プラークアッセイ:プラーク形成単位(PFU)としてウイルス定数を決定する方法
概要
ソース: ティルデ・アンダーソン1, ロルフ・ルード1
1臨床科学ルンド, 感染医学の部門, ルンド大学生物医学センター, 221 00 ルンド, スウェーデン
バクテリオファージまたは単にファージと呼ばれる原核生物に感染するウイルスは、20世紀初頭にTwort(1)とデレル(2)によって独立して同定された。ファージは、その治療価値(3)とヒト(4)、ならびにグローバルな生態系(5)への影響について広く認識されています。現在の懸念は、感染症の治療における現代の抗生物質の代替としてファージの使用に新たな関心を高めています(6)。本質的にすべてのファージ研究は、ウイルスを精製し、ウイルス力を定量する能力に依存しています, また、ウイルス力中剤として知られています.最初に1952年に記載された、これはプラークアッセイ(7)の目的であった。数十年および複数の技術の進歩後、プラークアッセイは、ウイルス力付き(8)の決定のための最も信頼性の高い方法の一つである。
バクテリオファージは、宿主細胞に遺伝物質を注入し、新しいファージ粒子の製造のための機械を乗っ取り、最終的に細胞リシスを通じて多数の子孫を放出させることによって、生き残る。その微小な大きさのために、バクテリオファージは、単に光顕微鏡を使用して観察することはできません。そのため、走査型電子顕微鏡検査が必要です(図1)。さらに、ファージは、餌食に宿主細胞を必要とするので、細菌のような栄養寒天プレート上で栽培することはできません。
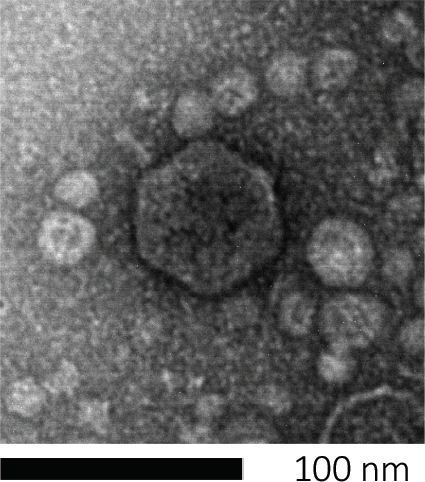
図1:バクテリオファージの形態は、ここで大腸菌ファージによって例示され、走査型電子顕微鏡を用いて研究することができる。ほとんどのバクテリオファージは、カウドビレア(尾のバクテリオファージ)に属しています。この特定のファージは、非常に短い尾構造とイコサヘドラルヘッドを有し、ポドウイルスのファミリーに置きます。
プラークアッセイ(図2)は、宿主細胞の組み込みに基づいており、優先的に対相増殖において、培地に。これは、ウイルスの成長を維持することができる細菌の密な、濁った層を作成します。単離されたファージは、その後感染し、内で複製し、1つの細胞をlyseすることができます。各細胞がlysed細胞で、複数の隣接する細胞が直ちに感染する。いくつかのサイクルで、クリアゾーン(プラーク)は、そうでなければ濁板(図2B/図3A)で観察することができ、最初に単一のバクテリオファージ粒子であったものの存在を示す。試料の体積当たりのプラーク形成単位数(すなわちPFU/mL)は、生成されるプラークの数から決定することができる。
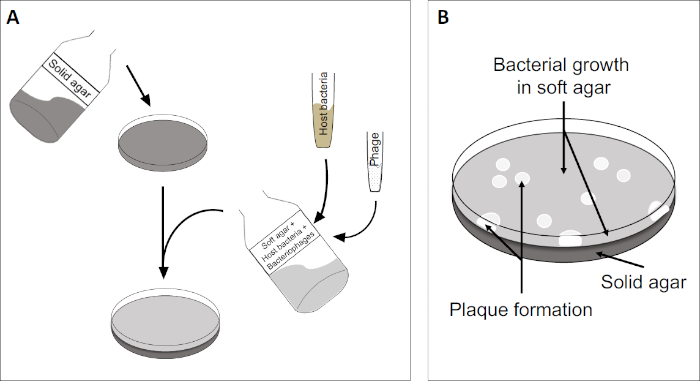
図2:プラーク形成単位(PFU)の試験は、サンプル中のバクテリオファージの数を決定するための一般的な方法である。(A)滅菌ペトリ皿のベースは、適切な固体栄養培地で覆われ、続いて、ソフト培地、感受性宿主細胞および元のバクテリオファージサンプルの希釈が混合される。ファージ懸濁液は、場合によっては、既に固化した柔らかい寒天の表面全体に均等に広がる可能性があることに注意してください。(B)宿主細菌の増殖は、上寒天層に細胞の芝生を形成する。バクテリオファージ複製は、宿主細胞リシスによって引き起こされるクリアゾーン、またはプラークを生成します。
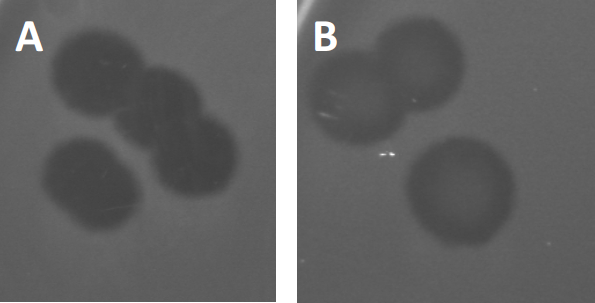
図3:PFU試験の結果は、バクテリオファージによって生成された複数のプラークを示す。感受性宿主細胞のリシスにより、プラークは(A)完全クリアランス、または(B)耐性細菌の生成によって引き起こされる部分的な再増殖(または場合には、または温帯ファージによって、リソジェニックサイクル)。
特定の温帯ファージは、以前に記載された溶解性成長に加えて、リソゲン性のライフサイクルと呼ばれるものを採用することができます。リソジェニーでは、ウイルスは、宿主細胞(9)のゲノムにその遺伝物質を取り込むことを通じて潜伏状態を仮定し、多くの場合、さらなるファージ感染に対する耐性を与える。これは時々プラークのわずかな曇りを通して明らかにされる(図3B)。しかし、以前のファージ感染とは無関係にファージに対する耐性を進化させた細菌の再増殖のためにプラークがぼやけて見えることも注目に値する。
ウイルスは、限られた範囲の宿主細菌にのみ付着または吸着することができる(10)。宿主範囲は、CRISPR-Casシステム(11)のような細胞内抗ウイルス戦略によってさらに制限される。細菌サブグループによって表示される特定のファージに対する耐性/感受性は、歴史的に異なるファージタイプに細菌株を分類するために使用されてきた(図4)。この方法の有効性は、新しいシーケンシング技術によって上回っていますが、ファージタイピングは、例えば、臨床使用のためのファージカクテルの設計を容易にする、細菌とファージの相互作用に関する貴重な情報を提供することができます.
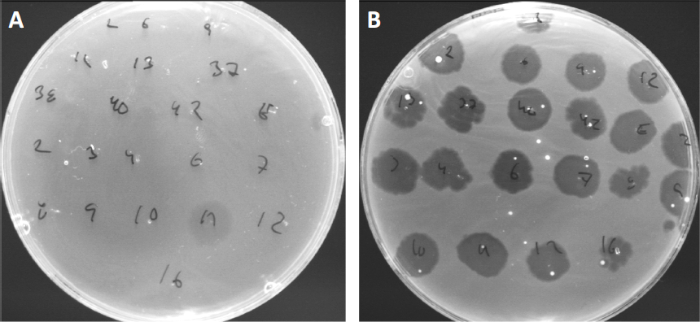
図4:異なる細菌株のファージ感受性。キューチバクテリウム・ニキビ株(A)AD27および(B)AD35を有する柔らかい寒天板を、21種類のC.にきび細菌食症で発見した。唯一のファージ11は、株AD35がすべてのファージに対する感受性を示しながら、AD27に感染し、殺すことができました。ファージタイピングと呼なこの技術は、ファージ感受性に基づいて細菌種と株を異なるサブグループに分割するために使用することができる。
手順
1. セットアップ
- 微生物を含む作業を開始する前に、作業スペースが殺菌されていることを確認してください(例えば、70%のエタノールで拭き取ります)。常にラボのコートと手袋を着用し、長い髪を縛り付け、傷が特によく保護されていることを確認してください。
- 完成したら、すべての表面を殺菌し、手や手首を十分に洗浄/殺菌します。
2. プロトコル
- LB メディアの準備
注:宿主細菌株およびバクテリオファージに応じて、異なる液体培地は、宿主細菌株または異なる固体培地の初期培養に対してより適してもよいし、その後の増殖に対してより適してもよい。リソジェニースープ(LB)は、スープと寒天のためにこのプロトコルで使用されます。- 200 mL蒸留水に4g LBを混ぜ、三量三回に、LBスープ、固体底寒天、柔らかいトップ寒天を混ぜます。オートクレーブ中のオーバーフローを防ぐために、すべてのソリューションは、最終ボリュームの2倍を保持できる容器に用意する必要があります。
注:三重にアッセイを行う場合は、LB底寒天の量を2倍に調べなさい。 - NaOH または HCl を必要に応じて、3 つのソリューションすべての pH を 7.4 に調整します。
- 底寒天ボトルに寒天の電力を3g加え、固体寒天の1.5%の寒天溶液を作る
- 上部の寒天ボトルに1.2gの寒天力を加え、柔らかい寒天の0.6%の寒天溶液を作ります。
- ボトルを半締め付けキャップ付きで、121°Cに設定したオートクレーブに20分間置き、溶液を殺菌します。汚染を防ぐために、実行が完了したらすぐにキャップを閉じます。
- LB培地が約45~50°Cの温度に達したら、200 mL LB溶液の3つすべてに450 μLの無菌1M CaCl2を加え、最終的な濃度を2.25mMにします。
- 固体寒天培地の15mLアリコットを無菌ペトリ皿に注ぎ(発泡を防ぐために揺れを避ける)、寒天が室温で数時間または一晩固まるようにします(図4A)。プレートは、数日間4°Cで逆さまに保存することができます。
- 200 mL蒸留水に4g LBを混ぜ、三量三回に、LBスープ、固体底寒天、柔らかいトップ寒天を混ぜます。オートクレーブ中のオーバーフローを防ぐために、すべてのソリューションは、最終ボリュームの2倍を保持できる容器に用意する必要があります。
- 宿主細胞の培養
- アッセイの1日前に、大腸菌培養の10μLをLBスープの10mLに加える。
- 振盪インキュベーターで160 rpmで一晩37°Cで細菌をインキュベートします。
- アッセイの朝、新鮮なLBの10mLに一晩培養の0.5mLを加える。
注:三重にアッセイを行う場合は、この培養量の2倍を準備します。 - 細菌培養が対数相増殖になるまで、振盪インキュベーターで160rpmで37°Cでインキュベートします。これは0.5-0.7のOD600によって分光光度計で決定することができる。
- 細菌が上部LB寒天に追加されるまで、室温で培養してください。
- バクテリオファージの10倍のシリアル希釈
- LBスープの180μLを96ウェルプレートの最初の行の7つのウェルに追加
注:その統計的信頼性を高めるために三量体で希釈を行うことが示唆される。これを行うには、プレートの2番目と3番目の行にバクテリオファージの追加の希釈を準備します。 - 元のバクテリオファージサンプルを慎重にボルテックスして均質性を確保し、20 μLを最初のウェルに移します。
- ピペッティングとダウンでサンプルをよく混ぜます。
- 得られた懸濁液の20μLを第2ウェルに移す。
- 第2井戸の溶液の20μLを第3ウェルに移してシリアル希釈を続け、6番目の井戸まで、ファージ懸濁液を加えない負の制御を残します。これにより、10-1 -1-10-6の希釈範囲が作成されます。
- LBスープの180μLを96ウェルプレートの最初の行の7つのウェルに追加
- めっき
- 名前、日付、短いサンプル説明(ファージサンプル希釈係数を含む)で、ペトリ皿のベース(以前はステップ2.1.7で調製)にラベルを付けます。ペトリ皿を37°Cのインキュベーターで予熱し、アッセイの1時間前に加熱します。
- 固化した柔らかい寒天媒体(通常は電子レンジを使用して行われ、固化寒天は85°Cで溶融)を溶かし、45°Cの周りに来るようにします。加熱された媒体は、適切な温度に達するために〜1時間の45°Cの水浴に置くことができます。それは液体の形で残るのに十分な熱いはずですが、追加された細菌を殺さないほど冷やします。
- 4 mL細菌培養(ステップ2.2から)を35 mL LB軟寒天(45°C)と混合します。細胞を均等に分配するために旋回するが、発泡を防ぐために揺れを避ける(図4A)。
- シリアル希釈ステップのそれぞれに対して1本の無菌試験管と、制御サンプル用に1本の無菌試験管をラベル付けし、合計7本の標識試験管にラベルを付けます。ステップ2.4.3から細菌培養/柔らかい寒天混合物の5 mLアリコートを7管に入れます。このような少量の寒天ベースのメディアは室温で素早く固まるため、ステップ2.4.6を通して迅速に作業します。
- 制御試料の100 μL(ステップ2.3から)をコントロール試験管に追加し、慎重に旋回します。使用したピペットチップを廃棄し、連続的に希釈されたバクテリオファージサンプル(ステップ2.3)のそれぞれから同じ体積をそれぞれの試験管に移し、混ぜ合わせます。
- 5 mL混合物をラベル付けされた、予熱された固体寒天プレートに直ちに移す(図4A)。プレートをそっと旋回して混合物を広げます。
注:アッセイを三回転で実行する場合は、手順 2.4.3- 6 をもう 2 回繰り返します。 - 実験室用フィルムで各プレートをシールし、両方の層を室温(約15分)で適切に固化させてから、37°Cのインキュベーターを上下に置き、細菌とファージの両方の成長を刺激し、24時間またはプラークまでプラークが現れるのに通常約1〜5日かかりますが(図4B)、インキュベーション条件、培地、細菌種によってタイミングが大きく異なります。
3. データ分析と結果
-
プラークのカウント
- これはウイルス汚染を示すので、「制御」とマークされたプレートにプラークが見えないことを確認してください。
- 最も希釈されたバクテリオファージサンプルを含む10-6のラベルのプレートから始めます。ふたを取り外さずにプラークを数え、ペンで印を付けて、どのプラークが既にカウントされているかを示します。
- 残りのプレートを数えます。一部のプレートには、数えるプラークが少なすぎたり多すぎたりすることがあります。さらなる分析のために10-150プラークとプレートを使用してください。
-
PFU の計算
- プラークの数を希釈係数で除算し(最も希釈されたサンプルの場合は10-6)、ファージ混合物の100μLでプラーク形成単位(PFU)の数を得る。
注:三量体でアッセイを行う場合は、3枚のプレートからプラークの平均数を使用します。 - 元のサンプルの濃度(PFU/mL)を決定するために、サンプルの100 μLのみがめっきされたので、10の追加の希釈係数を乗算します。(すなわち)
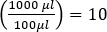
- より信頼性の高い結果を得るために、10~ 150 プラークの間にあるすべての希釈に対する PFU/mL の平均値を計算します。
- プラークの数を希釈係数で除算し(最も希釈されたサンプルの場合は10-6)、ファージ混合物の100μLでプラーク形成単位(PFU)の数を得る。
申請書と概要
複数の技術の進歩にもかかわらず、プラークアッセイはウイルス力確定のためのゴールドスタンダード(PFUとして)であり、純粋なバクテリオファージ集団の単離に不可欠です。感受性の高い宿主細胞は、2層寒天板のトップコートで培養され、ウイルス複製を可能にする均質なベッドを形成する。溶解ライフサイクル中の単離されたバクテリオファージが細胞に感染し、細胞内で複製し、最終的にそれを溶解する最初のイベントは、観察するには小さすぎます。しかし、放出されたビリオンは隣接する細胞に感染し、その後、単一のPFUの存在を示すクリアリングゾーン、またはプラークを生み出す。
参考文献
- Twort, F. An investigation on the nature of ultra-microscopic viruses. Lancet. 186 (4814): 1241-1243. (1915)
- d'Hérelle, F. An invisible antagonist microbe of dysentery bacillus. Comptes Rendus Hebdomadaires Des Seances De L Academie Des Sciences. 165: 373-375. (1917)
- Cisek AA, Dąbrowska I, Gregorczyk KP, Wyżewski Z. Phage Therapy in Bacterial Infections Treatment: One Hundred Years After the Discovery of Bacteriophages. Current Microbiology. 74 (2):277-283. (2017)
- Mirzaei MK, Maurice CF. Ménage à trois in the human gut: interactions between host, bacteria and phages. Nature Reviews Microbiology. 15 (7):397. (2017)
- Breitbart M, Bonnain C, Malki K, Sawaya NA. Phage puppet masters of the marine microbial realm. Nature Microbiology. 3 (7):754-766. (2018)
- Leung CY, Weitz JS. Modeling the synergistic elimination of bacteria by phage and the innate immune system. Journal of Theoretical Biology. 429:241-252. (2017)
- Dulbecco R. Production of Plaques in Monolayer Tissue Cultures by Single Particles of an Animal Virus. Proceedings of the National Academy of Sciences of the United States of America. 38 (8):747-752. (1952)
- Juarez D, Long KC, Aguilar P, Kochel TJ, Halsey ES. Assessment of plaque assay methods for alphaviruses. J Virol Methods. 187 (1):185-9. (2013)
- Clokie MRJ, Millard AD, Letarov AV, Heaphy S. 2011. Phages in nature. Bacteriophage. 1 (1):31-45. (2011)
- Moldovan R, Chapman-McQuiston E, Wu XL. On kinetics of phage adsorption. Biophys J. 93 (1):303-15. (2007)
- Garneau JE, Dupuis M-È, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadán AH, Moineau S.. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature. 468 (7320):67. (2010)
タグ
スキップ先...
このコレクションのビデオ:

Now Playing
プラークアッセイ:プラーク形成単位(PFU)としてウイルス定数を決定する方法
Microbiology
186.4K 閲覧数

ウィノグラツキーカラムの作成:堆積物サンプル中の微生物種を濃縮する方法
Microbiology
129.7K 閲覧数

シリアル希釈とめっき:微生物列挙
Microbiology
316.7K 閲覧数

エンリッチメント培養:選択的および差動媒体における好気性微生物と嫌気性微生物の培養
Microbiology
132.2K 閲覧数

純粋な培養物と縞めっき:混合サンプルからの単一細菌コロニーの単一の分離
Microbiology
166.3K 閲覧数

16S rRNAシーケンシング:細菌種を同定するPCRベースの技術
Microbiology
189.4K 閲覧数

成長曲線:コロニー形成単位と光学密度測定を用いて成長曲線を生成する
Microbiology
297.2K 閲覧数

抗生物質感受性試験:2つの抗生物質のMIC値を決定し、抗生物質の相乗効果を評価するエプシロメーター試験
Microbiology
93.9K 閲覧数

顕微鏡検査と染色:グラム、カプセル、内胞染色
Microbiology
363.7K 閲覧数

適応塩化カルシウム手順を用いて大腸菌細胞の形質転換
Microbiology
87.0K 閲覧数

結合:アンピシリン耐性をドナーからレシピエント大腸菌に移す方法
Microbiology
38.3K 閲覧数

ファージトランスダクション:アンピシリン耐性をドナーからレシピエント大腸菌に伝達する方法
Microbiology
29.1K 閲覧数
Copyright © 2023 MyJoVE Corporation. All rights reserved