A subscription to JoVE is required to view this content. Sign in or start your free trial.
A Photopolymerizable Hyaluronic Acid-Collagen Model of the Invasive Glioma Microenvironment with Interstitial Flow
In This Article
Summary
We present a method for replicating the glioma tumor microenvironment at the invasive front that incorporates interstitial fluid flow. This model is a hyaluronan-collagen I hydrogel in a tissue culture insert where a fluid pressure head can be applied. Invasion can be quantified, and cells can be isolated or lysed.
Abstract
Glioblastoma recurrence is a major hindrance to treatment success and is driven by the invasion of glioma stem cells (GSCs) into healthy tissue that are inaccessible to surgical resection and are resistant to existing chemotherapies. Tissue-level fluid movement, or interstitial fluid flow (IFF), regulates GSC invasion in a manner dependent on the tumor microenvironment (TME), highlighting the need for model systems that incorporate both IFF and the TME. We present an accessible method for replicating the invasive TME in glioblastoma: a hyaluronan-collagen I hydrogel composed of human GSCs, astrocytes, and microglia seeded in a tissue culture insert. Elevated IFF can be represented by applying a fluid pressure head to the hydrogel. Additionally, this model can be tuned to replicate inter- or intra-patient differences in cellular ratios, flow rates, or matrix stiffnesses. Invasion can be quantified, while gels can be harvested for a variety of outcomes, including GSC invasion, flow cytometry, protein or RNA extraction, or imaging.
Introduction
Glioblastoma is a devastating disease, marked by short survival1 that is extended only moderately by clinically-available treatments2,3. This impediment to effective therapy is largely driven by the highly infiltrative nature of chemoresistant glioma stem cells (GSCs) that are inaccessible to resection and seed recurrent tumors4. Concurrently, GSCs are highly plastic, responding to diverse stimuli in the tumor microenvironment (TME) in order to survive and invade5,6. Specifically, the densely populated tumor and leaky vasculature produce a steep pressure differential at the tumor border, increasing interstitial fluid flow (IFF) in distinct regions that correlate with GSC invasion7. This increased IFF influences8,9,10 and often enhances8,9 GSC invasion via molecular pathways that are amenable to pharmacological inhibition. However, this process is confounded by the influence of glia in the TME on glioma invasion; indeed, glia have been described to modulate glioma invasion in numerous contexts11, and preliminary evidence in our lab indicates a similar influence of glia on IFF-enhanced glioma invasion12. Thus, our lab has developed a tunable, 3D TME model that replicates these invasive tumor border regions by incorporating both IFF and glia-GSC interactions to quantify GSC invasion and other outcomes (i.e., surface or intracellular protein expression, RNA expression, etc.) as influenced by IFF, glia, and/or candidate therapeutics12(Figure 1).
The TME model is similar to other tissue culture insert-based interstitial fluid flow assays (i.e., static/flow assays) that have been widely published8,9,10,13,14,15,16,17,18. The main distinctions of this model are the incorporation of a nonproprietary and tunable collagen-hyaluronic acid matrix, inclusion of astrocytes and microglia into the matrix at ratios representative of patient TMEs12, and the system is entirely enclosed in a well-plate, lacking external tubing, reservoirs, or pumps. Hyaluronic acid is among the principal components of the brain extracellular matrix19, but it is not sufficient to permit fluid flow; thus, collagen I is included in the mixture. Crosslinking occurs in two steps: a free-radical-initiated chain growth polymerization whereby upon exposure to ultraviolet (UV) light, free radicals are produced from the photoinitiator and generate chemical crosslinks between methacrylated hyaluronic acid molecules20, and neutralization of acid-preserved collagen and exposure to heat to form collagen fibers21,22,23.
Various parameters that were optimized for this assay have been described in previously published work in thiol-modified hyaluronan-collagen hydrogels8,12. Specifically, the cells (primary human GSCs24, primary human cortical astrocytes, and immortalized human microglia) used for this assay are at least 60% viable for 3 days in gels composed of astrobasal media supplemented with GSC growth factors, 0.5% v/v N-2 and 1% v/v B-27 without vitamin A12. In cases where the gels must be degraded (e.g., for protein extraction or flow cytometry), collagenase and dispase are used and maintain sufficient viability (Figure 2).
To quantify invasion, GSCs must be fluorescently labeled before they are combined with glia in the matrix. This may be achieved by labeling with Hoechst 33342 (as described below), cell trackers12, genetic modification, or transduction. Regardless, the viability of cells labeled with new markers should be measured before incorporating them into the TME model. Alternatively, if invasion will not be quantified but the gels will be harvested for protein extractions, RNA extractions, or other outcomes, cells often do not need to be fluorescently labeled during the gel preparation process. In that case, gels can be either formalin-fixed for imaging, degraded and cells lysed for RNA extraction, or degraded using proteases for flow cytometry and subsequently lysed with protein lysis buffers for Western blots or other protein work.
Here, we present a protocol for incorporating human astrocytes, microglia, and patient-derived GSCs at a patient-defined ratio, 4:1:112, into a 1.2 mg/mL collagen I, 4 mg/mL hyaluronan matrix under static of flow conditions. The specific parameters used for an assay (i.e., cellular ratios, stiffness, cell types, etc.) can be altered to represent different contexts if viability is adequate.
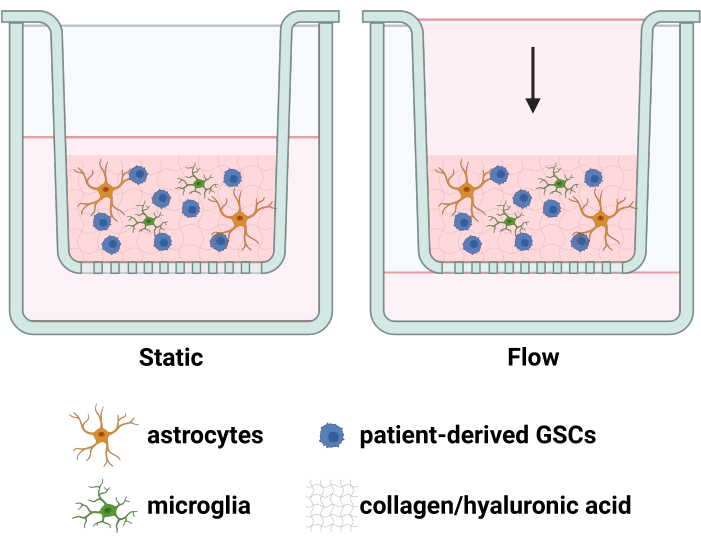
Figure 1: Diagram of the TME model consisting of GSCs, astrocytes, and microglia in a hydrogel under a fluid pressure head. Cells are embedded in a collagen-hyaluronic acid hydrogel inside a tissue culture insert. Gravitational force drives the net downward flow. Pores in the tissue culture insert membrane allow cells invading downward to attach to the bottom surface of the tissue culture insert, and these cells can be fixed, imaged, and quantified. Created with BioRender.com. Please click here to view a larger version of this figure.
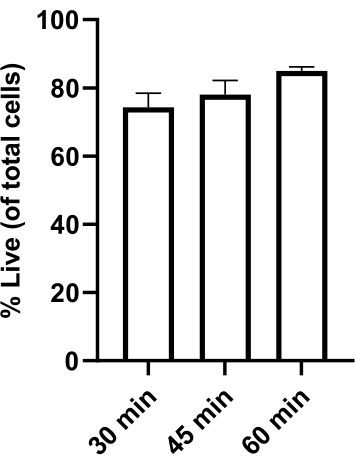
Figure 2: Cellular viability following hydrogel degradation. G34s (GSCs), astrocytes, and microglia were seeded in the TME model at 4:1:1 and incubated for 21 h at 37 °C. Gels were removed and degraded using 0.3 mg/mL collagenase + 0.02 mg/mL dispase for a total of either 30, 45, or 60 min, pipetting to mix every 15 min (30 min condition) or at 30 min then every 15 min (45 min and 60 min conditions). After the gels were degraded, the cells were stained with acridine orange (all cells) and propidium iodide (dead cells) and counted using an automated cell counter. Viability is reported for all cells within the TME model. Cells maintained at least 70% viability over 30-60 min. Data is presented as mean ± SEM; n = 3 biological replicates. Please click here to view a larger version of this figure.
Protocol
This protocol was developed in compliance with the guidelines set forth by the Virginia Tech Institutional Biosafety Committee.
NOTE: Perform all procedures in a BSL-2 cell culture cabinet unless otherwise specified. Please consult the institution's biosafety committee for guidance on the use of human cells.
1. Calculations for gel preparation
- Calculate the number of conditions needed for the experiment.
NOTE: A "condition" is defined by the contents of the solution (e.g., cell concentrations, collagen concentrations, etc.), not externally applied variables, like the fluid pressure head used in this example. - Calculate the total volume of gel needed for each condition solution.
NOTE: The volume required for 12-well gels that use 12 mm diameter tissue culture inserts is 100-150 µL per insert. At least three technical replicates are recommended when quantifying invasion. Additionally, to account for volume loss from pipetting error, add the volume of an additional replicate to each condition solution. - Calculate the total volume of gel solution required for the experiment from step 1.2.
- Calculate the total volume of 3 mg/mL collagen solution required for the experiment. Add at least 10% to this volume to account for volume loss when pipetting.
Total volume 3 mg/mL collagen = 0.4 × Total volume of gel solution - Calculate the volumes of high-concentration rat tail collagen I, sterile 1 N NaOH, sterile 10x PBS, and sterile water for each component in the 3 mg/mL collagen solution at the given final concentrations (cf): collagen I cf = 3 mg/mL, NaOH cf = 2.3% v/v of the collagen I volume, and PBS cf =1x. Dilute the solutions in sterile ultrapure (18.2 MΩ·cm) water.
- Calculate the volumes of 3 mg/mL collagen solution, 1% w/v methacrylated hyaluronic acid, and the photoinitiator, 17 mg/mL lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP), and cell culture media for each of the condition solutions at the given final concentrations (cf): collagen cf = 1.2 mg/mL, hyaluronic acid cf = 0.4% w/v, LAP cf = 2% v/v of the hyaluronic acid volume. Dilute the solutions in cell culture media containing cells.
- Calculate the number of cells for each condition solution at the desired ratio (e.g., 100,000 GSCs per 100 uL hydrogel at a ratio of 4:1:1 GSCs:astrocytes:microglia).
2. Preparing materials
- In the week prior to the experiment, plate GSCs, astrocytes, and microglia.
NOTE: Plate to yield approximately double the number of cells than what is needed for gels to account for both passaging and the loss of cells during the procedure. - Prepare a working solution of the photoinitiator, LAP, at 17 mg/mL in sterile 1x PBS. Protect it from light.
- Reconstitute the methacrylated hyaluronic acid in 1x PBS per the manufacturer's instructions. Seal the lid and rock the suspension gently for 1 h, or until dissolved, at 4 °C.
NOTE: The methacrylated hyaluronic acid used in this protocol is stable for approximately 1 month at 4 °C as specified by the manufacturer. Protect it from light. - Set up a UV lamp using a collimated light-emitting diode (LED) with a nominal wavelength of 385 nm at an intensity of 50 mW/cm2 at the surface of the gel. Place the lamp in the cell culture cabinet.
CAUTION: UV light can damage the eyes. Wear protective eyewear, and do not look directly into the lamp.
NOTE: Conduct separate experiments to ensure that viability or the functional outcome being measured is not significantly impacted by 385 nm light exposure at 50 mW/cm2 over the exposure period (45 s). - Prepare media (astrobasal media + 0.5% v/v N-2 + 1% v/v B-27 without vitamin A) for the experiment.
NOTE: Combine media components and filter through a 0.22 µm pore vacuum system to ensure sterility. Avoid freeze-thaw cycles of the growth factors (N-2 and B-27) to prevent degradation. - Prepare 1 N NaOH in a fume hood with the splash guard in place.
CAUTION: 1 N NaOH is highly corrosive. Handle with care, wear eye protection, or use a splash guard, and keep the solution under a fume hood. - Prepare sterile ultrapure (18.2 MΩ·cm) water.
3. Passaging cells and labeling GSCs
- Passage astrocytes, microglia, and GSCs according to the manufacturer's instructions.
- Label the GSCs with Hoechst 33342 following the manufacturer’s recommendation.
CAUTION: Hoescht 33342 can cause serious eye irritation and can induce DNA damage. Handle with care. - Centrifuge all cells at 200 x g for 5 min at RT.
- Gently dump the supernatant and remove excess media with a pipette.
- Resuspend the GSCs in 1x PBS.
- Centrifuge the GSCs at 200 x g for 5 min.
- Resuspend all cells in the media (astrobasal media + 0.5% N-2 v/v + 1% v/v B-27) for the experiment.
- Count all the cells.
- Add the absolute number of cells needed for each condition solution into one vial per condition solution. Place these at 37 °C, 5% CO2 until use.
4. Preparing hydrogels
NOTE: Perform the steps after opening the methacrylated hyaluronic acid and photoinitiator in the dark until the gels have been photo-crosslinked.
- Place the 8-11 mg/mL (high concentration) collagen, 1 N NaOH, sterile ultrapure (18.2 MΩ·cm) water, sterile 10x PBS, one microcentrifuge tube (for the 3 mg/mL collagen solution), methacrylated hyaluronic acid, and the photoinitiator on ice.
NOTE: All materials that come into contact with the collagen solution must be kept on ice to prevent premature crosslinking. - Place the tissue culture inserts in the plate and label the plate.
- Using the calculated values, prepare the 3 mg/mL collagen solution by combining high-concentration collagen, 1 N NaOH, ultrapure (18.2 MΩ·cm) water, and 10x PBS. Add components dropwise and keep the pipette tip submerged when mixing to prevent bubble formation.
NOTE: If bubbles do form, centrifugation or tapping the tube on a hard surface often removes the bubbles. Furthermore, media containing phenol red will be yellow if the pH is too acidic; more NaOH should be added in this case. - Centrifuge cells at 200 x g for 5 min at RT.
- Gently remove excess media from the top of each cell condition solution, leaving the volume of media required for the condition. Place these solutions on ice.
NOTE: Do not transfer the cells out of this vial to avoid loss of cells. Instead, add all gel condition components to this vial, then mix gently and distribute the gels in the plate. - Add the calculated volume of 3 mg/mL collagen solution for a final concentration of 1.2 mg/mL to each cell condition solution from step 4.5.
- Add the calculated volume of 1% w/v methacrylated hyaluronic acid for a final concentration of 0.4% w/v to each cell condition solution from step 4.5.
- Add the calculated volume of 17 mg/mL LAP to each cell condition solution from step 4.5.
NOTE: Add this solution just prior to distributing gels in the plate to maximize viability; LAP in liquid form is cytotoxic. - One at a time, mix each condition tube and add 100-150 µL to each tissue culture insert. Keep the pipette tip submerged to avoid bubble formation.
- Turn on the 385 nm UV lamp (set up at 50 mW/cm2) at a constant current.
- Photo-crosslink each gel, one at a time, by exposing each well to the UV light for 45 s each.
- Crosslink the collagen by placing the plate at 37 °C for 30-45 min.
- Apply the fluid pressure head to gels using astrobasal + 0.5% N-2 v/v + 1% v/v B-27.
- Net zero flow (static): Add 700 µL of media in the well-plate and 100 µL in the tissue culture insert.
- Net flow: Add 100 µL of media in the well-plate and 700 µL in the tissue culture insert.
NOTE: Use capillary action to add 100 µL underneath the tissue culture insert; this volume will not fill the well plate.
- Place the plate at 37 °C, 5% CO2, for at least 18 h or up to 5 days.
NOTE: Systems can incubate for up to 5 days depending on the needs of the assay, but an 18-24 h timepoint is sufficient for initial invasion, viability, or proliferation experiments.
5. Fixing cells for invasion analysis
NOTE: Do not allow the tissue culture insert membrane to dry out, and do not apply the solutions directly to the membrane to avoid detachment of cells from the membrane.
- Aspirate media from the tissue culture inserts and well-plate.
- Suction the gels from the tissue culture inserts with a P1000 tip.
NOTE: Gels can be retained for imaging by leaving the gels in the well during this step and following the alternative fixation protocol in the note of step 5.4. - Gently wash the well-plate well with 1x PBS so that each tissue culture insert membrane is covered.
- Add 4% v/v formalin in PBS to the well-plate so that each tissue culture insert membrane is covered and incubate for 15-20 min at RT.
CAUTION: Formalin is a suspected carcinogen and can cause serious eye damage. Use formalin in a fume hood with the splash guard in place.
NOTE: This incubation length will only fix cells on the tissue culture insert membrane. If gels are kept for imaging, fix gels for 1 h at 4 °C, gently rocking, and add enough formalin to the tissue culture insert to permeate the gel. After fixing, thoroughly wash gels with PBS for at least 20 min. - Wash each well with 1x PBS. Afterward, add an additional 1x PBS to each well-plate well and tissue culture insert.
- Use a cotton swab to remove gel fragments by wiping the upper side of each tissue culture insert membrane. Return any lost 1x PBS to each well after wiping to avoid drying out the membranes.
- Wash each well and tissue culture insert in 1x PBS.
NOTE: The plate can be sealed and placed at 4 °C if needed, but image membranes as soon as possible to avoid loss of fluorescence.
6. Imaging and quantification
- On the DAPI channel, image each membrane. If possible, take a tiled fluorescence image of the entire well. If this is not possible, at 20x, image the center of the well and four regions in a cross formation that intersects the center point.
NOTE: Exclude the edges of the well to avoid gel fragments and do not overlap regions of interest (ROIs). - Count the number of cells in each image.
- Calculate the percent of seeded GSCs that invaded for each well:
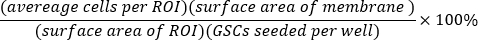
- Calculate the mean and standard deviation of percent invasion across technical replicates.
7. Alternative endpoint: flow cytometry
NOTE: In addition to or instead of analyzing invasion, unfixed gels may be degraded and the cells harvested for flow cytometry or other endpoint analyses. An example protocol analyzing cell viability (fixable live/dead stain), proliferation (Ki67 antibody), and stemness (CD71 antibody) is provided. Other markers of interest can be substituted in this protocol after validating staining efficacy. In particular, a wide variety of markers exist that target GSCs (reviewed in25). This protocol begins after step 4.14.
- Degrade the hydrogels.
NOTE: A 2x concentration of collagenase and dispase solutions are prepared first with experimental media (astrobasal + 0.5% v/v N-2 + 1% v/v B-27 without vitamin A), which is then added into an equal volume of hydrogel to reach a final concentration of 1x. As different cells have different responses to enzymatic degradation, checking cell viability and yield (Figure 2) is strongly recommended. The range of the final collagenase concentration should be 0.2-0.4 mg/mL, and the range of final dispase concentration should be 0.01-0.04 mg/mL; in this example, 0.3 mg/mL of collagenase and 0.02 mg/mL of dispase is used to degrade the hydrogels.- Prepare 2x collagenase and dispase solutions (gel degradation media solution). Dilute stock collagenase to 0.6 mg/mL and stock dispase to 0.04 mg/mL with experimental media (astrobasal + 0.5% N-2 v/v + 1% v/v B-27 without vitamin A).
- Remove the gels from the tissue culture inserts and transfer them into a V-bottom 96-well plate.
- Add the equivalent volume of gel degradation media solution per hydrogel to each well. For example, add 100 µL of gel degradation media solution to 100 µL of hydrogel.
- Incubate the plate on a shaker at 300-500 RPM (orbit: 2 mm) at 37 °C for 15-30 min.
NOTE: Try the minimum interval, 15 min incubation, if cells are sensitive to enzymatic degradation. - Pipette each hydrogel up and down 30-50 times to break up the hydrogels.
NOTE: Hydrogels will be easy to pipette once they are completely degraded. If gels cannot be easily pipetted, the gels must be incubated on a shaker at 300-500 RPM (orbit: 2 mm) at 37 °C for another 15 min. Then, pipette each hydrogel up and down 30-50 times to break up the gels. Importantly, the incubation time for degrading hydrogels should not exceed 1 h; this could lead to low cell viability. - Centrifuge the plate at 625 x g for 1 min at 4 °C, and discard the supernatant.
NOTE: The supernatant may be removed by either pipetting or by quickly inverting the plate over a waste tray. - Resuspend the cells with 50 µL of 1x PBS and pool three technical replicates into one well to yield a single biological replicate.
- Centrifuge the plate at 625 x g for 1 min at 4 °C, and discard the supernatant to continue onto flow cytometry.
- Block non-specific binding.
- Wash the cells with 200 µL of 1x PBS/well, centrifuge the plate at 625 x g for 1 min at 4 °C, and discard the supernatant. Perform all of the following steps on ice unless otherwise specified.
- Resuspend the cells with 200 µL of blocking solution (1x PBS with 10% v/v FBS) and incubate the plate for 15 min at RT.
- Wash the cells with 200 µL of 1x PBS/well, centrifuge the plate at 625 x g for 1 min at 4 °C, and discard the supernatant.
- Label cells with a live/dead stain.
- Test the cell viability using a fixable live/dead near-infrared stain. Dilute the live/dead stain with 1x PBS to 1:1000.
- Resuspend the cells in one well with 200 µL of 70% ethanol for a negative control.
- Resuspend the cells in the remaining wells with 200 µL of 1x PBS.
- Incubate the plate for 5-10 min on ice, centrifuge the plate at 625 x g for 1 min at 4 °C, and discard the supernatant.
- Add 50 µL of live/dead solution to the respective wells and incubate the plate on ice for 15 min.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Wash the cells twice with 200 µL of flow buffer (HBSS with 2% w/v BSA) per well.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Perform surface immunostaining for cellular markers of interest (CD71 antibody).
- Dilute the CD71 antibody with flow buffer (HBSS with 2% w/v BSA) at 1:50.
- Add 50 µL of antibody solution to the respective wells and incubate the plate on ice for 15 min.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Wash the cells once with 200 µL of flow buffer (HBSS with 2% w/v BSA) per well.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Perform intracellular immunostaining for cellular markers of interest (Ki67 antibody).
- Use the Foxp3 transcription factor staining buffer set to fix and permeabilize the cells. Add 100 µL of Foxp3 fixation/permeabilization working solution to each well. Resuspend the cell pellets by pipetting up and down.
- Incubate the plate on ice or at RT for 30-60 min in the dark.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Wash the cells twice with 200 µL of 1x permeabilization buffer per well.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Add 100 µL of blocking solution (1x PBS with 2% v/v FBS) for 15 min at RT.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Dilute the Ki67 antibody with blocking solution (1x PBS with 2% v/v FBS) at 1:25.
- Add 50 µL of 1:25 Ki67 antibody solution to the respective wells and incubate the plate for 30 min at RT in the dark.
- Centrifuge the plate at 625 x g for 1 min at 4 °C and discard the supernatant.
- Wash the cells twice with 200 µL of 1x permeabilization buffer per well.
- Resuspend the cells with 200 µL of flow buffer.
- Process the samples using a flow cytometer. An example gating strategy is detailed in Cornelison et al. 202212.
Results
Representative data for invasion (Figure 3), viability, and the expression of Ki67 and CD71 via flow cytometry (Figure 4) are provided for GSC lines as previously published for a thio-modified hyaluronan-collagen hydrogel12. The presence of astrocytes and microglia within the TME model has a differential effect on GSC invasion dependent on the cell line (Figure 4). Specifically, the GSCs G44 and G62
Discussion
Assembling the TME model includes six fundamental steps: 1) passaging cells and separating the cells between like conditions, 2) assembling a concentrated collagen solution for all conditions, 3) combining the gel components (cells, collagen, methacrylated hyaluronic acid, and the photoinitiator) for each condition 4) plating gels, 5) crosslinking by UV exposure and heat, and 6) adding the fluid pressure head. After 18 h or more, the gels can be harvested for flow cytometry, RNA extraction, protein extraction, imaging, o...
Disclosures
The authors have no relevant conflicts of interest to disclose.
Acknowledgements
We would like to thank the funding sources for this work: the National Institutes of Health National Cancer Institute (R37 CA222563 to J.M.), the Coulter Foundation (J.M.), and Virginia Tech ICTAS-CEH (J.M. & J.H.). The GSCs used in this assay were derived by Jakub Godlewski, Ph.D. (Harvard Medical School).
Materials
| Name | Company | Catalog Number | Comments |
| 12 Well Tissue Culture Plate, Sterile | Celltreat Scientific Products | 229112 | |
| 250 mL Filter System, PES Filter Material, 0.22 µm, 50 mm, Sterile | DOT Scientific | 667706 | |
| 385 nm, 1650 mW (Min) Mounted LED, 1700 mA | Thorlabs | M385LP1-C1 | |
| 75cm2 Tissue Culture Flask - Vent Cap, Sterile | Celltreat Scientific Products | 229341 | |
| 8.0 μm Cell Culture Plate Insert 12 mm Diameter | Millicell | PI8P01250 | |
| Absolute Ethanol, 200 proof, Molecular Biology Grade | Thermo Fisher Scientific | T038181000CS | Ethanol for flow cytometry dead cell control. |
| Astrocyte Medium (Astrofull) | ScienCell Research Laboratories | 1801 | Contains astrobasal, FBS, and penicillin/streptomycin. |
| B-27 Supplement (50x), minus vitamin A (Gibco) | Thermo Fisher Scientific | 12587010 | |
| BSA (MACS) | Miltenyi Biotec | 130-091-376 | |
| CD71 antibody (eBioscience, Invitrogen) | Fisher Scientific | 25-0719-41 | |
| Cell Counting Chambered Slides | Nexcelom Bioscience | CHT4-PD100-002 | |
| Cell Scrapers | Biologix USA | 70-1250 | |
| Cellometer K2 Fluorescent Cell Counter (Nexelcom Bioscience) | VWR | NEXCCMK2-SK150-FCS | |
| Centrifuge - Low-Speed | Eppendorf | 5702 R | Centrifuge for cell culture. |
| Clear Polystyrene 96-Well Microplates, Corning | Fisher Scientific | 07-200-108 | V-bottom plates for flow cytometry staining. |
| CO2 Incubator, 150L, Heracell 150i (Thermo Scientific) | Thermo Fisher Scientific | 50116047 | |
| Collagen I, High Concentration, Rat Tail | Corning | 354249 | |
| Collagen I, Rat Tail | Corning | 354236 | "Low" concentration for coating adherent flasks. |
| Collagenase (CAS# 9001-12-1) | United States Biological | C7511-30 | |
| Collimation Adapter for Olympus BX & IX, AR Coating: 350 - 700 nm | Thorlabs | COP1-A | |
| Cotton Swabs, Q-tips Precision Tips | Amazon | B01KCJB3R2 | |
| Dispase (CAS# 9001-92-7) | United States Biological | D3760 | |
| DMEM, high glucose (Gibco) | Thermo Fisher Scientific | 11330032 | |
| EVOS FL | Invitrogen | AMF4300 | |
| Fetal Bovine Serum (Gibco) | Thermo Fisher Scientific | 26140079 | For microglia culture. |
| Formalin solution, neutral buffered, 10% (Sigma-Aldrich) | Millipore Sigma | HT501128 | |
| Foxp3 / Transcription Factor Staining Buffer Set (eBioscience, Invitrogen) | Thermo Fisher Scientific | 00-5523-00 | |
| Glioma stem cells | n/a | n/a | Can be patient derived or commercial glioma stem cell lines. |
| Guava easyCyte HT System | Millipore Sigma | 0500-4008 | Flow cytometer. |
| HBSS (Sigma-Aldrich) | Millipore Sigma | H6648 | |
| HEPES (1 M) (Gibco) | Thermo Fisher Scientific | 15630080 | |
| High-Power 1-Channel LED Driver with Pulse Modulation, 10.0 A Max, 50.0 V Max | Thorlabs | DC2200 | Interface for UV Lamp. |
| Hoechst 33342 Solution (20 mM) (Thermo Scientific) | Thermo Fisher Scientific | 62249 | |
| Human Astrocytes | ScienCell Research Laboratories | 1800 | Primary astrocytes derived from the cerebral cortex. |
| Human EGF Recombinant Protein (Gibco) | Thermo Fisher Scientific | PHG0311 | |
| Human FGF-basic (FGF-2/bFGF) (aa 10-155) Recombinant Protein (Gibco) | Thermo Fisher Scientific | PHG0021 | |
| Immortalized Human Microglia - hTERT | Applied Biological Materials | T0251 | |
| Incu-mixer MP Heated Microplate Vortexer, 2 position | Benchmark Scientific | H6002 | |
| Ki67 REAfinity, PerCP-Vio 700 antibody | Miltenyi Biotec | 130-120-420 | |
| LED UV Curing Meter | Gigahertz-Optik | X1-RCH-116 | Optometer to measure UV light intensity. |
| Lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP) | Advanced Biomatrix | 5269-100MG | Photoinitiator. |
| LIVE/DEAD Fixable Near-IR (Invitrogen) | Thermo Fisher Scientific | L24975 | |
| Methacrylated hyaluronic acid (photoHA) | Advanced Biomatrix | 5212-100MG | |
| Microcentrifuge, Sorvall ST8R (Thermo Scientific) | Fisher Scientific | 75-997-203 | Centrifuge for flow cytometry staining. |
| N-2 Supplement (100X) (Gibco) | Thermo Fisher Scientific | 17502048 | |
| Neurobasal-A Medium (Gibco) | Thermo Fisher Scientific | 10888022 | |
| PBS (10X), pH 7.4 without Ca & Mg | Quality Biological | 119-069-101 | |
| Sodium hydroxide, pellets ACS (CAS# 1310-73-2) | VWR | 97064-476 | |
| Synergy Ultrapure Water Purification System (MilliporeSigma) | Fisher Scientific | SYNS0HFUS | |
| Trypsin-EDTA (0.25%), phenol red (Gibco) | Thermo Fisher Scientific | 25200056 | |
| ViaStain AOPI Staining Solution | Nexcelom Bioscience | CS2-0106 |
References
- Ostrom, Q. T., et al. CBTRUS statistical report: Primary brain and other central nervous system tumors diagnosed in the United States in 2016-2020. Neuro Oncol. 25, 1-99 (2023).
- Stupp, R., et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 352 (10), 987-996 (2005).
- Stupp, R., et al. Effect of tumor-treating fields plus maintenance temozolomide vs maintenance temozolomide alone on survival in patients with glioblastoma: A randomized clinical trial. JAMA. 318 (23), 2306-2316 (2017).
- Lathia, J. D., Mack, S. C., Mulkearns-Hubert, E. E., Valentim, C. L. L., Rich, J. N. Cancer stem cells in glioblastoma. Genes Dev. 29 (12), 1203-1217 (2015).
- Roos, A., Ding, Z., Loftus, J. C., Tran, N. L. Molecular and microenvironmental determinants of glioma stem-like cell survival and invasion. Front Oncol. 7, 120 (2017).
- Yabo, Y. A., Niclou, S. P., Golebiewska, A. Cancer cell heterogeneity and plasticity: A paradigm shift in glioblastoma. Neuro Oncol. 24 (5), 669-682 (2022).
- Geer, C. P., Grossman, S. A. Interstitial fluid flow along white matter tracts: A potentially important mechanism for the dissemination of primary brain tumors. J Neurooncol. 32 (3), 193-201 (1997).
- Munson, J. M., Bellamkonda, R. V., Swartz, M. A. Interstitial flow in a 3D microenvironment increases glioma invasion by a CXCR4-dependent mechanism. Cancer Res. 73 (5), 1536-1546 (2013).
- Kingsmore, K. M., et al. Interstitial flow differentially increases patient-derived glioblastoma stem cell invasion via CXCR4, CXCL12, and CD44-mediated mechanisms. Integr Biol. 8 (12), 1246-1260 (2016).
- Qazi, H., Shi, Z. -. D., Tarbell, J. M. Fluid shear stress regulates the invasive potential of glioma cells via modulation of migratory activity and matrix metalloproteinase expression. PLoS One. 6 (5), e20348 (2011).
- Parmigiani, E., Scalera, M., Mori, E., Tantillo, E., Vannini, E. Old stars and new players in the brain tumor microenvironment. Front Cell Neurosci. 15, 709917 (2021).
- Cornelison, R. C., et al. A patient-designed tissue-engineered model of the infiltrative glioblastoma microenvironment. NPJ Precis Oncol. 6 (1), 54 (2022).
- Tchafa, A. M., Shah, A. D., Wang, S., Duong, M. T., Shieh, A. C. Three-dimensional cell culture model for measuring the effects of interstitial fluid flow on tumor cell invasion. J Vis Exp. (65), e4159 (2012).
- Shields, J. D., et al. Autologous chemotaxis as a mechanism of tumor cell homing to lymphatics via interstitial flow and autocrine CCR7 signaling. Cancer Cell. 11 (6), 526-538 (2007).
- Shieh, A. C., Rozansky, H. A., Hinz, B., Swartz, M. A. Tumor cell invasion is promoted by interstitial flow-induced matrix priming by stromal fibroblasts. Cancer Res. 71 (3), 790-800 (2011).
- Miteva, D. O., et al. Transmural flow modulates cell and fluid transport functions of lymphatic endothelium. Circ Res. 106 (5), 920-931 (2010).
- Kingsmore, K. M., et al. MRI analysis to map interstitial flow in the brain tumor microenvironment. APL Bioeng. 2 (3), 031905 (2018).
- Roberts, L. M., et al. Myeloid derived suppressor cells migrate in response to flow and lymphatic endothelial cell interaction in the breast tumor microenvironment. Cancers. 14 (12), 3008 (2022).
- Bellail, A. C., Hunter, S. B., Brat, D. J., Tan, C., Van Meir, E. G. Microregional extracellular matrix heterogeneity in brain modulates glioma cell invasion. Int J Biochem Cell Biol. 36 (6), 1046-1069 (2004).
- Choi, J. R., Yong, K. W., Choi, J. Y., Cowie, A. C. Recent advances in photo-crosslinkable hydrogels for biomedical applications. Biotechniques. 66 (1), 40-53 (2019).
- Artym, V. V., Matsumoto, K. Imaging cells in three-dimensional collagen matrix. Curr Protoc Cell Biol. , 11-20 (2010).
- Gelman, R. A., Williams, B. R., Piez, K. A. Collagen fibril formation. Evidence for a multistep process. J Biol Chem. 254 (1), 180-186 (1979).
- Williams, B. R., Gelman, R. A., Poppke, D. C., Piez, K. A. Collagen fibril formation. Optimal in vitro conditions and preliminary kinetic results. J Biol Chem. 253 (18), 6578-6585 (1978).
- Mineo, M., et al. The long non-coding RNA HIF1A-AS2 facilitates the maintenance of mesenchymal glioblastoma stem-like cells in hypoxic niches. Cell Rep. 15 (11), 2500-2509 (2016).
- Tang, X., et al. Targeting glioblastoma stem cells: A review on biomarkers, signal pathways and targeted therapy. Front Oncol. 11, 701291 (2021).
- Dirkse, A., et al. Stem cell-associated heterogeneity in glioblastoma results from intrinsic tumor plasticity shaped by the microenvironment. Nat Commun. 10 (1), 1787 (2019).
- Sofroniew, M. V. Astrocyte reactivity: Subtypes, states, and functions in CNS innate immunity. Trends Immunol. 41 (9), 758-770 (2020).
- Tate, K. M., Munson, J. M. Assessing drug response in engineered brain microenvironments. Brain Res Bull. 150, 21-34 (2019).
- Cai, X., et al. Application of microfluidic devices for glioblastoma study: Current status and future directions. Biomed Microdevices. 22 (3), 60 (2020).
- Kleinman, H. K., Martin, G. R. Matrigel: Basement membrane matrix with biological activity. Semin Cancer Biol. 15 (5), 378-386 (2005).
- Jadin, L., et al. Hyaluronan expression in primary and secondary brain tumors. Ann Transl Med. 3 (6), 80 (2015).
- Cornelison, R. C., Munson, J. M. Perspective on translating biomaterials into glioma therapy: Lessons from in vitro models. Front Mater. 5, 27 (2018).
- Kapałczyńska, M., et al. 2D and 3D cell cultures - a comparison of different types of cancer cell cultures. Arch Med Sci. 14 (4), 910-919 (2018).
- Nelson, C. M., Bissell, M. J. Of extracellular matrix, scaffolds, and signaling: Tissue architecture regulates development, homeostasis, and cancer. Annu Rev Cell Dev Biol. 22, 287-309 (2006).
- Phon, B. W. S., Kamarudin, M. N. A., Bhuvanendran, S., Radhakrishnan, A. K. Transitioning pre-clinical glioblastoma models to clinical settings with biomarkers identified in 3D cell-based models: A systematic scoping review. Biomed Pharmacother. 145, 112396 (2022).
- Pampaloni, F., Reynaud, E. G., Stelzer, E. H. K. The third dimension bridges the gap between cell culture and live tissue. Nat Rev Mol Cell Biol. 8 (10), 839-845 (2007).
- Ortiz-Cárdenas, J. E., et al. Towards spatially-organized organs-on-chip: Photopatterning cell-laden thiol-ene and methacryloyl hydrogels in a microfluidic device. Organs Chip. 4, 100018 (2022).
- Ozulumba, T., et al. Mitigating reactive oxygen species production and increasing gel porosity improves lymphocyte motility and fibroblast spreading in photocrosslinked gelatin-thiol hydrogels. bioRxiv. , (2024).
- Lim, K. S., et al. Fundamentals and applications of photo-cross-linking in bioprinting. Chem Rev. 120 (19), 10662-10694 (2020).
- Ghosh, R. N., et al. An insight into synthesis, properties and applications of gelatin methacryloyl hydrogel for 3D bioprinting. Mater Adv. 4 (22), 5496-5529 (2023).
- Venkataramani, V., et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature. 573 (7775), 532-538 (2019).
- Venkataramani, V., et al. Glioblastoma hijacks neuronal mechanisms for brain invasion. Cell. 185 (16), 2899-2917 (2022).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved