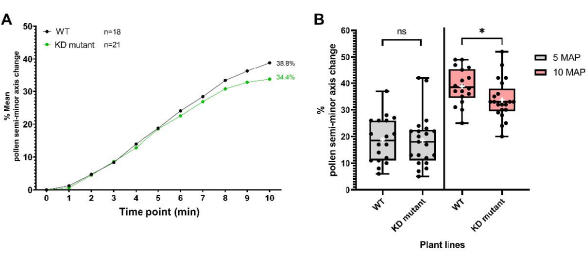
Figure 8: Pollen hydration profiles for WT and a 'knockdown' pollen coat protein mutant line (KD mutant). (A) Hydration profiles over a 10 min time period for WT and mutant pollen. The pollen parents were the Col-0 accession of A. thaliana and the pollen coat protein KD mutant (also in the Col-0 background). In both cases, the pistil parent was the pA9-barnase male sterile A. thaliana (Col-0) line. (B) Box and whisker plots showing the extent of pollen hydration (in terms of percentage change of the semi-minor axis) at 5 MAP and 10 MAP for WT and mutant pollen datasets. Whiskers represent sample minimum and maximum values. Boxes depict the lower quartile, median, and upper quartile of the dataset. White crosses represent the mean of the dataset. Unpaired t-test analysis show that the mean percentage of pollen hydration is significantly different between the two plant lines at 10 MAP. One asterisk indicates p < 0.05 (unpaired t-test). Abbreviations: WT = wild type; KD = knockdown; MAP = min-after-pollination. Please click here to view a larger version of this figure.
Supplementary Figure S1: A cropped pollen hydration bioassay time-series of a WT pollen grain hydrating on a stigmatic papilla cell of a pA9-barnase male sterile plant over the course of 10 min. Images were taken at 1 min intervals. The images at 0 MAP and 10 MAP were used in Figure 6 (separately attached). Scale bar = 50 µm. Abbreviations: WT = wild type; MAP = min-after-pollination. Please click here to download this File.
Supplementary Figure S2: Box and whisker plot showing the extent of pollen hydration (in terms of percentage change of the semi-minor axis) over a 10 min time period for the three WT pollen datasets described in Figure 7. The pistil parent was the pA9-barnase male sterile A. thaliana (Col-0) line. Whiskers represent sample minimum and maximum values. Boxes depict the lower quartile (lower hinge), median (middle hinge), and upper quartile (upper hinge) of the dataset. Individual datapoints are shown. One-way ANOVA shows that the mean percentage of pollen hydration values between the three datasets were not statistically significantly different from one another throughout the 10 min time period. The significant threshold is p < 0.05 (one-way ANOVA). Abbreviation: WT = wild type. Please click here to download this File.
Supplementary Table S1: Raw pollen hydration data used to construct Figure 7 (A. thaliana WT Col-0 pollen on pA9-barnase male sterile stigma). Please click here to download this File.
Supplementary Video S1: A video demonstrating the transfer of a single WT (Col-0 accession) pollen grain on the tip of a pair of forceps to a 'virgin' stigmatic papilla cell (pA9-barnase male sterile line). To facilitate the accessibility of the video, the image quality was intentionally lowered. Please click here to download this File.
Supplementary Video S2: A video demonstrating the transfer of a single WT (Col-0 accession) pollen from a monolayer of pollen grains at the tip of a pair of forceps to a 'virgin' stigmatic papilla cell (pA9-barnase male sterile line). To facilitate the accessibility of the video, the image quality was intentionally lowered. Please click here to download this File.
Discussion
For flowering plants, the very early stages of sexual reproduction are arguably the most important. At the level of the pollen-stigma interaction, molecular decisions are made that determine the 'compatibility' of the interacting partners. Such decisions, if made correctly, avoid wastage of resources that could impact reproductive fitness21. Thus, permitting only compatible pollen to effect fertilization is one important component of maintaining well adapted genotypes, and thus the evolutionary success of species. Research conducted with the model plant A. thaliana has been extremely valuable in deepening our understanding of this process. A number of studies over the past few decades have revealed the presence of factors in the pollen coat that act at the first compatibility 'checkpoint', where the pollen gains access to stigmatic water to permit pollen hydration13. Despite these first insights into the mechanisms that regulate pollen-stigma compatibility, there are still many gaps in our understanding of this process. To date, no mutants of pollen-borne ligands or stigmatic receptors known to impact pollen hydration can completely block compatible pollination, suggesting the presence of other undiscovered pollen hydration determinants. By being able to readily observe the phenotype of interest, the pollen hydration bioassay described here is one of the most straightforward techniques to study potential mutants that regulate pollination.
Existing methodologies for measuring pollen hydration commonly utilize bulk pollinations and report fewer timepoints14,22,23, and thus may miss important subtle hydration profile phenotypes. For example, the study by Wang et al.13, along with work on other pollen coat protein mutants in our lab (unpublished observations), have revealed intriguing differences in hydration profiles between mutants. Such subtle differences may hold important clues to the regulatory mechanisms underlying compatible pollination.
The method described here focuses on the acquisition of relatively small numbers of measurement between mutant and WT plant lines, with an emphasis on methodological precision to reduce variation within the datasets. Whilst this method is highly reproducible (as shown in Figure 7), assuming that temperature and humidity are adequately controlled, it is important to gather hydration data for nearly equal numbers of WT and mutant pollen on the same day to further reduce the potential for variation. Data can be then pooled across different days if required. In addition, selecting the appropriate WT control plants is vital for correct interpretation of the hydration results. For the pollen recipient, the same plant line should be used for receiving both WT control and mutant pollen grains.
For example, we use the pA9-barnase male sterile plant line, which is also featured in the video protocol, as the pollen recipient for both WT (control) and mutant (experimental) pollen when investigating T-DNA pollen mutant lines (such as the 'KD' mutant described in Figure 8). The mixing of data from such a male sterile line, which need not be emasculated, with that gathered from a manually emasculated control line should be avoided as these stigmas will likely behave differently. Likewise, emasculated mutant lines should be utilized in conjunction with an emasculated WT (control) line whenever possible. The same caution should also be applied when considering the genetic background of the plants under study. While most popular T-DNA mutant collections were generated in the Col-0 background, others, such as the FLAG collection from Institut national de la Recherche Agronomique (INRA), are available in the Wassilewskija (WS) genetic background24,25. In such cases, it is advisable to use the respective ecotype's WT plant lines as controls.
Although here we have focused on pollen hydration over the first 10 min of the pollen-stigma interaction, this method can also be adapted to encompass hydration profiles that cover a longer time period. A key feature of the protocol is that flowers remain attached to the parent plant-current published protocols typically require excision of the pistil and placement in media to sustain the tissue for the duration of the experiment14,18,26. Although there is no direct evidence to suggest such a semi in vivo approach impacts pollen hydration or indeed alters the in vivo regulation of this process, it is conceivable that excision of the flowers from the parent plant could impact pollination. Thus, this protocol achieves a true in vivo environment for the study of the pollen-stigma interaction, where the structural integrity of the plant is preserved.
The transfer of single pollen grains to 'virgin' stigmatic papillae is arguably one of the most challenging operations described in this protocol. It is not uncommon to transfer clusters of pollen grains in error. However, the chance of this occurring can be greatly reduced by ensuring that only a monolayer of pollen is present on the forceps (Figure 3A) (or even just a single pollen grain; Figure 5), and/or by utilizing pollen grains that are already orientated, such that they 'protrude' from others on the tip of the forceps. We have found that an experienced operator can successfully complete the transfer of a single pollen to a stigmatic papilla cell in approximately 3 min and record data for up to five pollen grains over a 1 h period. Thus, over a period of 2-4 days, enough data can be accumulated for meaningful statistical analysis of the plant lines under study.
Human error is potentially the biggest source of variation in the analysis of datasets derived from studies utilizing this protocol. For example, the definition of the 'pollen boundary' during image analysis comes down to the judgement of the individual researcher. Thus, there is the potential that measurements made by different researchers, even on the same dataset, may generate variation. Wherever possible, a single researcher should carry out the measurements to minimize sampling errors. In addition, coupling the analysis of WT and mutant datasets by the same operator negates the potentially subjective definition of the 'pollen boundary' and interoperator variation.
In conclusion, a sophisticated yet accurate method to measure pollen hydration profiles in the model organism A. thaliana is described. We have demonstrated that, by utilizing this protocol, highly consistent pollen hydration data for A. thaliana can be readily acquired. Three independent batches of data for WT pollinations acquired on different days showed consistent small deviations of <3% across all timepoints (Figure 7 and Supplementary Table S1). Although the bioassay presented here is slightly more complex than most existing protocols, the resolution of the data generated is superior and suitable for the identification and characterization of novel mutants that impact pathways regulating compatible pollination.
Disclosures
The authors have no conflicts of interest to declare.
Acknowledgements
This research was supported by University of Bath (University of Bath, Bath, UK, BA2 7AY) postgraduate scholarships to Y.-L.L. and L.W. Figure 1 was created with BioRender.com (https://biorender.com/).
Materials
| Name | Company | Catalog Number | Comments |
| A9-barnase line | University of Bath | Courtsey of Prof. Rod Scott | Male sterile Arabidopsis thaliana wildtype equivalent line of the ecotype Columbia-0 |
| Dumont Tweezer, Dumont #5 Inox 11cm | Fisher | Dumont 500342 | Tweezer uses for transfer of pollen grain |
| GraphPad Prsim (version 8.0.2) | Dotmatics | Prism | Comprehensive data analysis, graphing and statistics software |
| JMP (version 17) | JMP Statistical Discovery LLC | JMP 17 | Statistical analysis software |
| Levington F2S seed & modular compost (with sand) | Levington | LEV75F2SMS | General-purpose compost for plant growth |
| Micromanipulator | Singer instrument Co. LTD. | Singer Micromanipulator | Micromanipulator to aid transfer of pollen grain |
| Nikon Digit sight DS-U1 | Nikon | DS-U1 | Microscope camera (coupletd to SMZ1500) |
| Nikon Eclipse TE2000-S Inverted Microscope | Nikon | TE2000-S | Inverted microscope |
| Nikon SMZ1500 Stereomicroscope | Nikon | SMZ1500 | Stereomicroscope |
| Nikon DS-Fi3 microscope camera | Nikon | DS-Fi3 | Microscope camera (coupletd to TE2000-S) |
| Nikon NIS-Elements Basic Research | Nikon | NIS-Elements BR | Image accquisition and analysis software (for DS-Fi3) |
| Nikon NIS-Elements F | Nikon | NIS-Elements F | Image accquisition and analysis software (for DS-U1) |
| WT Col-0 plant line | NASC | N700000 | Wildtype Arabidopsis thaliana, ecotype Columbia-0 |
References
- Rieseberg, L. H., Willis, J. H. Plant speciation. Science. 317 (5840), 910-914 (2007).
- Hiscock, S. J., Allen, A. M. Diverse cell signalling pathways regulate pollen-stigma interactions: the search for consensus. New Phytologist. 179 (2), 286-317 (2008).
- Kandasamy, M. K., Nasrallah, J. B., Nasrallah, M. E. Pollen pistil interactions and developmental regulation of pollen-tube growth in Arabidopsis. Development. 120 (12), 3405-3418 (1994).
- Bosch, M., Wang, L. Pollen-stigma interactions in Brassicaceae: complex communication events regulating pollen hydration. Journal of Experimental Botany. 71 (9), 2465-2468 (2020).
- Rozier, F., et al. Live-cell imaging of early events following pollen perception in self-incompatible Arabidopsis thaliana. Journal of Experimental Botany. 71 (9), 2513-2526 (2020).
- Dickinson, H. Dry stigmas, water and self-incompatibility in Brassica. Sexual Plant Reproduction. 8, 1-10 (1995).
- Takasaki, T., et al. The S receptor kinase determines self-incompatibility in Brassica stigma. Nature. 403 (6772), 913-916 (2000).
- Schopfer, C. R., Nasrallah, M. E., Nasrallah, J. B. The male determinant of self-incompatibility in Brassica. Science. 286 (5445), 1697-1700 (1999).
- Takayama, S., et al. Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature. 413 (6855), 534-538 (2001).
- Shiba, H., et al. A pollen coat protein, SP11/SCR, determines the pollen S-specificity in the self-incompatibility of Brassica species. Plant Physiology. 125 (4), 2095-2103 (2001).
- Broz, A. K., Bedinger, P. A. Pollen-pistil interactions as reproductive barriers. Annual Review of Plant Biology. 72 (1), 615-639 (2021).
- Cheung, A. Y., Duan, Q., Li, C., James Liu, M. -. C., Wu, H. -. M. Pollen-pistil interactions: It takes two to tangle but a molecular cast of many to deliver. Current Opinion in Plant Biology. 69, 102279 (2022).
- Wang, L. D., et al. PCP-B class pollen coat proteins are key regulators of the hydration checkpoint in Arabidopsis thaliana pollen-stigma interactions. New Phytologist. 213 (2), 764-777 (2017).
- Liu, C., et al. Pollen PCP-B peptides unlock a stigma peptide-receptor kinase gating mechanism for pollination. Science. 372 (6538), 171-175 (2021).
- Bordeleau, S. J., Sanchez, L. E. C., Goring, D. R. Finding new Arabidopsis receptor kinases that regulate compatible pollen-pistil interactions. Frontiers in Plant Science. 13, 1022684 (2022).
- Suwabe, K., et al. Double-locking mechanism of self-compatibility in Arabidopsis thaliana: the synergistic effect of transcriptional depression and disruption of coding region in the male specificity gene. Frontiers in Plant Science. 11, 576140 (2020).
- Smyth, D. R., Bowman, J. L., Meyerowitz, E. M. Early flower development in Arabidopsis. Plant Cell. 2 (8), 755-767 (1990).
- Lee, H. K., Macgregor, S., Goring, D. R. A toolkit for teasing apart the early stages of pollen-stigma interactions in Arabidopsis thaliana. Pollen and Pollen Tube Biology. 2160, 13-28 (2020).
- Dilkes, B. P., et al. The maternally expressed WRKY transcription factor TTG2 controls lethality in interploidy crosses of Arabidopsis. PLoS Biology. 6 (12), 2707-2720 (2008).
- Riglet, L., et al. KATANIN-dependent mechanical properties of the stigmatic cell wall mediate the pollen tube path in Arabidopsis. eLife. 9, e57282 (2020).
- Zhou, L. Z., Dresselhaus, T. Friend or foe: Signaling mechanisms during double fertilization in flowering seed plants. Plant Development and Evolution. 131, 453-496 (2019).
- Gao, X. -. Q., et al. The Arabidopsis KINβγ subunit of the SnRK1 complex regulates pollen hydration on the stigma by mediating the level of reactive oxygen species in pollen. PLoS Genetics. 12 (7), e1006228 (2016).
- Lee, H. K., Goring, D. R. Two subgroups of receptor-like kinases promote early compatible pollen responses in the Arabidopsis thaliana pistil. Journal of Experimental Botany. 72 (4), 1198-1211 (2021).
- O'Malley, R. C., Barragan, C. C., Ecker, J. R. A user's guide to the Arabidopsis T-DNA insertion mutant collections. Pollen and Pollen Tube Biology. 1284, 323-342 (2015).
- Samson, F., et al. FLAGdb++: a database for the functional analysis of the Arabidopsis genome. Nucleic Acids Research. 32, D347-D350 (2004).
- Doucet, J., et al. Investigations into a putative role for the novel BRASSIKIN pseudokinases in compatible pollen-stigma interactions in Arabidopsis thaliana. BMC Plant Biology. 19 (1), 549 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved