Quantification of Mouse Heart Left Ventricular Function, Myocardial Strain, and Hemodynamic Forces by Cardiovascular Magnetic Resonance Imaging
In This Article
Summary
This study describes a comprehensive cardiovascular magnetic resonance imaging (CMR) protocol to quantify the left ventricular functional parameters of the mouse heart. The protocol describes the acquisition, post-processing, and analysis of the CMR images as well as assessment of different cardiac functional parameters.
Abstract
Mouse models have contributed significantly to understanding genetic and physiological factors involved in healthy cardiac function, how perturbations result in pathology, and how myocardial diseases may be treated. Cardiovascular magnetic resonance imaging (CMR) has become an indispensable tool for a comprehensive in vivo assessment of cardiac anatomy and function. This protocol shows detailed measurements of mouse heart left ventricular function, myocardial strain, and hemodynamic forces using 7-Tesla CMR. First, animal preparation and positioning in the scanner are demonstrated. Survey scans are performed for planning imaging slices in various short- and long-axis views. A series of prospective ECG-triggered short-axis (SA) movies (or CINE images) are acquired covering the heart from apex to base, capturing end-systolic and end-diastolic phases. Subsequently, single-slice, retrospectively gated CINE images are acquired in a midventricular SA view, and in 2-, 3-, and 4-chamber views, to be reconstructed into high-temporal resolution CINE images using custom-built and open-source software. CINE images are subsequently analyzed using dedicated CMR image analysis software.
Delineating endomyocardial and epicardial borders in SA end-systolic and end-diastolic CINE images allows for the calculation of end-systolic and end-diastolic volumes, ejection fraction, and cardiac output. The midventricular SA CINE images are delineated for all cardiac time frames to extract a detailed volume-time curve. Its time derivative allows for the calculation of the diastolic function as the ratio of the early filling and atrial contraction waves. Finally, left ventricular endocardial walls in the 2-, 3-, and 4-chamber views are delineated using feature-tracking, from which longitudinal myocardial strain parameters and left ventricular hemodynamic forces are calculated. In conclusion, this protocol provides detailed in vivo quantification of the mouse cardiac parameters, which can be used to study temporal alterations in cardiac function in various mouse models of heart disease.
Introduction
Cardiovascular magnetic resonance (CMR) in small animals provides an accurate in vivo measurement of myocardial function, making CMR an optimal tool for preclinical research in cardiovascular disease. Due to the high spatial resolution and high contrast between blood and myocardium in CMR images, it is possible to delineate the endo- and epicardial contours and calculate myocardial mass and ventricular volumes1,2. Despite the high heart rates of up to 600 beats/min, the use of electrocardiogram (ECG) and respiratory triggering allows high-quality measurements of different cardiac phases (also called CINE images) without respiratory motion artifacts. In this way, multiple slices can be used to cover the heart from apex to base to extract systolic function parameters such as ejection fraction (EF), end-systolic volume (ESV), end-diastolic volume (EDV), and cardiac output (CO)3. Aside from basic systolic function assessment, additional CMR techniques have been recently developed to assess diastolic dysfunction4, myocardial strain5, and hemodynamic forces (HDF)6.
ECG gating allows synchronization to the cardiac cycle by starting MR signal acquisition after detection of the R-peak and recording a defined number of cardiac phases during the R-R interval. However, the number of cardiac phases (frame rate) that can be acquired this way depends on the lowest possible repetition time (TR) the system can reach while maintaining an acceptable signal-to-noise ratio (SNR) and spatial resolution4. Furthermore, because the use of high magnetic field gradients can temporarily distort the ECG signal, the acquisition is usually stopped before the end-diastolic phase. Both factors limit the use of such scans to systolic function assessments, as the calculation of other cardiac functional parameters requires a better definition of the left ventricular (LV) volume-time curve.
High-frame-rate CINE images can be acquired by retrospective gating, whereby MR signal is continuously acquired during scanning, and an incorporated navigator echo after radiofrequency (RF) excitation detects cardiac and respiratory motion. Because the CMR acquisition is performed asynchronously with the cardiac motion, the acquired MR signals can then be assigned to a retrospectively chosen number of cardiac frames. In this way, if sufficient data is collected, high-frame-rate CINE images can be reconstructed4,7. This then allows for diastolic function assessment, represented by the ratio between the peak early filling rate (E') and peak late filling rate from atrial contraction (A').
In clinical research, CINE images can be analyzed with CMR feature-tracking to assess the myocardial strain and HDF6,8. Myocardial strain is a cardiac deformation parameter that measures the difference in percentages between the initial length (usually in end-diastolic length) and maximum length (usually in end-systole) of a myocardial segment9. Myocardial strain measurements can be of incremental value for assessing the LV function as strain values quantify myocardial wall shortening and thickening. A reduction in shortening function might be an indication of subendocardial fiber damage10. Alterations in myocardial strain can occur independent of EF and could be a precursor for underlying complications.
Specifically, the global longitudinal strain (GLS) and global circumferential strain (GCS) have been shown to be of added value in characterizing heart disease10,11,12. Similarly, HDF has been suggested to be a potential novel parameter to indicate altered cardiac function6,13. These HDF or interventricular pressure gradients (IVPG) drive the blood motion during ejection and filling of the heart and are affected by the momentum exchange between blood and myocardium, including the aortic and mitral valve14,15.
