A subscription to JoVE is required to view this content. Sign in or start your free trial.
Detection of Nuclear Blebbing and DNA Leakage in Mammalian Cells by Immunofluorescence
In This Article
Summary
Laminopathy disorders often lead to changes in the nuclear envelope, which may cause nuclear blebbing and leakage. This study presents an immunofluorescence method to visualize the nuclear lamina along with double-stranded DNA (dsDNA), providing a means to assess nuclear structure and integrity in mammalian cells.
Abstract
The nuclear lamina is a network of filaments underlying the nuclear membrane, composed of lamins and lamin-associated proteins. It plays critical roles in nuclear architecture, nuclear pore positioning, gene expression regulation, chromatin organization, DNA replication, and DNA repair. Mutations in genes involved in the expression or post-translational processing of lamin proteins result in genetic disorders known as laminopathies. Specifically, mutations in the LMNA or ZMPSTE24 genes can lead to the accumulation of incompletely processed forms of lamin A that retain farnesyl and methyl groups, which are absent in fully processed lamin A. These incompletely processed lamin A proteins localize to the inner nuclear membrane instead of the nuclear lamina, where mature lamin A resides. Mislocalized lamin proteins profoundly disrupt nuclear function and structure, often resulting in nuclear blebbing. In severe cases, nuclear rupture can occur, causing a loss of compartmentalization and leakage of genomic DNA into the cytosol. Abnormal nuclear structure and compartmentalization loss can be identified through indirect immunofluorescence (IF) on fixed cells. This study outlines such a method, employing specific antibodies against a lamin protein and double-stranded DNA (dsDNA) to simultaneously visualize the nuclear envelope and DNA. This approach enables a rapid assessment of nuclear structural integrity and the potential leakage of nuclear DNA into the cytosol.
Introduction
The nuclear lamina is a network of filaments that underlies the nuclear membrane and is made up of proteins called lamins. The nuclear lamina plays essential roles in nuclear architecture, positioning of nuclear pores, gene expression regulation, chromatin organization, DNA replication, and DNA repair1,2,3. Mutations in genes that play a role in the expression of the lamin proteins lead to genetic disorders called laminopathies3.
Restrictive dermopathy (RD) is a severe laminopathy disorder predominantly caused by compound heterozygous mutations that create premature termination codons in the ZMPSTE24 gene4, leading to an absence of the metalloprotease ZMPSTE24. This laminopathy is characterized by intrauterine growth retardation, tight, rigid skin, small-sized mouth, thin hair, and bone mineralization defects1. RD patients usually do not live past their first week of life due to pulmonary insufficiency1. Two other laminopathy disorders known as atypical Hutchinson-Gilford Progeria Syndrome (AT-HGPS) and Mandibuloacral dysplasia type B (MAD-B) involve reduced ZMPSTE24 expression and are associated with reduced lifespan and show similarities to premature aging disorders5. The ZMPSTE24 protease that is impacted in these laminopathies is vital in the post-translational modification of lamin A, which is a critical component of the nuclear lamina. ZMPSTE24 deficiency results in the accumulation of an incompletely processed farnesylated form of lamin A, known as prelamin A2.
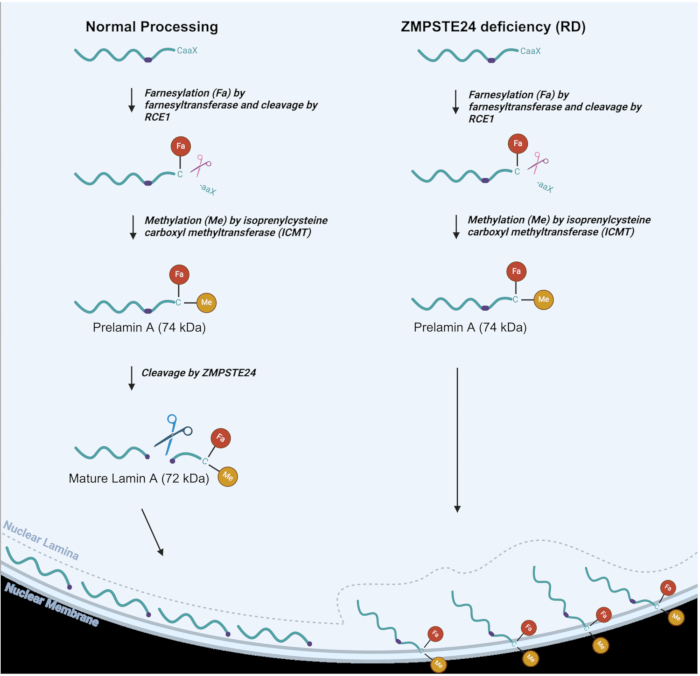
Figure 1: Lamin A processing in normal vs. RD cells. Lamin A processing pathway in normal cells (left) and altered lamin A processing in RD, or ZMPSTE deficient cells (right). Farnesyl (Fa) and methyl (Me) groups are indicated. Please click here to view a larger version of this figure.
As seen in Figure 1, in normal lamin A processing, a farnesyl (lipid) group is attached to a cysteine residue near the C-terminus, followed by proteolytic cleavage of three C-terminal amino acids3,6. The cysteine residue is then methylated. These two modifications allow prelamin A to be targeted to the inner nuclear membrane2. ZMPSTE24 then carries out a cleavage reaction whereby the last 15 C-terminal amino acids, along with the farnesyl and methyl groups, are removed to produce mature Lamin A protein, which is delivered to the nuclear lamina6,7,8. In RD, AT-HGPS, and MAD-B, the final step of processing does not occur effectively since the ZMPSTE24 metalloproteinase is lacking or not fully functional8,9. This results in the accumulation of prelamin A, which remains permanently farnesylated and methylated9. These latter groups cause the prelamin A to stick to the inner nuclear membrane instead of localizing to the nuclear lamina, where mature lamin A should ultimately reside, as seen in Figure 1. Thus, ZMPSTE deficiency has profound effects on a variety of nuclear functions, along with nuclear structure10,11,12,13,14,15. These alterations to nuclear structure may include nuclear blebbing and even nuclear rupturing which can result in the leakage of DNA into the cytosol as well as ingress of cytosol into the nucleus. Aberrantly shaped nuclei are indeed a hallmark of laminopathy disorders, along with numerous other phenotypes that are caused by defective prelamin A processing16,17. Defects in the nuclear lamina lead to a multitude of detrimental impacts, including the mislocalization of nuclear protein quality control and DNA repair proteins in the nucleoplasm, which, in turn, results in numerous defects in nuclear function18. It is important to have a simple technique that can help to identify and monitor these hallmarks of laminopathies to facilitate research into the development of therapeutic approaches aimed at ameliorating laminopathy phenotypes. For mice with laminopathies, it has been shown that the elimination of nuclear blebbing correlates with the generalized elimination of laminopathy phenotypes16. A technique that allows monitoring of nuclear integrity in human cells could enhance the study of potential treatments for eliminating or significantly improving disease phenotypes.
Indirect immunofluorescence (IF) is a sensitive and widely used technique that uses both a primary unlabeled antibody and a fluorophore-labeled secondary antibody that recognizes the primary antibody to detect a target of interest19. IF methods can provide a powerful means for visualizing specific intracellular components and structures. It is also possible that more than one secondary antibody molecule may interact with the primary antibody, resulting in amplification of the signal19. Indirect IF is a versatile technique that also allows the detection of multiple primary antibodies with a relatively small set of secondary antibodies since secondary antibodies are raised against the Fc domain of the primary antibody, which is conserved within species19.
This article describes an indirect (IF) method to assess nuclear blebbing and DNA leakage in cells deficient in ZMPSTE24, using antibodies against double-stranded DNA (dsDNA) and lamin B1 to detect DNA and the nuclear lamina, respectively. To demonstrate the utility of this approach, the procedure was applied to a HeLa cell line with knocked-out ZMPSTE24 expression, as well as a HeLa cell line that expresses ZMPSTE24, and the outcomes for the two cell lines were compared.
Protocol
Details of the reagents and the equipment used are listed in the Table of Materials.
1. Preparation of materials
- Autoclave at least ten 22 mm x 22 mm square glass coverslips and forceps in advance of plating cells onto the coverslips.
2. Preparation of the solutions
- Supplemented media: Prepare 500 mL of Dulbecco's Modified Eagle Medium (DMEM) with 10% fetal bovine serum (FBS), 100 units/mL of penicillin, and 100 µg/mL streptomycin.
- Antibody dilution buffer (ADB): Dissolve 2 g of bovine serum albumin (BSA) and 1 g of Fish gelatin in 80 mL of 1x phosphate-buffered saline (PBS). Bring to 100 mL with PBS once BSA and fish gelatin are dissolved.
- 1x PBS-Tween (PBS-T): Prepare 500 mL of PBS and 0.5 mL of Tween. Place in a 37 °C water bath until Tween can be easily mixed into the solution.
3. Growing the desired cells
NOTE: HeLa cells were grown in this protocol to confluency within a T75 flask.
- Spray six coverslips with 70% ethanol and allow them to air dry on a lint-free wipe. Place coverslips into a sterile 6-well plate using forceps, one coverslip per well.
- Add 2 mL of supplemented DMEM to each well in the 6-well plate.
- Aspirate media from the cells and wash the cells by adding 15 mL of PBS to the flask, gently swirling. Aspirate the PBS and add 2 mL of trypsin-EDTA (trypsin).
- Coat the cells with trypsin and place the flask back into a 37 °C and 5% CO2 humidified incubator for 2 min to allow cells to detach from the flask.
- Add 8 mL of supplemented DMEM to the flask to collect the trypsinized cells. Pipette up and down a few times to separate clumping cells. Collect the cell suspension in the flask and place it into a 15 mL polystyrene tube.
- Count the cells using a hemocytometer.
- Add 500,000 cells, drop by drop, directly onto each of the coverslips in the 6-well plate. Allow cells to grow overnight at 37 °C and 5% CO2 in a humidified incubator.
4. Cell fixation and permeabilization
- Prepare the 4% formaldehyde solution fresh before use: Add 2 mL of 37.5% formaldehyde solution into 18 mL of PBS. Mix thoroughly.
- Prepare the permeabilization solution fresh before use: Add 100 µL of Triton-X into 20 mL of PBS. Allow to incubate in a 37 °C water bath for a few minutes until the Triton-X dissolves into the solution. Mix thoroughly.
- Aspirate the media in the wells of the plate. Wash cells once with 2 mL of PBS. Aspirate all PBS from the wells.
- Add 2 mL of 4% formaldehyde solution to each well. Allow to sit for 10 min at room temperature (RT). Aspirate 4% formaldehyde completely from the wells.
- Wash cells 3 times with 2 mL of PBS. Remove PBS completely from the wells after each wash.
- Add 2 mL of permeabilization solution to each well. Allow to sit for 10 min at RT. Aspirate permeabilization solution completely.
- Wash cells 3 times with 2 mL of PBS. Aspirate PBS completely after the first two washes, leaving the PBS in the wells after the third wash.
NOTE: Cells can be stored in the last PBS wash at 4 °C for a few weeks after this step, ensuring the plate is sealed with paraffin film to avoid liquid evaporation. If the procedure can be continued, this step can be skipped.
5. Immunofluorescence staining
- Remove all PBS from wells. Block cells by adding 2 mL of ADB to each well and incubate at RT, rocking for 30 min.
- Towards the end of the 30 min blocking time, mix the primary antibody dilution in ADB (1:1,000 dilution for both Lamin B1 and dsDNA antibodies, Table 1). 75 µL is needed for each coverslip, therefore 600 µL of ADB, 0.6 µL of Lamin B primary antibody, and 0.6 µL of dsDNA antibody should be mixed for six coverslips. Vortex thoroughly.
- Remove ADB from the wells. On a flat surface, tape down a piece of paraffin film that is large enough in area to lay down all the coverslips being processed.
- For each coverslip, add 75 µL of primary antibody dilution onto the paraffin film, ensuring no bubbles are pipetted in the ADB. Then, place the coverslips cell side down onto the droplet of ADB using forceps.
- Cover the incubating coverslips with the well plate lid and allow to incubate for 1 h at RT.
- Remove the coverslips from the paraffin film using forceps and place the coverslips back into the 6-well plate, cell side facing upward.
- Wash the coverslips 3 times with 2 mL of PBS-T, rocking for 3-5 min each time. Remove all PBS-T from the wells after each wash.
- During the last wash, prepare the secondary antibody dilution (1:1,000 for both respective secondary antibodies, Table 1). 75 µL is needed for each coverslip, therefore 600 µL of ADB and 0.6 µL of both secondary Alexa Fluor tagged antibodies should be mixed for six coverslips. Vortex thoroughly. Cover the secondary antibody dilution from light as much as possible.
| Antibody | Source | Company | Reference | ||
| dsDNA Marker Antibody (HYB331-01) | Mouse monoclonal | Santa Cruz | sc-58749 | ||
| Goat anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor Plus 488 | Goat | ThermoFisher | A32723 | ||
| Goat anti-Mouse IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor Plus 405 | Goat | Invitrogen | A31553 | ||
| Lamin B1 Polyclonal Antibody | Rabbit | Proteintech | 12987-1-AP | ||
| Goat anti-Rabbit IgG (H+L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor 594 | Goat | Invitrogen | A11037 | ||
Table 1: Antibodies. List of all antibodies used in this protocol.
- Remove all PBS-T from the last wash. Incubate the coverslips in the secondary antibody dilution on paraffin film, as in step 5, for 30 min at RT. Protect from light for this step and all remaining steps. A small box can be used to shield light. Ensure to work quickly when needed to add or remove solutions from the wells.
- Remove coverslips from the paraffin film using forceps and place the coverslips back into the 6-well plate, cell side up.
- Wash the coverslips 3 times with 2 mL of PBS-T. Remove all PBS-T from the wells after each wash.
- Dehydrate the slides with an ethanol series (70%, 90%, 100%). Start by adding 2 mL of 70% ethanol to each well. Allow to sit for 1-2 min, and remove all ethanol from the wells. Repeat with the remaining ethanol percentages.
- Remove the coverslips from the wells using forceps and let them air-dry, protected from light, on a lint-free wipe.
- Mount the coverslip's cell side down onto glass microscope slides using 20 µL of mounting medium per coverslip. Allow to dry overnight, protected from light.
6. Image acquisition
- Before imaging, coat the edges of coverslips with a thin layer of clear nail polish (any brand or type will suffice). Allow to air dry, protected from light.
NOTE: The instructions for using a microscope to image will vary, depending on the microscope used. The resolution of the camera on the microscope used here is 1280 x 960 pixels. Similar or better resolution will suffice. - Before putting a slide on the microscope stage, the file-saving settings must be adjusted. Turn on the microscope and insert a USB drive. Adjust the save settings within the microscope settings and ensure that all channels (GFP and RFP) will be saved separately and that the images are going to the correct location on the USB drive.
NOTE: Create a folder on the USB drive and give each coverslip its own name for future reference. - Prior to imaging, spray a lint-free wipe with 70% ethanol and gently clean off the back of the slides. Gently clean the front of the slides, ensuring very minimal pressure is exerted.
- Place a clean slide onto the microscope, coverslips facing downward. Cover with the light shield box.
- Using a 40x objective lens, use the appropriate channel (RFP for the lamin B1 antibody employed here) to locate the nuclei on the microscope screen. Adjust the light intensity to around 50% to start.
- To check for oversaturated areas, select the Color Off option and ensure no areas across the coverslip have a red signal in the black and white setting. If there is a lot of red signal, lower the light intensity until the minimal red signal is shown.
NOTE: A red signal indicates oversaturation. If the nuclei appear barely visible, increase the intensity by 10% increments until an appropriate intensity is achieved without oversaturation. - Check all the other coverslips with the same intensity to ensure they are not oversaturated. Check all coverslips with the other channel using the same process as above (GFP for the dsDNA antibody used here).
- Record light intensities for each channel. Ensure all the light intensities for each channel are kept constant throughout the imaging process.
- Image each coverslip individually. Start off with the first channel, focus the nuclei, and capture the image.
- Carefully switch channels and bring the cells into focus again. When focused, capture the image.
- After capturing images in the necessary channels (GFP and RFP), select Overlay to observe the merged image and ensure the slide did not move when taking the image. If the channels do not overlap, redo the necessary channel until the images line up.
- Save the image by clicking on the flash drive icon. Move to a different place on the coverslip to continue imaging.
NOTE: Ensure that a different area of the coverslip is imaged each time to avoid capturing the same region repeatedly. It is recommended to start imaging at the upper left-hand corner of the coverslip and move in an "S" pattern to ensure complete coverage without duplication.
7. Data analysis
- Data obtained with this method is visual. Examine the images for signs of DNA leakage or nuclear blebbing. Examples of blebbing and leakage are shown in the representative results section.
Results
This study introduces an IF method for visualizing the nuclear lamina in conjunction with double-stranded DNA (dsDNA). Once IF is performed, the images captured can be examined for signs of nuclear blebbing and DNA leakage.
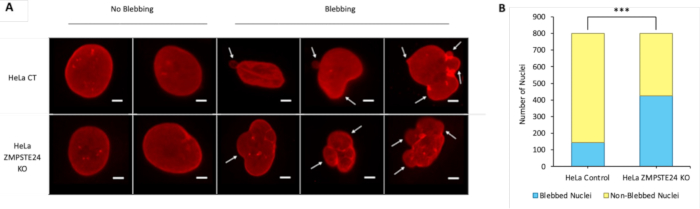
Figure 2: Nuclear blebbing in HeLa CT and ZMPSTE24<...
Discussion
The protocol presented contains several critical steps, the most important being the treatment of the fixed cells with primary and secondary antibodies. Ensuring the use of a good quality primary antibody against the target of interest with a correct corresponding secondary antibody with fluorophores within the range of the microscope used will yield optimal results19. The quality of the antibody can greatly impact the results of this technique, as faulty antibodies can interact with nonspecific t...
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by the National Institute on Aging grant R03AG064525 to ASW. We would like to thank Dr. Jason A. Stewart for help in technique mentorship and the Hui Chen laboratory for providing the microscope used. We would also like to thank Fabio Martinon for the HeLa cell lines used in these experiments.
Materials
| Name | Company | Catalog Number | Comments |
| 22 mm Square Glass Coverslips | Propper Manufacturing Company | M8710 | Any size or shape can be used as long as they can be fixed to a standard microscope slide. |
| 36.5% Formaldehyde | Sigma Aldrich | F8775 | Fixation Reagent. |
| 75 cm 2 Flasks | Corning | 430725U | For cell culture. |
| Bovine Serum Albinum (BSA) | Fisher Scientific | 9048-46-8 | Antibody dilution buffer. |
| DMEM with 1 g/L glucose, L-glutamine & sodium pyruvate | Corning | 10-014-CV | For cell culture. |
| Ethanol (200 Proof) | Decon Laboratories | 2701 | For dehydration of samples before mounting, diluted to make multiple concentrations. |
| EVOS FL Digital Inverted Fluorescence Microscope | Fisher Scientific | 12-563-460 | Imaging. |
| Fish Gelatin | Sigma Aldrich | G7041 | Antibody dilution buffer. |
| Fluoromount-G | Southern Biotech | 0100-01 | Mounting medium to prevent photobleaching. |
| Parafilm (4 in) | Fisher Scientific | 13-374-12 | Any size can be substituted, as long as the coverslips being used can fit. |
| Penicillin-Streptomycin | Gibco | 15-140-122 | For cell culture. |
| Phosphate Buffered Saline (PBS) | N/A | N/A | Made and sterilized in the lab for tissue culture and solutions. |
| Premium Fetal Bovine Serum (FBS) | Atlanta Biologicals | S11150 | For cell culture. |
| Superfrost Premium Microscope Slides | Fisher Scientific | 12-544-7 | Any standard microscope slides can be used. |
| Tissue Culture Treated 6-well Flat Bottom Plates | Falcon | 353046 | For cell culture. |
| TritonX-100 | Thermofisher Scientific | A16046.AP | For washing. |
| Trypsin-EDTA (0.5%) | Gibco | 15-400-054 | For cell culture. |
| Tween 20 | Fisher Scientific | 9005-64-5 | Cell Permeation reagent. |
References
- Navarro, C. L., et al. New ZMPSTE24 (FACE1) mutations in patients affected with restrictive dermopathy or related progeroid syndromes and mutation update. Eur J Hum Genet. 22 (8), 1002-1011 (2014).
- Dittmer, T. A., Misteli, T. The lamin protein family. Genome Biol. 12 (222), 1-16 (2011).
- Gruenbaum, Y., Margalit, A., Goldman, R. D., Shumaker, D. K., Wilson, K. L. The nuclear lamina comes of age. Nat Rev Mol Cell Biol. 6, 21-31 (2005).
- Navarro, C. L., et al. Lamin A and ZMPSTE24 (FACE-1) defects cause nuclear disorganization and identify restrictive dermopathy as a lethal neonatal laminopathy. Hum Mol Genet. 13 (20), 2493-2503 (2004).
- Barrowman, J., Wiley, P. A., Hudon-Miller, S. E., Hrycyna, C. A., Michaelis, S. Human ZMPSTE24 disease mutations: Residual proteolytic activity correlates with disease severity. Hum Mol Genet. 21 (18), 4084-4093 (2012).
- Richards, S., Muter, J., Ritchie, P., Lattanzi, G., Hutchison, C. The accumulation of un-repairable DNA damage in laminopathy progeria fibroblasts is caused by ROS generation and is prevented by treatment with N-acetyl cysteine. Hum Mol Genet. 20 (20), 3997-4004 (2011).
- Davies, B. S., Fong, L. G., Yang, S. H., Coffinier, C., Young, S. G. The post-translational processing of prelamin A and disease. Annu Rev Genomics Hum Genet. 10, 153-174 (2009).
- Barrowman, J., Hamblet, C., George, C. M., Michaelis, S. Analysis of prelamin A biogenesis reveals the nucleus to be a CaaX processing compartment. Mol Biol Cell. 19, 5398-5408 (2008).
- Young, S., Meta, M., Yang, S. H., Fong, L. G. Prelamin A farnesylation and progeroid syndromes. J Biol Chem. 281 (52), 39741-39745 (2006).
- Houthaeve, G., Robijns, J., Braeckmans, K., De Vos, W. H. Bypassing border control: Nuclear envelope rupture in disease. Physiol. 33 (1), 39-49 (2017).
- Schreiber, K. H., Kennedy, B. K. When lamins go bad: Nuclear structure and disease. Cell. 152 (6), 1365-1375 (2013).
- Fong, L. G., et al. Heterozygosity for Lmna deficiency eliminates the progeria-like phenotypes in Zmpste24-deficient mice. Proc Natl Acad Sci USA. 101 (52), 18111-18116 (2004).
- Pendas, A. M., et al. Defective prelamin A processing and muscular and adipocyte alterations in Zmpste24 metalloproteinase-deficient mice. Nat Genet. 31, 94-99 (2002).
- Goldman, R. D., et al. Accumulation of mutant lamin A causes progressive changes in nuclear architecture in Hutchinson-Gilford progeria syndrome. Proc Natl Acad Sci USA. 101, 8963-8968 (2004).
- Eriksson, M., et al. Recurrent de novo point mutations in lamin A cause Hutchinson–Gilford progeria syndrome. Nature. 423, 293-298 (2003).
- Young, S. G., Fong, L. G., Michaelis, S. Thematic review series: Lipid post-translational modifications. Prelamin A, Zmpste24, misshapen cell nuclei, and progeria—new evidence suggesting that protein farnesylation could be important for disease pathogenesis. J Lipid Res. 46 (12), 2531-2558 (2005).
- Worman, H. J., Michaelis, S. Prelamin A and ZMPSTE24 in premature and physiological aging. Nucleus. 14 (1), 1-17 (2023).
- Pande, S., Ghosh, D. K. Nuclear proteostasis imbalance in laminopathy-associated premature aging diseases. FASEB J. 37 (8), 1025-1035 (2023).
- Odell, I. D., Cook, D. Immunofluorescence techniques. J Invest Dermatol. 133 (4), 1-12 (2013).
- Kumar, V. Immunofluorescence and enzyme immunomicroscopy methods. J Immunoassay Immunochem. 21 (2-3), 235-253 (2000).
- Tjaden, A., Giessmann, R. T., Knapp, S., Schroder, M., Muller, S. High-content live-cell multiplex screen for chemogenomic compound annotation based on nuclear morphology. STAR Protoc. 3 (101791), 1-15 (2022).
- Gunn, A. L., Yashchenko, A. I., Dubrulle, J., Johnson, J., Hatch, E. M. A high-content screen reveals new regulators of nuclear membrane stability. Sci Rep. 14, 6013 (2024).
- Yu, L., Liu, P. Cytosolic DNA sensing by cGAS: Regulation, function, and human diseases. Nat Rev Mol Cell Biol. 6 (170), 1-14 (2021).
- Odinammadu, K. O., et al. The farnesyl transferase inhibitor (FTI) lonafarnib improves nuclear morphology in ZMPSTE24-deficient fibroblasts from patients with the progeroid disorder MAD-B. Nucleus. 14 (1), 1-14 (2023).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved