Method Article
Using the Protozoan Paramecium caudatum as a Vehicle for Food-borne Infections in Zebrafish Larvae
In This Article
Summary
Zebrafish (Danio rerio) are becoming a widely-used vertebrate animal model for microbial colonization and pathogenesis. This protocol describes the use of the protozoan Paramecium caudatum as a vehicle for food-borne infection in zebrafish larvae. P. caudatum readily internalizes bacteria and get taken up by larval zebrafish through natural preying behavior.
Abstract
Due to their transparency, genetic tractability, and ease of maintenance, zebrafish (Danio rerio) have become a widely-used vertebrate model for infectious diseases. Larval zebrafish naturally prey on the unicellular protozoan Paramecium caudatum. This protocol describes the use of P. caudatum as a vehicle for food-borne infection in larval zebrafish. P. caudatum internalize a wide range of bacteria and bacterial cells remain viable for several hours. Zebrafish then prey on P. caudatum, the bacterial load is released in the foregut upon digestion of the paramecium vehicle, and the bacteria colonize the intestinal tract. The protocol includes a detailed description of paramecia maintenance, loading with bacteria, determination of bacterial degradation and dose, as well as infection of zebrafish by feeding with paramecia. The advantage of using this method of food-borne infection is that it closely mimics the mode of infection observed in human disease, leads to more robust colonization compared to immersion protocols, and allows the study of a wide range of pathogens. Food-borne infection in the zebrafish model can be used to investigate bacterial gene expression within the host, host-pathogen interactions, and hallmarks of pathogenicity including bacterial burden, localization, dissemination and morbidity.
Introduction
Zebrafish share morphologically and functionally conserved features with mammals, including granulocytic lineages (e.g., neutrophils), monocyte/macrophage-like cells, Toll-like receptors, pro-inflammatory cytokines, and antimicrobial peptides1. The intestinal tract in zebrafish is fully developed at 6 days post fertilization (dpf) and shows morphological and functional conservation with the mammalian gastrointestinal tract, such as conserved transcriptional regulation in intestinal epithelial cells2. This makes zebrafish an excellent model for intestinal microbial colonization and pathogenesis. A wide range of enteric microbes has been studied in the zebrafish model, including enterohemorrhagic Escherichia coli3, Vibrio cholerae4,5, Salmonella enterica6, the zebrafish microbiota7,8, and the role of probiotics in intestinal immunity9. A distinct advantage of the zebrafish model is that it is colonized by many microbes without disrupting the endogenous microbiota, which allows the investigation of microbial behavior in the context of mixed microbial populations3,6. Currently, most zebrafish models of gastrointestinal colonization and disease rely on the administration of microbes by bath immersion, where zebrafish are incubated in a bacterial suspension for a specific amount of time10. However, this makes it difficult to determine the exact dose of bacteria administered, and leads to limited colonization with some microbes, particularly with non-pathogenic bacteria. Alternatively, a bacterial suspension is administered to fish via oral gavage11, but this is technically challenging and limited to older larvae and adult fish.
This protocol describes the use of the unicellular protozoan Paramecium caudatum as a vehicle for food-borne delivery of microbes to the gastrointestinal tract of zebrafish larvae. Paramecia are easy and cheap to maintain and are capable of feeding on a wide variety of microbes, including algae, fungi, and bacteria, which they internalize through a ciliated oral groove12,13,14. Once internalized, bacteria are held in vacuoles, which eventually acidify and contents are degraded over a time frame of several hours15. Larval zebrafish capture paramecia as natural prey soon after hatching, around 3–4 dpf depending on temperature16, and take them up with high efficiency. The process of prey capture takes on average 1.2 s from detection to capture17, and captured paramecia are quickly digested in the zebrafish foregut, such that internalized viable bacteria are released into the intestinal tract3. As a result, paramecia can be used as a quick and easy method to deliver a high and consistent dose of bacteria into the gastrointestinal tract of zebrafish. The delivered bacteria can either be transformed to express a fluorescent protein, such as mCherry as described here, or, in the case of genetically intractable bacteria, they can be pre-stained with a fluorescent dye to allow visualization within the gastrointestinal tract.
This protocol describes the food-borne delivery of enteropathogenic E. coli (enterohemorrhagic E. coli [EHEC] and adherent invasive E. coli [AIEC]), and Salmonella enterica ssp. Typhimurium. Both pathogenic E. coli and S. typhimurium are transmitted via the fecal-oral route18,19, and can be acquired via contaminated food, such as meat, vegetables, and dairy. Using P. caudatum as a vehicle, E. coli and S. typhimurium successfully colonize the zebrafish larvae within 30–60 min of co-incubation with the paramecium vehicle. The achieved bacterial burden is robust enough to visualize colonization and determine burden by plating tissue homogenates.
Protocol
Zebrafish care, breeding, and experiments described here are in accordance with the Guide for the Care and Use of Laboratory Animals, and have been approved by the Institutional Animal Welfare Committee of the University of Texas Health Science Center, protocol number AWC-16-0127.
1. Growth and Maintenance of Paramecia
- Obtain live paramecia cultures from sources such as the Zebrafish International Resource Center (ZIRC).
- From a live growing culture, add 1 mL of the paramecia culture, 1 mL of an E. coli MG1655 culture (optical density OD600 1.0–2.0) resuspended in 1x E3 (0.29 g/L NaCl, 13 mg/L KCl, 44 mg/L CaCl2, 81 mg/L MgSO4, 0.48 g/L HEPES, pH 7.0, sterile), and 8 mL of 1x E3 into a 10 mL tissue culture flask. Swirl lightly and then store at 22 °C.
- To maintain a culture, every two weeks, passage 1 mL of a paramecia culture into a 10 mL tissue culture flask with 9 mL of fresh 1x E3 medium containing 108 CFU/mL of E. coli MG1655 resuspended in 1x E3.
2. Determination of Bacterial Dose Administered to Zebrafish
- Determine bacterial half-life within Paramecium
NOTE: The half-life of bacteria within paramecia is determined by plating viable E. coli recovered from paramecium, as described below.- Following a 2 h incubation of bacteria (OD600 of 1.0) with paramecia at 22 °C, combine the paramecia and bacterial co-culture into a 50 mL conical tube, wash the paramecia with 1x E3, and count the number of paramecia per milliliter (as described below in steps 3.2.7 and 3.2.8).
- After counting the number of paramecia, remove 50 µL of the paramecia and bacterial co-culture every hour (for 6 h post-exposure) and add each sample into a fresh 1.5 mL tube.
- Place 950 µL of 1% nonionic surfactant (see Table of Materials) in phosphate buffered saline (PBS) solution into each of the 1.5 mL tubes and vortex for 1 min to lyse the paramecia. Perform 1:10 dilutions of each sample using sterile PBS as a diluent (i.e., 100 µL in 900 µL).
- Plate 100 µL of each dilution onto selective plates (LB tet medium: 1 g/L tryptone, 0.5 g/L yeast extract, 1 g/L NaCl, 30 µg/mL tetracycline) and incubate at 37 °C for 16 h. The next day, count and record the number of bacterial colonies on the plate (colony forming units, CFUs) by counting only the isolated and distinct individual colonies. Identify a plate with a dilution that gives 30–300 CFU.
- Determine the dilution factor used. For example, if 1 µL of the bacterial culture is mixed with 99 mL of sterile PBS, this is a 1:100 (0.01) dilution. This number will be the number of CFU in the dilution. Perform the calculation below, for every time point, to determine the CFU in the original sample:
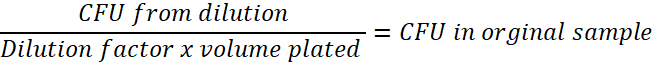
- Calculate and graph the number of viable E. coli/paramecium corresponding to the hours post exposure using the paramecia concentration calculated in step 3.3.1 and determine the half-life within the paramecia (Figure 1).
- Using the CFU number calculated for each time point, calculate and graph the number of viable E. coli per paramecium on the y-axis versus the hours post incubation on the x-axis, using the paramecia concentration calculated in step 3.3.1. Determine the half-life within the paramecia.
- Determine bacterial dosing from bacterial half-life and preying rate.
NOTE: To determine the bacterial dosage, the half-life of the bacteria inside the paramecia and the preying rate of zebrafish on paramecia (see step 3.4) have to be taken into account.- Use the following formula to determine bacterial decay within paramecia:
(1)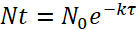
Where N0 is the initial quantity of bacteria per paramecia after the 2 h incubation, Nt is the remaining quantity after time t, τ is the time after which the number of viable bacteria has halved, and k is the decay constant. - From the degradation experiment, determine the decay constant k using the bacterial half-life (i.e., the time after which the amount of viable bacteria/paramecium has halved).
(2)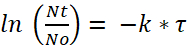
For the determination of half-life, the term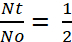 and
and
(3)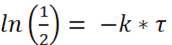 or
or
(4)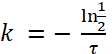
NOTE: Based on Figure 1, the half-life of E. coli in paramecia is approximately 2.3 h. Thus, using formula (4), the decay rate, k, for E. coli is:
(5)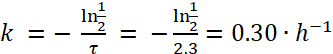
- Determine the decay rate (k) from the half-life experiment to find the dose of viable bacteria (Nt) taken up by the zebrafish larvae after preying time (t), where (P) is the preying rate or the number of paramecia eaten by one fish per hour:
(6)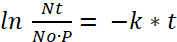 or
or
(7)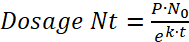
NOTE: Per Figure 1, the initial quantity of bacteria per paramecia (N0) after a 2 h incubation (t) is 790 CFU. Per Movie 1 and Figure 2, the preying rate (P) is 1539. - Using these values to substitute into equation (7), calculate the bacterial dosage consumed by a zebrafish following a 2 h incubation as:
(8)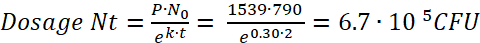
- Use the following formula to determine bacterial decay within paramecia:
3. Food-borne Infection of Zebrafish
- Incubate bacteria with Paramecia.
- Prepare a co-culture of paramecia and E. coli MG1655 the night prior to infection. Combine 8 mL of E3 media, 1 mL of an ongoing paramecia culture, and 1 mL of an E. coli MG1655 culture (OD600 = 1.0) resuspended in 1x E3 in T25 tissue culture flasks. Incubate the flasks at room temperature (RT) overnight. For each treatment condition, prepare two flasks of paramecia.
- Inoculate bacterial growth media (LB: 1 g/L tryptone, 0.5 g/L yeast extract, 1 g/L NaCl) with infectious strain of bacteria, by picking an individual bacterial colony from a plate using a sterile inoculation loop. Incubate the liquid culture at 37 °C and leave shaking at 110 rotations per minute (rpm) overnight.
NOTE: Personal protective equipment (a laboratory coat and gloves) should be worn and biosafety level 2 facilities should be used when handling infectious agents. - On the next day, measure the OD600 of the overnight culture. Calculate the volume of culture required to achieve an OD600 of 1 when resuspended in 11 mL of media.
- Harvest the volume of bacteria from step 3.1.2 via centrifugation at 6,000 x g for 5 min, one volume for each flask of paramecia. Discard the supernatant and resuspend the bacterial pellet in 1 mL of E3 media.
- Optionally, pre-stain the bacteria with a fluorescent dye.
- Add 1 µL of FM 4-64FX bacterial stain (5 mg/mL stock solution). Cover tube with foil to protect from photobleaching and incubate rotating end-over-end at RT for 15 min.
- Remove excess dye by washing with 1x E3: Pellet bacteria via centrifugation at 6,000 x g for 1.5 min, then resuspend pellet in 1 mL of E3 media. Repeat wash step two times.
- Harvest stained bacteria via centrifugation at 6,000 x g for 5 min. Discard the supernatant and resuspend the bacterial pellet in 1 mL of E3 media.
- Add 1 mL of the bacterial suspensions to each of the two flasks of fresh paramecia. Incubate at RT for 2 h.
NOTE: If working with stained bacteria, incubate in the dark at RT for 2 h.
- Wash bacteria/paramecia co-culture.
- Combine the contents of both flasks of paramecia/bacteria co-culture into a 50 mL conical tube. Centrifuge samples at 300 x g at 15 °C for 10 min. Make sure that the centrifuge is pre-cooled prior to this step.
- Remove approximately 10 mL of the E3 supernatant using a serological pipette and add approximately 10 mL of fresh 1x E3 to the conical tube.
NOTE: During all wash steps, it is essential to be very quick when removing the supernatant, as the paramecia will begin to swim out of the pellet. Spin and remove supernatant from one tube at a time to ensure quick enough handling at this step, and avoid loss of paramecia in the supernatant. - Spin samples via centrifugation at 300 x g at 15 °C for 5 min. Remove approximately 10 mL of the E3 supernatant using a serological pipette, and add approximately 10 mL of fresh 1x E3 to the conical tube. Repeat this step twice.
- Centrifuge samples at 300 x g at 15 °C for 5 min. Remove approximately 10 mL of the E3 supernatant, taking care not to disrupt the pellet.
- Resuspend pellet into the remaining 10 mL of E3 media and transfer 500 µL of the suspension into a new 1.5 mL microcentrifuge tube. Pellet the 500 µL of paramecia by centrifuging at 300 x g for 5 min to count the number of paramecia.
- Remove 400 µL of the E3 supernatant from the 500 µL sample. Add 20 µL of 36.5% formaldehyde solution to the remaining 100 µL of paramecia and gently resuspend, and incubate for 5 min at 22 °C.
NOTE: This step kills the paramecia to allow for counting. - Measure actual total volume using the pipette and record. Dilute the paramecia suspension 1:1 v/v with 0.4% trypan blue solution.
- Use a cell counter or hemocytometer to count the number of dead paramecia/mL.
NOTE: Because of the prior fixation step, most paramecia will be dead at this point, but this number reflect the number of live paramecia for the co-incubation experiment. The authors have not found significant paramecia death due to bacterial co-incubation, so this can be disregarded as a factor here.
- Co-incubate Paramecia and zebrafish larvae.
- Calculate the concentration of paramecia:
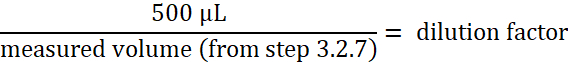

NOTE: This calculation gives the concentration of paramecia in the 50 mL conical tube from step 3.2.5. - Calculate the volume of washed paramecia required for a concentration of 2 x 105 paramecia/mL in a final volume of 3 mL of E3.
NOTE: The concentration of paramecia can be adjusted based on the desired bacterial dosage, which is subject to optimization. - Anesthetize zebrafish by adding tricaine in 100 mM Tris pH 8.0 to a final concentration of 100 mg/L. Transfer 10 zebrafish into each well of a 6-well plate into a total volume of 3 mL of fresh E3 containing the appropriate concentration of paramecia (calculated in step 3.3.2). Ensure to transfer larvae in a minimal amount of liquid, to ensure they recover from anesthesia in the recipient well.
- Incubate at 30 °C for 2 h in a diurnal incubator under day-light conditions, to ensure optimal lightning conditions for preying.
- Wash zebrafish at least 5 times by transferring fish into a new well containing 3 mL of fresh E3 containing 100 mg/L tricaine each time.
NOTE: Do not attempt to omit the tricaine during the washing step. Transferring mobile larvae without anesthesia increases the risk of damage and distress to the animal. - Optionally, prepare zebrafish for imaging by embedding zebrafish in 3 mL of 1% low-melt agarose in a black-walled 6-well plate: Low-melt agarose is made up in 1xE3 and heated in a microwave. Once molten, add tricaine to a final concentration of 160 mg/mL. Position fish under a stereomicroscope, using a clipped gel loading tip, making sure that the head is on the left and the tail is on the right (Figure 3). Wait for 5 minutes for the agarose to set, then overlay the embedded fish with 1x E3 containing 160 mg/mL tricaine for imaging.
- Calculate the concentration of paramecia:
- Determine the preying rate.
NOTE: No separate experiment needs to be set up to determine the preying rate. Rather, this can be done during step 3.3.4, as described below.- During step 3.3.4, view preying zebrafish on a stereomicroscope and capture video footage of the prey capture.
- Score the video footage. Prey capture is characterized by striking of zebrafish toward the prey. Count each strike as one prey capture event, although this is only an approximation (see Discussion).
- Calculate the average number of prey capture events per hour from multiple video clips, each representing a different zebrafish larva (Figure 2).
Results
Paramecium caudatum readily internalizes a wide range of bacteria into storage vacuoles. The intracellular bacterial density depends on the densities of bacteria and paramecia in the co-culture, as well as the bacterial species used. Over time, the vacuoles acidify and bacterial degradation ensues. The rate of degradation has to be individually determined for all strains used. For pathogenic E. coli, the initial bacterial density is 790 bacteria/paramecium, and bacteria are degraded with a half-life of approximately 2.3 h (Figure 1).
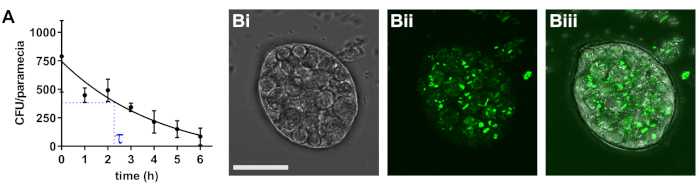
Figure 1: Determination of bacterial half-life in paramecia. (A) Following 2 h of co-incubation with infectious E. coli, P. caudatum was washed and transferred to medium without bacteria. At the indicated time points, numbers of viable E. coli cells were determined by dilution plating on selective agar. Results are means ± standard error of the mean (SEM; n = 3). (B) Typical image of paramecium carrying internalized bacteria, with bright field (Bi), fluorescent bacteria (Bii), and merged channels (Biii). Scale bar = 20 μm. Please click here to view a larger version of this figure.
Further, the zebrafish preying rate, that is, the rate at which zebrafish internalize bacteria-loaded paramecia upon co-incubation, was studied. Larval zebrafish start to hunt and capture live prey from 5 dpf20, although it was found that, when raised at 30 °C, larval development is accelerated and animals display preying behavior from 4 dpf. Preying is accompanied by a characteristic striking behavior20 (Figure 2A), and the determination of the preying rate is based on the assumption that each strike leads to internalization of one paramecium, although this can only be regarded an approximation (see Discussion). Based on the herein described observations of preying zebrafish larvae, the rate of paramecia uptake is approximately 1,539 per h (Figure 2B).
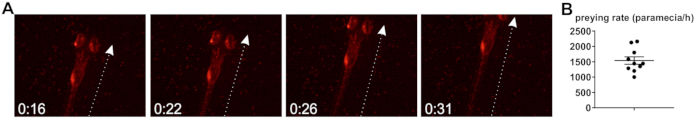
Figure 2: Determination of zebrafish preying rate. (A) Still images from a preying video, showing a zebrafish larvae (5 dpf) preying on paramecia carrying fluorescent bacteria. Time in [seconds]. Arrow indicates the main axis of movement during striking. (B) Quantification of preying rate (paramecia intake per hour), based on n = 10 videos taken over the full 2 h exposure time. Please click here to view a larger version of this figure.
Following internalization of paramecia, the zebrafish efficiently degrades the prey in the foregut, releasing infectious bacteria into the digestive system. As described herein, paramecia degradation proceeds quickly, and free bacteria can be detected in the intestinal tract within 30 minutes of preying. Free bacteria then move from the foregut to the mid- and posterior intestine, where they are detected approximately 1–2 h after the beginning of preying (Figure 3). Bacterial persistence in the intestine depends on species and dose but ranges from several h to several days in the case of E. coli and S. enterica. S. enterica localizes primarily in the intestinal mucosae, with some epithelial invasion (Figure 3D), leading to infiltration of neutrophils into the epithelium (Figure 3C).
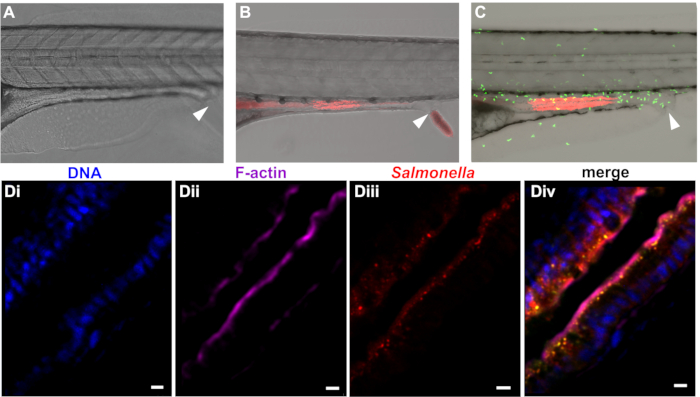
Figure 3: Colonization of zebrafish with bacteria. Zebrafish at 5 dpf were left uninfected (A) or colonized with mCherry expressing (B) E. coli or (C) S. enterica. Infection experiments may be performed in wild type (A and B) fish or transgenic lines (e.g., the line Tg(MPO::EGFP)i114 expressing green fluorescent neutrophils shown in (C). The rectal opening is marked by an arrow. (D) Higher magnification of intestinal section from whole-mount embedded larvae infected with Salmonella enterica infection. (Di) Blue = Hoechst marking nuclei, (Dii) Purple = phalloidin marking F-actin, (Diii) Red = Salmonella, (Div) merge. Scale bar = 5 μm. Please click here to view a larger version of this figure.
Movie 1: Video footage of the prey capture. Please click here to download this video.
Discussion
The basic protocol described here has been optimized for pathogenic E. coli, and has been successfully adapted for other bacterial species, including Salmonella enterica and Vibrio cholerae. For some species that do not colonize the zebrafish gut following bath immersion, including some Salmonella enterica strains and some anaerobes, food-borne infection as described here can be used to successfully establish colonization. Compared to microgavage, which is also used to establish high bacterial burdens in the larval intestinal tract, food-borne infection is technically less challenging and requires less specialized equipment. However, critical parameters should be optimized for the bacterial species and strains to be used. Such factors include bacterial and paramecium density for the bacteria-paramecium co-culture step: If bacterial numbers within paramecia are low, this could be improved by increasing the bacterial density in the co-culture step. Some bacterial species may cause damage to the paramecium host, and this should be assessed by microscopy.
Another important factor in this protocol is prey capture by zebrafish. The preying rate as described here is based on the assumption that every prey capture strike results in the ingestion of one paramecium. High densities of paramecia per fish are used in the protocol to ensure high preying rates. However, prey capture is dependent on the density of paramecia in the system, and in very dilute paramecium cultures, preying rates may be as low as 13–15 paramecia per hour21,22. A limitation is that prey capture rates are also strongly dependent on lighting conditions and in the dark, capture rates are 80% lower than in light conditions21 and this should be taken into account when setting up experiments. If exposure times to prey have to be expanded to optimize colonization, consideration has to be given to secondary exposure to bacteria through feces. Under the conditions described above – 2 h of prey exposure – this exposure is negligible, since gut passage time of bacteria is more than 1 h and the concentration of bacteria in the vehicle is much higher than in feces. However, if prey exposure time is significantly increased, this may become a significant factor.
Appropriate controls should be included in this protocol, including colonization of zebrafish following feeding with paramecia containing non-pathogenic E. coli MG1655. If multiple bacterial strains are compared for their ability to colonize the zebrafish host, it is important to test whether their half-life within paramecia is comparable. Bacterial mutations, including those compromising cell wall integrity or acid sensing, may compromise bacterial stability within paramecia. In such cases, zebrafish feeding has to be adjusted to account for the differences in dosage.
The protocol described here can be used to investigate bacterial colonization and its consequences, including by imaging bacterial colonization of zebrafish as described above, as well as by determining CFU per zebrafish from tissue homogenate3, or investigating infection-associated morbidity and mortality. Ideally, for bacterial visualization, bacterial strains expressing fluorescent proteins such as mCherry or red fluorescent protein (RFP) should be used. This will allow the visualization of growing bacterial populations. If the bacterial strain is not genetically tractable or the use of fluorescent protein expression is precluded for other reasons, bacteria may be stained with a fluorescent dye, such as FM 4-64FX, prior to co-culture with paramecia. When using the protocol described here, co-culture with paramecia does not decrease the brightness of the dye and stained bacteria are clearly visible in the zebrafish intestine. However, the dye will be diluted over time should significant bacterial proliferation occur within the zebrafish host. In either case, red-fluorescent bacteria are preferable over green-fluorescent bacteria, since tissue autofluorescence can be higher in the green than in the red channel.
It has been found that this protocol can be adapted for aerobic and microaerophilic bacterial species. It may be possible to adapt it for the feeding of spores and fungal species, although this remains to be tested experimentally.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We would like to thank members of the Krachler group for critical reading and comments on the manuscript. This work was funded by a UT Systems STAR award, the BBSRC, and the NIH (R01AI132354).
Materials
| Name | Company | Catalog Number | Comments |
| Paramecium caudatum, live | Carolina | 131554 | no not store growing cultures below room temperature |
| 0.4% Trypan Blue Solution | Sigma | T8154-20ML | liquid, sterile-filtered, suitable for cell culture; prepared in 0.81% sodium chloride and 0.06% potassium phosphate, dibasic |
| Dimethyl sulfoxide (DMSO) | Sigma | 276855-100ML | store in a solvent safety cabinet |
| Escherichia coli, MG1655 | ATCC | ATCC 700926 | can be replaced by any other non-pathogenic E. coli strain |
| FM 4-64FX stain | Thermo Fisher | F34653 | aliquot and store frozen |
| Formaldehyde | Sigma | F8775-4X25ML | |
| LB Broth | Sigma | L3397-1KG | |
| Phosphate buffered saline tablets | Thermo Fisher | 18912014 | |
| Tetracycline | Sigma | 87128-25G | toxic, irritant |
| Tricaine (Ethyl 3-aminobenzoate methanesulfonate) | Sigma | E10521-10G | |
| Triton X-100 | Sigma | X100-100ML | |
| Trypan Blue Solution, 0.4% | Sigma | 93595-50ML | |
| UltraPure Low Melting Point Agarose | Thermo Fisher | 16520050 | |
| hemocytometer or cell counter | any | ||
| stereomicroscope | any | ||
| table-top centrifuge | |||
| microwave | |||
| rotator wheel | |||
| heated shaking incubator | |||
| aquatics facilities | |||
| breeding tanks |
References
- Broz, P., Ohlson, M. B., Monack, D. M. Innate immune response to Salmonella Typhimurium, a model enteric pathogen. Gut Microbes. 3 (2), 62-70 (2012).
- Lickwar, C. R., et al. Genomic dissection of conserved transcriptional regulation in intestinal epithelial cells. PLoS Biology. 15 (8), e2002054(2017).
- Stones, D. H., et al. Zebrafish (Danio rerio) as a Vertebrate Model Host To Study Colonization, Pathogenesis, and Transmission of Foodborne Escherichia coli O157. mSphere. 2 (5), (2017).
- Mitchell, K. C., Breen, P., Britton, S., Neely, M. N., Withey, J. H. Quantifying Vibrio cholerae enterotoxicity in a zebrafish infection model. Applied and Environmental Microbiology. , 00783(2017).
- Logan, S. L., et al. The Vibrio cholerae type VI secretion system can modulate host intestinal mechanics to displace gut bacterial symbionts. Proceedings of the National Academy of Sciences of the United States of America. 115 (16), E3779-E3787 (2018).
- Howlader, D. R., et al. Zebrafish as a novel model for non-typhoidal Salmonella pathogenesis, transmission and vaccine efficacy. Vaccine. 34 (42), 5099-5106 (2016).
- Troll, J. V., et al. Microbiota promote secretory cell determination in the intestinal epithelium by modulating host Notch signaling. Development. 145 (4), (2018).
- Wiles, T. J., et al. Host Gut Motility Promotes Competitive Exclusion within a Model Intestinal Microbiota. PLoS Biology. 14 (7), e1002517(2016).
- Rendueles, O., et al. A new zebrafish model of oro-intestinal pathogen colonization reveals a key role for adhesion in protection by probiotic bacteria. PLoS Pathogens. 8 (7), e1002815(2012).
- Varas, M., et al. Salmonella Typhimurium induces cloacitis-like symptomsin zebrafish larvae. Microbial Pathogenesis. 107, 317-320 (2017).
- Runft, D. L., et al. Zebrafish as a natural host model for Vibrio cholerae colonization and transmission. Applied and Environmental Microbiology. 80 (5), 1710-1717 (2014).
- Meier, R., Wiessner, W. Infection of algae-free Paramecium bursaria with symbiotic Chlorella sp. Isolated from green paramecia: I. Effect of the incubation period. European Journal of Protistology. 24 (1), 69-74 (1988).
- Miura, T., Moriya, H., Iwai, S. Assessing phagotrophy in the mixotrophic ciliate Paramecium bursaria using GFP-expressing yeast cells. FEMS Microbiology Letters. 364 (12), (2017).
- Watanabe, K., et al. Ciliate Paramecium is a natural reservoir of Legionella pneumophila. Scientific Reports. 6, 24322(2016).
- Bragg, A. N., Hulpieu, H. A Method of Demonstrating Acidity of Food Vacuoles in Paramecium. Science. 61 (1580), 392(1925).
- Borla, M. A., Palecek, B., Budick, S., O'Malley, D. M. Prey capture by larval zebrafish: evidence for fine axial motor control. Brain, Behavior and Evolution. 60 (4), 207-229 (2002).
- Patterson, B. W., Abraham, A. O., MacIver, M. A., McLean, D. L. Visually guided gradation of prey capture movements in larval zebrafish. Journal of Experimental Biology. 216 (Pt 16), 3071-3083 (2013).
- Megraud, F. Transmission of Helicobacter pylori: faecal-oral versus oral-oral route. Alimentary Pharmacology & Therapeutics. 9 Suppl 2, 85-91 (1995).
- Spears, K. J., Roe, A. J., Gally, D. L. A comparison of enteropathogenic and enterohaemorrhagic Escherichia coli pathogenesis. FEMS Microbiology Letters. 255 (2), 187-202 (2006).
- Bianco, I. H., Kampff, A. R., Engert, F. Prey capture behavior evoked by simple visual stimuli in larval zebrafish. Frontiers in Systems Neuroscience. 5, 101(2011).
- Gahtan, E., Tanger, P., Baier, H. Visual prey capture in larval zebrafish is controlled by identified reticulospinal neurons downstream of the tectum. Journal of Neuroscience. 25 (40), 9294-9303 (2005).
- Westphal, R. E., O'Malley, D. M. Fusion of locomotor maneuvers, and improving sensory capabilities, give rise to the flexible homing strikes of juvenile zebrafish. Frontiers in Neural Circuits. 7, 108(2013).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved