A subscription to JoVE is required to view this content. Sign in or start your free trial.
An Efficient Transgenesis Approach for Gene Delivery in the Mouse Embryonic Heart
In This Article
Summary
This protocol presents a detailed methodological framework for electroporation-based transgenesis of cardiac cells in developing mouse hearts. The video assets provided here will facilitate learning of this versatile technique.
Abstract
The mammalian heart is a complex organ formed during development via highly diverse populations of progenitor cells. The origin, timing of recruitment, and fate of these progenitors are vital for the proper development of this organ. The molecular mechanisms that govern the morphogenesis of the heart are essential for understanding the pathogenesis of congenital heart diseases and embryonic cardiac regeneration. Classical approaches to investigate these mechanisms employed the generation of transgenic mice to assess the function of specific genes during cardiac development. However, mouse transgenesis is a complex, time-consuming process that often cannot be performed to assess the role of specific genes during heart development. To address this, we have developed a protocol for efficient electroporation and culture of mouse embryonic hearts, enabling transient transgenesis to rapidly assess the effect of gain- or loss-of-function of genes involved in cardiac development. Using this methodology, we successfully overexpressed Meis1 in the embryonic heart, with a preference for epicardial cell transfection, demonstrating the capabilities of the technique.
Introduction
The heart is the first organ formed during embryonic development. This process involves the spatiotemporal coordination of various populations of progenitor cells from distinct areas of the embryo. All this occurs while the developing heart continues to beat and function, emphasizing the remarkable coordination required for its formation1,2,3. Given the crucial role of the heart, tight regulation at the cellular and molecular levels is essential for its proper formation4,5. Identifying the mechanisms that control heart development has been of great interest, as they are crucial for unraveling congenital heart disorders, which impact a substantial number of patients worldwide6. Furthermore, comprehending heart development is pivotal in deciphering cardiac regeneration, as postnatal mammalian hearts retain a regenerative capacity that is lost or hindered in adulthood7,8. Consequently, dissecting molecular regulators of heart development is imperative to advance research efforts on congenital heart disease and cardiac regeneration.
In pursuit of this objective, there has been a growing focus on investigating the role of the epicardium in cardiac development and regeneration9. The epicardium is a thin layer of mesothelial tissue that comprises the outermost layer of the mammalian heart (Figure 1). Recent studies have shown the importance of the epicardium during cardiac injury, revealing that this tissue is able to send proliferation signals to cardiomyocytes in the affected area to mitigate the damage10,11. Despite the importance of the epicardium, conducting further molecular investigations has been challenged by its immense heterogeneity. Single-cell RNAseq experiments have revealed the epicardium's heterogeneity, housing multiple cell subpopulations with distinct transcriptomic signatures12,13,14,15,16. Thus, a strategy to screen potential regulators of cardiac development should accommodate the diversity of epicardial progenitor cells.
In this sense, the mouse model's amenability to genetic modification has facilitated the identification of numerous genes crucial for heart development, allowing the generation of mutant lines with gain-of-function (GOF) or loss-of-function (LOF) of specific genes. However, these approaches imply a considerable investment of time and experimental resources; therefore, they are impractical when assessing the roles of a large number of candidate genes. Besides, developmental genes often exert pleiotropic functions in different tissues or are required for early embryonic development, hampering the interpretation of their contribution to development in a specific process. While it is possible to target gene function at specific structures or developmental time points, this usually requires the use of more complex genetic constructions, which can be difficult to generate or are generally unavailable.
To overcome these limitations, we present a methodology to electroporate mouse embryonic hearts for transient transgenesis (Figure 2). Paired with ex vivo culture and fluorescence-activated cell sorting (FACS), this strategy demonstrates its capabilities through transient GOF of Meis1, a well-characterized gene implicated in heart development and regeneration17,18,19. In this article, other potential applications of this methodology are also explored, and its advantages and limitations are discussed, as well as compared to existing protocols for transiently modulating gene expression. We believe the framework and visual examples presented will enhance the understanding of epicardium biology during development and disease.
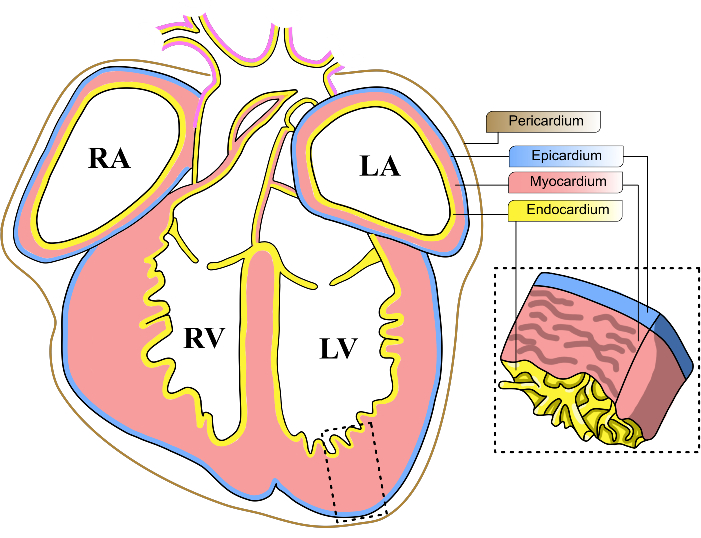
Figure 1: Mouse embryonic heart layers. Schematic diagram of a coronal view of an E13-14 mouse embryonic heart. The three main cellular layers of the heart are represented in yellow (endocardium), red (myocardium), and blue (epicardium). The pericardium is represented in a brown line. The four chambers of the heart are abbreviated as LV, left ventricle; RV, right ventricle; LA, left atrium; RA, right atrium. Please click here to view a larger version of this figure.
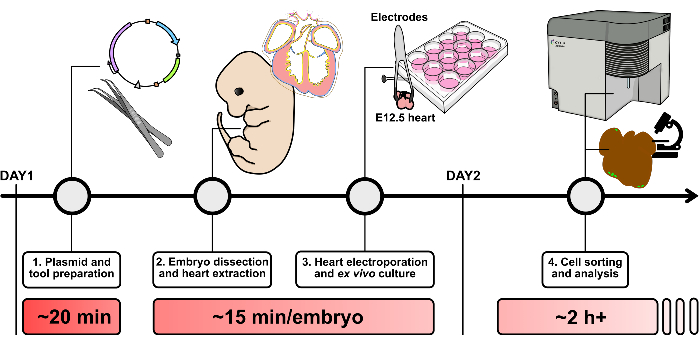
Figure 2: Schematic overview of the heart electroporation protocol. Please click here to view a larger version of this figure.
Protocol
All animal procedures were approved by the CNIC Animal Experimentation Ethics Committee and conformed to current legislation, including EU Directive 2010/63EU and Recommendation 2007/526/EC, as enforced by Spanish Law under Real Decreto 53/2013. For this protocol, female wild-type CD-1 mice aged 15-21 weeks were employed. Details regarding the animals, reagents, and equipment used are listed in the Table of Materials.
1. Plasmid and tool preparation
- First, prepare the electroporation mix by adding the desired plasmid DNA in ice at a final concentration of 1 µg/µL. Then, complete the mix by adding sucrose at 6% final concentration (dissolved in PBS), a biocompatible dye at 0.01%, and complete with PBS until a final volume of 30 µL.
NOTE: Approximately 1-2 µL of the electroporation mix is used per sample. Scale the proportions of the electroporation mix according to the number of samples in the experiment. Please remember that DNA for electroporation should be purified from bacterial culture, assuring optimal stability and minimal contaminants. For more details on the plasmids used for this study, please refer to the representative results section. - Prepare Growth Medium by supplementing Dulbecco's Modified Eagle Medium (DMEM) with 10% fetal bovine serum (FBS), 2% Hepes buffer, and 1% penicillin/streptomycin. Keep the medium in an incubator at 37 °C until use. Additionally, warm 1x PBS at 37 °C, together with the growth medium.
- Prepare the needles by stretching a 20 µm diameter glass capillary using a pipette puller with the following conditions: heat = 550; pull = 100; velocity = 250; time = 120. The final diameter after pulling should be approximately between 5-8 µm.
- Set up the electroporator parameters to five 50 ms cycles at 40 V with intervals of 900 ms. Prepare the electrodes, mouth pipet, and blunt needle (Figure 3A). Equip the pipet with the needle to load the DNA (Figure 3A).
2. Embryo dissection and heart extraction
- Set up a cross between wild-type CD-1 mice. The next morning, select the females that exhibit a vaginal plug and note this time as embryonic day E0.5. Euthanize the animals at embryonic day E12-12.5 by cervical dislocation (following institutionally approved protocols).
- Carefully extract the uterus from the mouse and place it in a dry paper towel. Following this, cut the uterus using fine scissors by inserting the tip of the scissors in the mesometrial side and slowly running the edge of the blade through the whole length. Expose the deciduae and transfer them to the growth medium. For more details, please see the work by Behringer et al.20.
- Dissect the deciduae to extract the embryos, then place the embryos in a 60 mm Petri dish with warm growth medium. Using fine forceps, carefully cut the tail and head of the embryo so that only the trunk of the body remains. The stage of the embryos may be verified by assessing anatomical features, as described by Wong et al.21.
- Then, remove with caution as much tissue from the trunk of the embryo as possible without damaging the heart or the pulmonary and aorta arteries. Once the heart has been exposed, immediately place it in a new, warm growth medium and keep it in an incubator at 37 °C until electroporation.
NOTE: Maintaining the heart's temperature is crucial for its survival. Ensure that the medium is always warm and replace it every 5-10 min. On the other hand, samples may be handled without maintaining sterility, as this will not affect the efficiency of the procedure. Please note that the growth medium contains antibiotics.
3. Heart electroporation and ex vivo culture
- Load approximately 10 µL of plasmid mix (prepared in step 1.1) into the needle by carefully aspirating up with the mouth pipette.
- Prepare a single heart for electroporation by placing it in a clean 60 mm Petri dish with sterile 1x PBS warmed to 37 °C. Place the dish under the dissecting microscope (Figure 3A).
- Prepare the electrodes by adjusting the distance between the positive and negative poles to approximately 4 mm (Figure 3B).
NOTE: 4 mm is the aperture distance in which E12.5 mouse hearts fit in the electrodes used in this work. Please consider changing this distance based on different sample sizes or electrode specifications. - Gently puncture the most superficial layer of the heart with the needle in the mouth pipet. Carefully pipet 1-2 µL of the plasmid mix and remove the needle. Multiple injections may be performed to ensure optimal penetration, as well as to expand the number of electroporated cells.
- Hold the electrodes in place so that the heart is located in between both poles (Figure 3C). Then, electroporate the heart with the conditions specified above.
NOTE: To ensure maximum electroporation efficiency, it is possible to switch the polarity of the electrodes and give another pulse. - Transfer the electroporated heart to a new 12-well plate containing 1 mL of growth medium and place it at 37 °C, 5% CO2.
- Repeat steps 3.1 to 3.6 until all the samples in the experiment are electroporated. Finally, incubate all the hearts in an incubator at 37 °C, 5% CO2, for 24 h or until analysis.
NOTE: In our hands, hearts could be left in culture for up to 48 h without apparent signs of deterioration (Supplementary Movie 1; Supplementary Movie 2; Supplementary Figure 1).
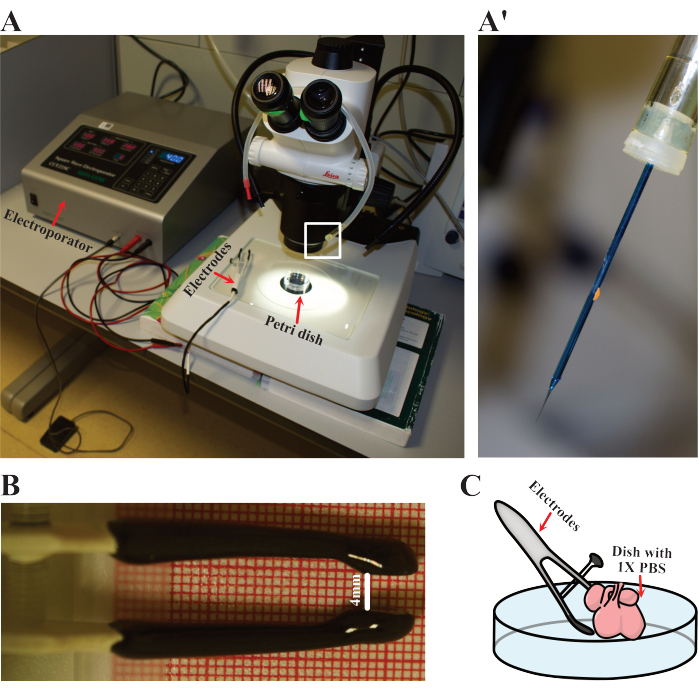
Figure 3: Electroporation setup. (A) Setup used for electroporation. Red arrows indicate the electroporator, electrodes, and Petri dish where hearts are electroporated. (A') Close detail of the electroporation needle as highlighted in (A) by the white rectangle. (B) Detail of the adjusted distance of the electrodes used in E12.5 hearts. (C) Schematic representation of the electroporation. Please click here to view a larger version of this figure.
4. Cell sorting and immunohistochemistry analysis
- For the cell sorting analysis of the samples, prepare a digestion medium to dissociate the tissue by mixing together 400 µL of liberase (5 mg/ml), 300 µL of dispase (0.1 g/mL), 4 µL of CaCl2(0.4 µM), 50 µL of MgCl2(5 µM) and complete until 10 mL of DMEM. Keep on ice until use. Each heart needs approximately 1 mL of Digestion Medium. Adjust the volumes according to the number of samples.
NOTE: For immunohistochemistry, directly fix the hearts in 1.5 mL of 4% PFA for 2 h at RT or overnight at 4 °C. The next day, wash the hearts 3x with PBS and proceed to downstream histological and immunostaining procedures. For a detailed immunohistochemistry protocol, refer to Morris et al.22. - Transfer each heart in culture to a new tube containing 1 mL of ice-cold digestion medium.
- To facilitate dissociation, chop the tissue by gently applying pleasure with a syringe 3-4 times and incubate immediately in a hot plate at 37 °C, 600 rpm for 45 min.
- Filter the digestion mix through a 70 µm cell strainer into a new tube and let it empty by gravity flow. Then, rescue the pass-through volume and filter it again using a 40 µm cell strainer into a new 50 mL tube.
- Add 500 µL of fetal bovine serum (FBS) and complete up to 30 mL using cold DMEM.
- Centrifuge the mix at 240 x g for 10 min at room temperature.
- Discard the supernatant and place the 50 mL tube upside down on a paper towel to dry completely.
NOTE: This is a convenient point to pause the experiment by freezing the cell pellet. If so, resuspend the cells with 10% DMSO in FBS and store at -80 °C. - For cell sorting, resuspend the cells in 300 µL of sorting buffer (1% FBS, 1% penicillin/streptomycin in PBS) and add 0.3 µL of DAPI (1000x). Perform cell sorting.
Results
To demonstrate the effectiveness of this technique in performing gain-of-function (GOF) experiments for relevant heart developmental regulators, a construct was electroporated overexpressing the Meis1 transcription factor. To achieve this, RNA was extracted from E9.5 embryos, and reverse transcription was performed to obtain complementary DNA (cDNA). Using the cDNA as a template, the Meis1 coding sequence was cloned (Supplementary Table 1) into a pCAG expression plasmid (hereafter referr...
Discussion
Overall, the methodology described here offers a robust framework for expressing transgenic constructs in the developing epicardium (Figure 4B), as demonstrated by Meis1 overexpression (Figure 4C). With the appropriate constructs, this protocol can be used to transiently assess the impact of either gain-of-function (GOF) or loss-of-function (LOF) of a specific gene. LOF can be implemented into the technique by transfecting a plasmid targeting a candidate gene th...
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
This study was supported by grant RTI2018-097617-J-I00 from the Spanish Ministerio de Ciencia e Innovación and Acción 9 from Universidad de Jaén to O.H.O. Grant PGC2018-096486-B-I00 from the Spanish Ministerio de Ciencia e Innovación and grant H2020-MSCA-ITN-2016-722427 from the EU Horizon 2020 program to M.T. JMG was supported by a PhD fellowship from the Spanish Ministry of Science and the Fundación Severo Ochoa (PRE2022-101884). Both the CNIC and CBMSO are supported by the Spanish Ministry of Science, and the CNIC is supported by the ProCNIC Foundation.
Materials
| Name | Company | Catalog Number | Comments |
| #55 Forceps | Dumont | 11295-51 | |
| 12-well Clear Flat Bottom Multiwell Cell Culture Plate | BD Falcon | 353043 | |
| 35 mm vise table | Grandado | SKU 8798771617573 | |
| 40 µm Cell Strainer | Fischer Scientific | 08-771-1 | |
| 50 mL tubes | BD Falcon | 352070 | |
| 70 µm Cell Strainer | Corning | CLS431751 | |
| Anti-GFP Policlonal Antibody | Invitrogen | A10262 | 1:1000 dilution used |
| Anti-Myosin 4 (MF20) Monoclonal Antibody | Invitrogen | 14-6503-82 | 1:500 dilution used |
| CD1 Wild Type mice | Provided by Animalary Unit (CNIC) | ||
| Cleaved Caspase-3 (Asp175) Antibody | Cell Signalling Technologies | 9661 | 1:400 dilution used |
| DAPI | Cell Signalling Technologies | 4083 | 1:1000 dilution used |
| Dispase/collagenase | Roche | 10269638001 | |
| Distilled water | |||
| DMEM - Dulbecco's Modified Eagle Medium | Gibco | 10313021 | |
| Fetal Bovine Serum | Invitrogen | 10438-026 | |
| Heracell 150i CO2 Incubator | Thermo Scientific | 51032720 | |
| Leica Stereoscopic Microscope S8AP0 | Leica | 11524102 | |
| Liberase | Roche | 5401119001 | |
| Micropipette Puller Model P-97 | Sutter Instrument | SU-P-97 | |
| pCAG expression plasmid | Addgene | #89689 | |
| Penicillin-streptomycin | Invitrogen | 15070-063 | |
| Petri dishes 35 × 10 mm | BD Falcon | 351008 | |
| Petri dishes 60 × 15 mm | BD Falcon | 353002 | |
| Phenol Red | Merck | P3532 | |
| Pipette tips | Reused from old laboratory equipment | ||
| Rat Serum culture embryo, male rats SPRAGUE DAWLEY RjHan SD | Janvier Labs | 9979 | |
| Recombinant anti-Wilms Tumor Protein 1 (WT1) Antibody | Abcam | ab89901 | 1:300 dilution used |
| Square Wave Electroporator CUY21SC | Nepa Gene | CUY664-10X15 | |
| Sterile PBS | Provided and autoclaved by technical unit | ||
| Sucrose | Millipore | 84100 | |
| Tweezer electrodes with variable gap | Nepa Gene | CUY650P5 |
References
- Tyser, R. C., et al. Calcium handling precedes cardiac differentiation to initiate the first heartbeat. eLife. 5, e17113 (2016).
- Tyser, R. C. V., et al. Characterization of a common progenitor pool of the epicardium and myocardium. Science. 371 (6533), 2986 (2021).
- Sendra, M., Domínguez, J., Torres, M., Ocaña, O. Dissecting the complexity of early heart progenitor cells. J Cardiovasc Dev Dis. 9 (1), 5 (2021).
- Ivanovitch, K., Temiño, S., Torres, M. Live imaging of heart tube development in mouse reveals alternating phases of cardiac differentiation and morphogenesis. eLife. 6, e30668 (2017).
- Ai, D., et al. Canonical Wnt signaling functions in second heart field to promote right ventricular growth. PNAS. 104 (22), 9319-9324 (2007).
- Zimmerman, M. S., et al. regional, and national burden of congenital heart disease, 1990-2017: A systematic analysis for the Global Burden of Disease Study 2017. Lancet Chil Adolesc Heal. 4 (3), 185-200 (2020).
- Xin, M., Olson, E. N., Bassel-Duby, R. Mending broken hearts: Cardiac development as a basis for adult heart regeneration and repair. Nat Rev Mol Cell Biol. 14 (8), 529-541 (2013).
- Porrello, E. R., et al. Transient regenerative potential of the neonatal mouse heart. Science. 331 (6020), 1078-1080 (2011).
- Cao, J., Poss, K. D. The epicardium as a hub for heart regeneration. Nat Rev Cardiol. 15 (10), 631-647 (2018).
- Zhou, B., et al. Adult mouse epicardium modulates myocardial injury by secreting paracrine factors. JCI. 121 (5), 1894-1904 (2011).
- Van Wijk, B., Gunst, Q. D., Moorman, A. F. M., Van Den Hoff, M. J. B. Cardiac regeneration from activated epicardium. PLOS One. 7 (9), e44692 (2012).
- Hesse, J., et al. Single-cell transcriptomics defines heterogeneity of epicardial cells and fibroblasts within the infarcted murine heart. eLife. 10, e65921 (2021).
- Streef, T. J., Smits, A. M. Epicardial contribution to the developing and injured heart: Exploring the Cellular composition of the epicardium. Front Cardiovasc Med. 8, 750243 (2021).
- Sanchez-Fernandez, C., et al. Understanding epicardial cell heterogeneity during cardiogenesis and heart regeneration. J Cardiovasc Dev Dis. 10 (9), 376 (2023).
- Quijada, P., et al. Coordination of endothelial cell positioning and fate specification by the epicardium. Nat Commun. 12 (1), 4155 (2021).
- Mantri, M., et al. Spatiotemporal single-cell RNA sequencing of developing chicken hearts identifies interplay between cellular differentiation and morphogenesis. Nat Commun. 12 (1), 1771 (2021).
- Paul, S., Zhang, X., He, J. Q. Homeobox gene Meis1 modulates cardiovascular regeneration. Semin Cell Dev Biol. 100, 52-61 (2020).
- Stankunas, K., et al. Pbx/Meis deficiencies demonstrate multigenetic origins of congenital heart disease. Circ Res. 103 (7), 702-709 (2008).
- Liu, Y., et al. Transcription factor Meis1 act as a new regulator of ischemic arrhythmias in mice. J Adv Res. 39, 275-289 (2022).
- Behringer, R. . Manipulating the mouse embryo: A laboratory manual. , (2014).
- Wong, M. D., et al. 4D atlas of the mouse embryo for precise morphological staging. Development. 142 (20), 3583-3591 (2015).
- Morris, L., Klanke, C., Lang, S., Lim, F. Y., Crombleholme, T. TdTomato and EGFP identification in histological sections: Insight and alternatives. Biotech Histochem. 85 (6), 379-387 (2010).
- Schiaffino, S., Rossi, A. C., Smerdu, V., Leinwand, L. A., Reggiani, C. Developmental myosins: expression patterns and functional significance. Skelet. Muscle. 5 (1), 22 (2015).
- Eissa, N., et al. Stability of reference genes for messenger RNA quantification by real-time pcr in mouse dextran sodium sulfate experimental colitis. PLOS One. 11 (5), e0156289 (2016).
- Livak, K. J., Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 25 (4), 402-408 (2001).
- Lai, S. R., Andrews, L. G., Tollefsbol, T. O. RNA interference using a plasmid construct expressing short-hairpin RNA. Methods Mol Biol. 405, 31-37 (2007).
- Carmona, R., Barrena, S., López Gambero, A. J., Rojas, A., Muñoz-Chápuli, R. Epicardial cell lineages and the origin of the coronary endothelium. FASEB J. 34 (4), 5223-5239 (2020).
- Gittenberger-de Groot, A. C., Vrancken Peeters, M. P. F. M., Mentink, M. M. T., Gourdie, R. G., Poelmann, R. E. Epicardium-derived cells contribute a novel population to the myocardial wall and the atrioventricular cushions. Circ Res. 82 (10), 1043-1052 (1998).
- Chong, Z. X., Yeap, S. K., Ho, W. Y. Transfection types, methods and strategies: A technical review. Peer J. 9, e11165 (2021).
- Kałużna, E., Nadel, A., Zimna, A., Rozwadowska, N., Kolanowski, T. Modeling the human heart ex vivo-Current possibilities and strive for future applications. JTERM. 16 (10), 853-874 (2022).
- Aguilera-Castrejon, A., et al. Ex utero mouse embryogenesis from pre-gastrulation to late organogenesis. Nature. 593 (7857), 119-124 (2021).
- Dyer, L. A., Patterson, C. A novel ex vivo culture method for the embryonic mouse heart. J Vis Exp. (75), e50359 (2013).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved