Method Article
Метод для измерения РНК
В этой статье
Резюме
Модифицированный северный блоттинг метод измерения N 6 -methyladenosine (м 6 а) изменения в РНК описана. Текущий метод может обнаружить изменения в различных РНК и элементов управления в различных экспериментальных конструкций.
Аннотация
N6-Methyladenosine (m6A) modifications of RNA are diverse and ubiquitous amongst eukaryotes. They occur in mRNA, rRNA, tRNA, and microRNA. Recent studies have revealed that these reversible RNA modifications affect RNA splicing, translation, degradation, and localization. Multiple physiological processes, like circadian rhythms, stem cell pluripotency, fibrosis, triglyceride metabolism, and obesity are also controlled by m6A modifications. Immunoprecipitation/sequencing, mass spectrometry, and modified northern blotting are some of the methods commonly employed to measure m6A modifications. Herein, we present a northeastern blotting technique for measuring m6A modifications. The current protocol provides good size separation of RNA, better accommodation and standardization for various experimental designs, and clear delineation of m6A modifications in various sources of RNA. While m6A modifications are known to have a crucial impact on human physiology relating to circadian rhythms and obesity, their roles in other (patho)physiological states are unclear. Therefore, investigations on m6A modifications have immense possibility to provide key insights into molecular physiology.
Введение
Dynamic and reversible RNA modifications have important roles in RNA homeostasis. Four decades ago, N6-methyladenosine (m6A) modifications were found to be abundant in eukaryotic transcriptomes1. They have diverse functions in messenger RNA (mRNA), ribosomal RNA (rRNA), small nucleolar RNA, transfer RNA, and microRNA2. The m6A modifications of mRNA influence their splicing3, translation4, degradation5, and localization2. Moreover, they affect ribosome biogenesis and microRNA function6. The evolutionary conservation of m6A modifications of RNA is noted in unicellular bacteria to multi-cellular humans7. Delineation of the roles of m6A modifications is currently under extensive exploration. The efforts are expected to provide new insights into transcription control. Recent studies reveal that other chemical modifications of mRNA8 also play critical roles in RNA metabolism.
Circadian rhythms9, stem cell pluripotency10, triglyceride metabolism4, fibrosis11, obesity12, and major depression13 are a few examples of processes where m6A modifications are known to control outcomes. Many circadian clock gene transcripts have m6A sites14. Modulation of m6A methylase or demethylase elicits circadian period changes15. Mettl3, an m6A transferase, is a regulator for stem cell pluripotency. A deficiency of Mettl3 leads to early embryonic lethality and aberrant lineage priming at the post-implantation stage10. A deficiency of fat mass- and obesity-associated (FTO), an m6A demethylase, in adipocytes affects fatty acid mobilization and body weight through posttranscriptional regulation of Angptl44. These studies reveal that m6A not only controls mRNA processing, but also plays critical roles in embryological development and patho-physiology. The function of m6A modifications holds implications for therapeutic considerations in the future.
Several methods are available to measure m6A modifications of RNA16-18. Traditionally, thin layer chromatography (TLC) and high-performance liquid chromatography (HPLC) are used to study the distribution of m6A in several RNAs19,20. Mass spectrometry is a sensitive tool for the detection of m6A modifications in RNA. However, the RNA needs to be excised by RNase into short fragments before analysis by mass spectrometry21. Methyl-RNA immunoprecipitation and sequencing (m6A-Seq)22 immunoprecipitates fragmented RNAs with m6A-specific antibodies and performs parallel RNA sequencing. This method generates transcriptome-wide m6A landscapes. High-resolution mapping of m6A individual-nucleotide-resolution cross-linking and immunoprecipitation (miCLIP) further maps m6A modifications at a single-nucleotide resolution23. Both methods provide details of m6A modification across the whole transcriptome, with specific genes' information. However, quantifications and standardization in both methods are difficult if experiments require the comparison of multiple conditions. Moreover, fragmentations of RNA for m6A-Seq alter the original RNA structure, which may affect native m6A levels. To detect global m6A modifications of RNA and their changes under different experimental conditions, we report a method that employs a modified northern blotting protocol. This method resolves RNA by molecular weight, using gel electrophoresis18. This procedure provides better standardization and quantifications for experiments that involve multiple conditions or samples. It also provides specific m6A modification information for different RNAs, whether rRNA, mRNA, or microRNA.
протокол
Примечание: Общую РНК м 6 Уровень включает рРНК, мРНК и другие малые РНК. Так как рибосомальной РНК имеет обильные м 6 модификаций А, измерение м 6 уровней А необходимо будет рассмотреть этот факт.
Выделение 1. РНК
- Используя решение рибонуклеазы дезактивации (разбрызгивают на бумажные полотенца), протрите пипеток и лабораторного стола поверхности. Влажные бумажные полотенца с нуклеазы без воды и протирать пипеток и лабораторного стола поверхности снова.
- Экстракция РНК
- Общая Выделение РНК
- С помощью верхней мешалки, гомогенизируют 50-100 мг ткани или 5-10 х 10 6 клеток в 1 мл раствора для выделения РНК выдерживали при 4 ° С.
Примечание: больше ткани (100-150 мг), которые могут потребоваться для образцов жировой ткани мышей из-за более низкой концентрации РНК в них. Образцы получены из сердца мыши, печени, скелетных мышцах, легких, головного мозга, и макрофаги были использованы ранее 4. - Вcubate выборочные гомогенатов в течение 5 мин при комнатной температуре.
- Добавляют 100 мкл 1-бром-3-хлорпропана (БКП) на 1 мл образца гомогената. Встряхнуть пробирки энергично в течение 15 секунд и приступить к изоляции общей РНК в соответствии с инструкциями изготовителя 24.
- С помощью верхней мешалки, гомогенизируют 50-100 мг ткани или 5-10 х 10 6 клеток в 1 мл раствора для выделения РНК выдерживали при 4 ° С.
- Полиаденилированная мРНК Очистка от изолированной Общую РНК
- Очищают Полиаденилированная мРНК с использованием доступных наборов или протоколов.
- Тщательно взболтать, чтобы вновь приостановить каждое из следующих решений: 2x связывающего раствора, олиго (дТ) полистирольные шарики и промывочный раствор. Убедитесь, что олиго (дТ) бусин нагреться до комнатной температуры перед использованием.
- Перенести 120 мкл элюирующего раствора на препарата в пробирку и нагревают до 70 ° С в нагревательном блоке.
- Пипетировать до 500 мкг общей РНК в микроцентрифужных трубки. С помощью нуклеазы без воды, регулировать громкость до 250 мкл.
- Добавьте 250 мкл раствора связывающего 2x до полной РНК, таклюция и вихрь коротко перемешать содержимое.
- Добавьте 15 мкл олиго (дТ) бусы и вихрь тщательно перемешать.
- Выдержите смесь при температуре 70 ° С в течение 3 мин.
- Извлеките образец из нагревательного блока и дать постоять при комнатной температуре в течение 10 мин.
- Центрифуга при 15000 мкг в течение 2 мин.
- Осторожно удалите супернатант, оставляя позади приблизительно 50 мкл.
- Добавьте 500 мкл промывочного раствора повторно приостанавливать осадок пипетированием.
- Пипетировать суспензии в набор спинового фильтра / сбора трубки.
- Центрифуга при 15000 мкг в течение 2 мин. Удалить столбец из коллекторной трубки и отбросить проточные. Вернуть колонку в пробирку.
- Пипетка 500 мкл промывочного раствора на спиновый фильтр.
- Центрифуга при 15000 х г в течение 2 мин.
- Перенести спиновый фильтр в новую пробирку для сбора.
- Пипетка 50 мкл элюирующего раствора нагреваютдо 70 ° С на центр спинового фильтра.
- Выдержите в течение 2-5 мин при температуре 70 ° С. Центрифуга при 15000 мкг в течение 1 мин.
- Пипетировать дополнительно 50 мкл элюирующего раствора нагревают до 70 ° С на центр спинового фильтра.
- Повторите шаг 1.2.2.1.18.
- Добавить 100 мкг / мл гликогена, 0,1 объема 3 М буфере ацетата натрия (рН 5,2) и 2,5 объема абсолютного этанола. Осадок в течение ночи при -80 ° С.
- Центрифуга при 15000 х г в течение 25 мин при температуре 4 ° С. Удалите супернатант.
- Промывают осадок с 1 мл 75% -ного этанола. Центрифуга при 15000 х г в течение 15 мин при температуре 4 ° С. Осторожно удалите этанол.
- Сушить в воздухе в течение 3-5 мин.
- Добавьте 10 мкл нуклеазы без воды, чтобы растворить осадок.
- Хранить при температуре -70 ° С.
- Очищают Полиаденилированная мРНК с использованием доступных наборов или протоколов.
- Общая Выделение РНК
- РНК Квалификация и Количественная
- С помощью спектрофотометра, определяют концентрацию РНК в присутствии NotIнг оптическую плотность при длине волны 260 нм и 280 нм. Отношение А 260 / А 280 должно быть 1,8-2,2.
- Проверка качества образцов РНК с использованием 0,8% агарозном геле 25.
ПРИМЕЧАНИЕ: неповрежденными эукариотической суммарную РНК должны показать интенсивные полосы 28S и 18S рРНК. Отношение 28S в сравнении интенсивности полос 18S рРНК для хорошей подготовки РНК составляет ~ 2.
2. Гель электрофорез и передачи
Примечание: Протоколы подготовки буфера приведены в таблице 1.
- Формальдегид гель Приготовление (1%)
- Промыть все оборудование электрофорез с диэтилпирокарбонатом (DEPC) водой.
- Melt 2,5 г агарозы в 215 мл воды DEPC полностью в микроволновой печи.
- Добавить 12,5 мл 10х 3- (N - морфолино) пропансульфоновой кислоты (MOPS) буфер и 22,5 мл 37% -ного формальдегида.
- Налейте его в аппарате для электрофореза с толстым гребнем 1,5 мм.
- Устранить пузырьки или толкать трубчик к краям геля с чистым гребнем.
- Дайте гель затвердеть при комнатной температуре.
- Базовые приготовления
- Смешайте 1-10 мкг общей РНК и 11,3 мкл буфера для образцов.
- Смешайте 2 мкг РНК-маркера (1 мкг / мкл) 11,3 мкл буфера для образцов.
- Добавить нуклеазы без воды к образцам из 2.2.1, 2.2.2 до общего объема 16 мкл.
- Нагревают при 60 ° С в течение 5 минут, а затем охладить на льду.
- Смешайте 16 мкл образца из 2.2.3 с 4 мкл отслеживания красителя, который содержит 0,1 мкг / мкл этидийбромид на льду.
ВНИМАНИЕ: этидий бромид, возможно, тератогенным и токсичен при вдыхании. Read этидий бромид технический паспорт безопасности.
- электрофорез
- Добавить 200 мл 10х MOPS буфера до 1800 мл DDH 2 O , чтобы сделать 1x MOPS работает буфера.
- Используйте 1x MOPS работает буфера для промывки лунок геля.
- Предварительно запуск тон гель при 20 В в течение 5 мин.
- Загрузка образцов РНК в лунки геля.
- Покрытие с фольгой, чтобы избежать попадания света. Запуск на 35 В в течение приблизительно 17 ч (в течение ночи)
- Осмотрите и сфотографируйте гель под УФ-светом.
- Перевод
- Вырезать гель для удаления неиспользованную часть.
- Приготовьте 500 мл 10х SSC-буфере (250 мл 20x SSC буфера и 250 мл DEPC воды).
- Вымойте гель дважды с 10-кратным SSC буфером в течение 20 мин при встряхивании при 50 оборотах в минуту.
- Вырезать 1 бумажный фильтр лист , достаточно большой , чтобы служить в качестве фитиля бумаги, впитывающей буфера переноса с передающего лотка (рисунок 1).
- Вырезать 4 куска фильтровальной бумаги такого же размера, что и гель.
- Порезать 1 лист положительно заряженную нейлоновую мембрану в тот же размер, что и гель.
- Отметить насечку на геле и мембрану в качестве маркера, чтобы обеспечить правильную ориентацию.
- Замочите мембрану в 2х буфера SSC в течение 15 мин.
- Залить 500 мл оF 10x SSC буфера в передаточную лоток.
- Положите большой фильтр бумажного листа через стеклянную пластину и смочить фильтровальную бумагу буфера переноса.
- Раскатать все пузырьки с пипеткой.
- Замочить 2 куска фильтровальной бумаги предварительно вырезанное в буфере передачи и поместить их в центр фильтровальной бумаги со стадии 2.4.5. Удалите все пузырьки.
- Уложить гель верхняя сторона была обращена вниз на фильтровальную бумагу.
- Lay мембрану поверх геля. Совместите вырезы геля и мембраны.
- Поместите 2 куска фильтровальной бумаги предварительно вырезанное на верхней стороне мембраны и смочить фильтровальную бумагу буфера переноса.
- Удалите все пузырьки между слоями.
- Нанести пластмассовую обертку вокруг геля, чтобы гарантировать, что полотенца только впитывать буфер, проходящий через гель и мембрану.
- Стек бумажные полотенца на верхней части фильтровальной бумаги.
- Поместите негабаритный стеклянную пластину поверх полотенца.
- Поместите вес на верхней части стека.
- Держите в течение ночи, чтобы позволить РНК перенести из геля на мембрану.
3. Обнаружение N 6 -methyladenosine
- Мембрана Сшивание
- Держите мембрану влажной в буфер 2x SSC.
- Поместите 2 листа фильтровальной бумаги, погруженной с 10x SSC буфером в УФ сшивателя.
- Поместите мембрану в верхней части фильтровальной бумаги, чтобы сторона с РНК-адсорбированной лицевой стороной вверх.
- Выберите режим "autocrosslink" (120000 мкДж) и нажмите кнопку "Пуск", чтобы начать облучение.
- Осмотрите мембрану и гель с УФ-уловителя изображение геля, чтобы подтвердить перевод РНК из геля на мембрану.
- иммуноблоттинг
- Блок мембраны в 5% молоке обезжиренном в Трис-буферном солевом растворе с Tween 20 (TBST) буфере при комнатной температуре в течение 1 часа.
- Промыть трижды TBST буфером в течение 15 мин.
- Выдержите в течение ночи в мембрану м 6 А ( N 6 -methyladenosine) раствор антитела (1: 1000 в 5% обезжиренное молоко TBST буфер) при 4 ° С.
- Промывают мембрану трижды TBST буфером в течение 15 мин.
- Инкубируйте мембраны в растворе конъюгированным с пероксидазой хрена ослиного антителам кролика (1: 2000 в 5% обезжиренном буфера молока TBST) в течение 1 ч при комнатной температуре.
- Промывают мембрану трижды TBST буфером в течение 15 мин.
- Примените расширенный хемилюминесцентный субстрат (0,125 мл на см 2 мембраны).
- Захват хемилюминесценции с оптимальными настройками цифрового формирования изображений, в соответствии с инструкциями изготовителя 26.
- м 6 A Количественное
- Измерьте относительную м 6 хемилюминесцентный интенсивности с использованием программного обеспечения ImageJ.
- В меню программного обеспечения ImageJ, выберите опцию "Файл", чтобы открыть соответствующую файл.
- Выберите "Прямоугольник" инструмент из ImageJ и нарисуйте рамку вокругсигналы.
- Если рибосомной РНК не является предметом экспериментов, избежать полос 18S и 28S рРНК м 6 А для расчета.
- Нажмите кнопку "Command" и "1", чтобы подтвердить выбранную полосу.
- Нажмите кнопку "Command" и "3", чтобы показать выбранный участок.
- Нажмите на "прямой" инструмент и рисовать линии, чтобы отделить сегментированный область.
- Нажмите кнопку "Wand" инструмент для записи измерений.
- Экспорт данных.
- Нормализовать уровень м 6 А с 18S рибосомальной РНК из группы окрашивания бромистым этидием.
- Измерьте относительную м 6 хемилюминесцентный интенсивности с использованием программного обеспечения ImageJ.
Результаты
Через 14 дней в нормальной свет-темнота циркадного фазы, мыши дикого типа, были помещены в постоянной темноте. РНК из печени отбирали каждые 4 ч и изучали с модифицированной северной блоттинга. Метилирование рРНК, мРНК, и малых РНК были четко обнаружены (рисунок 2). Сравнение между различными циркадных раза (КТ) могут быть точно вычислены со стандартом 18S рРНК. Был прочный циркадные колебания м 6 уровней А в рРНК, мРНК, и малых РНК.
Для того, чтобы избежать помех протянутую рРНК, Полиаденилированная РНК могут быть очищены, как на этапе 1.2.2. После очистки рРНК может быть в значительной степени устранены , чтобы позволить для лучшей визуализации других РНК (рисунок 3).
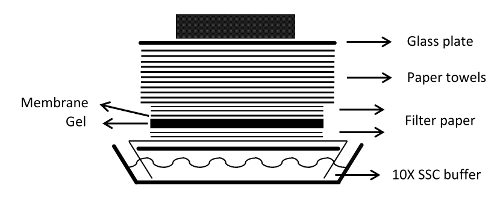
г> Рисунок 1: Сборка блока переноса блот. Блок переноса капиллярная для блоттинга РНК к мембране показана. Пожалуйста , нажмите здесь , чтобы посмотреть увеличенную версию этой фигуры.

Рисунок 2: Суточный ритм уровней м 6 А в C57BL / 6J. Эта цифра показывает 6 м пятном тотальной РНК из печени мышей дикого типа , забитых на первый день темно-темно - фазы через 14 дней нормального светло-темной фазы через 4 ч интервалами. Количественное было сделано с использованием м 6 разностное изображение плотности между 18S и 28S рРНК. М 6 А обилие нормализовалось к группе 18S рРНК из геля формальдегида в нижней части.и др = "_blank"> Пожалуйста, нажмите здесь, чтобы посмотреть увеличенную версию этой фигуры.
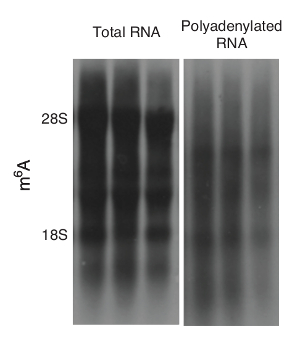
Рисунок 3: 6 м A кляксы с или без рРНК. Представитель м 6 A - блоты полной РНК и РНК полиаденилированой из печени мышей дикого типа показаны. Пожалуйста , нажмите здесь , чтобы посмотреть увеличенную версию этой фигуры.
| 1 | 10x МОПС буфер (рН 7,0, защищенном от света месте) | |||||
| МОПС | 41.85 | г | 0,2 М | |||
| Ацетат натрия | 4.1 | г | 0,05 М | |||
| ЭДТА, динатриевой соли | 3.7 | г | 0,01 М | |||
| DEPC воды | ||||||
| Всего | 1 | L | ||||
| Перемешивают при комнатной температуре | ||||||
| 2 | 20x SSC буфер (рН 7,0) | |||||
| хлористый натрий | 175,3 | г | 3 М | |||
| тринатрийцитрат | 88,3 | г | 0,3 М | |||
| DEPC воды | ||||||
| Всего | 1 | L | ||||
| Перемешивают при комнатной температуре, а затем автоклав | ||||||
| 3 | Отслеживание Dye | |||||
| 10x МОПС буфер | 500 | мкл | 1x | |||
| Ficoll 400 | 0,75 | г | 15% | |||
| бромфеноловый синий | 0,01 | г | 0,2% | |||
| ксиленцианол | 0,01 | г | 0,2% | |||
| DEPC воды | ||||||
| Всего | 5 | мл | ||||
| Хранить при температуре -20 ° C | ||||||
| 4 | DEPC воды | |||||
| Развести соотношение 1: 1000 из DEPC в DDH 2 O | ||||||
| Перемешивают в течение ночи при комнатной температуре, а затем автоклав | ||||||
| 5 | Пример буфера (защиты от света) | |||||
| 10x МОПС буфер | 200 | мкл | ||||
| 37% формальдегида | 270 | мкл | ||||
| формамид | 660 | мкл | ||||
| Хранить при температуре -20 ° C | ||||||
Таблица 1: Буферы и растворы.
Обсуждение
Modifications of RNA have important roles in cellular function and physiology. The current understanding of the regulation, function, and homeostasis of these modifications is still being explored and expanded8. Therefore, a precise and gold-standard method to evaluate the modifications of RNA is needed. The modified northern blotting method provides precise quantification of RNA modifications and clear delineation of the modifications in diverse RNAs. Although the method requires at least 3 days, it can be standardized and can be used in various experimental designs. Moreover, with different antibodies, it can detect different RNA modifications27.
It is important to separate different RNAs when analyzing RNA modifications. Ribosomal RNA comprises a large portion of the total amount of RNA28,29. The results from analyzing RNA modifications only in total RNA will represent mostly the changes of rRNA. Methylation and other such modifications of rRNA could potentially mask the changes in other RNAs. With the procedure of gel separation, the modifications of mRNA and other small RNAs can be more accurately analyzed.
Transcriptome-wide mapping with m6A immunoprecipitation and sequencing provides detailed insight into the modification of each type of RNA22. It provides information on the specific RNAs and a resolution of around 80-120 bp. Although m6A-Seq can compare the modifications between different experimental conditions, the selection of proper standards and controls for such experiments is difficult18. Immunoprecipitation is difficult to reproduce, often giving significant variations amongst repeats. Moreover, m6A-Seq requires the fragmentation of RNA samples before immunoprecipitation and sequencing30. The fragmentation process could potentially induce undue influences on the original RNA modifications. If the experiment does not need the specific gene's information but requires different conditions for comparison, the current method provides better visualization and control for diverse experimental setups.
Modification of RNA is an important step in regulating transcriptional control31. However, the homeostasis and the regulatory mechanisms of various RNA modifications under diverse physiological realms are still unclear. Using the present modified northern blotting method, different RNA modifications can be quantified and compared. Furthermore, the changes and regulations of RNA modifications can be investigated in greater detail. In the future, it could also be possible to combine the experimental data from both the classical northern blotting and the modified northern blotting protocols, providing greater insights into RNA biology.
The most important factor determining the success of the modified northern blotting protocol is the integrity of the RNA sample. RNAs with some amount of degradation may yield good classical northern blotting results, but this could potentially have significant impact on the modified northern blotting results. The modifications of RNA in different tissues or cell lines could also vary significantly. It is important to test the suitable RNA sample loads for different tissues before performing the final experiments.
As blotting procedures have been traditionally named after Dr. Southern and different geographical directions, we propose the name "northeastern" blotting for the current technique.
Раскрытие информации
Авторы не имеют ничего раскрывать.
Благодарности
C.Y.W. received support from the National Health Research Institute (NHRI-EX101-9925SC), the National Science Council (101-2314-B-182-100-MY3, 101-2314-B-182A-009), and Chang Gung Memorial Hospital (CMRPG3B1643, CMRPG3D1002, CMRPG3D0581, CMRPG380091, and CMRPG3C1763).
Материалы
| Name | Company | Catalog Number | Comments |
| RNaseZap solution | Ambion | AM9782 | Protocol 1.1 |
| TRI Reagent solution | Ambion | AM9738 | Protocol 1.2.1.1 |
| 1-Bromo-3-chloropropane (BCP) | Sigma | B9673 | Protocol 1.2.1.3 |
| Ethanol | JTbaker | 8006 | Protocol 1.2.1.3 |
| Isopropanol | Sigma | I9516 | Protocol 1.2.1.3 |
| GenElute mRNA Miniprep Kit | Sigma | MRN70 | Protocol 1.2.2.1 |
| Glycogen | Ambion | AM9510 | Protocol 1.2.2.2 |
| Sodium acetate | Fluka | 71183 | Protocol 1.2.2.2 |
| Nuclease-free water | Ambion | AM9930 | Protocol 1.2.2.6 |
| DEPC | Sigma | D5758 | Protocol 2.1.1 |
| Agarose | JT Baker | A426 | Protocol 2.1.2 |
| MOPS | Sigma | M1254 | Protocol 2.1.3 |
| 37% formaldehyde Solution | Sigma | F8775 | Protocol 2.1.3 |
| EDTA, Disodium Salt | JT Baker | 8993 | Protocol 2.1.3 10X MOPS buffer |
| formamide | Sigma | F7503 | Protocol 2.2.1 |
| RNA Millennium Marker | Ambion | AM7150 | Protocol 2.2.2 |
| Ethidium Bromide | Amresco | X328 | Protocol 2.2.5 |
| Ficoll 400 | GE Healthcare | 17-0300-10 | Protocol 2.2.5 tracking dye |
| Bromophenol blue | Sigma | 114391 | Protocol 2.2.5 tracking dye |
| Xylene Cyanol | Sigma | X4126 | Protocol 2.2.5 tracking dye |
| Sodium chloride | JT Baker | 3624 | Protocol 2.4.2 20X SSC buffer |
| Trisodium citrate | Sigma | S1804 | Protocol 2.4.2 20X SSC buffer |
| filter paper | GE Healthcare | RPN6101M | Protocol 2.4.4 |
| GE Hybond-N+ membrane | GE Healthcare | RPN303B | Protocol 2.4.6 |
| Stratalinker UV Crosslinker 2400 | Stratagene | 400075 | Protocol 3.1.2 |
| Gel Catcher 1500 | ANT Technology | Gel Catcher 1500 | Protocol 3.1.5 |
| Anti-m6A (N6-methyladenosine) | Synaptic Systems | 202003 | Protocol 3.2.3 |
| Amersham ECL Anti-Rabbit IgG, HRP-linkd whole Ab | GE Healthcare | NA934 | Protocol 3.2.5 |
| ChemiDoc MP System | BIO-RAD | 1708280 | Protocol 3.2.8 |
Ссылки
- Thammana, P., Held, W. A. Methylation of 16S RNA during ribosome assembly in vitro. Nature. 251 (5477), 682-686 (1974).
- Meyer, K. D., Jaffrey, S. R. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nat Rev Mol Cell Biol. 15 (5), 313-326 (2014).
- Niu, Y., et al. N6-methyl-adenosine (m6A) in RNA: an old modification with a novel epigenetic function. Genomics Proteomics Bioinformatics. 11 (1), 8-17 (2013).
- Wang, C. Y., et al. Loss of FTO in adipose tissue decreases Angptl4 translation and alters triglyceride metabolism. Sci Signal. 8 (407), 127 (2015).
- Wang, X., et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 505 (7481), 117-120 (2014).
- Berulava, T., Rahmann, S., Rademacher, K., Klein-Hitpass, L., Horsthemke, B. N6-adenosine methylation in MiRNAs. PLoS One. 10 (2), 0118438 (2015).
- Deng, X., et al. Widespread occurrence of N6-methyladenosine in bacterial mRNA. Nucleic Acids Res. 43 (13), 6557-6567 (2015).
- Dominissini, D., et al. The dynamic N-methyladenosine methylome in eukaryotic messenger RNA. Nature. , (2016).
- Fustin, J. M., et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell. 155 (4), 793-806 (2013).
- Geula, S., et al. Stem cells. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science. 347 (6225), 1002-1006 (2015).
- Wang, C. Y., et al. FTO modulates fibrogenic responses in obstructive nephropathy. Sci Rep. , (2016).
- Jia, G., et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol. 7 (12), 885-887 (2011).
- Du, T., et al. An association study of the m6A genes with major depressive disorder in Chinese Han population. J Affect Disord. 183, 279-286 (2015).
- Wang, C. Y., Yeh, J. K., Shie, S. S., Hsieh, I. C., Wen, M. S. Circadian rhythm of RNA N6-methyladenosine and the role of cryptochrome. Biochem Biophys Res Commun. 465 (1), 88-94 (2015).
- Wang, C. Y., Shie, S. S., Hsieh, I. C., Tsai, M. L., Wen, M. S. FTO modulates circadian rhythms and inhibits the CLOCK-BMAL1-induced transcription. Biochem Biophys Res Commun. 464 (3), 826-832 (2015).
- Heyn, H., Esteller, M. An Adenine Code for DNA: A Second Life for N6-Methyladenine. Cell. 161 (4), 710-713 (2015).
- Xiao, W., et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol Cell. 61 (4), 507-519 (2016).
- Meyer, K. D., et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons. Cell. 149 (7), 1635-1646 (2012).
- Horowitz, S., Horowitz, A., Nilsen, T. W., Munns, T. W., Rottman, F. M. Mapping of N6-methyladenosine residues in bovine prolactin mRNA. Proc Natl Acad Sci U S A. 81 (18), 5667-5671 (1984).
- Resnick, R. J., Noreen, D., Munns, T. W., Perdue, M. L. Role of N6-methyladenosine in expression of Rous sarcoma virus RNA: analyses utilizing immunoglobulin specific for N6-methyladenosine. Prog Nucleic Acid Res Mol Biol. 29, 214-218 (1983).
- Golovina, A. Y., et al. Method for site-specific detection of m6A nucleoside presence in RNA based on high-resolution melting (HRM) analysis. Nucleic Acids Res. 42 (4), 27 (2014).
- Dominissini, D., et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 485 (7397), 201-206 (2012).
- Linder, B., et al. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods. 12 (8), 767-772 (2015).
- Fu, L., et al. Simultaneous Quantification of Methylated Cytidine and Adenosine in Cellular and Tissue RNA by Nano-Flow Liquid Chromatography-Tandem Mass Spectrometry Coupled with the Stable Isotope-Dilution Method. Anal Chem. 87 (15), 7653-7659 (2015).
- Rio, D. C., Ares, M., Hannon, G. J., Nilsen, T. W. Nondenaturing agarose gel electrophoresis of RNA. Cold Spring Harb Protoc. 2010 (6), 5445 (2010).
- Wang, C. Y., et al. FTO modulates fibrogenic responses in obstructive nephropathy. Sci Rep. 6, 18874 (2016).
- Li, X., et al. Transcriptome-wide mapping reveals reversible and dynamic N-methyladenosine methylome. Nat Chem Biol. , (2016).
- Sanschagrin, S., Yergeau, E. Next-generation sequencing of 16S ribosomal RNA gene amplicons. J Vis Exp. (90), (2014).
- Kukutla, P., Steritz, M., Xu, J. Depletion of ribosomal RNA for mosquito gut metagenomic RNA-seq. J Vis Exp. (74), (2013).
- Mishima, E., et al. Immuno-Northern Blotting: Detection of RNA Modifications by Using Antibodies against Modified Nucleosides. PLoS One. 10 (11), 0143756 (2015).
- Klungland, A., Dahl, J. A. Dynamic RNA modifications in disease. Curr Opin Genet Dev. 26, 47-52 (2014).
Перепечатки и разрешения
Запросить разрешение на использование текста или рисунков этого JoVE статьи
Запросить разрешениеСмотреть дополнительные статьи
This article has been published
Video Coming Soon
Авторские права © 2025 MyJoVE Corporation. Все права защищены