Method Article
Covalent Attachment of Single Molecules for AFM-based Force Spectroscopy
In This Article
Summary
Covalent attachment of probe molecules to atomic force microscopy (AFM) cantilever tips is an essential technique for the investigation of their physical properties. This allows us to determine the stretching force, desorption force and length of polymers via AFM-based single molecule force spectroscopy with high reproducibility.
Abstract
Atomic force microscopy (AFM)-based single molecule force spectroscopy is an ideal tool for investigating the interactions between a single polymer and surfaces. For a true single molecule experiment, covalent attachment of the probe molecule is essential because only then can hundreds of force-extension traces with one and the same single molecule be obtained. Many traces are in turn necessary to prove that a single molecule alone is probed. Additionally, passivation is crucial for preventing unwanted interactions between the single probe molecule and the AFM cantilever tip as well as between the AFM cantilever tip and the underlying surface. The functionalization protocol presented here is reliable and can easily be applied to a variety of polymers. Characteristic single molecule events (i.e., stretches and plateaus) are detected in the force-extension traces. From these events, physical parameters such as stretching force, desorption force and desorption length can be obtained. This is particularly important for the precise investigation of stimuli-responsive systems at the single molecule level. As exemplary systems poly(ethylene glycol) (PEG), poly(N-isopropylacrylamide) (PNiPAM) and polystyrene (PS) are stretched and desorbed from SiOx (for PEG and PNiPAM) and from hydrophobic self-assembled monolayer surfaces (for PS) in aqueous environment.
Introduction
Since its invention in the 1980s1, the atomic force microscope (AFM) has become one of the most important imaging techniques in natural science featuring sub-nanometer spatial resolution, sub-piconewton force resolution and the possibility of measuring in various solvent and temperature conditions2,3,4,5,6,7.
Apart from imaging8,9, AFM is used to perform single molecule force spectroscopy (SMFS) giving insight into adhesive interactions between a single polymer and surfaces, physical properties of single polymers and unfolding mechanisms of proteins7,10,11,12,13,14,15,16. In a regular SMFS experiment, the functionalized cantilever tip is brought into contact with a surface so that the polymer at the AFM cantilever tip physisorbs to this surface. By retracting the AFM cantilever tip from the surface, a change in the deflection of the AFM cantilever is converted into a force leading to a force-extension curve4. Physical parameters such as stretching force, desorption force and desorption length can be determined as dependent on different parameters such as pulling velocity, dwell time on surface, indentation depth into the surface, temperature, solvent17,18 and different surfaces like solid substrates, polymer films or supported lipid bilayers19,20,21,22. Furthermore, a polymer can be probed in different spatial directions, thus investigating the frictional properties of the polymer23,24,25,26.
A covalent attachment of the investigated polymer to an AFM cantilever tip is essential for such studies. Thus, a high yield of single molecule events with one and the same polymer bound to an AFM cantilever tip prevents any bias of the results due to calibration of the spring constant of the AFM cantilever27,28, varying attachment points29 or varying polymers (with different contour lengths) such as in the case of nanofishing experiments30,31,32. Also, interactions with other polymers as well as averaging effects can be widely prevented18,28. For the covalent attachment of a polymer to the AFM cantilever tip, different types of chemical modifications can be applied, many of which are summarized in the book by Hermanson33. Amine and thiol-based linking reactions as well as click chemistry represent the most commonly used methods in AFM cantilever tip functionalization34,35,36,37,38,39,40,41,42. Becke et al.40 show how to use 1-ethyl-3-(3-dimethylaminopropyl)carbodiimid (EDC)/NHS chemistry to attach a protein to an AFM cantilever tip. However, the said functional groups tend to crosslink, thus leading to a loss of functionality43,44. Also, carbodiimides show a tendency to fast hydrolysis in solution43. Maleimide and thiol groups are generally more stable and do not show crosslinking reactions. The presented protocol is an optimization of the previously published protocols given in references35,39.
Here, a reliable functionalization protocol is presented that can be easily adjusted to a large number of different polymers, irrespective of properties such as contour length or hydrophobicity. Three different polymers were chosen by way of example: hydrophilic polyethylene glycol (PEG) and poly(N-isopropylacrylamide) (PNiPAM) as well as high molar mass hydrophobic polystyrene (PS). In order to provide for a covalent binding capability with an appropriate linker molecule, the three polymers were selected for featuring a telechelic thiol moiety as functional end group. The linker molecule itself is typically a short PEG polymer with two active sites, a silane group at one end and a maleimide group at the other end. The former enables a covalent attachment to the AFM cantilever tip and the latter a binding reaction with the thiol group of the functionalized high molar mass polymer. Furthermore, inactive PEG linker molecules serve as a passivation layer to prevent unwanted interactions between the probe polymer and the AFM cantilever tip as well as between the AFM cantilever tip and the underlying surface.
Protocol
NOTE: See Figure 1 for a schematic overview.
1. Reagent setup
NOTE: The polymers used for this protocol are: maleimide-polyethylene glycol-triethoxysilane (silane-PEG-mal, 5 kDa), thiol-polyethylene glycol-thiol (HS-PEG-SH, 35 kDa), thiol terminated poly(N-isopropylacrylamide) (PNiPAM-SH, 637 kDa) and thiol terminated polystyrene (PS-SH, 1.3 mDa).
- Prepare the well-defined and high molar mass PNiPAM-SH via atom transfer radical polymerization, followed by conversion and reduction of the functional end group for the introduction of a thiol moiety, as described in the literature18. Please see Figure 1 for the detailed structures.
- For storing of the chemicals, prepare smaller aliquots within a dry glovebox system with nitrogen atmosphere to avoid exposure to atmospheric oxygen and moisture. PEG and PNiPAM are hygroscopic45,46 and the functional end groups of PEG, PNiPAM and PS are known to become easily oxidized when stored at ambient conditions33,47,48. All chemicals have to be stored at -20 °C.
- Use analytical grade solvents or higher. Moreover, use ultrapure water to rinse AFM cantilevers chips and glassware because single molecule experiments are very sensitive to all contamination.
2. Equipment Setup
NOTE: Use tweezers and beakers made of stainless steel or glass. Use inverted tweezers for a safe grip (e.g., model R3 SA having a low spring constant).
- Prepare RCA (ultrapure water, hydrogen peroxide and ammonia (5:1:1)) solution to clean glassware and tweezers.
- Put the vessels in a beaker and fill it with RCA until glassware or tweezers are fully covered.
- Heat the beaker from step 2.2 for 1 h at 80 °C.
- Rinse the vessels subsequently with ultrapure water until no pungent smell is ascertainable anymore (at least three times).
- Dry glassware and tweezers in an oven (120 °C).
3. Tip functionalization
NOTE: All steps should be carried out in a fume hood to avoid inhalation of organic vapors. Additionally, gloves, lab coat and eye protection are required. Use nitrile or latex gloves for every step to avoid contamination. Wear solvent resistant gloves when using toluene. All steps, unless specified otherwise, are done at RT. Use fresh equipment and gloves for every step to avoid possible cross-contamination.
- Perform surface activation by applying oxygen plasma to the AFM cantilever chip MLCT-Bio-DC.
NOTE: The efficiency of the plasma treatment for further functionalization steps scales with the content of oxygen in the plasma chamber.- Use freshly cleaned tweezers to place AFM cantilever chips in a plasma chamber (40 kHz, 600 W).
- Use custom-modified activation program: evacuation (0.1 mbar) – flooding with oxygen to a pressure of: 0.2 mbar (4 min) – plasma process (power: 40%, duration: 2 min, process pressure: 0.2 mbar).
- Ventilate chamber and carry on with step 3.2.2 immediately in order to prevent any adsorption of contaminants to AFM cantilever chips from air.
- Silanization and PEGylation
NOTE: Timing is a critical parameter between the steps. Prepare solutions as fresh as possible during the waiting times. Maleimide groups are subject to hydrolysis in aqueous media and thiols easily become oxidized to disulfides in solution33,47 impeding AFM tip functionalization reactions.- Prepare a silane-PEG-mal solution in toluene (1.25 mg/mL) in solvent resistant plastic or glass tubes and pour 6 mL of the solution in flat Petri dishes, 3 mL each.
NOTE: If binding of multiple probe polymers is observed in the SMFS experiment, mixing silane-PEG-mal with non-functional silane-PEG can reduce the number of anchoring points. For the adjustment of the passivation layer PEG with different masses (i.e., contour lengths) can be used27. - Incubate the AFM cantilever chips immediately after step 3.1.3 in the silane-PEG-mal solution (up to 10 chips per Petri dish) for 3 h at 60 °C35.
- Take Petri dishes out of the oven and let the solution cool down for at least 10 min.
- Rinse each AFM cantilever chip carefully. Reduce the impact of capillary forces on the AFM cantilever when passing the air-solvent interface, for example by tilting these chips slightly when immersing into solution.
- For PEG and PS polymers, rinse three times with toluene.
- For PNiPAM polymer, rinse once with toluene and twice with ethanol.
- Choose at least two AFM cantilever chips as control AFM cantilever chips, skipping step 3.3 and rinse them as follows to increase the polarity of the solvent:
- For PEG and PS polymers, rinse twice with ethanol and once with ultrapure water.
- For PNiPAM polymer, rinse twice with ultrapure water.
NOTE: Control AFM cantilever chips have gone through all functionalization steps except the polymer attachment (step 3.3). They serve to prove the cleanliness of the functionalization process, the AFM cantilever chip holder system, the surfaces and the solvents used for the SMFS experiment.
- Prepare a silane-PEG-mal solution in toluene (1.25 mg/mL) in solvent resistant plastic or glass tubes and pour 6 mL of the solution in flat Petri dishes, 3 mL each.
- Covalent polymer attachment
NOTE: Even though the AFM cantilever tip is expected to be completely covered with maleimide groups, there are just a few binding sites for the single probe polymer, because maleimide undergoes hydrolysis in water leading to inactive PEGs47. These inactive PEGs act as a passivation layer, as described above.- Incubate AFM cantilever chips directly after step 3.2.5 in one of the following polymer solutions in 3 mL Petri dishes. If the respective polymer is not dissolved properly, use a 40 °C water bath and stir the solution well.
NOTE: As the use of thiol terminated polymers might lead to the formation of disulfide bonds hampering the reaction with the maleimide groups of silane-PEG-mal, a reducing agent is recommended, in particular if step 3.3 is applied in aqueous buffers for water soluble polymers33.- For PEG and PS polymers, use a concentration of 1.25 mg/mL in toluene for 1 h at 60 °C.
- For PNiPAM polymers, use a concentration of 1.25 mg/mL in ethanol for 3 h at RT.
NOTE: If binding of multiple probe polymers is observed in the SMFS experiment, the concentration of the polymer should be reduced.
- Carefully rinse each AFM cantilever chip.
- For PEG and PS polymers, rinse twice with toluene, twice with ethanol and once with ultrapure water after a 10 min cool down.
- For PNiPAM polymers, rinse twice with ethanol and twice with ultrapure water.
- Store each AFM cantilever chip separately in a small (1 mL) Petri dish filled with ultrapure water at 4 °C until use in an experiment.
- Incubate AFM cantilever chips directly after step 3.2.5 in one of the following polymer solutions in 3 mL Petri dishes. If the respective polymer is not dissolved properly, use a 40 °C water bath and stir the solution well.
4. Surface preparation
- Silicon oxide wafer
NOTE: This surface was used for SMFS with PEG and PNiPAM.- Cut a silicon oxide wafer in small pieces using a diamond knife.
- Put the silicon oxide pieces separately in microcentrifuge tubes and fill these tubes with ethanol.
- Sonicate the silicon oxide pieces for 10 min.
- Rinse the silicon oxide pieces with ethanol twice and dry them under a nitrogen flow carefully. Use the silicon oxide pieces immediately.
- Self-assembled monolayer of hydrophobic alkane thiol on gold (SAM)
NOTE: This surface was used for SMFS with PS. See literature39,49 for more information about SAMs.- Use a gold-coated silicon wafer (A [100], 5 nm titanium, 100 nm gold) to perform steps 4.1.1 – 4.1.4.
- Incubate the surface pieces in a 1-dodecanthiol solution (2 mM) for 18 h.
- Rinse the freshly prepared SAMs in ethanol twice.
- Dry SAMs with nitrogen flow for direct use or store them in ethanol for up to 4 days for later use.
5. Data acquisition
NOTE: All measurements shown here were performed in ultrapure water with a Cypher ES AFM using a heating and cooling sample stage for temperature variation. Generally, all AFMs providing the capability to measure in liquids can be used.
- Insert the functionalized AFM cantilever chip into the AFM.
- Glue the prepared surface into a sample holder that is suitable for measuring in liquids (e.g., High Resolution Replicating Compound 101RF or an UV curable adhesive).
NOTE: These bonding agents are highly inert and resistant to a large number of polar solvents. The resistance of the adhesive to nonpolar solvents (e.g., toluene or hexane) or high temperatures should be checked prior to use. - Immerse the AFM cantilever chip and the probe sample in the liquid, here: ultrapure water.
NOTE: A solvent drop (about 100 µL) can be deposited on the AFM cantilever chip holder. Covering the AFM cantilever chip with solvent reduces capillary forces, which would otherwise act on the AFM cantilever when approaching the sample surface passing through the air-solvent interface. - If required, adjust the temperature and let the system equilibrate.
NOTE: Temperature changes may result in a deflection of the AFM cantilever due to a bimetallic effect for AFM cantilevers with a reflective coating like aluminum or gold. Equilibration should be performed away from the surface (several µm) until no further change of the deflection signal is observed (up to 15 min for MLCT-Bio-DC). - Vary the temperature randomly to exclude any effects of ageing of the functionalization. Make sure that the temperatures applied do not lead to an irreversible bending of the AFM cantilever.
NOTE: Any temperature effects on solvent properties (such as evaporation or changes in viscosity) might hamper your experiments. In the presented examples, the temperature was varied over a range of up to 40 K in steps of 10 K taking water as a solvent (e.g., from 278 K to 318 K). - Approach the surface to determine the InvOLS (inverse optical lever sensitivity) by taking force-extension curves on a hard surface (such as silicon oxide). For this, take the deflection signal of the photodetector (in V) vs piezo distance and determine the slope of the part representing the indentation of the AFM cantilever tip into the underlying surface (repulsive regime) using a linear function. In order to reduce errors, take the average of at least five values to obtain the final InvOLS value. For further details, see the literature4,39.
NOTE: The InvOLS can only be reliably determined on hard surfaces. In the case of experiments on soft surfaces or interfaces make sure you place a hard surface close to your soft surfaces. Then, the InvOLS calibration can be done before or after your soft surface experiments without the need of disassembling the AFM setup. - For spring constant determination, move the AFM cantilever to a height with neither attractive nor repulsive interactions between AFM cantilever tip and surface (several µm). Then, record a thermal noise spectrum where the power spectral density (PSD) vs frequency is plotted. The following steps are usually performed by automated built-in functions in commercial AFM software: first, the acquired thermal noise spectrum is analyzed by fitting a function to the PSD, e.g., a simple harmonic oscillator (SHO). The fit is done up to the minimum between the first and second resonance. Second, the area under the fitted part of the PSD vs frequency plot is determined representing the mean square displacement of the AFM cantilever in vertical direction. Finally, the equipartition theorem is used to obtain the AFM cantilever force constant28,50.
NOTE: An appropriate frequency range should be used comprising the first resonance peak of the AFM cantilever. To get a satisfactory signal-to-noise ratio, at least 10 PSDs should be accumulated with the highest possible frequency resolution. - Start the experiment. Record force maps by taking force-extension curves in a grid-like fashion (e.g., 10 x 10 points for an area of 20 x 20 µm2) to avoid any local surface effects (e.g., impurities, dislocations) and to average different surface areas.
NOTE: Typical parameters are a pulling velocity of 1 µm/s and a sampling rate of 5 kHz to ensure sufficient resolution. The sampling rate should be adapted when the pulling velocity is varied. The retract distance should be adapted to the contour or desorption length of the measured polymer (approx. twice the expected length). - Use and vary the dwell time towards the surface to allow the single polymer to adhere to the surface (typically 0 – 5 s).
- Repeat the determination of the InvOLS and the spring constant at the end of the experiment to check the consistency and the stability of the system.
NOTE: For strong adhesion between polymer and surface, the calibration can be done after the actual experiment to preserve the functionalization.
6. Data evaluation
NOTE: For data evaluation, a custom-written software based on Igor Pro was used for performing the following steps.
- Convert the raw deflection signal (in Volts) into force values (in Newtons) by multiplication with the recorded InvOLS and the determined spring constant.
- Subtract the deflection of the AFM cantilever (after multiplication of the raw deflection signal with the InvOLS) from the distance driven by the piezo elements in vertical direction in order to obtain the true extension (tip-surface distance)4.
- Correct the force-extension curves obtained for drift by fitting a linear function to the baseline after the last event and subtracting the same from the force-extension curve. The fitted part should represent a sufficient extension from the surface where neither attractive nor repulsive interactions are observed between AFM cantilever tip and underlying surface. Then, the baseline is set to the zero-axis.
NOTE: In the case of measurements on highly reflective surfaces like gold, interferences might appear. These result from partial reflection of the laser beam from the surface and from the backside of the AFM cantilever. So, the obtained force-extension curves might show a sinusoidal force signal artifact along the vertical extension. This is an artifact that hampers the final force values. In order to still take these force-extension curves into account, a correction is possible (Figure 2). - If the interferences appear in the force-extension curves, select a representative force-extension curve (retraction curve) showing no other events than possibly a peak of unspecific adhesion and the same sinusoidal artifact (i.e., amplitude and phase) (Figure 2A).
NOTE: Smooth the representative force-extension curve in order to obtain the low frequency pattern of the interference. - Select a force-extension curve that is to be corrected (Figure 2B).
- Overlay both force-extension curves from steps 6.4. and 6.5. to make sure that both show the same sinusoidal artifact (i.e., amplitude and phase) (Figure 2C).
- Subtract the (smoothed) representative force-extension curve from the force-extension curve to be corrected leading to a straight rather than a sinusoidal baseline (Figure 2D).
NOTE: Take care that the unspecific adhesion peak of the representative curve is distinct from any single molecule events appearing in the curves to be corrected. In fact, the selection of the representative curve is crucial for a proper correction.
Results
The following examples show results of single molecule stretching and desorption of the polymers PEG, PNiPAM and PS. All AFM cantilever tips were functionalized with the protocol given above. PEG and PNiPAM were measured on SiOx with temperature variation. For a detailed discussion of the resulting temperature-dependent stretching curves for PEG and PNiPAM, see Kolberg et al.18 A different force-extension motif is a plateau of constant force (e.g., when desorbing PS from self-assembled monolayers of methyl terminated alkane thiols on gold (SAM) in water4,27,39,51).
Example 1: Stretching of PEG and PNiPAM in water
The temperature-dependent stretching behavior in water was measured using single PNiPAM and PEG polymers covalently bound to an AFM cantilever tip at one end and physisorbed on a SiOx surface at the other end. After the calibration and clean control experiments (less than 2% of the force-extension curves show single molecule events), at least two force maps were recorded for each AFM cantilever. The temperature-dependent experiment was performed by recording at least one force map at each temperature. When only few stretching events appeared, the respective AFM cantilever was discarded and the next AFM cantilever of the chip was taken (usually in the order C, B, D and E of MLCT-Bio-DC). For the exemplary data of PEG, a single stretching event was observed in 95 out of 500 measured force-extension curves (19%). For PNiPAM, 252 out of 600 force-extension curves showed a stretching pattern (42%). For a better comparison of the force-extension curves, a single master curve for every temperature was generated. For this purpose, only those curves with a stretching event to at least 500 pN, where conformational fluctuations and solvent effects are negligible, were chosen52. The final number of stretches taken into account was 3 at 278 K, 7 at 298 K and 4 at 318 K for PEG and 4 at 278 K, 3 at 298 K and 3 at 318 K for PNiPAM18.
The procedure for generating master curves is given in Figure 3. The force-extension curves chosen (Figure 3A) are rescaled to a length L0 (extension at a force of 500 pN), see Figure 3B. The adhesion peak shows a large variation of unspecific adhesion between the surface and the AFM cantilever tip, but does not influence the polymer stretching behavior. After merging the rescaled force-extension curves they are averaged by a binominal smoothing as presented in Figure 3C. For this, a Gaussian filter convolves the data with normalized coefficients derived from Pascal's triangle at a level equal to the smoothing parameter 2053. Finally, a master curve is obtained for every temperature as given in Figure 3D. The zoom-in shows the range where the temperature effect on the force-extension behavior is most pronounced.
A comparison of the temperature behavior of PEG (A) and PNiPAM (B) can be found in Figure 4. For PEG a decrease of the stretching force with increasing temperature was observed. An increase of approximately 5% of rescaled extension at 100 pN was observed when increasing the temperature from 278 to 318 K. For PNiPAM, an opposite temperature-dependent shift could be revealed. A decrease of approximately 1% of rescaled extension at 100 pN was observed when the temperature was increased from 278 to 328 K. Additionally, the stretching free energy could be obtained from the force-extension master curves by determining the area under the curve for any given force value. This could be used for extracting energetic and entropic contributions of the stretching free energy with the help of molecular dynamics (MD) simulations18.
Example 2: Desorption of PS from a SAM surface in water
The desorption of PS from a SAM surface in water could be used to determine the desorption force and length and thereby quantify the hydrophobic interaction. After calibration, at least two force maps were recorded at two different spots of the surface. When the polymer attachment was successful, the force-extension curves showed plateaus of constant force, as characteristic feature, see Figure 5A and Figure 5C. Plateau-like desorption is observed when the dynamics of the probed bonds are much faster than the pulling rate of the AFM cantilever tip (quasi-equilibrium). Desorption forces of plateau-like force-extension curves directly provide adhesion free energies by integrating the force-extension trace54. They have been used to determine electrostatic, dispersive and hydrophobic interactions as well as friction properties of single polymers on surfaces in liquid environment2,4,23,51,54,55.
Each plateau of constant force was fitted with a sigmoidal curve to determine the desorption force and desorption length, which were then plotted in histograms. The histograms were fitted with a Gaussian to extract the maximum value and standard deviation. For a better overview, the desorption force and length values were displayed together in a scatter plot, as given in Figure 5B and Figure 5D.
For polystyrene on SAM in water, the determined desorption forces correspond to previously obtained values19,23. As the desorption length correlates with the polymer contour length51, the desorption length distribution can be used as a proof of the covalent binding of the respective polymer to the AFM cantilever tip via its functional end group. Thus, the desorption length serves as a fingerprint.
For more than one polymer attached to the AFM cantilever tip, cascades of plateaus (discrete steps) can be observed in the force-extension curves56. Each plateau represents the desorption of a polymer at a different extension. The experiment given in Figure 5C and Figure 5D showed a typical case of two polymers attached to the AFM cantilever tip at the same time. By fitting the final rupture, a bimodal distribution could be found for the desorption length, while the desorption force showed a narrow distribution. In this case, the smaller desorption length could be found in 90% of the force-extension curves, either as a single plateau or as an additional plateau on the longer plateau, as given in Figure 5C. The higher desorption length was found in 37% of the obtained force-extension curves. Thus, the desorption length distribution could be used to determine the number of different polymers attached to the AFM cantilever tip. In general, a narrow distribution of the desorption length values is a good indication that one and the same single polymer was probed in the obtained force-extension curves. At the same time, a superposition of the respective forces-extension can be used to decide whether one and the same single polymer has been measured.
After proving covalent binding of a single PS polymer, further experiments with this PS polymer can be performed varying substrate (solid surface as well as polymer films), solvent conditions, temperature, pulling velocity or dwell time.
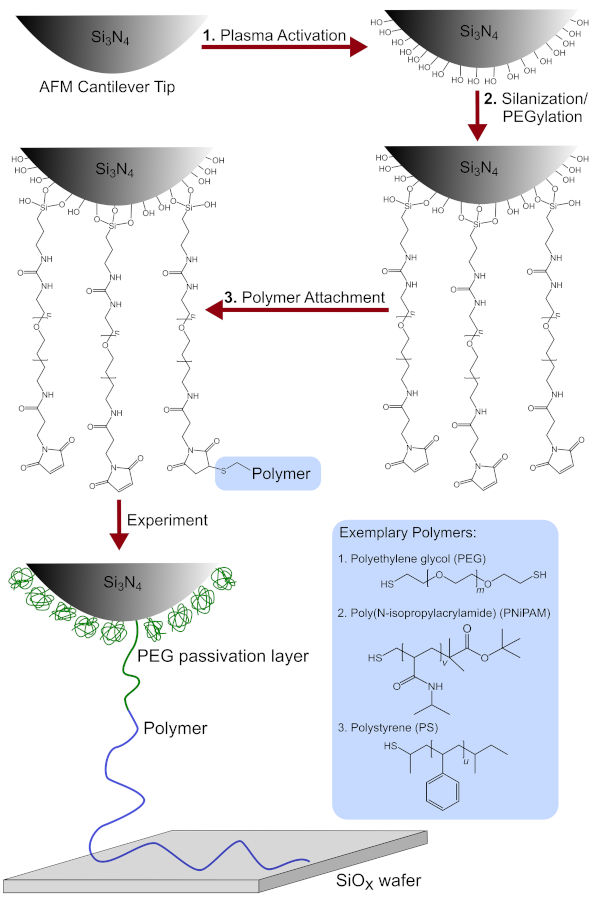
Figure 1: Schematic overview of the tip functionalization process. Includes the chemical modification of the AFM cantilever tip after (1) plasma activation (2) silanization/PEGylation and (3) polymer attachment. Additionally, the detailed chemical structures of the polymers used, namely PEG, PNiPAM and PS are shown. Please click here to view a larger version of this figure.
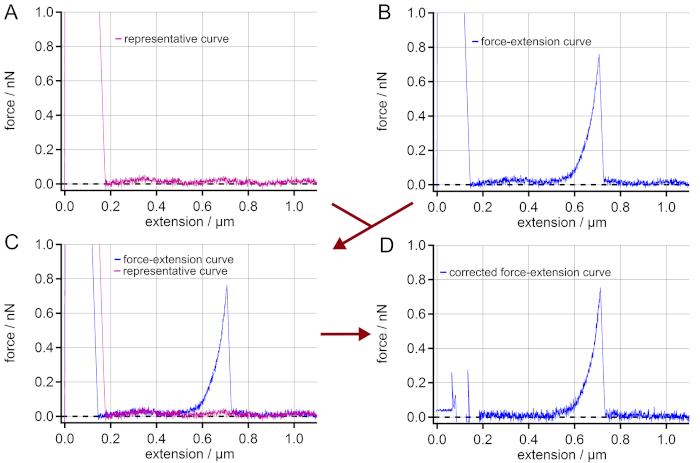
Figure 2: Elimination of interferences in force-extension curves. (A) Find a force-extension curve showing a sinusoidal force signal artifact along the extension but having no single molecule stretching event. (B) Choose a force-extension curve with a single molecule event, which is to be corrected from the sinusoidal artifact. (C) Superimpose the curves to control if the sinusoidal artifacts of the curves really match. (D) By subtracting the force-extension curve (A) from (B) a force-extension curve with a straight baseline is obtained. Although the adhesion peak cannot be used for further analysis, the force-extension curve is now corrected for the artifact leading to much more accurate force values in the region of the single molecule event (here: > 0.2 µm of extension). Please click here to view a larger version of this figure.
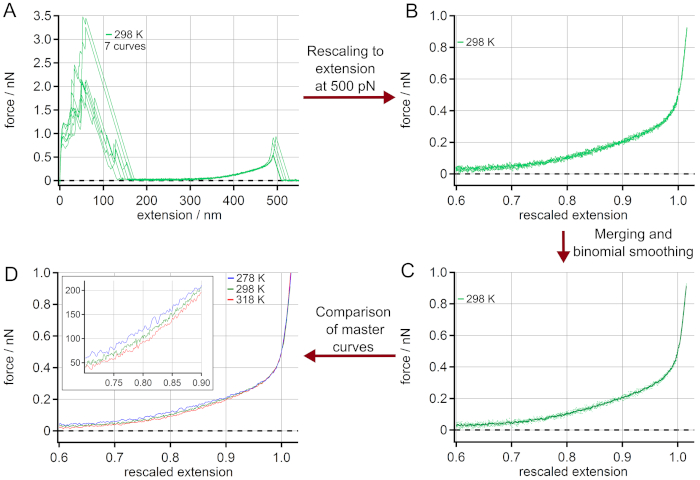
Figure 3: Determination of master curves from force-extension curves of PEG at 298 K. (A) Experimental data at 298 K, using 7 force-extension curves. After rescaling to a length L0 at a force of 500 pN (B), the force-extension curves can be merged and averaged by binominal smoothing obtaining a master curve (C). The rescaled curves are given as dots while the master curve is shown as a solid line. Finally, the obtained master curves for different temperatures can be compared (D). The zoom-in indicates the range where the temperature effect on the force-extension behavior is most pronounced. Please click here to view a larger version of this figure.
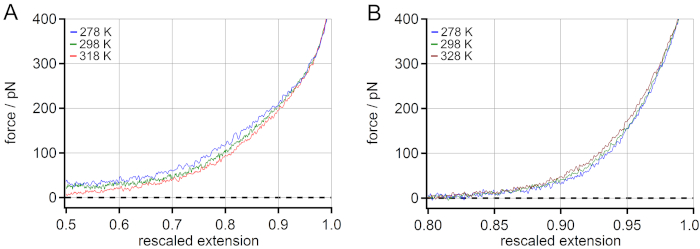
Figure 4: Comparison of the temperature-dependent master curves of PNiPAM and PEG. For PEG an increase of rescaled extension at 100 pN (mid-force range) is observed when increasing the temperature (A), while for PNiPAM an opposite temperature-dependent shift is revealed (B). Please click here to view a larger version of this figure.
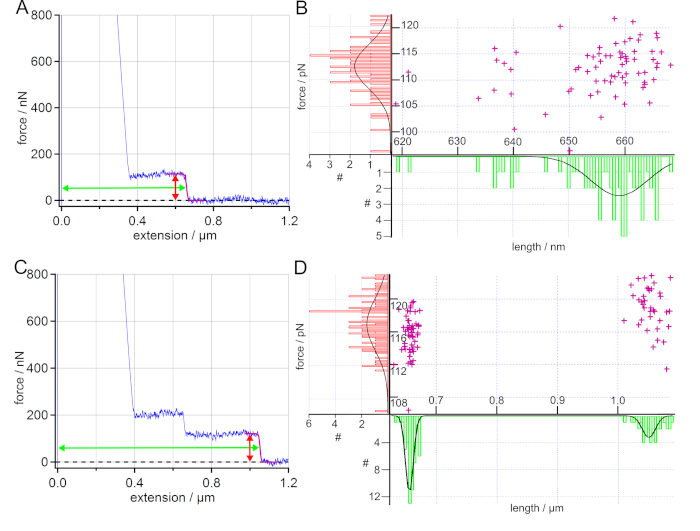
Figure 5: Analysis of force-extension curves of PS on SAM in water. (A) Exemplary force-extension curve (blue) with a sigmoidal fit of the plateau (purple). Additionally, the arrows mark the determined force (red) and length (green) of the plateau. The desorption force and desorption length values obtained by sigmoidal fits are displayed in a scatter plot and the resulting histograms are fitted with a Gaussian. (B) The determined average desorption force and desorption length values are (112 ± 6) pN and (659 ± 7) nm, wherein 93% of the force-extension curves show such single plateau events. (C) Exemplary force-extension curve (blue) for two polymers attached to the AFM cantilever tip at the same time. Here, the desorption force shows a unimodal distribution with an average force value of (117 ± 5) pN, while a bimodal distribution can be found for the desorption length leading to average length values of (656 ± 9) nm and (1050 ± 16) nm. (D) 90% of the sampled force-extension curves show only single plateau events. Please click here to view a larger version of this figure.
Discussion
AFM-based SMFS is one of the major tools for investigating single molecule interactions in polymer physics. For a true single molecule experiment, covalent attachment of the probe polymer to an AFM cantilever tip is essential.
Many previous works are based on nanofishing experiments, in particular for PNiPAM, where polymers are adsorbed onto a surface and then stretched by randomly picking them from the substrate using an AFM cantilever tip30,31. This might alter the results and lead to misinterpretation of the single molecule behavior. There, cooperative effects might dominate the results because interactions with neighboring polymers cannot be excluded. This has a large impact on the results, especially for polymers that show significantly different behavior in bulk compared to single isolated molecules57,58.
The functionalization protocol presented here is reliable and can be easily applied to different polymers, irrespective of their contour length, hydrophobicity or the steric hindrance of the monomers. Additionally, a passivation is provided to prevent unwanted interactions between the single probe polymer and the AFM cantilever tip as well as between the AFM cantilever tip and the underlying surface. Furthermore, the evaluation of force-extension curves showing stretching events is shown. There, a procedure is proposed for the determination of master force-extension curves. This offers a better means of revealing, e.g., temperature-related effects on the force-extension behavior. Furthermore, the analysis of single molecule desorption events featuring constant force plateaus is provided. Also, a simple way of correcting sinusoidal force signal artifacts in force-extension curves is given which might otherwise impair the results of the experiment.
Compared to Stetter et al.39, the functionalization procedure presented here is reduced to three steps instead of four and the robustness of the procedure is improved. The major benefit of performing PEGylation and silanization in one step is to have a better-controlled reaction and to increase the yield. Furthermore, fewer solutions need to be prepared and fewer rinsing steps are required. This reduces the effort and time for preparation and increases the reproducibility. Furthermore, moving AFM cantilevers is always a critical part of the functionalization process. A transfer from one solution to the other always runs the risk of strongly influencing the functionalization quality due to transfers through the air-water interface or of losing AFM cantilevers by improper use of tweezers.
In order to prove proper covalent attachment of a single polymer to an AFM cantilever tip different conditions have to be met. First, control AFM cantilevers are of significant importance and should be prepared for every functionalization. The functionalization process and the fluid cell for performing the experiments are only considered to be clean, if a small number of force-extension curves show stretches or plateaus in the control experiment (in the presented examples less than 2%).
A clear stretching pattern with no further drops or maxima is essential for having proper single molecule stretching events. Additionally, the dependence of rupture force on the force loading rate at rupture or the complete elastic response of the stretching curve should be analyzed in order to exclude simultaneous desorption of multiple polymers59,60. For PEG and PNiPAM, 19% and 42% of the force-extension curves taken at different positions of the surface showed such a stretching pattern, respectively. In order to obtain stretching events, the physisorption of the polymer to the respective underlying surface must be strong. Otherwise a plateau-like desorption event is observed. This is even more decisive for the detection of stretching events at high forces (up to 500 pN or more). As this strong physisorption is not met for every force-extension curve, the yield of such events is less than for pure plateau-like desorption events. As an alternative, strongly adhering groups such as catechols or chemisorption between polymer and underlying surface can be used. However, this requires the introduction of further functional groups or coupling sites at the polymer61,62.
In fact, the mass (i.e., contour length) of the polymer provides a valuable fingerprint. Although the mass cannot directly be translated into the measured contour length for the following reasons, the length distribution is very valuable to define single-molecule events. In the case of a PNiPAM polymer with a low polydispersity (Ɖ = 1.28), we found significant differences in the extension values for the obtained stretching events (and thus in the polymer length) in the experiments. One reason for this could be the determination of the polymer length and its distribution. In size-exclusion chromatography (SEC), a relative weight of the target polymer is determined in comparison with standards like PS or poly (methyl methacrylate) (PMMA)63. The presumed relative weight is expected to deviate from the absolute molecular weight because the hydrodynamic radius of the target polymer and the standard can significantly differ. Additionally, the silane layer might be oligomerized by spurious water in toluene during the functionalization process. The attachment of such oligomers to the AFM cantilever tip leads to a more flexible layer with fewer anchor points64. Also, the attachment point of the polymer to the silicon layer might not necessarily be at the apex leading to a shift of the detected length values29. While a polymer model such as the wormlike chain (WLC) or the freely jointed chain (FJC) model cannot reproduce the respective force-extension behavior for PEG or PNiPAM properly over the entire extension range18,29,41,65,66, such a polymer model might be valuable for other polymeric and protein systems10,15,67,68.
The covalent attachment of a single PS polymer (with a contour length of more than 1 µm) is only considered to be successful, when a considerable number of force-extension curves show a long enough plateau of constant force (Figure 5). A plateau resulting from desorbing a single polymer is defined by a single sharp drop of a constant force to the baseline at a certain extension, as given in Figure 5A. If more polymers are attached to the AFM cantilever tip, a cascade of plateaus is observed56 (Figure 5C). The plateau length (desorption length), correlating with the polymer contour length51, has to be significantly longer than any adhesion peak due to unspecific adhesion of the AFM cantilever tip to the underlying surface (here around 200 nm). Features appearing solely in a single force-extension curve, should not be interpreted. In the presented experiments, at least 80 out of 100 curves showed a plateau longer than 200 nm in at least two force maps at two different spots on the surface. Furthermore, the distribution of desorption lengths, using scatter plots such as given in Figure 5B and 5D, reveal if and how many polymers are bound to the AFM cantilever tip. In the case of PS, a narrow distribution of desorption force and length taken from plateaus of the force-extension curves served as evidence of a successful covalent attachment. This finally proved the success of the functionalization protocol. Thus, we strongly recommend to present such force and length distributions in publications.
Evaluating force-extension curves using built-in algorithms that comprise many pre-set parameters should be done with care. Reasons are for example that a fixed sampling rate is not appropriate for every applied pulling velocity or that an automated smoothing of the force-extension curves might average out important details. Usually a proper understanding of the respective evaluation procedure can prevent errors in the evaluation procedure, which can in turn strongly influence the final findings of an AFM-based SMFS experiment.
In summary, we present a functionalization protocol that is reliable and can be easily applied to a variety of polymers. Furthermore, proper evaluation of single molecule force-extension curves is presented, allowing the determination of physical parameters such as stretching force, desorption force and desorption length. The presented protocols and procedures are valuable for the investigation of stimuli-responsive systems at the single molecule level.
Disclosures
The authors declare that they have no competing financial interest.
Acknowledgements
B.N.B. and T.H. acknowledge funding by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy – EXC-2193/1 – 390951807, gefördert durch die Deutsche Forschungsgemeinschaft (DFG) im Rahmen der Exzellenzstrategie des Bundes und der Länder – EXC-2193/1 – 390951807, and grant HU 997/1-13 (project # 420798410). M.G. acknowledges partial support in the frame of the LOEWE Project iNAPO by the Hessen State Ministry of Higher Education, Research and the Arts. We thank Dr. Wolfgang Bronner and Dr. Agne Zukauskaite from Fraunhofer Institute for Applied Solid State Physics IAF for the donation of high quality gold coated silicon wafers.
Materials
| Name | Company | Catalog Number | Comments |
| 1-Dodecanethiol (≥98%) | Sigma-Aldrich, USA | 417364-500ML | Used for SAM |
| Ammonia solution (30%) | Roth, Germany | CP17.2 | Used for cleaning |
| Cypher ES | Asylum Research, an Oxford Instruments company, USA | - | AFM |
| Ethanol (≥99.9%) | Roth, Germany | PO76.1 | Solvent |
| Gold coated silicon wafer | Fraunhofer Institute for Applied Solid State Physics IAF, Germany | - | Used for SAM |
| High Resolution Replicating Compound | Microset Products Ltd, UK | 101RF | Bonding agent |
| Hydrogen peroxide solution | Sigma-Aldrich, USA | H1009 | Used for cleaning |
| Igor Pro | Wavemetrics, USA | - | Software environment |
| Tetra-30-LF-PC | Diener Electronic, Germany | - | Plasma chamber |
| Maleimide-polyethylene glycol-triethoxysilane | Creative PEG works, USA | PHB-1923 | Linker polymer |
| MLCT-Bio-DC | Bruker, USA | MLCT-Bio-DC | AFM cantilever |
| Prime CZ-Si wafer, n-type (Phosphor) TTV < 10 µm | MicroChemicals, Germany | WSA40600250 P1314SNN1 | Silicon wafer |
| Purelab Chorus 1, 18.2 MΩ cm | Elga LabWater, Germany | 10034-540 | Ultrapure water source |
| R3 SA | Vomm GmbH, Germany | 5803 Blank | Tweezers |
| Thiol terminated poly(N-isopropylacrylamide) | Gallei Group, Saarland University, Germany | - | PNiPAM probe polymer |
| Thiol terminated polystyrene | Polymer Source, Canada | P40722-SSH | PS probe polymer |
| Thiol-polyethylene glycol-thiol | Creative PEGWorks, USA | PSB-615 | PEG probe polymer |
| Toluene (99.99%) | Fisher Chemicals | T324-500 | Solvent |
References
- Binnig, G., Quate, C. F., Gerber, C. Atomic Force Microscope. Physical Review Letters. 56 (9), 930-933 (1986).
- Hugel, T., Seitz, M. The Study of Molecular Interactions by AFM Force Spectroscopy. Macromolecular Rapid Communications. 22 (13), 989-1016 (2001).
- Butt, H. -. J., Cappella, B., Kappl, M. Force measurements with the atomic force microscope: Technique, interpretation and applications. Surface Science Reports. 59 (1-6), 1 (2005).
- Balzer, B. N., Hugel, T., Hashmi, S. Single-Molecule Detection and Manipulation. Reference Module in Materials Science and Materials Engineering. , (2016).
- Krieg, M., et al. Atomic force microscopy-based mechanobiology. Nature Reviews Physics. 1 (1), 41-57 (2019).
- Edwards, D. T., Faulk, J. K., LeBlanc, M. -. A., Perkins, T. T. Force Spectroscopy with 9-μs Resolution and Sub-pN Stability by Tailoring AFM Cantilever Geometry. Biophysical journal. 113 (12), 2595-2600 (2017).
- Alsteens, D., et al. Nanomechanical mapping of first binding steps of a virus to animal cells. Nature Nanotechnology. 12 (2), 177-183 (2017).
- Kodera, N., Yamamoto, D., Ishikawa, R., Ando, T. Video imaging of walking myosin V by high-speed atomic force microscopy. Nature. 468, 72-76 (2010).
- Shibata, M., et al. Real-space and real-time dynamics of CRISPR-Cas9 visualized by high-speed atomic force microscopy. Nature Communications. 8 (1), 1-9 (2017).
- Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J. M., Gaub, H. E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science. 276 (5315), 1109-1112 (1997).
- Oesterhelt, F., Oesterhelt, D., Pfeiffer, M., Engel, A., Gaub, H. E., Müller, D. J. Unfolding pathways of individual bacteriorhodopsins. Science. 288 (5463), 143-146 (2000).
- Hugel, T., Holland, N. B., Cattani, A., Moroder, L., Seitz, M., Gaub, H. E. Single-molecule optomechanical cycle. Science. 296 (5570), 1103-1106 (2002).
- Yu, H., Siewny, M. G. W., Edwards, D. T., Sanders, A. W., Perkins, T. T. Hidden dynamics in the unfolding of individual bacteriorhodopsin proteins. Science. 355 (6328), 945-950 (2017).
- Erlich, K. R., Sedlak, S. M., Jobst, M. A., Milles, L. F., Gaub, H. E. DNA-free directed assembly in single-molecule cut-and-paste. Nanoscale. 11 (2), 407-411 (2019).
- Rico, F., Russek, A., González, L., Grubmüller, H., Scheuring, S. Heterogeneous and rate-dependent streptavidin-biotin unbinding revealed by high-speed force spectroscopy and atomistic simulations. Proceedings of the National Academy of Sciences of the United States of America. 116 (14), 6594-6601 (2019).
- Löf, A., et al. Multiplexed protein force spectroscopy reveals equilibrium protein folding dynamics and the low-force response of von Willebrand factor. Proceedings of the National Academy of Sciences of the United States of America. 116 (38), 18798-18807 (2019).
- Kienle, S., Liese, S., Schwierz, N., Netz, R. R., Hugel, T. The effect of temperature on single-polypeptide adsorption. Chemphyschem : a European journal of chemical physics and physical chemistry. 13 (4), 982-989 (2012).
- Kolberg, A., et al. Opposing Temperature Dependence of the Stretching Response of Single PEG and PNiPAM Polymers. Journal of the American Chemical Society. 141 (29), 11603-11613 (2019).
- Balzer, B. N., et al. Cohesion Mechanisms of Polystyrene-Based Thin Polymer Films. Macromolecules. 46 (18), 7406-7414 (2013).
- Balzer, B. N., et al. Adhesion property profiles of supported thin polymer films. ACS Applied Materials & Interfaces. 5 (13), 6300-6306 (2013).
- Stetter, F. W. S., Cwiklik, L., Jungwirth, P., Hugel, T. Single Lipid Extraction: The Anchoring Strength of Cholesterol in Liquid-Ordered and Liquid-Disordered Phases. Biophysical journal. 107 (5), 1167-1175 (2014).
- Schwierz, N., Krysiak, S., Hugel, T., Zacharias, M. Mechanism of Reversible Peptide-Bilayer Attachment: Combined Simulation and Experimental Single-Molecule Study. Langmuir. 32 (3), 810-821 (2016).
- Balzer, B. N., et al. Nanoscale Friction Mechanisms at Solid-Liquid Interfaces. Angewandte Chemie International Edition. 52 (25), 6541-6544 (2013).
- Balzer, B. N., Kienle, S., Gallei, M., von Klitzing, R., Rehahn, M., Hugel, T. Stick-Slip Mechanisms at the Nanoscale. Soft Materials. 12, 106-114 (2014).
- Kühner, F., Erdmann, M., Sonnenberg, L., Serr, A., Morfill, J., Gaub, H. E. Friction of single polymers at surfaces. Langmuir. 22 (26), 11180-11186 (2006).
- Grebíková, L., Gojzewski, H., Kieviet, B. D., Klein Gunnewiek, M., Vancso, G. J. Pulling angle-dependent force microscopy. The Review of Scientific Instruments. 88 (3), 33705 (2017).
- Geisler, M., et al. Hydrophobic and Hofmeister effects on the adhesion of spider silk proteins onto solid substrates: an AFM-based single-molecule study. Langmuir. 24 (4), 1350-1355 (2008).
- Pirzer, T., Hugel, T. Atomic force microscopy spring constant determination in viscous liquids. Review of Scientific Instruments. 80 (3), 35110 (2009).
- Liese, S., et al. Hydration Effects Turn a Highly Stretched Polymer from an Entropic into an Energetic Spring. ACS Nano. 11 (1), 702-712 (2017).
- Cui, S., Pang, X., Zhang, S., Yu, Y., Ma, H., Zhang, X. Unexpected Temperature-Dependent Single Chain Mechanics of Poly(N-isopropyl-acrylamide) in Water. Langmuir. 28 (11), 5151-5157 (2012).
- Liang, X., Nakajima, K. Nanofishing of a Single Polymer Chain: Temperature-Induced Coil-Globule Transition of Poly(N -isopropylacrylamide) Chain in Water. Macromolecular Chemistry and Physics. 219 (3), 1700394 (2018).
- Zhang, W., Zou, S., Wang, C., Zhang, X. Single Polymer Chain Elongation of Poly(N -isopropylacrylamide) and Poly(acrylamide) by Atomic Force Microscopy. The Journal of Physical Chemistry B. 104 (44), 10258-10264 (2000).
- Hermanson, G. T. . Bioconjugate techniques - 3rd Edition. , (2013).
- Leitner, M., et al. Single-molecule AFM characterization of individual chemically tagged DNA tetrahedra. ACS Nano. 5 (9), 7048-7054 (2011).
- Walder, R., et al. Rapid Characterization of a Mechanically Labile α-Helical Protein Enabled by Efficient Site-Specific Bioconjugation. Journal of the American Chemical Society. 139 (29), 9867-9875 (2017).
- Tang, J., et al. High-affinity tags fused to s-layer proteins probed by atomic force microscopy. Langmuir. 24 (4), 1324-1329 (2008).
- Wildling, L., et al. Linking of sensor molecules with amino groups to amino-functionalized AFM tips. Bioconjugate Chemistry. 22 (6), 1239-1248 (2011).
- Maity, S., Viazovkina, E., Gall, A., Lyubchenko, Y. A. A Metal-free Click Chemistry Approach for the Assembly and Probing of Biomolecules. Journal of Nature and Science. 2 (4), 187 (2016).
- Stetter, F. W. S., Kienle, S., Krysiak, S., Hugel, T. Investigating Single Molecule Adhesion by Atomic Force Spectroscopy. Journal of Visualized Experiments. (96), e52456 (2015).
- Becke, T. D., et al. Covalent Immobilization of Proteins for the Single Molecule Force Spectroscopy. Journal of Visualized Experiments. (138), e58167 (2018).
- Ott, W., et al. Elastin-like Polypeptide Linkers for Single-Molecule Force Spectroscopy. ACS Nano. 11 (6), 6346-6354 (2017).
- Newton, R., et al. Combining confocal and atomic force microscopy to quantify single-virus binding to mammalian cell surfaces. Nature Protocols. 12 (11), 2275-2292 (2017).
- Staros, J. V., Wright, R. W., Swingle, D. M. Enhancement by N-hydroxysulfosuccinimide of water-soluble carbodiimide-mediated coupling reactions. Analytical Biochemistry. 156 (1), 220-222 (1986).
- Grabarek, Z., Gergely, J. Zero-length crosslinking procedure with the use of active esters. Analytical Biochemistry. 185 (1), 131-135 (1990).
- Baird, J. A., Olayo-Valles, R., Rinaldi, C., Taylor, L. S. Effect of Molecular Weight, Temperature, and Additives on the Moisture Sorption Properties of Polyethylene Glycol. Journal of Pharmaceutical Sciences. 99 (1), 154-168 (2010).
- Halperin, A., Kröger, M., Winnik, F. M. Poly(N-isopropylacrylamide) Phase Diagrams: Fifty Years of Research. Angewandte Chemie International Edition. 54 (51), 15342-15367 (2015).
- Barradas, R. G., Fletcher, S., Porter, J. D. The hydrolysis of maleimide in alkaline solution. Canadian Journal of Chemistry. 54 (9), 1400-1404 (1976).
- Kharasch, N., Tarbell, D. S. Chapter 10 - The Mechanism of Oxidation of Thiols to Disulfides. Organic Sulfur Compounds. , 97-102 (1961).
- Folkers, J. P., Laibinis, P. E., Whitesides, G. M. Self-assembled monolayers of alkanethiols on gold: comparisons of monolayers containing mixtures of short- and long-chain constituents with methyl and hydroxymethyl terminal groups. Langmuir. 8 (5), 1330-1341 (1992).
- Hutter, J. L., Bechhoefer, J. Calibration of atomic-force microscope tips. Review of Scientific Instruments. 64 (7), 1868-1873 (1998).
- Krysiak, S., Liese, S., Netz, R. R., Hugel, T. Peptide desorption kinetics from single molecule force spectroscopy studies. Journal of the American Chemical Society. 136 (2), 688-697 (2014).
- Hugel, T., Rief, M., Seitz, M., Gaub, H. E., Netz, R. R. Highly Stretched Single Polymers: Atomic-Force-Microscope Experiments Versus Ab-Initio Theory. Physical Review Letters. 94 (4), 48301 (2005).
- Marchand, P., Marmet, L. Binomial smoothing filter: A way to avoid some pitfalls of least-squares polynomial smoothing. Review of Scientific Instruments. 54 (8), 1034-1041 (1983).
- Horinek, D., et al. Peptide adsorption on a hydrophobic surface results from an interplay of solvation, surface, and intrapeptide forces. Proceedings of the National Academy of Sciences of the United States of America. 105 (8), 2842-2847 (2008).
- Friedsam, C., Gaub, H. E., Netz, R. R. Adsorption energies of single charged polymers. EPL (Europhysics Letters). 72 (5), 844-850 (2005).
- Scherer, A., Zhou, C., Michaelis, J., Brauchle, C., Zumbusch, A. Intermolecular Interactions of Polymer Molecules Determined by Single-Molecule Force Spectroscopy. Macromolecules. 38 (23), 9821-9825 (2005).
- Abbott, L. J., Tucker, A. K., Stevens, M. J. Single Chain Structure of a Poly(N-isopropylacrylamide) Surfactant in Water. The Journal of Physical Chemistry B. 119 (9), 3837-3845 (2015).
- Okano, T., Bae, Y. H., Jacobs, H., Kim, S. W. Thermally on-off switching polymers for drug permeation and release. Journal of Controlled Release. 11 (1), 255-265 (1990).
- Sulchek, T., Friddle, R. W., Noy, A. Strength of multiple parallel biological bonds. Biophysical journal. 90 (12), 4686-4691 (2006).
- Sulchek, T. A., et al. Dynamic force spectroscopy of parallel individual Mucin1-antibody bonds. Proceedings of the National Academy of Sciences of the United States of America. 102 (46), 16638-16643 (2005).
- Krysiak, S., Wei, Q., Rischka, K., Hartwig, A., Haag, R., Hugel, T. Adsorption mechanism and valency of catechol-functionalized hyperbranched polyglycerols. Beilstein Journal of Organic Chemistry. 11, 828-836 (2015).
- Jobst, M. A., Schoeler, C., Malinowska, K., Nash, M. A. Investigating receptor-ligand systems of the cellulosome with AFM-based single-molecule force spectroscopy. Journal of Visualized Experiments. (82), e50950 (2013).
- Trathnigg, B. Determination of MWD and chemical composition of polymers by chromatographic techniques. Progress in Polymer Science. 20 (4), 615-650 (1995).
- Blass, J., Albrecht, M., Wenz, G., Zang, Y. N., Bennewitz, R. Single-molecule force spectroscopy of fast reversible bonds. Physical Chemistry Chemical Physics. 19 (7), 5239-5245 (2017).
- Oesterhelt, F., Rief, M., Gaub, H. E. Single molecule force spectroscopy by AFM indicates helical structure of poly(ethylene-glycol) in water. New Journal of Physics. 1, 1-11 (1999).
- Xue, Y., Li, X., Li, H., Zhang, W. Quantifying thiol-gold interactions towards the efficient strength control. Nature Communications. 5, 4348 (2014).
- Lyu, X., Song, Y., Feng, W., Zhang, W. Direct Observation of Single-Molecule Stick-Slip Motion in Polyamide Single Crystals. ACS Macro Letters. 7 (6), 762-766 (2018).
- Hugel, T., et al. Elasticity of Single Polyelectrolyte Chains and Their Desorption from Solid Supports Studied by AFM Based Single Molecule Force Spectroscopy. Macromolecules. 34 (4), 1039-1047 (2001).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved