A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Using Three-color Single-molecule FRET to Study the Correlation of Protein Interactions
In This Article
Summary
Here, we present a protocol to obtain three-color smFRET data and its analysis with a 3D ensemble Hidden Markov Model. With this approach, scientists can extract kinetic information from complex protein systems, including cooperativity or correlated interactions.
Abstract
Single-molecule Förster resonance energy transfer (smFRET) has become a widely used biophysical technique to study the dynamics of biomolecules. For many molecular machines in a cell proteins have to act together with interaction partners in a functional cycle to fulfill their task. The extension of two-color to multi-color smFRET makes it possible to simultaneously probe more than one interaction or conformational change. This not only adds a new dimension to smFRET experiments but it also offers the unique possibility to directly study the sequence of events and to detect correlated interactions when using an immobilized sample and a total internal reflection fluorescence microscope (TIRFM). Therefore, multi-color smFRET is a versatile tool for studying biomolecular complexes in a quantitative manner and in a previously unachievable detail.
Here, we demonstrate how to overcome the special challenges of multi-color smFRET experiments on proteins. We present detailed protocols for obtaining the data and for extracting kinetic information. This includes trace selection criteria, state separation, and the recovery of state trajectories from the noisy data using a 3D ensemble Hidden Markov Model (HMM). Compared to other methods, the kinetic information is not recovered from dwell time histograms but directly from the HMM. The maximum likelihood framework allows us to critically evaluate the kinetic model and to provide meaningful uncertainties for the rates.
By applying our method to the heat shock protein 90 (Hsp90), we are able to disentangle the nucleotide binding and the global conformational changes of the protein. This allows us to directly observe the cooperativity between the two nucleotide binding pockets of the Hsp90 dimer.
Introduction
Many proteins fulfill their function in dynamic complexes with other molecules, mediated by conformational changes and transient associations on a broad range of timescales1,2,3. Coupled to an external energy source (e.g., ATP) these dynamic interactions can lead to directionality in a functional cycle and ultimately maintain the non-equilibrium steady-state in a cell, the prerequisite for life.
In order to fully understand these molecular machines, a static description guided by structural studies is not sufficient. In addition, it is essential to have knowledge of the underlying kinetic model and to determine the kinetic rate constants. Several existing methods allow researchers to study the dynamics of binary interactions between two molecules of interest, e.g., surface plasmon resonance, relaxation methods with a spectroscopic readout (e.g., jump or stopped-flow techniques), and nuclear magnetic resonance. However, their applicability is in most cases limited to simple two-state systems (e.g., one bound and one unbound state) due to the averaging inherent to bulk experiments. In cases where more states or intermediates are involved, they yield only a complex mixture of the rate constants. Single-molecule methods such as optical or magnetic tweezers or two-color smFRET, i.e., one donor and one acceptor fluorophore, with a surface-immobilized sample can recover the rate constants for all observed conformational changes. However, when it comes to interactions affecting more than one binding site, these methods remain limited and the information on the possible correlation of the two (or more) interactions will only be accessible via indirect conclusions from a set of experiments.
Multi-color smFRET4,5,6,7,8,9 offers the opportunity to study the interaction between these components directly, at real time and under near-physiological conditions10. This permits one to investigate for example, the conformation-dependent binding of a ligand or another protein8,9,11. The overall approach presented here is to label the protein(s) of interest at specific positions, to attach one protein to the surface of the measurement chamber, and to track the fluorescence intensity over time on a prism-type TIRFM (for details see 9,12). The spatial proximity of the different dyes can then be determined from the energy transfer between them. Labeling strategies may vary from protein to protein (reviewed in 13) and guidelines to avoid artifacts in smFRET measurements exist14.
Since a donor dye may transfer energy to different acceptor dyes in a multi-color smFRET experiment, the relative position of all dyes is not accessible from excitation of one dye alone15,16. But in combination with alternating laser excitation (ALEX17, and reviewed in 18) this method provides all spatio-temporal information at sub-second and sub-nanometer resolution.
In principal, high resolution structural information can be achieved by using the inter-dye distances calculated from the combination of all fluorescence intensities in a multi-color smFRET experiment with ALEX. However, here we focus on state identification and separation as well as the extraction of kinetic models, where multi-color smFRET is indispensable. When "only" structure determination by triangulation is desired, a set of simpler two-color smFRET experiments with high signal-to-noise ratio can be performed12,19.
We use the partial fluorescence ( ) as a proxy for the energy transfer between two fluorophores7. The PF is calculated from the fluorescence intensity analogous to the FRET efficiency of a two-color experiment:
) as a proxy for the energy transfer between two fluorophores7. The PF is calculated from the fluorescence intensity analogous to the FRET efficiency of a two-color experiment:

Where, 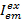 is the intensity in emission channel em after excitation with color ex, and c is the acceptor with the longest wavelength. Detection channels represent the same position in the sample chamber but record different spectral ranges of the fluorescence light. The same identifier for excitation and emission are used in this protocol (i.e., "blue," "green," and "red").
is the intensity in emission channel em after excitation with color ex, and c is the acceptor with the longest wavelength. Detection channels represent the same position in the sample chamber but record different spectral ranges of the fluorescence light. The same identifier for excitation and emission are used in this protocol (i.e., "blue," "green," and "red").
Because of experimental shortcomings the measured fluorescence intensities depend not only on the energy transfer but also on fluorophore and setup properties. In order to obtain the true energy transfer efficiency between two fluorophores, the measured intensities have to be corrected. The following procedure is based on reference9. Correction factors for apparent leakage (lk, i.e., the detection of photons from a fluorophore in a channel designated for another dye) and apparent gamma (ag, i.e., the fluorescence quantum yield of the dye and the detection efficiency of the channel) are obtained from single-molecule traces that show an acceptor bleaching event.
The leakage of the donor dye into every possible acceptor channel is calculated from all data points in the recorded fluorescence traces where the acceptor dye bleached but the donor is still fluorescent (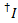 ):
):
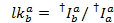
The median of the leakage histogram is used as the apparent leakage factor. After correction for leakage, the apparent gamma factor is determined from the same set of traces. It is calculated by dividing the change of fluorescence in the acceptor channel by the change of fluorescence in the donor channel upon bleaching of the acceptor dye:

Where c again is the detection channel for the acceptor with the longest wavelength. The median of the resulting distribution is used as the apparent correction factor.
The corrected intensities in each channel are obtained by:
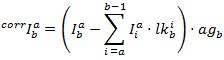
The PF is then calculated according to:

Different populations can be separated in the multi-dimensional space spanned by the PFs. The position and width of each state is determined by fitting the data with multi-dimensional Gaussian functions. Subsequent optimization of one global HMM based on all PF traces provides a quantitative description of the observed kinetics. Even small changes of the rates are detectable.
HMMs provide a way of inferring a state model from a collection of noisy time traces. The system is considered to be in one of a set of discrete, hidden states at any given time and the actual observation (i.e., the emission) is a probabilistic function of this hidden state20. In the case of TIRFM smFRET data, the emission probabilities bi per state i can be modeled by continuous Gaussian probability density functions. At regularly spaced discrete time points, transitions from one to another state can occur according to the transition probability that is time-invariant and only depends on the current state. The transition matrix A contains these transition probabilities aij between all hidden states. The initial state distribution  gives the state-specific probabilities
gives the state-specific probabilities 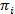 for the first time point of a time trace. Using a maximum-likelihood approach, these parameters can be optimized to best describe the data with the Forward-Backward and Baum-Welch algorithms20,21. This yields the maximum likelihood estimators (MLE). Finally, the state sequence that most likely produced the trajectory of observations can be inferred with the Viterbi algorithm. In contrast to other HMM analyses of smFRET data24,25,26 we do not use the HMM as a mere "smoothing" of the data but extract the kinetic state model from the data set without the need for fitting dwell time histograms27. HMM analysis is done with in-house scripts using Igor Pro. Implementation of the code is based on reference21. We provide a software kit and exemplary data on our webpage in order to follow sections 5 and 6 of this protocol (https://www.singlemolecule.uni-freiburg.de/software/3d-fret). Full software is available upon request.
for the first time point of a time trace. Using a maximum-likelihood approach, these parameters can be optimized to best describe the data with the Forward-Backward and Baum-Welch algorithms20,21. This yields the maximum likelihood estimators (MLE). Finally, the state sequence that most likely produced the trajectory of observations can be inferred with the Viterbi algorithm. In contrast to other HMM analyses of smFRET data24,25,26 we do not use the HMM as a mere "smoothing" of the data but extract the kinetic state model from the data set without the need for fitting dwell time histograms27. HMM analysis is done with in-house scripts using Igor Pro. Implementation of the code is based on reference21. We provide a software kit and exemplary data on our webpage in order to follow sections 5 and 6 of this protocol (https://www.singlemolecule.uni-freiburg.de/software/3d-fret). Full software is available upon request.
Time points in the data with PF <-1 or PF >2 in any detection channel are assigned the minimal emission probability for all states (10-200). This prevents artificial transitions at these data points.
The parameters for the emission probabilities are obtained from the fit of the 3D PF histogram with Gaussian functions as described in step 5.7. These parameters are kept fixed during the optimization of the HMM.
In the presented approach, the initial state distribution vector and the transition matrix are used globally to describe the entire ensemble of traces. They are updated based on all N molecules from the data set according to reference27.
Start parameters for the initial state distribution are determined from 2D projections of the PF histogram (step 5.3) and the transition probabilities are set to 0.05 with the exception of the probabilities to stay in the same state, which are chosen such that the probability to leave a certain state is normalized to unity.
A likelihood profiling method is used to give confidence intervals (CIs) for all transition rates21,22, which serve as meaningful estimates for their uncertainty. To calculate the bounds of the CI for a specific rate, the transition probability of interest is fixed to a value other than the MLE. This yields the test model λ'. A likelihood ratio (LR) test of the likelihood 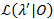 given the data set 0 is performed according to:
given the data set 0 is performed according to:
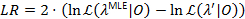
The 95% confidence bound for the parameter is reached when LR exceeds 3.841, the 95% quantile of a x2-distribution with one degree of freedom22,23.
The power of the method is demonstrated using the Hsp90. This abundant protein is found in bacteria and eukaryotes and is part of the cellular stress response28. It is a promising drug target in cancer treatment29. Hsp90 is a homodimer with one nucleotide binding pocket in the N-terminal domain of each subunit30. It can undergo transitions between at least two globally distinct conformations, one closed and one N-terminal open, V-shaped conformation19,31,32. The dimeric nature directly raises the question of the interplay between the two nucleotide binding sites in Hsp90.
In the following, we provide a step-by-step protocol for the data acquisition and analysis of a three-color smFRET experiment on yeast Hsp90 and nucleotide. The conformation-dependent binding of fluorescently labeled AMP-PNP (AMP-PNP*, a non-hydrolyzable analog of ATP) is analyzed. The application of the described procedure permits the study of the nucleotide binding and at the same time the conformational changes of Hsp90 and thereby reveals the cooperativity between the two nucleotide binding pockets of Hsp90.
Protocol
1. Setup and Prerequisites
- Perform the multi-color smFRET measurements on a prism-type TIRFM. A description of a two-color setup as a JoVE publication is given in reference12.
- Construct a multi-color TIRFM. A general layout is detailed in 9.
- Use switchable, diode pumped solid state continuous wave lasers, which render the use of mechanical shutters in the excitation paths unnecessary.
- Employ an asymmetric, elongated prism that prevents the back reflection of the excitation beam from the rear side to enter the objective.
- Use 2-inch achromatic aspheric fused silica lenses in the detection paths that collect as much light as possible and prevent auto-fluorescence and aberrations, e.g., distortions in off-center regions of the image.
- Focus each detection path on the chip of the EMCCD with a separate lens. This allows optimal focusing of each detection channel.
Caution: Class 3B lasers are used in the TIRFM. This means they are hazardous if the eye is exposed directly, but diffuse reflections are not harmful. Ensure compliance with laser safety precautions according to local government regulations before the system is operated.
- Determine the correction factors for setup and fluorophore properties beforehand using dsDNA samples.
- Use one high-FRET dsDNA sample for each dye in combination with the acceptor having the longest excitation wavelength (for the presented setup: Atto488-Atto647N, Atto550-Atto647N, Atto594-Atto647N). Make sure that the DNA is additionally modified with a biotin.
- Dilute the sample to 5 nM with TNM buffer (5 mM Tris pH 7.5, 5 mM NaCl, 20 mM MgCl2) and 2 mM Trolox (use this buffer also for measurement).
- Immobilize the dsDNA as described for Hsp90 in steps 2.5 and 2.7.
- Calculate the correction factors for apparent leakage (lk) and apparent gamma (ag) from single-molecule traces that show an acceptor bleaching event.
- Construct a flow chamber that is a sandwich of a PEG/biotin-PEG passivated quartz slide, a thin film that is adhesive on both sides, and a cover slip. For detail protocols for the cleaning of the quartz slides and passivation see reference9.
- Use thick (3 mm) quartz slides to geometrically prevent the collection of laser light scattered at the quartz-glycerol-quartz interface between the prism and the functionalized quartz slide.
- Use a thin (40 µm) sealing film that is sprayed with adhesive on the non-adhesive side. The thin film reduces the distance between the surface-attached molecules and objective. Heat to 80 °C and press on.
- Place a cover slip on the top. Heat to 80 °C and press on. Use a drop of glycerol when placing the flow chamber over the prism.
NOTE: The materials of the prism and the quartz slides as well as the glycerol are index-matched.
- Express Hsp90 from Saccharomyces cerevisiae in the form of two single cysteine point mutants at the positions D61 or Q385. Add a C-terminal coiled-coil motif to prevent dimer dissociation at picomolar concentrations. Label the mutant proteins separately and exchange the monomers to obtain heterodimers labeled with Atto488 at amino acid position 61 and Atto550 at amino acid position 3859.
2. Measurement
- Start the camera software and set the imaging parameters as specified below:
- Set the temperature for the sensor cooling as low as possible (-95 °C with external water cooling) to decrease the dark current noise.
- Use camera settings that are optimized for single-molecule recording: 3.3 µs vertical shift speed, normal vertical clock voltage, 17 MHz 16-bit horizontal read out, pre-amplification gain 3, gain of electron multiplier 1,000.
- Set the triggering of the acquisition to "External" and the exposure time to 70 ms. Record movies with a length of 750 acquisition cycles.
NOTE: Turn off the room light when the camera is acquiring to prevent saturation of the EMCCD sensor.
- Create a folder on the local SSD for the measurement. In the software settings go to the <Auto-Save> rider, enable <Auto-save>, and choose file format "Tiff" for movie acquisition. Select the folder on the SSD as auto-save location.
- Start the software that controls the acousto-optical tunable filter (AOTF), the software that controls the operation of the lasers and the trigger software that synchronizes the lasers, AOTF, shutters in the detection path, and cameras. Adjust the laser power with the AOTF (ca. 3 mW before entering the prism) and load the correct triggering pattern.
- Mount the sample holder with the prism and the flow chamber, attach tubing, place the inlet tubing in a microcentrifuge cup, and connect the outlet tubing to a syringe pump. Flush the chamber with approximately 150 µL buffer, align the excitation beam, and focus. Bleach any fluorescent contaminants on the surface by slowly moving along the complete detection range of the slide with a laser power of about 10 mW for all lasers. This takes about 1 h.
NOTE: If not stated otherwise, the used buffer contains 40 mM HEPES pH 7.5, 150 mM KCl, and 10 mM MgCl2. - Flush approximately 300 µL of a NeutrAvidin solution (0.25 mg/mL in buffer) into the chamber and incubate for 1 min. Flush out unbound NeutrAvidin with buffer and flush approximately 300 µL of a BSA solution (0.5 mg/mL in buffer) through the chamber.
NOTE: This block remaining surface functionalization defects by unspecific adsorption of BSA to the surface. - Immobilize the sample by loading the flow chamber with approximately 150 µL of the biotinylated and labeled Hsp90 at rising concentrations (diluted in buffer + 0.5 mg/mL BSA) until a sufficient surface density is reached, which is usually the case at a concentration of 5 - 10 pM. Wash out unbound protein with approximately 300 µL buffer + 0.5 mg/mL BSA.
- Flush in 150 µL of 25 nM AMP-PNP* in buffer + 0.5 mg/mL BSA. Let it incubate for 5 min and repeat this step once to ensure correct nucleotide concentration.
NOTE: For experiments in the presence of additional, unlabeled AMP-PNP, also add 250 µM AMP-PNP. - Begin with the data acquisition. For an appropriate amount of data acquire approximately 20 movies which takes about 1.5 h.
- Move the position of the sample chamber perpendicular to the excitation beam with a piezo stepper to change the field of view.
- Adjust the focus with the z-piezo that controls the height of the objective if needed. This should not be necessary too often when the measurement chamber is mounted without inclination.
- Prepare the recording of the cameras by pressing <Take Signal> in the camera software and start the excitation/acquisition cycles using the <Start> button in the trigger software. This starts the acquisition of the fluorescence intensity.
- Perform a channel registration by first recording a movie with fluorescent beads that show fluorescence emission in the spectral range of all detection channels of the setup. Then, detect bead positions in the calibration movie by searching for the brightest spots and determine the central position from a Gaussian fit to the intensity profile. Save the coordinates of beads that are found in all channels and fit both the mapping offset in x- and in y-direction with a 2D polynomial of degree three9.
3. Selection of Single-molecule Traces
- Data analysis is done with in-house scripts using Igor Pro. Load all necessary scripts by opening "iniTIRF.ipf" and select the correct type of experiment.
NOTE: In the following, <Button> specifies clickable elements in the menu or the user interface. Function calls are indicated in quotes, e.g., "Print "Hello World"". These commands can be pasted to the command line of Igor Pro (without the enclosing quotes). - Make sure the parameters for the detection channel registration are loaded.
- Start the GUI by clicking <smFRET new|Analysis GUI>.
- Load the movies (i.e., the sequence of frames with 512 x 512 pixels stored as 16 bit TIFF stacks) that hold the intensity in the respective channels. Do this by pressing the <Load Film> button and selecting the files from the so-called "master" and "slave" cameras one after the other.
- Identify the positions of potential single molecules by searching for the brightest spots in the sum of the first five frames in a certain detection channel. Calculate the corresponding positions in all other detection channels from the channel mapping. To obtain the fluorescence intensity trace, sum the intensity of a pixel square around the central position for each frame. Do this by pressing the button <Find Traces> in the GUI.
NOTE: The side length of the square (in pixel) is given by: 2 * <Sum Pxs> + 1. - For each molecule, calculate a joint raw intensity trace as the sum of all traces of this spot with the same excitation color. Evaluate the intensity profile of the molecule in all channels for the following criteria:
- A roughly flat plateau in the joint raw intensity and a single bleaching step for all excitation colors, anticorrelated behavior in the appropriate detection channels, detection of red fluorescence (indicating FRET to a bound AMP-PNP*) at least once within the trace, and no multiple steps in the red fluorescence, which would indicate the presence of two AMP-PNP* bound to one Hsp90 dimer.
- Save the fluorescence traces of the spot for further analysis if these criteria are fulfilled. Do this by selecting the trace with the cursor and then pressing the <Save> button in the "timelines" graph. Manually inspect the intensity traces in all three detection channels after blue excitation for about 200 molecules per movie.
4. Calculation of the Partial Fluorescence Traces
- Display all fluorescence intensity traces for one of the saved molecules. Use the cursors in the graph to select time ranges.
- Select a time interval where all fluorophores are bleached already. The mean background intensity is calculated from this range and subtracted from the intensity trace in each channel. Do this by pressing the <Background> button.
- Select the FRET efficiency range, where at least both the dyes attached to Hsp90 (Atto488 and Atto550) are present. Make sure to exclude traces that contain a blinking event (see Figure 3B). These events are characterized by a drop-in fluorescence intensity in one channel without an accompanying increase in any other channel.
- Calculate the PF traces. Do this by pressing the <PF Calc> button. The predefined correction factors for apparent leakage (lk) and apparent gamma (ag) are applied to the raw intensity in order to correct for photo-physical and setup properties.
5. Population Selection and 3D Histogram Fitting
- Remove molecules that show a low signal-to-noise ratio in the PF traces. Molecules that exceed the interval [-1;2] in any PF trace for more than 10% of the frames are removed from the data set. Do this by executing "RemoveTracesLowSNR()" from the Command Window.
- Calculate binned 2D projections of the PF data. Plot
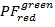 over
over 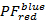 and
and 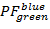 over
over 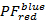 in the range [-0.5; 1.5] with a resolution of 100 x 100 bins. To do so, execute:
in the range [-0.5; 1.5] with a resolution of 100 x 100 bins. To do so, execute:
- "HistFret2D("r_b", "r_g", binHist=100)"
- "HistFret2D("r_b", "g_b", binHist=100); MoveWindow 553.5, 42.5, 1055.25, 508.25"
- Determine the relative population of each distinguishable state in the 2D projections.
- Bring the appropriate graph to the front and execute "panelHist2DCount()".
- Press the <Init> button and draw a free-hand polygon around the peak.
- Click the <Count> button. The number of data points in the polygon and the total number of data points in the projection are printed in the Command Window.
- Prepare a 3D histogram of the PF data by executing "HistFret3D("g_b", "r_b", "r_g")".
- Normalize the 3D histogram to an integral of 1. Execute the following:
- "NewDataFolder/S fit0"
- "Duplicate/O ::FRET:Hist3D, Hist3D"
- "Variable /G div = sum(Hist3D)*(DimDelta(Hist3D,0))^3"
- "Hist3D /= div; Print div"
- Provide initial parameters for the 3D Gaussian fit and prepare necessary data structures.
- Execute "Gauss3D_initParam(); edit W_coef_old."
- Add the state populations to the end of the parameter vector.
- Execute "Gauss3D_prepareFit()."
Note: W_coef_old is a vector that holds the initial parameters for the fit. Per state this means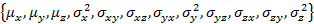 and the state population, which is concatenated at the end of the vector. Make sure that the covariance matrix is symmetric.
and the state population, which is concatenated at the end of the vector. Make sure that the covariance matrix is symmetric.
- Fit the sum of S 3D Gaussian functions to the 3D PF histogram, with S being the number of distinguishable states.
- Execute "do3D()." This may take an hour or more on a normal office PC, depending on the quality of the initial parameters.
- Execute "postprocessFitMultiGauss3D(); evalFitMultiGauss3D(); edit W_coef."
- Display the fit result. For each of the two 2D projections, use the following commands:
- "contourPF3D_new(0); contourPF3D_new(1); contourPF3D_new(2); contourPF3D_new(3); contourPF3D_new(4)"
- "contourPF3D_colorize()"
6. Kinetic Analysis with 3D Ensemble HMM
- Prepare an ensemble HMM run to extract the kinetic information. One HMM is optimized from all molecules in the data set. Use the information obtained in the previous step to define the position and width of each state in the 3D PF space.
- Initialize the HMM user interface (<HMM|Init HMM>) and choose the appropriate number of states (in the case of the Hsp90 data this means <NumStates> = 5), number of dimensions of the input signal (<NumDims> = 3), and the type of the input (<Input Type> = "FRET 3D bgr").
- Optimize the parameters of the HMM by letting the software converge the likelihood of the HMM (execute "prepENS_CONVERGE_gB(GetDataFolder(1), -14)") until the change in the transition matrix compared to the previous iteration falls below a threshold (10-14 for the sum of the absolute change for each transition probability). This yields the MLE for the transition probabilities in about an hour on a normal office PC.
- Repeat the population selection, Gaussian fitting, and the HMM optimization for subsets of the data (e.g., 75% of the full data set). If the full data set was merged from different experiments, repeat the optimization also for each of the single experiments. Analysis of subsets permits to estimate the uncertainty of the manual population selection and the variability within the data set.
- Calculate the CI for the transition probabilities, which report on the data set heterogeneity and the precision of the HMM.
- Get a rough estimate of the CI bounds by executing "cd $(root:path3Dimport + "HMM"); loop_getCI_estimate_limits()."
- Calculate the exact limits of the CI by executing:
- "loop_getCI_HMM_converge(1)"
- "CIresults_conv_new()"
- "cd ::HMM_CIresult; reportCI_conv()"
- "cd ::cmp_CI_conv; CI_plot2("HMM", doAppend=0)"
- Condense the available kinetic information to simplify the interpretation.
- Collect information about the time a labeled nucleotide stays bound to Hsp90. Do this by collapsing the states that are bound to AMP-PNP* (i.e., S1, S2 and S3, see also Figure 2) and compile the dwell time histogram. Execute "cd $(root:path3Dimport +"HMM"); collapse_states_get_DT({0,1,1,1,0})", which combines dwells of states S0 and S4 as well as S1, S2, and S3.
- Extract the dwell times from the Viterbi path for each state of interest and compare the resulting dwell time histogram for different experimental conditions. Execute "plot_collapsed_DT_Hist (wDTo_01110_record1)."
- For a more detailed picture, collapse states that have distinct PF but are functionally identical to ease the further data analysis, e.g., in the case of a three-color experiment with labeled Hsp90 and nucleotide, states S2 and S3 can be collapsed. Execute "collapse_states_get_DT({0,1,2,2,4})."
Results
Multi-color smFRET measurements allow the direct detection of correlation between two or more distinct interaction sites. This renders the technique unique to investigate multi-component systems, such as protein complexes. We focus on the presentation of a three-color smFRET experiment here, which serves as an illustrative example.
The general workflow of the method is shown in Figure 1. The first p...
Discussion
We present the experimental procedure to obtain three-color smFRET data for a complex protein system and a step-by-step description of the analysis of these measurements. This approach offers the unique possibility to directly assess the correlation between multiple interaction sites or conformational changes.
In order to obtain suitable multi-color single-molecule data on proteins it is important to perform reproducible measurements at a low noise level. This can be achieved by using an effic...
Disclosures
The authors declare no conflicts of interest.
Acknowledgements
This work is funded by the German Research Foundation (INST 39/969-1) and the European Research Council through the ERC Grant Agreement n. 681891.
Materials
| Name | Company | Catalog Number | Comments |
| Setup | |||
| vibration-damped optical table | Newport, Irvine, CA, USA | RS2000 | |
| OBIS 473nm LX 75mW LASER | Coherent Inc, Santa Clara, CA, USA | 1185052 | |
| OBIS 532nm LS 50mW LASER | Coherent Inc, Santa Clara, CA, USA | 1261779 | |
| OBIS 594nm LS 60mW LASER | Coherent Inc, Santa Clara, CA, USA | 1233470 | |
| OBIS 637nm LX 140mW LASER | Coherent Inc, Santa Clara, CA, USA | 1196625 | |
| laser control unit | Coherent Inc, Santa Clara, CA, USA | 1234465 | Scientific Remote |
| aspheric telescope lenses | Thorlabs Inc, Newton, New Jersey, USA | d=25.4mm, f=50mm and f=100mm | |
| CF ex1 | AHF analysentechnik AG, Tübingen, Germany | ZET 473/10 | cleanup filter excitation |
| CF ex2 | AHF analysentechnik AG, Tübingen, Germany | ZET 532/10 | cleanup filter excitation |
| CF ex3 | AHF analysentechnik AG, Tübingen, Germany | ZET 594/10 | cleanup filter excitation |
| CF ex4 | Thorlabs Inc, Newton, New Jersey, USA | FL635-10 | cleanup filter excitation |
| DM ex1 | AHF analysentechnik AG, Tübingen, Germany | ZQ594RDC | dichroic mirror excitation |
| DM ex2 | AHF analysentechnik AG, Tübingen, Germany | 570DCXR | dichroic mirror excitation |
| DM ex3 | AHF analysentechnik AG, Tübingen, Germany | ZQ491RDC | dichroic mirror excitation |
| AOTFnC-Vis | AA Opto-Electronic, Orsay, France | ||
| λ/4 plate | Thorlabs Inc, Newton, New Jersey, USA | AQWP05M-600 | |
| CFI Apo TIRF 100x | Nikon Instruments Inc, Melville, NY, USA | high-NA objective | |
| piezo focus positioner MIPOS 250 CAP | piezosystem jena GmbH, Jena, Germany | Piezo Controller NV 40/1 CLE | |
| piezo stepper | Newport, Irvine, CA, USA | PZA12 | PZC200-KT NanoPZ Actuator Kit |
| achromatic aspheric lenses | Qioptiq Photonics GmbH & Co. KG, Göttingen, Germany | G322-304-000 | d=50mm, f=200mm |
| adjustable optical slit | Owis GmbH, Staufen i. Br., Germany | 27.160.1212 | max. aperture 12 x 12 mm |
| DM det1 | AHF analysentechnik AG, Tübingen, Germany | T 600 LPXR | dichroic mirror detection |
| DM det2 | AHF analysentechnik AG, Tübingen, Germany | H 560 LPXR superflat | dichroic mirror detection |
| DM det3 | AHF analysentechnik AG, Tübingen, Germany | HC BS R635 | dichroic mirror detection |
| BP det1 | AHF analysentechnik AG, Tübingen, Germany | 525/40 BrightLine HC | bandpass filter detection |
| BP det2 | AHF analysentechnik AG, Tübingen, Germany | 586/20 BrightLine HC | bandpass filter detection |
| BP det3 | AHF analysentechnik AG, Tübingen, Germany | 631/36 BrightLine HC | bandpass filter detection |
| BP det4 | AHF analysentechnik AG, Tübingen, Germany | 700/75 ET Bandpass | bandpass filter detection |
| optical shutters detection | Vincent Associates, Rochester, NY, USA | Uniblitz VS25S2T0 | |
| EMCCD iXon Ultra 897 | Andor Technology Ltd, Belfast, Northern Ireland | ||
| digital I/O card, PCIe-6535 | National Instruments, Austin, Texas, USA | ||
| syringe pump | Harvard Apparatus, Holliston, MA, USA | PHD22/2000 | |
| Name | Company | Catalog Number | Comments |
| Flow chamber | |||
| quartz slides | G. Finkenbeiner Inc, Waltham, MA, USA | Spectrosil2000, h=3mm | |
| TEGADERM film | 3M Deutschland GmbH, Neuss, Germany | 1626W | 10 x 12cm |
| spray adhesive | 3M Deutschland GmbH, Neuss, Germany | Photo Mount 050777 | |
| glycerol | Carl Zeiss AG, Oberkochen, Germany | Immersol G | |
| immersion oil | OLYMPUS EUROPA SE & CO. KG, Hamburg, Germany | IMMOIL-F30CC | |
| prism | Vogelsberger Quarzglastechnik GmbH, Hauzenberg, Germany | Suprasil1 | |
| aluminium prism holder | custom built | ||
| hollow setscrews | Thorlabs Inc, Newton, New Jersey, USA | with custom drilling | |
| Tygon S3 E-3603 tubing | neoLab Migge GmbH, Heidelberg, Germany | 2-4450 | ACF00001 |
| PTFE tubing | Bohlender GmbH, Grünsfeld, Germany | S1810-08 | |
| Name | Company | Catalog Number | Comments |
| Sample | |||
| yeast Hsp90 D61C, Q385C_biotin | UniProt ID P02829 | ||
| Maleimide derivatives of Atto488, Atto550 | ATTO-TEC GmbH, Siegen, Germany | ||
| AMP-PNP* | Jena Bioscience, Jena, Germany | γ-[(6-Aminohexyl)-imido]-AMP-PNP-Atto647N | |
| Fluospheres | Thermo Fisher Scientific, Waltham, MA, USA | F8764 | amine-modified, 0.2 μm, yellow-green fluorescent |
| Name | Company | Catalog Number | Comments |
| Software | |||
| Andor Solis | Andor Technology Ltd, Belfast, Northern Ireland | version 4.30 | |
| LabVIEW | National Instruments, Austin, Texas, USA | version 2012, 32bit; misc. hardware control | |
| MDS control software | AA Opto-Electronic, Orsay, France | version 2.03a | |
| Coherent Connection | Coherent Inc, Santa Clara, CA, USA | version 3 | |
| Igor Pro | WaveMetrics Inc, Portland, OR, USA | version 6.37 |
References
- Nooren, I. M. A., Thornton, J. M. Diversity of protein-protein interactions. EMBO J. 22 (14), 3486-3492 (2003).
- Marsh, J. A., Teichmann, S. A. Structure, dynamics, assembly, and evolution of protein complexes. Annu Rev Biochem. 84, 551-575 (2015).
- Henzler-Wildman, K., Kern, D. Dynamic personalities of proteins. Nature. 450 (7172), 964-972 (2007).
- Hohng, S., Joo, C., Ha, T. Single-Molecule Three-Color FRET. Biophys J. 87 (2), 1328-1337 (2004).
- Person, B., Stein, I. H., Steinhauer, C., Vogelsang, J., Tinnefeld, P. Correlated movement and bending of nucleic acid structures visualized by multicolor single-molecule spectroscopy. ChemPhysChem. 10 (9-10), 1455-1460 (2009).
- Lee, J., Lee, S., Ragunathan, K., Joo, C., Ha, T., Hohng, S. Single-molecule four-color FRET. Angew Chem Int Ed. 49 (51), 9922-9925 (2010).
- Ratzke, C., Berkemeier, F., Hugel, T. Heat shock protein 90's mechanochemical cycle is dominated by thermal fluctuations. Proc Natl Acad Sci U S A. 109 (1), 161-166 (2012).
- Ratzke, C., Hellenkamp, B., Hugel, T. Four-colour FRET reveals directionality in the Hsp90 multicomponent machinery. Nat Commun. 5, 4192 (2014).
- Götz, M., Wortmann, P., Schmid, S., Hugel, T. A Multicolor Single-Molecule FRET Approach to Study Protein Dynamics and Interactions Simultaneously. Methods Enzymol. 581, 487-516 (2016).
- Yengo, C. M., Berger, C. L. Fluorescence anisotropy and resonance energy transfer: Powerful tools for measuring real time protein dynamics in a physiological environment. Curr Opin Pharmacol. 10 (6), 731-737 (2010).
- Wortmann,P , ., Götz M, ., Hugel T , . Cooperative Nucleotide Binding in Hsp90 and Its Regulation by Aha1. Biophys J. 113, 1711-1718 (2017).
- Dörfler, T., Eilert, T., Röcker, C., Nagy, J., Michaelis, J. Structural Information from Single-molecule FRET Experiments Using the Fast Nano-positioning System. J Vis Exp. (120), e54782 (2017).
- Stephanopoulos, N., Francis, M. B. Choosing an effective protein bioconjugation strategy. Nature chemical biology. 7 (12), 876-884 (2011).
- Sánchez-Rico, C., Voith von Voithenberg, L., Warner, L., Lamb, D. C., Sattler, M. Effects of Fluorophore Attachment on Protein Conformation and Dynamics Studied by spFRET and NMR Spectroscopy. Chemistry (Weinheim an der Bergstrasse, Germany). , (2017).
- Roy, R., Hohng, S., Ha, T. A practical guide to single-molecule FRET. Nat Methods. 5 (6), 507-516 (2008).
- Lee, N. K., et al. Three-color alternating-laser excitation of single molecules: monitoring multiple interactions and distances. Biophys J. 92 (1), 303-312 (2007).
- Kapanidis, A. N., Lee, N. K., Laurence, T. A., Doose, S., Margeat, E., Weiss, S. Fluorescence-aided molecule sorting: analysis of structure and interactions by alternating-laser excitation of single molecules. Proc Natl Acad Sci U S A. 101 (24), 8936-8941 (2004).
- Hohlbein, J., Craggs, T. D., Cordes, T. Alternating-laser excitation: single-molecule FRET and beyond. Chem Soc Rev. 43 (4), 1156-1171 (2014).
- Hellenkamp, B., Wortmann, P., Kandzia, F., Zacharias, M., Hugel, T. Multidomain structure and correlated dynamics determined by self-consistent FRET networks. Nat Methods. 14, 174-180 (2017).
- Rabiner, L. R. A tutorial on hidden Markov models and selected applications in speech recognition. Proc IEEE. 77 (2), 257-286 (1989).
- Fink, G. A. . Markov Models for Pattern Recognition. , (2014).
- Giudici, P., Ryden, T., Vandekerkhove, P. Likelihood-Ratio Tests for Hidden Markov Models. Biometrics. 56 (3), 742-747 (2000).
- Visser, I., Raijmakers, M. E. J., Molenaar, P. C. M. Confidence intervals for hidden Markov model parameters. Br J Math Stat Psychol. 53 (2), 317-327 (2000).
- McKinney, S. A., Joo, C., Ha, T. Analysis of Single-Molecule FRET Trajectories Using Hidden Markov Modeling. Biophys J. 91 (5), 1941-1951 (2006).
- Bronson, J. E., Fei, J., Hofman, J. M., Gonzalez, R. L., Wiggins, C. H. Learning Rates and States from Biophysical Time Series: A Bayesian Approach to Model Selection and Single-Molecule FRET Data. Biophys J. 97 (12), 3196-3205 (2009).
- Greenfeld, M., Pavlichin, D. S., Mabuchi, H., Herschlag, D. Single Molecule Analysis Research Tool (SMART): an integrated approach for analyzing single molecule data. PLoS ONE. 7 (2), e30024 (2012).
- Schmid, S., Götz, M., Hugel, T. Single-Molecule Analysis beyond Dwell Times: Demonstration and Assessment in and out of Equilibrium. Biophys J. 111 (7), 1375-1384 (2016).
- Taipale, M., Jarosz, D. F., Lindquist, S. HSP90 at the hub of protein homeostasis: emerging mechanistic insights. Nat Rev Mol Cell Biol. 11 (7), 515-528 (2010).
- Trepel, J., Mollapour, M., Giaccone, G., Neckers, L. Targeting the dynamic HSP90 complex in cancer. Nat Rev Cancer. 10 (8), 537-549 (2010).
- Wayne, N., Bolon, D. N. Dimerization of Hsp90 is required for in vivo function. Design and analysis of monomers and dimers. J Biol Chem. 282 (48), 35386-35395 (2007).
- Ali, M. M. U., et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature. 440 (7087), 1013-1017 (2006).
- Southworth, D. R., Agard, D. A. Species-dependent ensembles of conserved conformational states define the Hsp90 chaperone ATPase cycle. Mol Cell. 32 (5), 631-640 (2008).
- Blanchard, S. C., Kim, H. D., Gonzalez, R. L., Puglisi, J. D., Chu, S. tRNA dynamics on the ribosome during translation. Proc Natl Acad Sci U S A. 101 (35), 12893-12898 (2004).
- Aitken, C. E., Marshall, R. A., Puglisi, J. D. An Oxygen Scavenging System for Improvement of Dye Stability in Single-Molecule Fluorescence Experiments. Biophys J. 94 (5), 1826-1835 (2008).
- Swoboda, M., et al. Enzymatic oxygen scavenging for photostability without pH drop in single-molecule experiments. ACS Nano. 6 (7), 6364-6369 (2012).
- Rognoni, L., Stigler, J., Pelz, B., Ylänne, J., Rief, M. Dynamic force sensing of filamin revealed in single-molecule experiments. Proc Natl Acad Sci U S A. 109 (48), 19679-19684 (2012).
- Okumus, B., Wilson, T. J., Lilley, D. M. J., Ha, T. Vesicle encapsulation studies reveal that single molecule ribozyme heterogeneities are intrinsic. Biophys J. 87 (4), 2798-2806 (2004).
- Boukobza, E., Sonnenfeld, A., Haran, G. Immobilization in Surface-Tethered Lipid Vesicles as a New Tool for Single Biomolecule Spectroscopy. J Phys Chem B. 105 (48), 12165-12170 (2001).
- Levene, M. J., Korlach, J., Turner, S. W., Foquet, M., Craighead, H. G., Webb, W. W. Zero-mode waveguides for single-molecule analysis at high concentrations. Science. 299 (5607), 682-686 (2003).
- Panaretou, B., et al. ATP binding and hydrolysis are essential to the function of the Hsp90 molecular chaperone in vivo. EMBO J. 17 (16), 4829-4836 (1998).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved