Aby wyświetlić tę treść, wymagana jest subskrypcja JoVE. Zaloguj się lub rozpocznij bezpłatny okres próbny.
Method Article
Replication of the Ordered, Nonredundant Library of Pseudomonas aeruginosa strain PA14 Transposon Insertion Mutants
* Wspomniani autorzy wnieśli do projektu równy wkład.
W tym Artykule
Podsumowanie
Pseudomonas aeruginosa infection causes significant morbidity in vulnerable hosts. The nonredundant transposon insertion mutant library of P. aeruginosa strain PA14, designated as PA14NR Set, facilitates analysis of gene functionality in numerous processes. Presented here is a protocol to generate high-quality copies of the PA14NR Set mutant library.
Streszczenie
Pseudomonas aeruginosa is a phenotypically and genotypically diverse and adaptable Gram-negative bacterium ubiquitous in human environments. P. aeruginosa is able to form biofilms, develop antibiotic resistance, produce virulence factors, and rapidly evolve in the course of a chronic infection. Thus P. aeruginosa can cause both acute and chronic, difficult to treat infections, resulting in significant morbidity in certain patient populations. P. aeruginosa strain PA14 is a human clinical isolate with a conserved genome structure that infects a variety of mammalian and nonvertebrate hosts making PA14 an attractive strain for studying this pathogen. In 2006, a nonredundant transposon insertion mutant library containing 5,459 mutants corresponding to 4,596 predicted PA14 genes was generated. Since then, distribution of the PA14 library has allowed the research community to better understand the function of individual genes and complex pathways of P. aeruginosa. Maintenance of library integrity through the replication process requires proper handling and precise techniques. To that end, this manuscript presents protocols that describe in detail the steps involved in library replication, library quality control and proper storage of individual mutants.
Wprowadzenie
Pseudomonas aeruginosa is a phenotypically and genotypically diverse and adaptable Gram-negative bacterium present in soil, water, and most human environments, as well as skin microflora. Compared to many bacterial species, P. aeruginosa has a relatively large genome of 5.5-7 Mbp with high G+C content (65-67%). Furthermore, a significant proportion of its genes are involved in metabolic adaptability and are part of regulatory networks, allowing for great flexibility in response to environmental stress1. P. aeruginosa expresses a plethora of virulence factors, exhibits proclivity to form biofilms, possesses the ability to coordinate responses through multiple quorum sensing pathways, and displays a notable capacity to develop antibiotic resistance and tolerance2,3,4,5,6,7,8. These attributes present significant challenges for treating infections caused by P. aeruginosa.
Chronic P. aeruginosa infections can occur in numerous disease states. Cystic fibrosis (CF), a genetic disease caused by mutation of the Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) gene, results in inspissated, infected secretions within the airway, progressive bronchiectasis and, ultimately, death from respiratory failure9. By adulthood, the majority of patients with CF are chronically infected with P. aeruginosa, which plays a key role in the morbidity and mortality associated with this disease10. Additionally, patients with severe burn injuries11, tracheostomies12, joint replacements13, or indwelling catheters14 are at risk for P. aeruginosa infection related to the bacteria's ability to form biofilms and escape host inflammatory responses15. Further, colonization occurs without competition after a multi-antibiotic resistant or tolerant population is selected through broad-spectrum, sequential antimicrobial treatment12,16,17,18. Better understanding the pathogenesis of P. aeruginosa will have significant implications for numerous disease states.
Several P. aeruginosa clinical isolates, including strains PAO1, PA103, PA14 and PAK, have been extensively studied to investigate different features of P. aeruginosa pathogenesis. Strain PA14 is a clinical isolate that belongs to one of the most common clonal groups worldwide19,20 and has not been extensively passaged in the laboratory. PA14is highly virulent in vertebrate models of infection, with a notable endotoxin profile21, pili structure22, pathogenicity islands23, type III secretion system (TTSS), cytotoxicity towards mammalian cells24 and profiles in antibiotic resistance and persistence25. Furthermore, PA14 is also highly virulent in numerous host-pathogen model systems, including plant leaf infiltration models26,27,Caenorhabditis elegans infection models28,29, insect models30,31, as well as mouse pneumonia models32,33 and skin burn models34.
Genome-wide mutant libraries are collections of isogenic mutants in nonessential genes that constitute very powerful tools to understand the biology of an organism by allowing analysis of gene function on a genomic scale. Two near-saturation transposon insertion mutant libraries constructed in P. aeruginosa are currently available for distribution. The insertion sites of the transposons have been determined for both libraries. These so-called nonredundant libraries facilitate genome-wide studies of bacterial strains by considerably decreasing the time and cost involved in screening uncharacterized random transposon mutants. The P. aeruginosa PAO1 transposon mutant library, constructed in the MPAO1 isolate of strain PAO1 using transposons ISphoA/hah and ISlacZ/hah35, is curated by the Manoil lab, University of Washington. The library consists of a sequence-verified collection of 9,437 transposon mutants that provides wide genome coverage and includes two mutants for most genes36. Information about the P. aeruginosa PAO1 transposon mutant library is available at the public, internet-accessible Manoil lab website at http://www.gs.washington.edu/labs/manoil/libraryindex.htm. The P. aeruginosa strain PA14 nonredundant transposon insertion mutant library (PA14NR Set) constructed in strain PA14 using transposons MAR2xT7 and TnphoA37 is currently distributed by the Department of Pediatrics at Massachusetts General Hospital. The PA14NR Set comprises a collection of more than 5,800 mutants with single transposon insertions in nonessential genes37. Details on the construction of the PA14NR Set are described in the public, internet-accessible site http://pa14.mgh.harvard.edu/cgi-bin/pa14/home.cgi?section=NR_LIB, which also contains a variety of online search tools to facilitate the use of the PA14NR Set.
The original PA14NR Set comprised 5,459 mutants, selected from a comprehensive library of approximately 34,000 random transposon insertion mutants, that correspond to 4,596 predicted PA14 genes representing 77% of all predicted PA14 genes37. Since the construction of the library in 2006 new mutants were added, and presently the PA14NR Set includes more than 5,800 mutants38 that represent approximately 4,600 PA14 genes. The majority of the PA14 transposon mutants were generated in the wild type background37. Details concerning each member of the mutant library, including genetic background, are available either through searching the online database, or by downloading the Nonredundant Library spreadsheet, both features available on the PA14 website (http://pa14.mgh.harvard.edu/cgi-bin/pa14/home.cgi). The majority of mutants were created using the MAR2xT7 (MrT7) transposon, with a small set created using the TnPhoA (phoA) transposon37. Each transposon has an antibiotic resistance cassette, which allows for mutant selection using gentamicin (MrT7) or kanamycin (phoA). The PA14NR set of mutants is stored in sixty-three 96-well plates and includes two additional 96-well control plates, which consist of wild type PA14 inoculated and uninoculated wells intercalated in a preset pattern. The 96-well plate format paired with the online search tools greatly facilitates the custom development of screening assays that allow users to easily identify genes associated with mutant phenotypes. The online search tools also facilitate the search and selection of additional relevant mutants required for further studies.
The PA14 and PAO1 transposon mutant libraries are very important global resources for the scientific community, and they complement each other in validating the function of unknown genes and pathways of this bacterial pathogen. Coincidentally, since the construction of the PAO1 and PA14 transposon mutation libraries, full-genome DNA sequencing analysis of many P. aeruginosa isolates has shown that PAO1 and PA14 belong to different major subclades of the P. aeruginosa phylogeny7,39,40,41. Because clinical P. aeruginosa isolates are found distributed throughout the phylogeny, the fact that PAO1 and PA14 belong to different P. aeruginosa subgroups enhances the value of the two transposon mutation libraries for comparative studies.
Publications describing the construction and screening of bacterial mutant libraries, including P. aeruginosa libraries35,37,42, are readily available in the literature. However, to the best of our knowledge, no published protocols describing detailed procedures and techniques used for replication, maintenance, and validation of bacterial mutant libraries are available.
The methodology outlined in this publication describes a set of three protocols that facilitate the use and maintenance of the PA14NR Set. The first protocol describes replication of the library as recommended to recipients of the PA14NR Set. The second protocol includes guidelines for streaking, growing, and storing individual mutants identified using the PA14NR Set. The third protocol describes quality control techniques, including PCR amplification of fragments from transposon mutants and subsequent sequencing to confirm mutant identity. This set of protocols may also be adapted for the replication and maintenance of other bacterial mutant libraries or collections. The replication of bacterial mutant libraries or collections is highly advised to preserve the integrity of the "master copy" (original copy received). Replication of several copies of the PA14NR Set for routine laboratory use minimizes the probability of interwell contamination of the master copy.
Protokół
CAUTION: Utilize standard BSL-2 safety measures when handling P. aeruginosa, a human pathogen. If you are an immunocompromised individual or have any medical condition that increases your susceptibility to bacterial infection, take special caution when working with P. aeruginosa. Consult the biosafety office in your institution and obtain approval from your physician before working with the the PA14 NR Set or mutant libraries of bacterial pathogens.

Figure 1: Overview of Protocol I: replication of the PA14NR set. Day 1: Replicate frozen mutant cultures from "master copy" of PA14NR Set into LB Agar media and grow mutants overnight at 37 °C. Day 2: Transfer mutant growth from LB Agar media to Deep Well Blocks containing LB liquid broth, grow overnight at 37 °C with shaking at 950 rpm. Day 3: Mix overnight LB cultures with glycerol, then transfer to 96-well destination plates for long-term storage. Place 96-well plates flat in -80 °C freezer. Please click here to view a larger version of this figure.
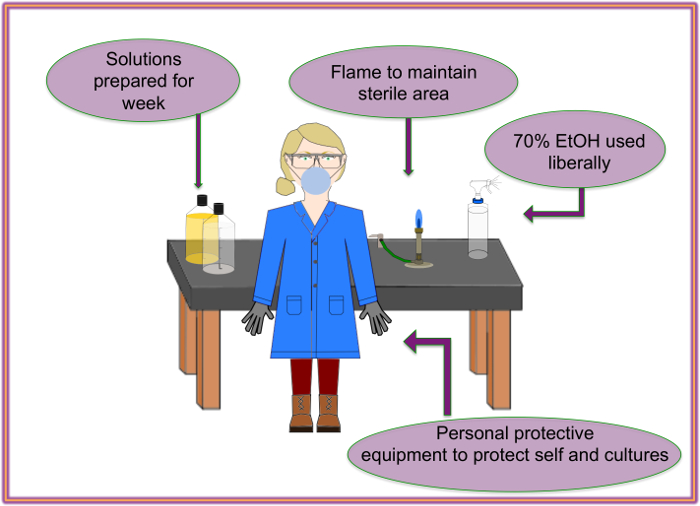
Figure 2: Recommended setup. Sterility and smooth workflow should be maintained through the use of appropriate precautions. Please click here to view a larger version of this figure.
1. Protocol I: Replication of the PA14NR Set
NOTE: Replication of the library can be achieved by dividing the PA14NR Set in four subsets of sixteen plates each that can be processed in four consecutive weeks. Generation of 1 to 6 copies adheres to the weekly workflow outlined in Table 1, while generation of more than 6 copies follows the weekly workflow outlined in Table 2. To generate 12 copies of the PA14NR Set, inoculate the same PA14NR subset into liquid LB media on Day 2 and again on Day 3 (from the same set of agar plates replicated on Day 1), and transfer overnight mutant cultures into copy plates on Day 3 and Day 4 respectively.
| Day 0 | Day 1 | Day 2 | Day 3 |
| Prep Day | Growth of PA14NR Set mutants on LB agar media | Growth of PA14NR Set mutants in liquid LB media | Transfer of PA14NR Set mutant cultures to destination plates |
Table 1: Schedule for replication of 1 - 6 copies of the PA14NR Set. Replication of a small number of copies may adhere to a weekly workflow.
| Day 0 | Day 1 | Day 2 | Day 3 | Day 4 |
| Prep Day | Growth of PA14NR Set mutants on LB agar | Grow of PA14NR Set mutants in liquid LB media | Transfer of Day 3 mutant cultures to generate 1st set of 6 copies of PA14NR Set | Transfer of Day 4 mutant cultures to generate 2nd set of 6 copies of PA14NR Set |
| Growth of PA14NR Set mutants in liquid LB media |
Table 2: Schedule for replication of up to 12 copies of the PA14NR Set. Replication of a larger number of copies will require layering within a weekly workflow.
- Day 1: Growth of PA14NR Set Master copy on LB agar plates
- Wear gloves, lab coat and mask to handle the PA14NR Set.
- Clear bench and wipe surface with 70% ethanol.
- Prepare LB agar media43 and sterilize in autoclave for 25 min. Cool media to 55 °C in water bath and add either 15 µg/mL gentamicin, for mutants containing MAR2xT7 transposon insertions, or 200 µg/mL kanamycin, for mutants containing TnphoA insertions.
- Pour molten LB agar in rectangular plates using approximately 60 mL of media per plate. Dry plates in a sterile hood for approximately 1 h before use. Make sure there is no water condensation on the surface of the agar. Store plates at 4 °C, if necessary.
- Create a sterile field with a Bunsen burner on ethanol-wiped bench top and set up containers with appropriate solutions for replicator pin sterilization (see step 1.1.9). Use open plastic containers (5.25"L x 4.25"W x 1.75"H approximate size) for replicator pin sterilization. Autoclave plastic containers before use.
NOTE: The heat of the Bunsen burner flame creates a convection current, which heats the space above the flame and lifts any particulates in the air up and away from the cooler air underneath, keeping the work area sterile. - Remove a maximum of four PA14NR Set master plates from the -80 °C freezer (to avoid unnecessary thawing), and place them on dry ice in 4 L ice pan.
- Take the 96-well master plate to be replicated from dry ice and place it on the bench to allow brief thawing, which will only take the time necessary for sterilization of the replicator pins (approximately 3-4 min) (see step 1.1.8). Mark the position of the A1 coordinate on the agar plate prior to stamping. Align master plate and agar plate with the A1 coordinate of both plates in the upper left corner.
- Sterilize the replicator "pins" by following the steps described below. Tap replicator slightly after each step to remove excess liquid.
- Immerse pins in 250 mL of 0.3-0.5% sodium hypochlorite (10% household bleach) for 30 seconds. Minimize contact with sodium hypochlorite solution, as it can lead to damage of pins. Immerse pins in 250 mL sterile ddH2O or sterile ultrapure water for 10 seconds, then in 250 mL 70% ethanol for 30 seconds, then in 250 mL of 95% ethanol for 2 min.
- Flame sterilize replicator pins by holding replicator perpendicularly to the Bunsen burner flame, slowly approaching flame until ethanol ignites, then immediately withdrawing it from flame. Flame will extinguish once all ethanol burns off. Keep a lid or similar container close by to suffocate burning ethanol, if necessary.
NOTE: Use extreme caution when working with ethanol near a flame. DO NOT hold replicator directly over flame. - Cool pins by pinning onto an unused sterile rectangular plate containing agar LB media for 30 seconds.
- Briefly warm aluminum seal from PA14NR Set master plate with your hand before peeling it off. Do this while cooling replicator pins. Remove aluminum seal carefully to prevent the seal from retouching the plate. Use tweezers to peel off any remnants of aluminum seal.
- Insert replicator pins into master plate, gently pushing and swinging replicator in all four directions to ensure that pins make contact with frozen bacterial cultures in each of the 96-wells.Pay extra attention to wells located in the outer edge of the plate by pushing replicator pins against frozen cultures in the wells located at the edge of the plate.
- Gently place replicator pins onto surface of the agar plate. Move replicator in a slight circular motion to form mini-lawns of approximately 4-5 mm for each mutant strain. Avoid the possibility that the mini-lawns may overlap, to avoid cross contamination.
- Seal PA14NR Set master plate with a new sterile aluminum seal. Do not touch the adhesive side of aluminum seal at any point to avoid contamination. Make sure that each well and the edges of plate are completely sealed by using a plate roller. Return 96-well plate to dry ice.
- Repeat procedure for each master plate.
- Wipe down all work surfaces with 70% ethanol after handling the PA14NR Set.
- Transfer replicated agar plates to 37 °C incubator and incubate overnight.
- Day 2: Growth of the PA14NR Set Master copy in LB liquid broth
- Prepare LB liquid broth43 containing either 15 µg/mL gentamicin, for mutants containing MAR2xT7 transposon insertions, or 200 µg/mL kanamycin, for mutants containing TnphoA insertions.
- Clear laminar flow hood of unnecessary equipment and turn hood blower on for a minimum of 10 min before beginning work. Wipe down hood surfaces and any items placed in the hood using 70% ethanol.
- Fill 2 mL deep-well blocks with 525 µL of LB liquid broth containing appropriate antibiotics in laminar flow hood using a 50-1200 µL 12-channel electronic pipette. Then, transfer media-filled deep-well blocks to ethanol-wiped bench top. Reuse tips as long as sterile conditions are maintained.
- Wear gloves, lab coat, and mask to handle PA14NR Set.
- Clear bench and wipe surface with 70% ethanol.
- Bring agar plates with overnight grown mutant strains to bench top.
- Create a sterile field with a Bunsen burner on ethanol-wiped bench top and set up containers with appropriate solutions for replicator pin sterilization (see next step).
- Sterilize the replicator "pins" following the steps described below. Tap replicator slightly after each immersion to remove excess liquid.
- Immerse pins in 250 mL of 0.3-0.5% sodium hypochlorite for 30 seconds. Minimize replicator pin contact with sodium hypochlorite solution, as it can lead to damage of pins. Then, immerse pins in 250 mL sterile ddH2O or ultrapure water for 10 seconds, then in in 250 mL of 70% ethanol for 30 seconds, then in 250 mL of 95% ethanol for 2 min.
- Flame sterilize replicator pins, then cool pins by pinning into unused rectangular plate containing agar LB media for 30 seconds.
NOTE: Use extreme caution when working with ethanol near a flame (see 1.1.8).
- Gently place replicator pins onto agar plate containing mutant growth and check that pins are in contact with all 96 mutants on agar plate, then submerge pins into deep wells containing LB liquid broth. Avoid touching the sides of the wells with the pins.
- Seal the deep well block with a sterile breathable sealing membrane. Use a plate roller to ensure that each individual well is properly sealed.
- Repeat procedure for each agar plate.
- Wipe down all work surfaces with 70% ethanol after handling the PA14NR Set.
- Grow inoculated liquid cultures for 15 - 16 h at 37°C at 950 rpm using a high speed shaker, if available.
NOTE: If a high speed shaker is not available, incubation of deep well blocks in shaker at 250-300 rpm is feasible. However, there is a greater chance of small colony variants (SCVs)25 emerging under low oxygenation conditions. Therefore, it is highly advisable to keep incubation times under 15 - 16 h when growing cultures in deep well blocks at lower shaker speeds. Unwanted proliferation of SCVs in mutant wells can alter mutant phenotypes when using the library to perform genetic screens.
Certain wells in the PA14NR Set contain slow-growing/non-growing mutants, missing clones, or contain uninoculated media. The location of these wells has been recorded and can be found in the Supplemental PA14NR Set Wells Information file included with this publication.
- Day 3: Transfer of PA14NR Set overnight cultures into destination plates
- Clear laminar flow hood and turn hood blower on for a minimum of 10 min before beginning work. Wipe down hood surfaces and any items placed in the hood using 70% ethanol.
- Print adhesive waterproof labels (see Supplemental PA14NR Set Labels file for template). Remove 96-well plates from plastic wrap inside a laminar flow hood. Peel-off plate label and place it along the plate's edge closest to A1 to H1 wells of the destination plate. Slightly lift the lid of the destination plate and place the label on the lower edge to display label when plate is covered with lid.
- Prepare 3.5 L of 60% glycerol (v/v) and sterilize 20 min in autoclave.
- Wear gloves, lab coat, and mask to handle the PA14NR Set. Remove deep well blocks from high speed shaker or regular shaker.
- Transfer deep-well blocks to sterile laminar flow hood and carefully remove breathable sealing membrane. Replace with aluminum seal.Spin-down the deep-well blocks at very low speed to collect condensation (30 seconds at 50-150 x g, then decelerate quicky by having centrifuge brake on).
- Transfer deep-well blocks back to sterile hood and using a 50-1200 µL 12-channel electronic pipette and sterile filtered tips add 525 µL of glycerol/LB liquid broth mix (equal parts LB liquid broth and 60% glycerol solution) to each well. Mix by gently pipetting 300 µL up and down 3 times with electronic pipette. Touch tips to side of well before ejecting tips to prevent dripping. Eject tips and continue with the next row until done with entire deep well block.
- Using the 12-channel electronic repetitive pipette and filtered tips, to prevent interwell contamination during aliquoting, pull up 900 µL of mutant culture and dispense 150 µL into each 96-well destination plates to generate 6 copies of the library plate. Prevent drips by touching the wall of the wells with tips before starting transfer of the culture into destination plates. If no repeat pipette is available, use a multichannel pipette to dispense 150 µL into each of the six 96-well plates, using the technique described above to prevent dripping.
- Use sterile aluminum seals to cover the plates and use a plate roller to completely seal plate edges and all wells. Make sure not to cover label identifier with the aluminum seal. Do not shake the plates, as culture may splash on sides of the wells or on aluminum seal.
- Remove sealed plates from hood and place plates on a flat, even surface in the -80 °C freezer.
- Wipe down all work surfaces with 70% ethanol after handling the library.
- Perform quality control checks after library replication and after using the library to perform genetic screens (see Protocol III).
2. Protocol II: Handling and Storage of Individual Mutants from the PA14NR Set
- Day 1: Streak mutant of interest
- Identify the location of the mutant of interest through the PA14NR Set link http://pa14.mgh.harvard.edu/cgi-bin/pa14/search.cgi?searchType=SEARCH_PLATE_POSITIONS or using the Nonredundant Library.xls file downloaded from the link http://pa14.mgh.harvard.edu/cgi-bin/pa14/downloads.cgi). Make a note of the antibiotic needed to select every particular mutant (gentamicin or kanamycin).
- Prepare LB agar media, sterilize in autoclave for 20-25 min, and cool to 55 °C in water bath. Add 15 µg/mL gentamicin for mutants containing MAR2xT7 transposon insertions, and 200 µg/mL kanamycin for mutants containing TnphoA transposon insertions. Pour LB agar media on plates (round or rectangular plates are adequate). Dry plates in sterile hood for approximately 30 min to 1 h before use.
- Autoclave all plasticware and non-sterile supplies before use.
- Wear gloves, lab coat, and mask to handle the PA14NR Set. Clear bench and wipe surface with 70% ethanol before working with the PA14NR Set.
- Create a sterile field with a Bunsen burner.
- Remove PA14NR Set 96-well plate with mutant of interest from -80 °C freezer and place it on dry ice, take dry ice container to the bench, and briefly place 96-well plate on top of bench to allow slight thawing (approximately 1-2 min).
NOTE: Keep a record of all PA14NR Set 96-well plates accessed to streak individual mutants, as more access to library plates is correlated with a greater risk for interwell contamination. - Warm aluminum seal with hand before peeling it off, being careful to prevent the seal from retouching the plate. Use tweezers to peel off any remnants of aluminum seal.
- Locate the mutant of interest on the 96-well plate. Use either sterile wooden stick or sterile pipette tip to pick a small amount of frozen culture from the individual well containing the mutant of interest.
- Streak frozen culture on agar plate for single mutant colonies as follows: Gently spread the bacteria over a section of the plate to create streak 1, using a fresh, sterile wooden stick or pipette tip, drag through streak 1 and spread the bacteria over a second section of the plate, to create streak 2. Using a third sterile wooden stick or pipette tip, drag through streak 2 and spread the bacteria over the last section of the plate, to create streak 3.
- Seal source plate with a new sterile aluminum seal. Do not touch adhesive side of aluminum seal at any point to avoid contamination. Make sure that each well and edges of plate are completely sealed by using a plate roller. Return 96-well plate to dry ice and then the -80 °C freezer.
- Incubate agar plate in 37 °C incubator overnight.
- Wipe down all work surfaces with 70% ethanol after handling the library.
- Day 2: Growth of mutant of interest in LB liquid broth
- Prepare liquid LB broth containing 15 µg/mL gentamicin or 200 µg/mL kanamycin, as per transposon insertion.
- Wear gloves, lab coat, and mask to handle P. aeruginosa.
- Clear bench and wipe surface with 70% ethanol. Create a sterile field with a Bunsen burner.
- Transfer 3 - 5 mL of LB broth with appropriate antibiotic into sterile culture tube with cap.
- Using a sterile applicator or a sterile pipette tip, pick a single colony of mutant strain and inoculate it into LB media.
- Incubate LB liquid cultures at 37 °C in a shaker at 225 - 250 rpm overnight.
- Wipe down all work surfaces with 70% ethanol after handling P. aeruginosa.
- Day 3: Store mutant of interest in -80 °C freezer
- Label cryovial with mutant name, antibiotic added to LB broth and date of storage.
- Prepare 500 mL of 50% Glycerol (v/v) and sterilize in autoclave.
- Wear gloves, lab coat, and mask to handle P. aeruginosa.
- Clear bench and/or laminar flood hood and wipe surface with 70% ethanol.
- Remove tube containing mutant culture from shaker.
- Prepare a small container with dry ice.
- Use a laminar flow hood or create a sterile field on ethanol-wiped bench top with Bunsen burner.
- Add equal amounts of bacterial culture and 50% glycerol to the labeled cryovial using sterile conditions, and gently mix with pipette (final volume 1 - 2 mL/vial depending on the size of the cryovials used). Place cryovial on dry ice to quick-freeze.
- Place cryovial in labeled box in -80 °C freezer.
- Wipe down all work surfaces with 70% ethanol after handling P. aeruginosa.
3. Protocol III: Quality Control of PA14NR Set
- Select random set of mutants from newly replicated plates to detect possible interwell contamination (testing of 30-40 mutants is recommended).
NOTE: In cases where it is necessary to confirm the identity of a mutant being used for characterization of a specific gene, it is recommended to perform PCR amplifications using gene-specific primers designed for the known sequence of the gene that contains the transposon insertion. Although more challenging, the benefits of using arbitrary PCR primers rather than gene-specific PCR primers when amplyfing DNA fragments from transposon mutants include ease of large-scale mutant confirmation and the ability to detect the presence of potential contaminants. For quality control purposes, it is not necessary to obtain high quality PCR sequencing data for all randomly slected mutants, as long as a sufficient number of mutants are screened to assess error rate. - Follow "Protocol II" to streak and grow mutant strains.
- Autoclave all plasticware and non-sterile supplies before use.
- Isolate genomic DNA by preferred method. For the analysis described in this work, a genomic DNA isolation kit was utilized following manufacturer's protocols. Multiple mutant strains may be analyzed simultaneously.
- Measure genomic DNA concentration using a microvolume spectrophotometer. Adjust genomic DNA concentration to approximately 100 ng/µL.
- Use genomic DNA as template to run "PCR1 reaction" as described in Step 1 of Table 3, generating Arb1 PCR fragments (Figure 3). Primers for this step are listed in Table 4.
- Add 0.5 µL of 10x loading buffer to 5 µL of PCR1 reaction, load into a 1.5-2% agarose gel and run the gel at 80-150 V.
NOTE: A specific fragment length or a particular fragment range is not expected, and more than one band may be present as the arbitrary primer may bind to multiple locations. - Use DNA from PCR1 reaction as template to run "PCR2 reaction" as described in Step 2 of Table 3, generating Arb2 PCR fragments (Figure 3). Primers for this step are listed in Table 4.
- Add 0.5 µL of 10x loading buffer to 5 µL of PCR2 reaction, load into a 1.5-2% agarose gel, and run the gel at 80-150 V.
NOTE: A specific fragment length or a particular fragment range is not expected, and more than one band may be present as the arbitrary primer may bind to multiple locations. - Send PCR2 reaction along with appropriate transposon-specific primer for sequencing.
- Analize sequencing results byBLASTing them against the complete PA14 genome using the BLAST link provided on the PA14NR Library website (http://pa14.mgh.harvard.edu/cgi-bin/pa14/blast.cgi) or by directly BLASTing them against the specific gene sequence of the mutant of interest.
NOTE: Information about selected mutants can be found by searching the PA14NR Set website (search http://pa14.mgh.harvard.edu/cgi-bin/pa14/search.cgi?searchType=SEARCH_PLATE_POSITIONS or download the Nonredundant Library.xls file from the link http://pa14.mgh.harvard.edu/cgi-bin/pa14/downloads.cgi).
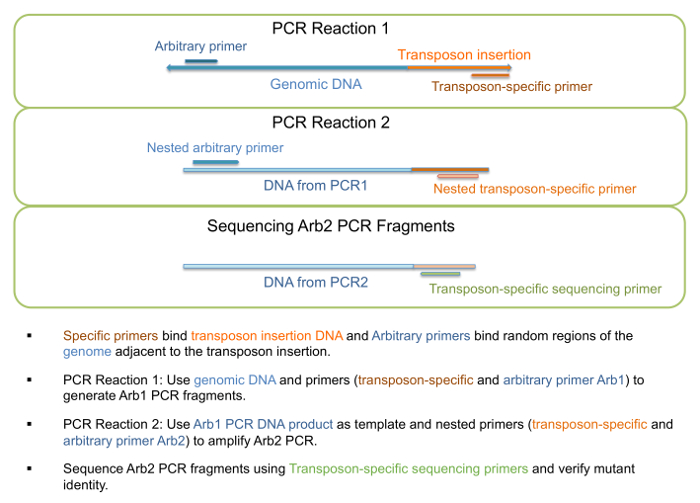
Figure 3: PCR amplification and sequencing of transposon insertion mutants. Schematic view of steps involved in PCR amplification and sequencing for verification of mutant identity. Please click here to view a larger version of this figure.
| PCR Reaction Set-up | |
| Step 1 | Step 2 |
| PCR1 reaction: | PCR2 reaction: |
| 23.25 µL water (Molecular Biology grade) | 19.15 µL Water (Molecular Biology Grade) |
| 3µL 10x Taq polymerase Buffer | 5 µL 10x Taq Polymerase Buffer |
| 0.5µL Taq polymerase | 0.6 µL Taq Polymerase |
| 0.625 µL 20 µM Primer Arb1D (Table 4) | 0.625 µL 20 µM Primer Arb2A (Table 4) |
| 0.625 µL 20 µM Transposon-specific Primer (PMFLGM.GB 3a or Tn5Ext ) (Table 4) | 0.625 mL 20 µM Transposon-specific Primer (PMFLGM.GB 2a or Tn5Int2) (Table 4) |
| 1 µL 10 mM dNTPs | 1 µL 10 mM dNTPs |
| 1 µL genomic DNA, 100 ng | 5 µL PCR1 reaction |
| 30 µL Final reaction volume | 30 µL Final reaction volume |
| PCR Reaction Settings | |
| PCR1 Thermocycler Conditions: | PCR2 Thermocycler Conditions: |
| 95 °C – 2 min | 95 °C – 2 min |
| Repeat 5 cycles: | Repeat 30 cycles: |
| 95 °C – 30 s | 95 °C – 30 s |
| 30 °C – 1 min | 54 °C – 30 s |
| 72 °C – 1 min | 72 °C – 1.5 min |
| Repeat 30 cycles: | 72 °C – 10 min |
| 95 °C – 30 s | 4 °C – Hold |
| 45 °C – 30 s | |
| 72 °C – 1 min | |
| 72 °C – 10 min | |
| 4°C – Hold |
Table 3: PCR reaction set-up and thermocycler conditions used for Arbitrary PCR. Arbitrary PCR reactions are performed sequentially, and fragments generated during PCR1 reaction are used as template in PCR2 reaction. Specific thermocycler settings are used for each set of reactions.
| Primer Name | Primer Sequence |
| MAR2xT7 Transposon-specific Primers | |
| PMFLGM.GB-3a | TACAGTTTACGAACCGAACAGGC |
| PMFLGM.GB-2a | TGTCAACTGGGTTCGTGCCTTCATCCG |
| MAR2xT7 Transposon Sequencing Primer | |
| PMFLGM.GB-4a | GACCGAGATAGGGTTGAGTG |
| TnphoA Transposon-specific Primers | |
| Tn5Ext | GAACGTTACCATGTTAGGAGGTC |
| Tn5Int2 | GGAGGTCACATGGAAGTCAGATCCTGG |
| TnphoA Transposon Sequencing Primer | |
| Tn5Int | CGGGAAAGGTTCCGTTCAGGACGC |
| Arbitrary Primers | |
| ARB1D | GGCCAGGCCTGCAGATGATGNNNNNNNNNNGTAT |
| ARB2A | GGCCAGGCCTGCAGATGATG |
Table 4: List of Primers used in Quality Control Experiments. Primers used for PCR amplification and sequencing of transposon insertion mutants to confirm mutant identity.
Wyniki
Twelve new copies of the PA14NR Set were replicated using Protocol I, and a quality control assessment of the new copies generated was conducted using Protocol III.
PA14NR Set mutant plates along with Control Plates, which consist of wild type PA14 inoculated and uninoculated wells intercalated in a preset pattern (Figure 4A), were replicated following methology described in Protocol I. Control plat...
Dyskusje
The P. aeruginosa PA14NR Set is a valuable resource for the scientific community. According to the March 2017 dataset from Clarivate Analytics' Essential Science Indicators database, Liberati et al. (2006)37, which describes the construction of the PA14NR Set, is ranked in the top 1% of microbiology publications. Google Scholar reports over 600 citations of the Liberati et al. (2006) original manuscript as of August 2017. The library has played an important ...
Ujawnienia
The authors report no financial conflicts of interest. Eliana Drenkard and Frederick Ausubel participated in the creation the PA14 nonredundant transposon mutant library. Bryan Hurley and Lael Yonker currently house and distribute the mutant library as part of the Department of Pediatrics at Massachusetts General Hospital.
Podziękowania
We would like to thank Lisa Philpotts of the MGH Treadwell Virtual Library for her guidance in the database search. This work was supported by the Cystic Fibrosis Foundation (YONKER16G0 and HURLEY16G0) and NIH NIAID (BPH and ADE: R01 A1095338).
Materiały
| Name | Company | Catalog Number | Comments |
| Materials for Library Replication | |||
| Sterile 96-well Tissue-culture treated, case of 50 | Corning Life Sciences | 353072 | via Fisher Scientific |
| Sterile 96 Well Clear V-Bottom 2000μL Deep Well Plates, case of 25 | Corning Life Sciences | 3960 | via Fisher Scientific |
| Nunc OmniTray (rectangular plates), case of 60 | Thermo Scientific Rochester | 242811 | via Fisher Scientific |
| Rectangular Ice Pan, Midi (4L) | Corning Life Sciences | 432104 | via Fisher Scientific |
| Secure-Gard Cone Mask, case of 300 | Cardinal Health | AT7509 | via Fisher Scientific |
| AluminaSeal, pack of 100 | Diversified Biotech | ALUM-100 | via Fisher Scientific |
| Breathe-Easy membrane, pack of 100 | Diversified Biotech | BEM-1 | via Sigma-Aldrich |
| Sterile, individually wrapped, 50mL Solution Trough/Reagent Reservoir, case of 100 | Sorenson | S50100 | via Westnet Incorporated |
| Plate roller | VWR | 60941-118 | via VWR |
| Cryo Laser Labels - CRYOLAZRTAG 2.64" x 0.277", pack of 16 sheets | GA International | RCL-11T1-WH | via Labtag.com (template for printing also available from Labtag.com) |
| 96-well replicator | V & P Scientific, Inc. | Custom 407C, 3.18mm pin diameter, 57mm long | via V & P Scientific, Inc. |
| Multitron Pro, 3mm Shaking incubator | Infors HT | l10003P | via Infors HT |
| Picus 12 Channel 50-1200μL Electronic Pipette | Sartorius | 735491PR | via Sartorius |
| Filter Tips 50-1200μL, pack of 960 | Biohit | 14-559-512 | via Fisher Scientific; use electronic multichannel-compatible tips |
| Dry Ice | User-specific vendor | ||
| Materials for Individual Mutant Storage | |||
| Fisherbrand Premium Microcentrifuge Tubes: 1.5mL | Fisher Scientific | 05-408-130 | via Fisher Scientific |
| Pipettes (P1000, P200, P20, P2) | Gilson | F167370 | via Gilson |
| Materials for Quality Control PCR | |||
| Fisherbrand Premium Microcentrifuge Tubes: 1.5mL | Fisher Scientific | 05-408-130 | via Fisher Scientific |
| NanoDrop | Thermo Scientific | ND-2000 | via ThermoFisher |
| PCR Thermocycler | |||
| Omnistrips PCR Tubes with domed lids | Thermo Scientific | AB0404 | via Fisher Scientific |
| ART Barrier low-retention pipette tips (10 uL, 100 uL, 1000 uL) | Molecular BioProducts, Inc. | Z676543 (10 uL), Z676713 (100 uL), Z676802 (1000 uL) | via Sigma-Aldrich |
| Pipettes (P1000, P200, P20, P2) | Gilson | F167370 | via Gilson |
| Fisherbrand Premium Microcentrifuge Tubes: 1.5mL | Fisher Scientific | 05-408-130 | via Fisher Scientific |
| MasterPure DNA Purification Kit | Epicentre | MCD85201 | via Epicentre Technologies Corp |
| GeneRuler 1 kb Plus DNA Ladder, ready-to-use | Thermo Scientific | SM1333 | via ThermoFisher |
| RediLoad Loading Buffer | Invitrogen | 750026 | via ThermoFisher |
| Chemicals | |||
| Chemicals for Library and Individual Mutant Storage | |||
| Glycerol MB Grade, 1L | Sigma Aldrich | G5516 | via Sigma-Aldrich |
| LB Broth | Per 1L dH2O: 10g tryptone, 5g yeast extract, 5g NaCl, 1ml 1N NaOH (Current Protocols in Molecular Biology. Wiley, 1994.) | ||
| Tryptone | Sigma Aldrich | T7293 | via Sigma-Aldrich |
| Yeast Extract | Sigma Aldrich | Y1625 | via Sigma-Aldrich |
| Sodium Chloride | Sigma Aldrich | S7653 | via Sigma-Aldrich |
| Sodium Hydroxide | Sigma Aldrich | S8045 | via Sigma-Aldrich |
| LB agar | See preparation above, add 15g Bacto Agar | ||
| Bacto Agar | Sigma Aldrich | A5306 | via Sigma-Aldrich |
| Gentamicin sulfate, 10g | BioReagent | 1405-41-0 | via Sigma-Aldrich |
| Kanamycin sulfate | Gibco | 11815024 | via ThermoFisher |
| Ethanol, 190 proof | Decon | 04-355-221 | via Fisher Scientific |
| Chemicals for Quality Control PCR | |||
| Primers | User-preferred vendor | See primers listed in Table 3 | |
| Corning cellgro Molecular Biology Grade Water | Corning | 46000CV | via Fisher Scientific |
| Taq Polymerase Buffer | Invitrogen | 10342020 | via ThermoFisher |
| Taq DNA Polymerase, recombinant | Invitrogen | 10342020 | via ThermoFisher |
| dNTPs | Invitrogen | 10297018 | via ThermoFisher |
| Agarose | Sigma | A9539 | via Sigma-Aldrich |
Odniesienia
- Moradali, M. F., Ghods, S., Rehm, B. H. Pseudomonas aeruginosa lifestyle: A paradigm for adaptation, survival, and persistence. Front Cell Infect Microbiol. 7, 39 (2017).
- Bleves, S., et al. Protein secretion systems in Pseudomonas aeruginosa: A wealth of pathogenic weapons. Int J Med Microbiol. 300 (8), 534-543 (2010).
- Breidenstein, E. B., de la Fuente-Nunez, C., Hancock, R. E. Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol. 19 (8), 419-426 (2011).
- Flynn, K. M., et al. Evolution of ecological diversity in biofilms of Pseudomonas aeruginosa by altered cyclic diguanylate signaling. J Bacteriol. 198 (19), 2608-2618 (2016).
- Hazan, R., Maura, D., Que, Y. A., Rahme, L. G. Assessing Pseudomonas aeruginosa persister/antibiotic tolerant cells. Methods Mol Biol. 1149, 699-707 (2014).
- Klockgether, J., et al. Genome diversity of Pseudomonas aeruginosa PAO1 laboratory strains. J Bacteriol. 192 (4), 1113-1121 (2010).
- Mathee, K., et al. Dynamics of Pseudomonas aeruginosa genome evolution. Proc Natl Acad Sci U S A. 105 (8), 3100-3105 (2008).
- Taylor, P. K., Yeung, A. T., Hancock, R. E. Antibiotic resistance in Pseudomonas aeruginosa biofilms: towards the development of novel anti-biofilm therapies. J Biotechnol. 191, 121-130 (2014).
- Flume, P. A., Van Devanter, D. R. State of progress in treating cystic fibrosis respiratory disease. BMC Med. 10, 88 (2012).
- . Cystic Fibrosis Foundation Patient Registry 2015 Annual Data Report Available from: https://www.cff.org/Our-Research/CF-Patient-Registry/2015-Patient-Registry-Annual-Data-Report.pdf (2016)
- Church, D., Elsayed, S., Reid, O., Winston, B., Lindsay, R. Burn wound infections. Clin Microbiol Rev. 19 (2), 403-434 (2006).
- Sadikot, R. T., Blackwell, T. S., Christman, J., Prince, A. Pathogen-host interactions in Pseudomonas aeruginosa pneumonia. Am J Respir Crit Care Med. 171 (11), 1209-1223 (2005).
- Song, Z., et al. Prosthesis infections after orthopedic joint replacement: the possible role of bacterial biofilms. Orthop Rev (Pavia). 5 (2), 65-71 (2013).
- National Nosocomial Infections Surveillance, S. National Nosocomial Infections Surveillance (NNIS) System Report, data summary from January 1992 through June 2004, issued October 2004. Am J Infect Control. 32 (8), 470-485 (2004).
- Cohen, T. S., Parker, D., Prince, A. . Pseudomonas aeruginosa Host Immune Evasion. 7, 3-23 (2014).
- Fernandes, A., Dias, M. The microbiological profiles of infected prosthetic implants with an emphasis on the organisms which form biofilms. J Clin Diagn Res. 7 (2), 219-223 (2013).
- Khosravi, A. D., Ahmadi, F., Salmanzadeh, S., Dashtbozorg, A., Montazeri, E. A. Study of Bacteria Isolated from Orthopedic Implant Infections and their Antimicrobial Susceptibility Pattern. Res J of Microbiol. 4 (4), 6 (2009).
- Roemhild, R., Barbosa, C., Beardmore, R. E., Jansen, G., Schulenburg, H. Temporal variation in antibiotic environments slows down resistance evolution in pathogenic Pseudomonas aeruginosa. Evol Appl. 8 (10), 945-955 (2015).
- Fischer, S., et al. Intraclonal genome diversity of the major Pseudomonas aeruginosa clones C and PA14. Environ Microbiol Rep. 8 (2), 227-234 (2016).
- Wiehlmann, L., et al. Population structure of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A. 104 (19), 8101-8106 (2007).
- Lam, J. S., Taylor, V. L., Islam, S. T., Hao, Y., Kocincova, D. Genetic and functional diversity of Pseudomonas aeruginosa lipopolysaccharide. Front Microbiol. 2, 118 (2011).
- Choi, J. Y., et al. Identification of virulence genes in a pathogenic strain of Pseudomonas aeruginosa by representational difference analysis. J Bacteriol. 184 (4), 952-961 (2002).
- He, J., et al. The broad host range pathogen Pseudomonas aeruginosa strain PA14 carries two pathogenicity islands harboring plant and animal virulence genes. Proc Natl Acad Sci U S A. 101 (8), 2530-2535 (2004).
- Mikkelsen, H., McMullan, R., Filloux, A. The Pseudomonas aeruginosa reference strain PA14 displays increased virulence due to a mutation in ladS. PLoS One. 6 (12), e29113 (2011).
- Drenkard, E., Ausubel, F. M. Pseudomonas biofilm formation and antibiotic resistance are linked to phenotypic variation. Nature. 416 (6882), 740-743 (2002).
- Rahme, L. G., et al. Common virulence factors for bacterial pathogenicity in plants and animals. Science. 268 (5219), 1899-1902 (1995).
- Rahme, L. G., et al. Use of model plant hosts to identify Pseudomonas aeruginosa virulence factors. Proc Natl Acad Sci U S A. 94 (24), 13245-13250 (1997).
- Kirienko, N. V., Cezairliyan, B. O., Ausubel, F. M., Powell, J. R. Pseudomonas aeruginosa PA14 pathogenesis in Caenorhabditis elegans. Methods Mol Biol. 1149, 653-669 (2014).
- Mahajan-Miklos, S., Tan, M. W., Rahme, L. G., Ausubel, F. M. Molecular mechanisms of bacterial virulence elucidated using a Pseudomonas aeruginosa-Caenorhabditis elegans pathogenesis model. Cell. 96 (1), 47-56 (1999).
- Limmer, S., et al. Pseudomonas aeruginosa RhlR is required to neutralize the cellular immune response in a Drosophila melanogaster oral infection model. Proc Natl Acad Sci U S A. 108 (42), 17378-17383 (2011).
- Miyata, S., Casey, M., Frank, D. W., Ausubel, F. M., Drenkard, E. Use of the Galleria mellonella caterpillar as a model host to study the role of the type III secretion system in Pseudomonas aeruginosa pathogenesis. Infect Immun. 71 (5), 2404-2413 (2003).
- Coleman, F. T., et al. Hypersusceptibility of cystic fibrosis mice to chronic Pseudomonas aeruginosa oropharyngeal colonization and lung infection. Proc Natl Acad Sci U S A. 100 (4), 1949-1954 (2003).
- Pazos, M. A., et al. Pseudomonas aeruginosa ExoU augments neutrophil transepithelial migration. PLoS Pathog. 13 (8), e1006548 (2017).
- Maura, D., Hazan, R., Kitao, T., Ballok, A. E., Rahme, L. G. Evidence for direct control of virulence and defense gene circuits by the Pseudomonas aeruginosa quorum sensing regulator, MvfR. Sci Rep. 6, 34083 (2016).
- Jacobs, M. A., et al. Comprehensive transposon mutant library of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A. 100 (24), 14339-14344 (2003).
- Held, K., Ramage, E., Jacobs, M., Gallagher, L., Manoil, C. Sequence-verified two-allele transposon mutant library for Pseudomonas aeruginosa PAO1. J Bacteriol. 194 (23), 6387-6389 (2012).
- Liberati, N. T., et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc Natl Acad Sci U S A. 103 (8), 2833-2838 (2006).
- Feinbaum, R. L., et al. Genome-wide identification of Pseudomonas aeruginosa virulence-related genes using a Caenorhabditis elegans infection model. PLoS Pathog. 8 (7), e1002813 (2012).
- Stewart, L., et al. Draft genomes of 12 host-adapted and environmental isolates of Pseudomonas aeruginosa and their positions in the core genome phylogeny. Pathog Dis. 71 (1), 20-25 (2014).
- Thrane, S. W., et al. The widespread multidrug-resistant serotype O12 Pseudomonas aeruginosa clone emerged through concomitant horizontal transfer of serotype antigen and antibiotic resistance gene clusters. MBio. 6 (5), e01396-e01315 (2015).
- van Belkum, A., et al. Phylogenetic Distribution of CRISPR-Cas Systems in Antibiotic-Resistant Pseudomonas aeruginosa. MBio. 6 (6), e01796-e01715 (2015).
- Lewenza, S., et al. Construction of a mini-Tn5-luxCDABE mutant library in Pseudomonas aeruginosa PAO1: a tool for identifying differentially regulated genes. Genome Res. 15 (4), 583-589 (2005).
- . . Current Protocols in Molecular Biology. , (1994).
- Breidenstein, E. B., Khaira, B. K., Wiegand, I., Overhage, J., Hancock, R. E. Complex ciprofloxacin resistome revealed by screening a Pseudomonas aeruginosa mutant library for altered susceptibility. Antimicrob Agents Chemother. 52 (12), 4486-4491 (2008).
- Musken, M., Di Fiore, S., Dotsch, A., Fischer, R., Haussler, S. Genetic determinants of Pseudomonas aeruginosa biofilm establishment. Microbiology. 156 (Pt 2), 431-441 (2010).
- Schurek, K. N., et al. Novel genetic determinants of low-level aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 52 (12), 4213-4219 (2008).
- Oumeraci, T., et al. Comprehensive MALDI-TOF biotyping of the non-redundant Harvard Pseudomonas aeruginosa PA14 transposon insertion mutant library. PLoS One. 10 (2), e0117144 (2015).
- Yeung, A. T., et al. Swarming of Pseudomonas aeruginosa is controlled by a broad spectrum of transcriptional regulators, including MetR. J Bacteriol. 191 (18), 5592-5602 (2009).
Przedruki i uprawnienia
Zapytaj o uprawnienia na użycie tekstu lub obrazów z tego artykułu JoVE
Zapytaj o uprawnieniaThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. Wszelkie prawa zastrzeżone