In this study, a comprehensive protocol is described for performing robust small animal CMR measurements to quantify LV function, myocardial strain, and HDF of mouse hearts. It contains the necessary steps for animal preparation, data acquisition using both prospectively and retrospectively gated CINE images of the heart, as well as analysis with dedicated software capable of calculating the volumetric measurements, E'/A' ratio, myocardial strain, and HDF of the heart. This protocol can be used for the extensive assessment of LV function in various mouse models of cardiovascular disease.
Protocol
The described animal experiments are conducted in accordance with European Union guidelines for the welfare of laboratory animals (Directive 2010/63/EU) and were approved by the Academic Medical Center Animal Ethics Committee.
1. Setup and animal preparation
- Prior to starting the experiment, make sure there is sufficient isoflurane anesthesia for at least 2 h, and that the battery available for ECG and respiratory monitoring is sufficiently charged. Make sure the scanner area is equipped with a working fume extraction tube to remove excess isoflurane.
- Prepare the mouse cradle (Figure 1A), and turn on the animal heating system with the temperature set to 40 °C. Prepare the ECG/respiratory interface module and battery setup (Figure 1B), and start the software for real-time monitoring of ECG and respiratory signals (Figure 1C).
- Remove the mouse from its housing cage and measure body weight.
- Place the mouse in an anesthesia induction chamber under a fume hood extraction arm, and provide 3-4% isoflurane in a mixture of 0.2 L/min O2 and 0.2 L/min medical air. After the animal is fully anesthetized, apply a small drop of eye ointment to each eye, and close the eyelids of the mouse.
- Place the mouse in the supine position on the mouse cradle. Hook the incisors of the mouse in the bite bar on the mouse cradle, and adjust the nose cone to fit properly (Figure 1A). Visually check if the breathing is stable below 100 breaths/min, and reduce the isoflurane to ~2% during animal preparation.
- Move the mouse cradle such that the heart is located in the part of the cradle holder that will end up in the center of the RF coil and iso-center of the magnet.
- Use petroleum jelly to insert the rectal temperature probe, and tape the fiber-optic cable of the temperature probe to the mouse cradle.
- Place the respiratory balloon on the lower abdomen of the mouse and secure it with tape. Insert two ECG electrode needles subcutaneously in the thorax at the height of the forepaws and gently tape them down to prevent movement (Figure 1A).
- Check whether respiration and ECG signals are of sufficient quality, and whether correct trigger points are detected by the software (Figure 1C).
- Make sure the respiratory rate is 50-80 breaths/min, heart rate ~400-600 beats/min, and body temperature around 37 °C. Adjust the isoflurane administration when the respiratory rate is outside of this range, and reduce the temperature of the animal heating system if the body temperature tends to exceed 37 °C.
- Place the RF coil over the mouse.
NOTE: Depending on the system, this might require temporary disconnection of the ECG electrodes and respiratory balloon plugs from the ECG/respiratory interface module. - Connect the coil cables, and place the cradle in the magnet bore. Check if the ECG signal is still stable.
- If the ECG signal is suboptimal, reposition the ECG electrodes for a better signal, as this cannot be done at a later stage without significantly changing the orientation of the animal.
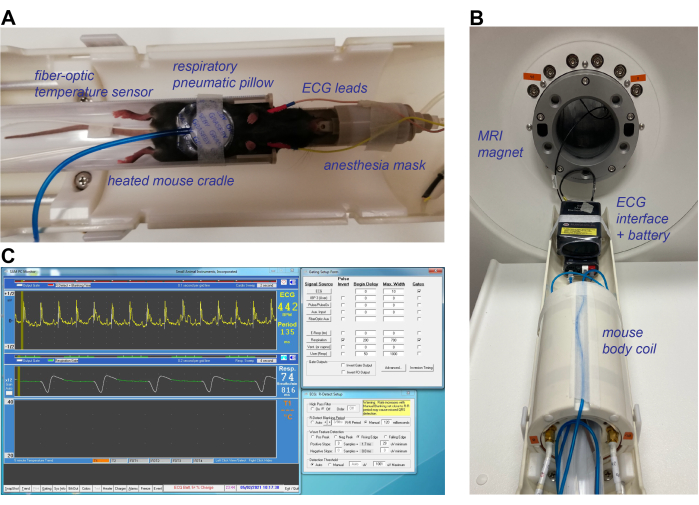
Figure 1: Animal preparation and equipment setup for CMR imaging of the mouse heart. (A) Fully anesthetized mouse in the supine position, placed in the heated mouse cradle with a respiratory pneumatic pillow placed on the abdomen, rectal fiber-optic temperature sensor, and subcutaneous ECG leads in the chest near the forepaws. (B) Mouse body coil placed over the mouse cradle, with ECG leads and respiratory pillow reconnected to the ECG and respiratory interface prior to placing the holder in the MRI magnet. (C) Depiction of the ECG and respiratory signals in dedicated small animal monitoring software. The R-peak of the ECG signal is detected and used as starting point for MRI signal acquisition. A blanking period between R-peaks can be manually adjusted based on the period of a heartbeat. Triggering can only occur during the respiratory plateau (green line in middle panel) for which the beginning delay and maximum width can be manually adjusted. Abbreviations: CMR = cardiovascular magnetic resonance imaging; ECG = electrocardiogram; MRI = magnetic resonance imaging. Please click here to view a larger version of this figure.
2. MRI scan calibration and triggering
- Adjust the ECG and respiratory gating parameters within the signal monitoring software such that trigger points are generated at the R-peaks and only during the flat portion of the respiratory signal. To minimize ECG gating errors, set a blanking period that is 10-15 ms shorter than the R-R interval.
NOTE: This blanking period should be adjusted during the entire experiment if changes in heart rate occur. - Perform a center frequency calibration and a standard (ungated) SCOUT scan with zero offset to determine the position of the mouse in the scanner in coronal, axial, and sagittal directions. If the heart is not positioned within 0.5-1 cm of the field-of-view (FOV) center, adjust the position of the cradle accordingly, and redo the SCOUT scan.
- Perform a manual shim and RF calibration using available vendor methods.
3. Scan planning and acquisition
NOTE: See Table 1 for detailed scan parameters of the following scans.
- Based on the initial SCOUT, perform a gated single-frame Gradient Echo (GRE) scout scan (Table 1, scan 1) with 5 slices in 3 orthogonal directions, and position each stack of slices on the approximate location of the heart to locate the exact position of the heart (Figure 2A).
- Perform a gated single-frame multi-slice SA scout scan (Table 1, scan 2).To this end, use the previous GRE scout to position 4-5 slices in a mid-left ventricular position, perpendicular to the long axis of the heart to find an initial estimate of the midventricular SA view, which is needed to plan the long-axis 2-chamber scout (Figure 2B).
- For the following prospective scans (steps 3.4-3.6), adjust the number of cardiac frames (Nframes) such that Nframes × TR is ~60-70% of the R-R interval.
NOTE: Acquisition for 60-70% of the R-R interval is enough to capture the end-diastolic phase of the cardiac cycle, while allowing additional T1 relaxation during end-diastole for improved SNR and preventing disturbance of the following R-peak by gradient switching. - Perform a gated single-slice GRE scan to generate the long-axis 2-chamber (2CH) scout, which combined with the SA scan, is needed to plan the 4-chamber (4CH) (Table 1, scan 3). To this end, position a slice perpendicular to the previous SA views running parallel to the connection points between the left and right ventricle. Move this slice to the middle of the left ventricle, and check in the coronal image of the GRE scout if the slice is aligned with the LV long axis such that it is placed through the apex (Figure 2C).
- Perform another gated single-slice GRE scan to generate the 4-chamber (4CH) scout scan, which is needed to plan the multi-slice SA and the 3-chamber scan (Table 1, scan 4). To this end, position a slice perpendicular to the 2CH scout scan, and align to the center of the long axis such that the slice goes through the mitral valve and the apex. In the SA views, adjust the slice such that it is placed parallel to the posterior and anterior ventricular wall and between the two papillary muscles (Figure 2D). Check if the slice remains in the center of the ventricle throughout the entire cardiac cycle.
- Perform a gated sequential multi-slice SA GRE scan (Table 1, scan 5) for systolic function measurements. To this end, position a midventricular slice perpendicular to the LV long axis in the 2CH and 4CH views in the center of the heart, and increase the number of slices (typically an odd number, e.g., 7 or 9 slices, no gap between the slices) to cover the heart from base to apex (Figure 2E).
- For the following retrospectively gated scans (steps 3.8-3.9), turn off all prospective cardiac and respiratory gating functionality. Make note of the cardiac and respiratory rate before and after each retrospectively gated scan, and use these values for reconstruction purposes later (step 5.2.2).
- Perform three sequential single-slice retrospectively gated GRE scans in the midventricular SA view (for quantification of the E'/A' ratio), 2CH, and 4CH view, the latter two which are necessary for the quantification of the myocardial strain and HDF values (Table 1, scan 6-8). If needed, optimize the final 2CH and 4CH slice orientations based on the multi-slice SA views as well as the available 2CH and 4CH scout scans.
- Perform an additional retrospectively gated single-slice GRE scan in a 3-chamber (3CH) view, which combined with the 2CH and 4CH view from step 3.8 is necessary for the quantification of the myocardial strain and HDF values (Table 1, scan 9). To this end, position a slice perpendicular to the midventricular SA view similar to the position of the final long-axis 4CH view, and turn the slice 45° to pass from the anterior wall to the papillary muscle closest to the posterior wall. Inspect the basal SA slice to see if the slice passes through the mitral and aortic valve. Inspect in the final long-axis 4CH view if the slice is going through the apex (Figure 2F).
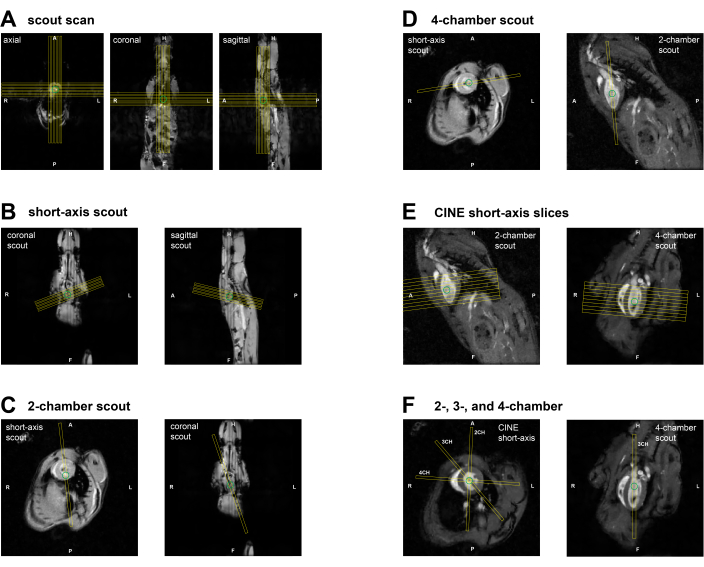
Figure 2: Slice planning for CMR imaging in a mouse. (A) GRE SCOUT planning through the heart in 3 orthogonal views using initial scout scan. (B) Short-axis scout planning on the GRE SCOUT coronal and sagittal slices. (C) Planning of 2CH scout view using the short-axis scout and the GRE SCOUT coronal slice. (D) Planning of 4CH scout view using the short-axis scout and the 2CH scout. (E) Planning of the multi-slice short-axis view using 2CH and 4CH scouts. (F) (left) Planning of final 2CH, 3CH, and 4CH views using the midventricular short-axis and 2CH/4CH scout views. Abbreviations: CMR = cardiovascular magnetic resonance imaging; GRE = Gradient Echo; CH = chamber. Please click here to view a larger version of this figure.
| Scan number(s) | 1 | 2 | 3 | 4 | 5 | 6-9 |
| Scan name(s) | GRE scout | multi-slice SA scout | 2CH scout | 4CH scout | multi-slice SA | SA, 2CH, 4CH, 3CH |
| total slices | 15 (3 x 5)* | 4-5 | 1 | 1 | 7-9 | 1 |
| Thickness (mm) | 1 | 1 | 1 | 1 | 1 | 1 |
| FOV (mm) | 60 | 35 | 30 | 30 | 35 | 30 |
| FOV Ratio | 1 | 1 | 1 | 1 | 1 | 1 |
| Flip Angle | 40 | 20 | 20 | 20 | 20 | 15 |
| TE (ms)** | 3.8 | 3.4 | 2.5 | 2.5 | 2.5 | 3.6 |
| TR (ms) | 200 | 1 R-R | 7 | 7 | 7 | 8 |
| Nframes | 1 | 1 | 12-14 | 12-14 | 12-14 | 32 *** |
| Matrix Size | 192 x 192 | 192 x 192 | 192 x 192 | 192 x 192 | 192 x 192 | 192 x 192 |
| ECG triggering | no | yes | yes | yes | yes | retrospective |
| Respiratory triggering | yes | yes | yes | yes | yes | retrospective |
| Averages | 1 | 3 | 5 | 5 | 5 | retrospective **** |
| Total imaging time (estimated *****) | 2 min | 2min | 3-4 min | 3-4 min | 20-25 min | 13 min / scan |
Table 1: Acquisition parameters for each sequence used during the CMR protocol. * Scans are performed in three different orthogonal orientations (axial, coronal, sagittal). **The shortest possible TE, given all other parameters are used, which depends on the specific scanner configuration. ***This is the number of cardiac frames after retrospective binning. **** The effective averaging depends on the random k-space filling during the total acquisition time. In total, 400 repetitions of all k-lines were performed. ***** Including ECG/respiratory triggering delays. Abbreviations: CMR = cardiovascular magnetic resonance imaging; ECG = electrocardiogram; GRE = gradient echo; FOV = field-of-view; TE = echo time; TR = repetition time; Nframes = number of cardiac frames; SA = short axis; CH = chamber. Please click here to download this Table.
4. Finalization of the experiment and data storage
- Remove the mouse from the cradle after detaching all other measurement equipment, and turn off the anesthesia. In the case of longitudinal experiments, place the mouse in a pre-warmed housing cage at 37 °C for recovery until the animal is awake and active.
- Clean all equipment that has been used with cleaning wipes or 70% alcohol.
- Generate Digital Imaging and Communication in Medicine (DICOM) files for the prospectively gated MRI data, and copy these together with the MRI raw data files of the retrospectively gated scans to a secure server for subsequent data analysis.
5. Offline reconstruction of the retrospectively acquired scans
NOTE: For the reconstruction of the retrospectively gated scans, a custom-built open-source software was used (Figure 3). Perform the following steps for each of the retrospectively triggered data separately.
- Open the reconstruction software Retrospective, and load the raw data file corresponding to a retrospectively gated MRI scan.
- Inspect the Raw navigator signal, and note that the higher signal peaks represent the respiratory frequency and the lower signal peaks represent the heart rate.
- If the peaks are registered upside down, flip the signal with the up/down switch.
- Additionally, check whether the automatically detected heart rate corresponds to 10% of the observed values during each scan. If not, manually adjust these values because automated detection failed.
- Choose an appropriate window percentage for exclusion of data during respiratory movement, usually 30%.
- Press Filter to perform the navigator analysis, and separate the heart navigator from the respiratory navigator.
- Set the number of CINE frames to 32 (value used in this study), and press sort k-space.
- Choose appropriate settings for compressed sensing (CS) regularization, and press reconstruct. Use the following typical regularization parameters: wavelet regularization parameter in the spatial (x, y, and z) dimensions (WVxyz) 0.001 or 0; total variation constraint in the CINE dimension (TVcine) 0.1; total variation constraint in spatial dimension (TVxyz) 0; and total variation constraint in the dynamics dimension (TVdyn) 0.05.
- Once the reconstruction is finished, preview the CINE movie to evaluate the reconstruction. Export DICOM images for further analysis with Export DCM.
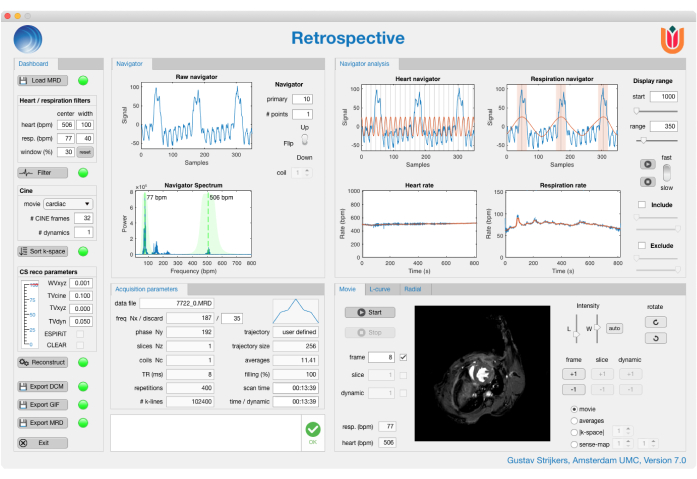
Figure 3: 'Retrospective' triggering graphical user interface. 'Retrospective' is a custom-built reconstruction application for retrospectively triggered cardiac magnetic resonance imaging scans. In the user interface, it is possible to evaluate the navigator signal, adjust the number of CINE frames to be reconstructed, adjust the compressed sensing parameters to improve reconstruction, preview the CINE images as a dynamic movie, and export the reconstructed data. Please click here to view a larger version of this figure.
6. Image analysis software
NOTE: The image analysis software (Figure 4) requires the use of DICOM images and has multiple plugins for different cardiovascular analysis applications, such as the plugin for volumetric measurements and the plugin for strain and HDF analysis.
- For volumetric assessment of the LV, select the multi-slice SA scan, and load it into the plugin for volumetric measurements.
- Assign end-systolic (ES) and end-diastolic (ED) labels to the corresponding cardiac frame.
- Use the contour tools to segment the endomyocardial borders in the ES and ED frames.
NOTE: The analysis software, used for this protocol, automatically displays the LV EF, EDV, ESV parameters when all necessary annotations have been made.
- For diastolic measurements, select the midventricular SA CINE images, and load these into the plugin for volumetric measurements.
- Assign the ED and ES labels to the corresponding cardiac frames.
- Use the contour tools to segment the endocardial border for all frames. Compare the segmentation of neighboring frames to ensure smooth transitions of the segmentation throughout the cardiac cycle.
- Export the time evolution from all cardiac frames and corresponding LV endomyocardial volumes (LV ENDO). Apply a custom-built script (see Supplemental Material) to calculate the E'/A' ratio.
NOTE: The script applies a Savitzky-Golay filter for robust calculation of the dV/dt curves and uses semi-automatic peak detection to find the E' and A' peaks.
- For strain and HDF calculations, select the 2CH, 3CH, and 4CH long-axis CINE images, and load them into the plugin for volumetric measurements.
- Assign the ED and ES labels to the corresponding cardiac frame in each slice orientation.
- Use the contour tools to segment the endocardial border for all frames in all 3 orientations. Compare the segmentation of neighboring frames to ensure smooth transitions of the segmentation throughout the cardiac cycle.
- Once the contours are drawn in the plugin for volumetric measurements, run the plugin for the strain and HDF analysis.
- Assign each of the acquired datasets to the corresponding labels for 2CH, 3CH, and 4CH views, and execute the strain analysis.
- For HDF analysis, draw the diameter of the mitral valve at the end-diastolic frame in all 3 orientations, and draw the diameter of the aorta in the 3-chamber long-axis image.
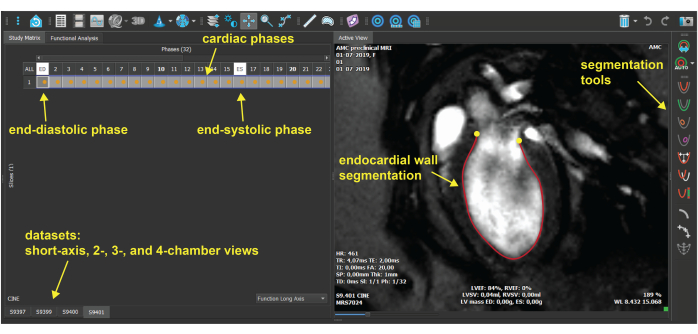
Figure 4: Image analysis software graphical user interface. The plugin for volumetric measurement in the image analysis software, which is used for contouring of the endomyocardial border. For each dataset, the end-diastolic and end-systolic cardiac phases are selected, and the endomyocardial border is segmented for all frames. Please click here to view a larger version of this figure.
Representative Results
Using the previously described protocol, a group of healthy C57BL/6 wildtype mice (n = 6, age 14 weeks) was scanned using a 7-Tesla MRI scanner using a 38-mm-diameter birdcage coil. During each scan session, the multi-slice CINE SA images were acquired using prospectively gated GRE sequences, while the single-slice midventricular SA, 2CH, 3CH, and 4CH views CINE images were acquired using retrospective gating. Representative high-frame rate reconstructions of retrospectively gated scans using a custom-built, post-processing software can be seen in Supplemental Video 1. From the resulting images, volume-time curves during the cardiac cycle (Figure 5A) were determined as well as the corresponding first-derivative curves (dV/dt) for calculation of systolic (EF = 72.4 ± 2.8%) and diastolic function parameters (E'/A' ratio = 1.5 ± 0.3) respectively.
The 2CH, 3CH, and 4CH view CINE images were analyzed using image analysis software to determine endocardial GLS (endoGLS) changes across the cardiac cycle (Figure 5B) and corresponding peak GLS values (-22.8 ± 2.4%) as a measure for myocardial strain. Additionally, the software calculates the root mean square (RMS) HDF in longitudinal (apex-base) (135.2 ± 31.7%) and transversal (inferolateral-anteroseptal) (12.9 ± 5.0%) directions. For each animal, it is also possible to produce an HDF time profile, which follows a consistent pattern of positive and negative peaks that represent the magnitude and direction of the HDF during the cardiac cycle (Figure 5C). Descriptive results of all outcome parameters are summarized in Figure 5D.
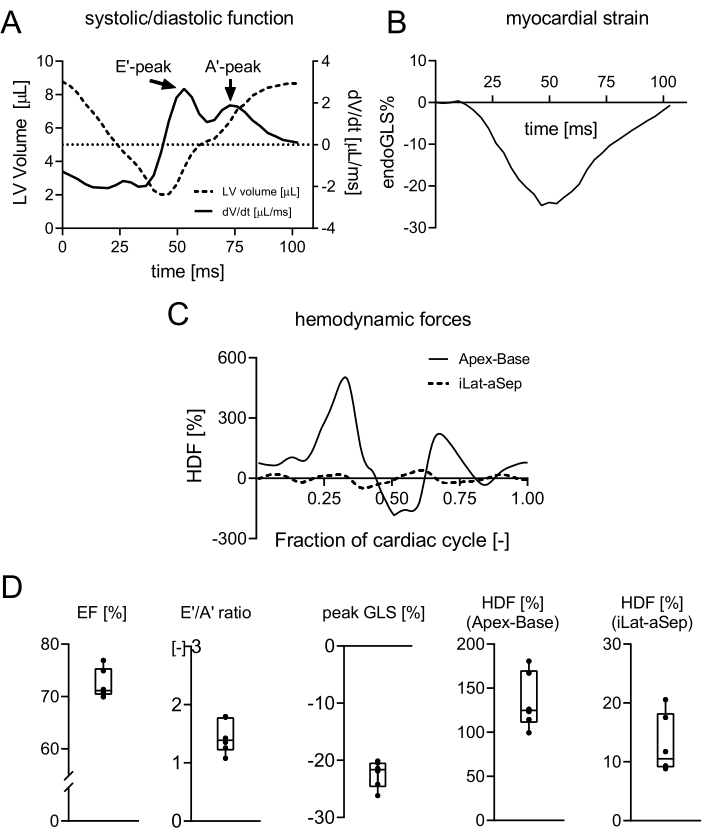
Figure 5: Quantification of LV functional parameters based on the mouse heart. (A) Representative volume-time curve and corresponding dV/dt curve. The latter depicts the flow velocity with distinct early filling peak (E') and atrial contraction (A') peak. (B) Representative GLS curve indicating strain deformation in the longitudinal direction throughout the cardiac cycle. (C) Representative HDF curve with distinct force peaks in the apex-base direction, starting with the systolic ejection force and followed by a downward force at the transition between systole and diastole, E-wave deceleration force, A-wave acceleration, and deceleration force. (D) Descriptive results of all animals for values of EF, E'/A' ratio, peak GLS, and root mean square of the HDF in apex-base and inferolateral-anteroseptal directions. Values are expressed as mean ± SD. Abbreviations: LV = left ventricle; V = volume; t = time; GLS = global longitudinal strain; HDF = hemodynamic forces; EF = ejection fraction. Please click here to view a larger version of this figure.
Supplemental Video 1: Representative reconstruction of retrospectively gated CINE images in SA, 2CH, 3CH, and 4CH views. Abbreviations: SA = short axis; CH = chamber. Please click here to download this Video.
Supplemental Material: Please click here to download this File.
Discussion
The presented protocol describes the use of CMR imaging for longitudinal, non-invasive, in vivo experiments to analyze heart function in mice. These results are examples of healthy animals to demonstrate the feasibility of using CINE images to quantify the cardiac parameters. However, the methods described can be used for various animal models. Although specific disease models might require small alterations to the protocol, its basic structure to assess the different cardiac functional parameters will be very similar. One particular case worth mentioning is a myocardial infarction model where part of the heart has significant loss in contractility. This can cause low quality of the cardiac navigator signal within this slice. In this case, an alternative option would be acquiring the navigator from a separate slice, as described in a previous study by Coolen et al.16. CINE images in different views are reconstructed from retrospectively gated data using CS algorithms and are analyzed using image analysis software to calculate the strain and HDF values.
The quality of the acquired images naturally depends on all preparation steps, which need to be carefully performed before starting the cardiac MRI protocol. For instance, if no clear ECG and respiratory signals are seen when placing the animal inside the MRI scanner, this will likely result in suboptimal acquisitions and even increased scan times due to the added effect of magnetohydrodynamic distortions17. It is important to realize that due to the sequential planning of the slice orientations, the animals cannot just be repositioned in between scans. It is therefore not possible to re-adjust the ECG leads in between scans, as this will alter the position of the mouse in the scanner. During scanning, temperature control is crucial for maintaining a constant cardiac and respiratory interval, which especially benefits the quality of the retrospectively gated scans that are acquired over a longer period of time. During this high-duty-cycle scan, the temperature of the animal might steadily increase, causing the heart rate and respiratory rate to increase. Adjusting the temperature of the heating system and the anesthesia could greatly contribute to stabilizing the respiratory rate prior to or during scanning.
A critical step during the analysis is the consistency in contour drawing. While automatic segmentation works well for clinical data, it does not perform robustly in the case of mouse cardiac data (not tested for rats). The high heart rate and high blood flow during specific cardiac phases, especially at the start of LV filling, may cause intravoxel dephasing and signal voids, compromising myocardial wall delineation. It is therefore not advised to analyze each frame independently, but visually inspect the motion of the myocardial wall between frames and take this into account when drawing the contours across all frames. It is advised to copy and adjust the endocardial contour between two consecutive frames to maintain a more natural contractile motion in the analysis. In this protocol, papillary muscles are excluded from the ventricular lumen volume in the SA images for systolic and diastolic function assessment, while they are included in the 2CH, 3CH, and 4CH views for strain and HDF analysis because the latter relies on knowledge of the precise motion of the myocardial wall, rather than the precise volume of the ventricular lumen.
Whereas systolic and diastolic function parameters are based on measuring LV volumes throughout the cardiac cycle, strain and HDF parameters depend on motion patterns within the myocardial wall as well. For this, feature-tracking techniques are used where the displacement of the myocardial segment can be assessed by recognizing distinct anatomical features and signal intensities between subsequent CINE phases. The strong contrast between blood pool and myocardium in CMR images facilitates the use of feature-tracking for subsequent strain and HDF analysis8. Prior to CMR feature-tracking, the myocardial strain was determined with speckle tracking echography and CMR tissue-tagging. CMR feature-tracking does not require additional scanning time compared to CMR tissue-tagging. However, despite the use of retrospective triggering, CMR still has a limited temporal resolution, which could make it difficult to correctly evaluate fast deformations within the cardiac cycle.
Assessment of HDF throughout the cardiac cycle requires measurements of the diameters of the mitral and aortic valves to calculate the HDF in apex-base and inferolateral-anteroseptal directions using previously described equations18. This method has shown consistent estimates of the HDF compared to the reference standard 4D-flow MRI, which has a limited availability in clinical use due to its complexity6. It is important to know that robust estimation of the valve diameters is difficult, and therefore, the valve diameters should be kept constant for a group of animals and across repeated measurements in a longitudinal study, as variations in this parameter by incorrect estimations could easily overshadow subtle changes in HDF parameters. The specific software used to calculate GLS and HDF parameters might not be available to all users. Therefore, one can refer to Voigt et al.19 (GLS) as well as Pedrizzetti et al.6,20 (HDF), which contain all mathematical descriptions that form the basis of the respective calculations as performed by the analysis software.
For the purpose of this study, the protocol was evaluated in healthy animals (N = 6). A representative set of time curves for LV volume, dV/dt, endoGLS, and HDF are shown in Figure 5A-C. Mean values of multiple cardiac functional parameters (EF, E'/A'-ratio, peak GLS, and HDF) are shown in Figure 5D. These agree well with comparable protocols used in the literature21. Literature on GLS and HDF data in mice is scarce. A mean GLS value of -22.8% was measured, which is in the same range as clinical data8, indicating that GLS measurements obtained with the described method are feasible in mice. HDF curves obtained in mice also show the same distinct phases as seen in human data, showing the successful translation of this technique to preclinical research. While HDF parameters are hypothesized to serve as early biomarkers of cardiac dysfunction, more studies are warranted to investigate the diagnostic and predictive value of this new parameter. The results in this protocol do show that HDF and GLS outcomes are expected to be more variable across animals, which needs to be taken into account when subtle differences in animal models or treatment effects are expected.
Disclosures
Ruslan Garipov is an employee of MR Solutions Ltd., Guildford, United Kingdom. David Hautemann is an employee of Medis medical imaging systems B.V., Leiden, The Netherlands.
Acknowledgements
The authors thank Dorita Dekkers and Fatimah Al Darwish for assistance with the mouse measurements and data analysis.
Materials
| Name | Company | Catalog Number | Comments |
| Equipment | |||
| AccuSens single and multi-channel signal conditioner | Opsens solutions inc., Canada | ACS-P4-N-62SC | Used with fiber optic temperature sensor to monitor body temperature |
| Duratears eye ointment | Alcon Nederland B.V., Netherlands | ||
| Mouse cell | Équipment Vétérinaire Minerve, France | referred to as mouse cradle | |
| MR-compatible Monitoring & Gating System for Small animals | SA Intuments, Inc., United States | Model 1030 | ERT Module (ECG/respiratory interface module) , ERT Control/Gating Module, battery pack and subdermal ECG Electrode Set |
| MRI scanner | MR Solutions Ltd., United Kingdom | Model: MRS-7024 | Preclinical MRI System 7.0T/24 cm |
| Multistation temperature control unit and High Flow PCA | Équipment Vétérinaire Minerve, France | Model: URT Multipostes | animal heating system |
| Respiration Sensor | Graseby Medical Limited, United Kingdom | Ref 2005100 | |
| RF coil | MR Solutions Ltd., United Kingdom | MRS-MVC | 38mm mouse volume RF coil for mouse body studies |
| SF flowmeter | flow-meter, Italy | SF 3 | |
| Vaporizer sigma delta Intermed | Penlon Ltd., United Kingdom | ||
| Materials | |||
| Isoflurane | AST farma, Netherlands | ||
| Vaseline petroleum jelly | Unilever, United Kingdom | ||
| Software | |||
| BART toolbox | https://mrirecon.github.io/bart/ | ||
| Mathematica 12.0 | Wolfram Research, Inc., United States | ||
| MATLAB 2019a | The MathWorks,Inc., United States | ||
| MEDIS Suite MR | Medis Medical Imaging Systems B.V. ,Netherlands | Image analysis software | |
| PC-SAM | SA Intuments, Inc., United States | ||
| Preclinical Scan | MR Solutions Ltd., United Kingdom | Scanning software | |
| Retrospective version 7.0 | Amsterdam UMC, the Netherlands | Reconstuction software: https://github.com/Moby1971?tab=repositories |
References
- Vallée, J. P., Ivancevic, M. K., Nguyen, D., Morel, D. R., Jaconi, M. Current status of cardiac MRI in small animals. Magnetic Resononance Materials in Physics, Biology and Medicine. 17 (3-6), 149-156 (2004).
- Bakermans, A. J., et al. Small animal cardiovascular MR imaging and spectroscopy. Progress in Nuclear Magnetic Resonance Spectroscopy. 88-89, 1-47 (2015).
- Wu, Y. L., Lo, C. W. Diverse application of MRI for mouse phenotyping. Birth Defects Research. 109 (10), 758-770 (2017).
- Coolen, B. F., et al. High frame rate retrospectively triggered Cine MRI for assessment of murine diastolic function. Magnetic Resonance in Medicine. 69 (3), 648-656 (2013).
- Lapinskas, T., et al. Cardiovascular magnetic resonance feature tracking in small animals - a preliminary study on reproducibility and sample size calculation. BMC Medical Imaging. 17 (1), 51 (2017).
- Pedrizzetti, G., et al. On estimating intraventricular hemodynamic forces from endocardial dynamics: A comparative study with 4D flow MRI. Journal of Biomechanics. 60, 203-210 (2017).
- Motaal, A. G., et al. Accelerated high-frame-rate mouse heart cine-MRI using compressed sensing reconstruction. NMR in Biomedicine. 26 (4), 451-457 (2013).
- Claus, P., Omar, A. M. S., Pedrizzetti, G., Sengupta, P. P., Nagel, E. Tissue tracking technology for assessing cardiac mechanics: principles, normal values, and clinical applications. JACC. Cardiovascular Imaging. 8 (12), 1444-1460 (2015).
- Scatteia, A., Baritussio, A., Bucciarelli-Ducci, C. Strain imaging using cardiac magnetic resonance. Heart Failure Reviews. 22 (4), 465-476 (2017).
- Modin, D., Andersen, D. M., Biering-Sørensen, T. Echo and heart failure: when do people need an echo, and when do they need natriuretic peptides. Echo Research and Practice. 5 (2), 65-79 (2018).
- Onishi, T., et al. Longitudinal strain and global circumferential strain by speckle-tracking echocardiography and feature-tracking cardiac magnetic resonance imaging: comparison with left ventricular ejection fraction. Journal of American Society of Echocardiography. 28 (5), 587-596 (2015).
- Faganello, G., et al. A new integrated approach to cardiac mechanics: reference values for normal left ventricle. The International Journal of Cardiovascular Imaging. 36, 2173-2185 (2020).
- Lapinskas, T., et al. The intraventricular hemodynamic forces estimated using routine CMR Cine images: a new marker of the failing heart. JACC. Cardiovascular Imaging. 12 (2), 377-379 (2019).
- Töger, J., et al. Intracardiac hemodynamic forces using 4D flow: a new reproducible method applied to healthy controls, elite athletes and heart failure patients. Journal of Cardiovascular Magnetic Resonance. 18, 61 (2016).
- Eriksson, J., Bolger, A. F., Ebbers, T., Carlhäll, C. J. Assessment of left ventricular hemodynamic forces in healthy subjects and patients with dilated cardiomyopathy using 4D flow MRI. Physiological Reports. 4 (3), 12685 (2016).
- Coolen, B. F., et al. Three-dimensional T1 mapping of the mouse heart using variable flip angle steady-state MR imaging. NMR in Biomedicine. 24 (2), 154-162 (2011).
- Nijm, G. M., Swiryn, S., Larson, A. C., Sahakian, A. V. Characterization of the magnetohydrodynamic effect as a signal from the surface electrocardiogram during cardiac magnetic resonance imaging. Computers in Cardiology. 33, 269-272 (2006).
- Domenichini, F., Pedrizzetti, G. Hemodynamic forces in a model left ventricle. Physical Review Fluids. 1, 083201 (2016).
- Voigt, J. U., et al. Definitions for a common standard for 2D speckle tracking echocardiography: consensus document of the EACVI/ASE/Industry Task Force to standardize deformation imaging. European Heart Journal - Cardiovascular Imaging. 16 (1), 1-11 (2015).
- Pedrizzetti, G. On the computation of hemodynamic forces in the heart chambers. Journal of Biomechanics. 95, 109323 (2019).
- Hoffman, M., et al. Myocardial strain and cardiac output are preferable measurements for cardiac dysfunction and can predict mortality in septic mice. Journal of American Heart Association. 8 (10), 012260 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
ISSN 2578-2614
ABOUT JoVE
Copyright © 2025 MyJoVE Corporation. All rights reserved
We use cookies to enhance your experience on our website.
By continuing to use our website or clicking “Continue”, you are agreeing to accept our cookies.