A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
NAD(P)H Fluorescence Lifetime Imaging for the Metabolic Analysis of the Murine Intestine and Parasites During Nematode Infection
In This Article
Summary
The present protocol describes the NAD(P)H fluorescence lifetime imaging of an explanted murine intestine infected with the natural parasite Heligmosomoides polygyrus, which allows one to investigate metabolic processes both in host and parasite tissues in a spatially resolved manner.
Abstract
Parasites generally have a negative effect on the health of their host. They represent a huge health burden, as they globally affect the health of the infested human or animal in the long term and, thus, impact agricultural and socio-economic outcomes. However, parasite-driven immune-regulatory effects have been described, with potential therapeutic relevance for autoimmune diseases. While the metabolism in both the host and parasites contributes to their defense and is the basis for nematode survival in the intestine, it has remained largely understudied due to a lack of adequate technologies. We have developed and applied NAD(P)H fluorescence lifetime imaging to explanted murine intestinal tissue during infection with the natural nematode Heligmosomoides polygyrus to study the metabolic processes in both the host and parasites in a spatially resolved manner. The exploitation of the fluorescence lifetime of the co-enzymes nicotinamide adenine dinucleotide (NADH) and nicotinamide adenine dinucleotide phosphate (NADPH), hereafter NAD(P)H, which are preserved across species, depends on their binding status and the binding site on the enzymes catalyzing metabolic processes. Focusing on the most abundantly expressed NAD(P)H-dependent enzymes, the metabolic pathways associated with anaerobic glycolysis, oxidative phosphorylation/aerobic glycolysis, and NOX-based oxidative burst, as a major defense mechanism, were distinguished, and the metabolic crosstalk between the host and parasite during infection was characterized.
Introduction
Parasitic infections impose a huge burden on human health1,2. A correlation between the rise in autoimmune diseases and the decline in parasitic infections has been observed in industrial countries. It is known that parasites can have beneficial effects by dampening excessive host immune responses. H. polygyrus is a natural parasite found in the intestine in rodents, and this parasite is known to induce immunoregulatory mechanisms that reduce the anti-parasitic immune response of the host via, among other mechanisms, the induction of regulatory T cells (Treg) in the infected host3,4,5,6,7,8,9,10,11. Those regulatory mechanisms are especially of interest in degenerative autoimmune diseases.
The analysis of the metabolic crosstalk between the host and intestinal nematodes remains widely neglected, although metabolism plays an important role in both the host and parasites for defense, survival, and function. We propose to adapt and apply NADH and NADPH fluorescence lifetime imaging upon two-photon excitation, a technology already widely used in different physiological and pathophysiological situations in mammalian cells and tissues12, to investigate host and nematode metabolism in living tissues correlatively.
NADH and NADPH, referred to as NAD(P)H, are ubiquitous molecules that are preserved in all cell-based lifeforms and play the role of co-enzymes in various metabolic pathways. For instance, they are involved in energy production, reductive biosynthesis, and NADPH oxidase-mediated reactive oxygen species (ROS) production, which are mainly linked to cell defense and cell communication13,14,15,16,17,20. Both co-enzymes emit fluorescence at ~450 nm upon two-photon excitation at 750 nm, thus allowing for marker-free metabolic imaging in cells and tissues19,21. Exciting both NADH and NADPH with only one wavelength is possible due to their similar and rather broad two-photon excitation spectra21.
The fluorescence lifetime of the co-enzyme NAD(P)H is directly dependent on the enzyme to which it binds18,21,22,23. Due to its chemical structure allowing for intramolecular energy transfer, the excited NADH or NADPH molecule loses energy through internal conversion processes, at a rate depending on its binding properties, to the enzymes (catalyst) before it relaxes and emits a fluorescence photon. This lifetime gives insight into the NAD(P)H binding site on the enzyme and, thus, the preferential biochemical reaction taking place19,21,22,23,24,25. The fluorescence lifetime of free NADH and NADPH molecules amounts to ~450 ps, whereas their fluorescence lifetime when bound to an enzyme is much longer (~2,000 ps) and depends on their binding site on the respective enzyme21.
There are more than 370 enzymes involved in NAD(P)H-linked processes; however, only the most abundant will be able to contribute to the resulting NAD(P)H fluorescence lifetime within the excitation range of the microscope. Using RNASeq data from mammal cells, we identified the most abundant NAD(P)H-dependent enzymes and generated a fluorescence lifetime reference to interpret the data generated in tissue and cell samples18. Thereby, this work distinguished for instance between the preferential activity of lactate dehydrogenase (LDH), which is associated with anaerobic glycolytic metabolic pathways, and isocitrate dehydrogenase (IDH) and pyruvate dehydrogenase (PDH) activity, which are mainly involved in aerobic glycolysis/oxidative phosphorylation metabolic pathways16,20. Additionally, NADPH binding to NADPH oxidases, which are the enzymes that are mainly responsible for oxidative burst, can be easily resolved due to the characteristic location of these enzymes in the cell (membrane-bound) and because of the particularly long NADPH fluorescence lifetime (3,650 ps)18,24,29,30,32. RNASeq data from H. polygyrus shows that the reference generated for mammalian cells also applies in adapted form to this nematode27.
Hence, in this work, by performing NAD(P)H fluorescence lifetime imaging (FLIM) in freshly explanted duodenum samples of mice infected with H. polygyrus, information on the ratio between free and enzyme-bound NAD(P)H was acquired, which depicted the general metabolic activity in all tissues, as well as the predominantly active enzyme in each pixel of the image (i.e., the enzyme to which NAD(P)H preferentially binds in that specific location). The success of these experiments relies on the accurate sample preparation of the explanted intestine, the reliable live imaging of the NAD(P)H fluorescence lifetime at subcellular resolution, and standardized data evaluation, as discussed in this protocol.
Protocol
All the experiments were performed in accordance with the National Animal Protection Guidelines and approved by the German Animal Ethics Committee for the protection of animals (G0176/16 and G0207/19). The protocol describes NAD(P)H fluorescence lifetime imaging data acquisition and data evaluation, which allow one to assess the general metabolic activity and specific metabolic pathways in both the host intestine and the parasites upon infection with the natural murine intestinal nematode, H. polygyrus. For this purpose, female C57BL/6 mice aged 10-12 weeks old were infected with 200 stage 3 larvae (L3). At different time points of the infection, the infected mice were sacrificed, and the duodenums were excised and prepared for imaging as previously described33. The duodenums of uninfected, age- and sex-matched mice were similarly prepared and imaged for control purposes. For maintaining the tissue properties necessary for further imaging and analysis, the samples must be processed immediately after explanting, and the next steps (steps 1.1-1.7) must be performed swiftly (Figure 1B).
1. Sample preparation
- Cut pieces of ~1 cm length from the initially excised duodenum.
- Glue the straightened tissue tube in the middle of a small Petri dish with fast-curing tissue glue (47 mm diameter, 10 mm rim height, see Table of Materials) (Figure 1A2).
- Apply a thin layer of additional glue to the bottom of the Petri dish over a large area around the tissue with a micro-brush (Figure 1A2).
- Use blunt scissors to make an incision along the entire length of the physically fixed intestine in the vicinity of the bottom of the Petri dish (Figure 1A3).
- Unfold the intestine with blunt tweezers so that the abluminal side comes into complete contact with the glue. Thus, the intestine lies fixed by the glue with the luminal side facing upward (Figure 1A4).
- In the case of an infected mouse, count the worms under a stereomicroscope (magnification 10x) to ensure the infection was successful (Figure 1A5).
- Seal the intestine with agarose to protect it from drying out. Here, 0.5%-0.9% agarose was used to protect the sensitive intestinal tissue from burning due to its low-temperature melting point of approximately 38 °C. Draw 1 mL of agarose into a pipette, and drizzle carefully onto the intestine with tender contact between the pipette tip and the tissue so that a thin layer of about 0.5 mm thickness completely encloses the tissue (Figure 1A6).
- Fill up the Petri dishes with the samples sealed in agarose with PBS (10%) at room temperature, and then close them with the lid. Either image the samples directly, or place them on ice in a thermally insulating box to queue for measuring. Place the first prepared sample under the microscope objective on a heating plate set to 37 °C (Figure 1A8).
2. Imaging
NOTE: The microscope system used to perform NAD(P)H-FLIM in infected and healthy duodenal tissue samples consists of the devices listed and described in Figure 2 and the Table of Materials. Use ImSpector 208 as the controlling software for all the modules used.
- To initially find the region of interest (ROI), place the sample in the Petri dish under the objective, and move along the x- and y-plane by hand to find a suitable ROI by visual inspection using the wide-field fluorescence microscopy mode.
- Switch the imaging system to two-photon excitation using either PMT detection or time-correlated single-photon counting (TCSPC) detection by switching modes in the software. Take special care to ensure that the environment is dark and vibration-free, to enclose the microscope in lightproof curtains, and to use a pneumatic suspended optical table, if possible.
- Immerse the objective lens in the Petri dish containing the sample with the controlling software by clicking on the lens-system icon and turning the mouse wheel to alter the z-position.
- Tune the laser to 765 nm, and set the maximum laser power to 10 %, which corresponds to 30 mW under the objective, by typing in the desired wavelength in the wavelength panel of the software. Adjust the laser power if needed. Measure below a gain of 40% (~30 mW to 100 mW) to avoid tissue photodamage and unwanted metabolic shifts.
- In the software, set the step size of the z-stage to 2 µm, which corresponds to the axial resolution of the microscope at 765 nm in tissue.
- Set the line scanning frequency to 400 Hz in the galvo mirror panel, resulting in a pixel dwell time of 4.95 µs.
- Set the image averaging over two to four images by choosing the averaging number from the dropdown menu in the software (i.e., scan the ROI two to four times to acquire smoother images). In this way, the total pixel dwell time is increased to 9.9-19.8 µs for the benefit of a higher signal-to-noise ratio (SNR) in the analysis process.
- Set the image field of view (FOV) to 505 pixels x 505 pixels (500 x 500 µm²) by choosing the parameters in the FOV panel in the software.
- For each measurement, calibrate the module time in advance to ensure the optimal function of the TCSPC electronics by clicking on initialize in the calibrating window in the hardware menu for the TCSPC.
- Adjust the fast photodiode externally to detect the Ti:Sa laser pulse train by using a fast oscilloscope.
NOTE: The signal of the photodiode is used to trigger the hPMT detection and photon counting by the TCSPC with respect to the excitation pulse. The detection is repeated every 12.5 ns (i.e., the time between two consecutive laser pulses), corresponding to the repetition rate of the Ti:Sa laser (80 MHz). - Define the range of the measurement depth by first setting the averaging to 1 and the gain of the laser power to ~10%, then starting the video mode (click on the video button), searching for the depth from which the signal can be still acquired by clicking the lens system icon again, and using the mouse wheel to progress into the sample while increasing the gain progressively.
- For the NAD(P)H-FLIM images of the tissue samples measured in FLIM mode, set the system to 765 nm at a maximum of 100 mW nominal power. Detect the fluorescence signal of NAD(P)H (466/60 nm) with the hybrid PMT at a gain of 97%, which is set in the software. The TCSPC module counts photons over 227 time windows (bins), each of 55 ps.
- Right-click on the displayed image during the measurement in the software, and choose display > T-PROFILE to visualize the decay curve acquired by the system.
- Note the number of bins before the laser pulse in the decay curve for later analysis.
- Acquire the data by clicking on start measurement.
NOTE: The data must match a specific format. Each slice, recorded at any tissue depth, is separated by a distance of 2 µm, and has an area of 500 µm x 500 µm, and the slices are acquired as a stack of 500 µm x 500 µm slices at 227 x 55 ps (Figure 3A) time points. Each voxel contains a spatially resolved (in the x-plane and y-plane) photon arrival time histogram. This represents the fluorescence decay (Figure 3B). A typically measured volume must be in the form of a hyper stack with dimensions of ~500 µm x 500 µm x 100-300 µm and 227 bins, where every slice in the z-plane contains the time-dependent intensity data as described above. For an exemplary set of measurement data, this results in (505 x 505 x 227) x 100 pixels (16 bit) and corresponds to about 4 GB.
3. Data analysis
NOTE: For the phasor analysis of the NAD(P)H-FLIM images, the program for calculating the lifetimes is a custom-written code in Python33.
- Use Anaconda with a Python 3.7 distribution on the Spyder IDE (see Table of Materials). The code uses Pythons' standard libraries.
- Load the code into the IDE, and execute. A file path dialog opens.
- Choose the folder with the raw data to analyze. The code is programmed to choose three parameters before the analysis through a Tkinter user input dialog with checkboxes, a dropdown menu, and text fields.
- Choose an offset within the input dialog. Determine the offset as the number of the first slices in the time stack that are not analyzed.
NOTE: The parameter cuts the time points before the excitation pulse. With the calibration of the system and trigger diode, this value must be approximately five slices or the number of dead bins before laser excitation, as described in step 2.14. - Choose the representation of the phasor plots within the input dialog's dropdown menu. Here, choose the appearance of the data points (cloud-like or topographic), as well as axis ticks of the semicircle time axis (enzymes or time [ps]), as options from the dropdown menu.
- Click on OK.
- Check that the code calculates two types of information from the NAD(P)H-FLIM z-stacks.
- First, measure the spatial information of each volume slice by the collapse of the TCSPC stacks, where the 227 time-resolved photon counting histograms are projected onto a single slice called the intensity projection image (Figure 3G), and from the Fourier-based analysis of the exponential decay curves (as described by Leben et al.18), and from the normalized real and imaginary part for each pixel in each volume slice (Figure 3C).
- From the real and imaginary images, obtain the phasor plots (Figure 3D) and the average fluorescence lifetime (t) image (color-coded, spatially resolved average decay constants for each voxel) (Figure 3E).
NOTE: As described in the introduction, the fluorescence lifetime of NAD(P)H, when bound to enzymes, is determined by the binding site of the co-enzyme to the respective enzyme. - Determine the contribution of the respective enzyme to the metabolic activity by generating the vector between the phase vector of each pixel and the phase vector of unbound NAD(P)H and projecting it onto the half-circle in the phasor plot. The half-circle represents all the possible mono-exponential decays of fluorescence lifetimes in pure compounds .
- Using the previously generated reference of fluorescence lifetimes of NAD(P)H bound to most abundant NAD(P)H-dependent enzymes18,33 (Supplementary Figure 1), and calculate the probability of the activation of these enzymes.
NOTE: A color code of the most dominant enzyme (i.e., of the enzyme for which the highest activation probability is calculated) is attributed to each pixel, thus producing an enzyme map (Figure 3F).
- From the ratio of free (unbound) to enzyme-bound NAD(P)H, specifically from the vectorial ratio in the phasor plot between free NAD(P)H at 450 ps and the enzyme-bound state, calculate the general metabolic activity as a percentage between 0% *only unbound NAD(P)H) and 100% (only enzyme-bound NAD(P)H). By attributing a metabolic activity (0%-100%) value to each pixel, a (color-coded) activity map is generated (Figure 3F).
- Overlay the resulting maps with the intensity images to gain additional morphologic information. Here, use an ImageJ macro (see Table of Materials) to overlay the hue and saturation of the enzyme or activity maps with the brightness of the intensity images.
- Ensure that the macro iterates over the whole stack. For every slice (depth) in the stack, it must split the intensity image and the map of choice respectively into HUE, SATURATION, and BRIGHTNESS (Image > type > HSB stack).
- Separate the main channels for both the HSB stacks (Image > stacks > stack to images).
- Close the HUE and SATURATION of the intensity stack, as well as the BRIGHTNESS of the map stack (close(slicename_HUE), close(slicename_BRIGHTNESS), ...).
- Recombine the remaining channels to form a new HSB stack consisting of the HUE and SATURATION from the slice of the map of interest and the BRIGHTNESS from the intensity image (Image > stacks > images to stack).
- Change the image type to RGB for a better visual appearance (Image > type > RGB color).
4. Tissue segmentation
NOTE: Use a pre-trained U-Net-based network (ILASTIK, see Table of Materials) for segmenting the intestinal host and nematode tissue, respectively, and furthermore, the epithelium and lamina propria in the host and the high NAD(P)H fluorescence signal areas and low NAD(P)H fluorescence signal areas in the nematode.
- Open ILASTIK, and choose a new project and pixel classification.
- Load the previously calculated intensity projection from step 3.7.1 by clicking on Add new > Add separate images. Load random slices from multiple measurements into the container.
- Click on Feature selection > select features. Add a sigma value of 50 (unitless weight), and tick all the features to be active.
NOTE: While the sigma value determines the weight of the features, the features determine the classes and the ability of the network to recognize edges, shapes, texture, or color. - Click on Training, and name the labels according to the tissue to segment (e.g., Epithelium, Background, etc.) by clicking on them. Select a label, and color all the pixels within the image corresponding to that label.
- Repeat for the other labels.
- Repeat for approximately half the images in the loaded dataset.
- Click on Live Update, and let the model predict the labels for the remaining unlabeled images.
- Correct approximately half of the solely predicted and unlabeled images by relabeling the falsely estimated segmentations, and repeat the Live Update.
- Repeat this step until the network has learned to estimate the desired tissue of interest correctly; ensure that the confidence of the network shown on the left close to the label names ranges between 95% and 98%.
- Load a new dataset as described in step 3.11, this time with all slices from one measurement, and click on Prediction Export.
- In the "Source dropdown" menu, choose Simple Segmentation; in "Format", choose the output to be in tiff format; and in "Choose Export Image Settings", select save path for the output. Click on export.
- This creates binary masks for the tissue type of interest. Since they consist of slices where the segmented tissue of interest has a pixel value of 1 and the rest have a pixel value of 0 (Figure 3H), simply multiply the binary masks with the generated data as listed in step 1.1 and steps 1.4-1.7. This leads to masked data (Figure 3 I, J, M). Use ImageJ for this step (Process > Image calculator > multiply).
- For each volume slice, calculate a signal-to-noise ratio (SNR) from the masked intensity images. Use an ImageJ macro for that purpose. Create your program such that it calculates the signal-to-noise ratio (SNR) from the segmented and masked images.
- Use the segmented background from step 3.21 as background (BG) and the segmented tissue of interest as signal (SIG) by iterating over every pixel of every slice with a value greater than 0 with two nested for-loops. Compute the SNR with the following formula33:
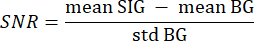
where mean SIG refers to the mean value of the signal histogram, mean BG refers to the mean value of the background histogram, and std BG refers to the standard deviation of the background histogram. - Discard volume slices with an SNR value lower than 5 from further analysis.
- Use the segmented background from step 3.21 as background (BG) and the segmented tissue of interest as signal (SIG) by iterating over every pixel of every slice with a value greater than 0 with two nested for-loops. Compute the SNR with the following formula33:
- With the Python script18,33, create an enzyme-frequency graph from the enzyme maps for each tissue type by summing and averaging the enzymes slice-wise and then normalizing them to a 100 µm³ tissue volume (Figure 3K, N).
- Code the script such that it loads the masked enzyme map.
- Iterate over every pixel, calculate the sum of pixels of the same enzyme, and then divide by the volume analyzed (the total of all pixels).
- Group the calculated abundances in their metabolic states, and average by dividing by the number of enzymes in each group. Write code that attributes metabolic states grouped around LDH to anaerobic glycolysis-like pathways, enzyme activity grouped around PDH/IDH/GAPDH to aerobic glycolysis/oxPhos-like pathways, and NADPH oxidase activity (NOX activity) to oxidative burst/oxidative stress used for defense (Figure 3L, O).
- Use the matplotlib library from Python to visualize the data with boxplots.
- Calculate the activity of the tissue from the masked activity maps (Figure 3F) as the mean value over each volume slice for all pixels with a value greater than 0 using an ImageJ macro by running Analyze > Measure.
- Visualize all the generated images side by side with ImageJ/FIJI.
Results
Using the current NAD(P)H-FLIM procedure28,29,33 combined with the described phasor analysis method, the metabolic activity and metabolic pathways in healthy and infected duodenums were measured at day 6, day 10, day 12, and day 14 post-infection with the murine intestinal nematode H. polygyrus.
Preserved intestinal tissue viability in the excised duodenum revealed by NAD(P)H-FLI...
Discussion
The critical steps within the protocol occur during the preparation and when finding the ROI. Fibers of partially digested food represent a challenge for imaging, mainly due to the endogenous luminescence of the fibers overlapping with the NAD(P)H fluorescence, but also due to their harmonic generation signal. It is of great importance to find ROIs that are free from feces. We aimed to avoid measuring areas containing feces. Washing was avoided because this affects the integrity of the fragile villi and influences the mu...
Disclosures
The authors declare no competing financial interests.
Acknowledgements
We thank Robert Günther for their excellent technical support. Financial support from the German Research Council (DFG) under Grant SPP2332 HA2542/12-1 (S.H.), NI1167/7-1 (R.A.N.), HA5354/11-1 (A.E.H.), and RA2544/1-1 (S.R.), under Grant SFB1444, P14 (R.A.N., A.E.H.), under Grant HA5354/8-2 (A.E.H.), and under Grant GRK2046 B4 and B5 (S.H., S.R.) and HA2542/8-1 (S.H.) are greatly acknowledged. W.L. received a PhD fellowship from the Berliner Hochschule für Technik, School of Applied Sciences, Berlin in Medical Physics/Physical Engineering.
Materials
| Name | Company | Catalog Number | Comments |
| Agarose | Thermo fisher | J32802.22 | ultra pure |
| Blunt scissors | FST fine science tools | 14108-09 | blund-blund 14 cm |
| Bodipy c12 | thermo fisher | D3822 | 1 mg solid |
| Control units, diode, TCSPC | LaVision Biontech | custom | TrimScope II |
| DMSO | Thermo fisher | D12345 | 3 mL |
| Filters | Chroma | 755 | 466 ± 20, 525 ± 25, 593 ± 20, 655 ± 20 nm |
| Foliodrape sheet | Hartmann | 277500 | |
| Gloves | Sigma-Aldrich | Z423262 | nitril |
| Halogen torch | Leica | This item has been phased out and is no longer available | KL 1500 LCD |
| hPMT | Hamamatsu, Germany | H7422 | GaAsP |
| Ilastik | Netlify | free Software | Java Backend |
| ImageJ | National Institutes of health | free Software | FIJI - standard plugins |
| Imspector | LaVision Biontech | - | Vers. 208 |
| Intravital stage | LaVision Biontech | custom | TrimScope II |
| Lens system 20x | Zeiss | custom | W-plan-apochom 20x Waterimmersion NA 1.05 |
| Mercury vapor torch | LaVision Biontech | custom | |
| microbrush | Fisher scientific | 22-020-002 | 85 mm |
| Microscope | LaVision Biontech | custom | TrimScope II |
| Oscilloscope | Rhode & Schwarz | 1326.2000.22 | |
| PBS | Sigma-Aldrich | AM9624 | 0.5 L |
| Petri dish | Sigma aldrich | P5606 | 40 x 15 mm |
| Pipette | thermo fisher | 4651280N | Einkanalpipette |
| Pipette tips | thermo fisher | 94056980 | Spitzen mit Filter |
| PMT | Hamamatsu, Japan | H7422 | GaAsP |
| Python | Python Software foundation | free Software | Anaconda 3.7 Spyder IDE, standard librarys with KYTE |
| Sterio microscope | Leica | This item has been phased out and is no longer available | M26, 6.3x zoom |
| Ti:Sa LASER CHAMELION ULTRA II | Coherent, APE | - | 690-1080 nm tunable, 80MHz |
| Tissueglue | 3M | 51115053603 | 3 mL |
| Tweezers | FST fine science tools | 11049-10 | blund, graefe, angeled |
| Tweezers | FST fine science tools | 91197-00 | Dumont, curved |
References
- Hotez, P. J., Fenwick, A., Savioli, L., Molyneux, D. H. Rescuing the bottom billion through control of neglected tropical diseases. Lancet. 373 (9674), 1570-1575 (2009).
- Sartorius, B., et al. Prevalence and intensity of soil-transmitted helminth infections of children in sub-Saharan Africa, 2000-18: A geospatial analysis. The Lancet Global Health. 9 (1), e52-e60 (2021).
- Affinass, N., et al. Manipulation of the balance between Th2 and Th2/1 hybrid cells affects parasite nematode fitness in mice. European Journal of Immunology. 48 (12), 1958-1964 (2018).
- Hartmann, S., et al. Gastrointestinal nematode infection interferes with experimental allergic airway inflammation but not atopic dermatitis. Clinical & Experimental Allergy. 39 (10), 1585-1596 (2009).
- Hepworth, M. R., et al. Mast cells orchestrate type 2 immunity to helminths through regulation of tissue-derived cytokines. Proceedings of the National Academy of Sciences of the United States of America. 109 (17), 6644-6649 (2012).
- Rausch, S., et al. Establishment of nematode infection despite increased Th2 responses and immunopathology after selective depletion of Foxp3+ cells. European Journal of Immunology. 39 (11), 3066-3077 (2009).
- Rausch, S., et al. Parasitic nematodes exert antimicrobial activity and benefit from microbiota-driven support for host immune regulation. Frontiers in Immunology. 9, 2282 (2018).
- Steinfelder, S., Rausch, S., Michael, D., Kuhl, A. A., Hartmann, S. Intestinal helminth infection induces highly functional resident memory CD4(+) T cells in mice. European Journal of Immunology. 47 (2), 353-363 (2017).
- Whelan, R. A., et al. A transgenic probiotic secreting a parasite immunomodulator for site-directed treatment of gut inflammation. Molecular Therapy. 22 (10), 1730-1740 (2014).
- Ziegler, T., et al. A novel regulatory macrophage induced by a helminth molecule instructs IL-10 in CD4+ T cells and protects against mucosal inflammation. The Journal of Immunology. 194 (4), 1555-1564 (2015).
- Grantham, B. D., Barrett, J. Amino acid catabolism in the nematodes Heligmosomoides polygyrus and Panagrellus redivivus. 2. Metabolism of the carbon skeleton. Parasitology. 93 (Pt 3), 495-504 (1986).
- Dmitriev, R. I., Intes, X., Barroso, M. M. Luminescence lifetime imaging of three-dimensional biological objects. Journal of Cell Science. 134 (9), 1-17 (2021).
- Mossakowski, A., et al. Tracking CNS and systemic sources of oxidative stress during the course of chronic neuroinflammation. Acta Neuropathologica. 130 (6), 799-814 (2015).
- Qian, T., et al. Label-free imaging for quality control of cardiomyocyte differentiation. Nature Communications. 12, 4580 (2021).
- Bayerl, S., et al. Time lapse in vivo microscopy reveals distinct dynamics of microglia-tumor environment interactions-a new role for the tumor perivascular space as highway for trafficking microglia. Glia. 64 (7), 1210-1226 (2016).
- Skala, M. C., et al. In vivo multiphoton microscopy of NADH and FAD redox states, fluorescence lifetimes, and cellular morphology in precancerous epithelia. Proceedings of the National Academy of Sciences of the United States of America. 104 (49), 19494-19499 (2007).
- Skala, M. C., et al. Multiphoton microscopy of endogenous fluorescence differentiates normal, precancerous, and cancerous squamous epithelial tissues. Cancer Research. 65 (4), 1180-1186 (2005).
- Leben, R., Kohler, M., Radbruch, H., Hauser, A. E., Niesner, R. A. Systematic enzyme mapping of cellular metabolism by phasor-analyzed label-free NAD(P)H fluorescence lifetime imaging. International Journal of Molecular Sciences. 20 (22), 5565 (2019).
- Chacko, J. V., Eliceiri, K. W. Autofluorescence lifetime imaging of cellular metabolism: Sensitivity toward cell density, pH, intracellular, and intercellular heterogeneity. Cytometry. 95 (1), 56-69 (2019).
- Stringari, C., et al. Phasor approach to fluorescence lifetime microscopy distinguishes different metabolic states of germ cells in a live tissue. Proceedings of the National Academy of Sciences of the United States of America. 108 (33), 13582-13587 (2011).
- Lakowicz, J. R., Szmacinski, H., Nowaczyk, K., Johnson, M. L. Fluorescence lifetime imaging of free and protein-bound NADH. Proceedings of the National Academy of Sciences of the United States of America. 89 (4), 1271-1275 (1992).
- Blacker, T. S., et al. Separating NADH and NADPH fluorescence in live cells and tissues using FLIM. Nature Communications. 5 (1), 3936 (2014).
- Blacker, T. S., Duchen, M. R. NAD(P)H binding configurations revealed by time-resolved fluorescence and two-photon absorption. Biophys. J. 122 (7), 1240-1253 (2023).
- Niesner, R., et al. Selective detection of NADPH oxidase in polymorphonuclear cells by means of NAD(P)H-based fluorescence lifetime imaging. Journal of Biophysics. 2008, 602639 (2008).
- Vishwasrao, H. D., Heikal, A. A., Kasischke, K. A., Webb, W. W. Conformational dependence of intracellular NADH on metabolic state revealed by associated fluorescence anisotropy. Journal of Biology and Chemistry. 280 (26), 25119-25126 (2005).
- Babior, B. M. NADPH oxidase: An update. Blood. 93 (5), 1464-1476 (1999).
- Hewitson, J. P., et al. Proteomic analysis of secretory products from the model gastrointestinal nematode Heligmosomoides polygyrus reveals dominance of venom allergen-like (VAL) proteins. Journal of Proteomics. 74 (9), 1573-1594 (2011).
- Rakhymzhan, A., et al. Synergistic strategy for multicolor two-photon microscopy: Application to the analysis of germinal center reactions in vivo. Scientific Reports. 7, 7101 (2017).
- Bremer, D., et al. Method to detect the cellular source of over-activated NADPH oxidases using NAD(P)H fluorescence lifetime imaging. Current Protocols in Cytometry. 80, 9.52.1-9.52.14 (2017).
- Leben, R., et al. Phasor-based endogenous NAD(P)H fluorescence lifetime imaging unravels specific enzymatic activity of neutrophil granulocytes preceding NETosis. International Journal of Molecular Sciences. 19 (4), 1018 (2018).
- Digman, M. A., Caiolfa, V. R., Zamai, M., Gratton, E. The phasor approach to fluorescence lifetime imaging analysis. Biophysics Journal. 94 (2), L14-L16 (2008).
- Lindquist, R. L., Bayat-Sarmadi, J., Leben, R., Niesner, R., Hauser, A. E. NAD(P)H oxidase activity in the small intestine is predominantly found in enterocytes, not professional phagocytes. International Journal of Molecular Sciences. 19 (5), 1365 (2018).
- Liublin, W., et al. NAD(P)H fluorescence lifetime imaging of live intestinal nematodes reveals metabolic crosstalk between parasite and host. Scientific Reports. 12, 7264 (2022).
- Nhu, N. T. Q., et al. Alkaline pH increases swimming speed and facilitates mucus penetration for Vibrio cholerae. Journal of Bacteriology. 203 (7), e00607 (2021).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved