A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Behavioral Assays for Optogenetic Manipulation of Neural Circuits in Drosophila melanogaster
* These authors contributed equally
In This Article
Summary
This paper presents methods for optogenetic manipulation in Drosophila melanogaster, utilizing CsChrimson and GtACR2 to activate and silence specific neurons. Four experiments are described to utilize optogenetics to explore thermotactic and gustatory behaviors, providing insights into the underlying neural mechanisms governing these processes.
Abstract
Optogenetics has become a fundamental technique in neuroscience, enabling precise control of neuronal activity through light stimulation. This study introduces easy-to-implement setups for applying optogenetic methods in Drosophila melanogaster. Two optogenetic tools, CsChrimson, a red-light-activated cation channel, and GtACR2, a blue-light-activated anion channel, were employed in four experimental approaches. Three of these approaches involve single-fly experiments: (1) a blue-light optogenetic thermotactic positional preference assay targeting temperature-sensitive heating cells, (2) a red-light optogenetic positional preference assay activating bitter sensing neurons, and (3) a proboscis extension response assay activating the sweet-sensing neurons. The fourth approach (4) is a fly maze setup to assess avoidance behaviors using multiple flies. The ability to manipulate neural activity temporally and spatially offers powerful insights into sensory processing and decision-making, underscoring the potential of optogenetics to advance our knowledge of neural function. These methods provide an accessible and robust framework for future research in neuroscience to enhance the understanding of specific neural pathways and their behavioral outcomes.
Introduction
Optogenetics has emerged as a powerful technique combining optics and genetics in neuroscience, providing precise, non-invasive control over neural activity through light stimulation1. In Drosophila melanogaster, a widely used model organism, optogenetic tools enable the activation and inhibition of specific neurons, allowing researchers to modulate neural circuits. Among the tools used, CsChrimson and GtACR (Guillardia theta anion channel rhodopsins) provide complementary approaches for neuronal targeting. CsChrimson channelrhodopsin, a red-light sensitive cation channel from green algae, facilitates neuronal activation through depolarization when exposed to red light, with peak activation at approximately 590 nm2. CsChrimson offers better tissue penetration than previous channelrhodopsins and reduces light-induced behavioral artifacts in Drosophila studies2. In contrast, GtACR, which includes variants such as GtACR2, is a light-gated chloride channel that silences neurons through hyperpolarization3,4. GtACR2 conducts anions and is activated by blue light with a peak activation around 470 nm4. CsChrimson and GtACR2 are activated by distinct wavelengths of light, ensuring precise and independent control of neuronal activity without cross-activation5.
Drosophila is an effective model for neuroscience research due to its cost-effectiveness, ease of rearing, and robust behavioral responses to environmental stimuli, including attractive and avoidance behaviors6. Its small size and semi-transparent cuticle enhance the penetration of light, especially long-wavelength red light, enabling efficient optogenetic manipulation7,8. While Drosophila cells cannot produce sufficient retinal, a crucial cofactor for the functionality of channelrhodopsins, adding retinal to their diet compensates for this limitation, ensuring effective activation of optogenetic tools9.
To explore the effects of optogenetic manipulation in Drosophila, we describe four experiments targeting different neural circuits and behaviors, each utilizing distinct modalities to assess either avoidance or attractive responses, ranging from single-fly assays to group-based evaluations. Heating cells (HC) in Drosophila are thermosensory neurons located in the arista, responding to temperature increases10. These neurons express warm-sensitive ion channels that trigger avoidance behavior, guiding flies away from heat sources10,11. In approach 1, we employed a single-fly blue-light optogenetic thermotactic positional preference assay to manipulate HC neurons. By expressing GtACR2 in these neurons, we inhibited their activity upon blue-light exposure. Flies were exposed to two temperature options: 25 °C and 31 °C. Under room light, flies avoided the 31 °C side, demonstrating a typical thermotactic response. However, blue-light activation of GtACR2 silenced the HC neurons. As a result, flies showed no significant temperature preference, suggesting successful optogenetic inhibition. In addition to assessing the function of sensory neurons, the expression of GtACR2 in downstream sensory neurons enables similar optogenetic manipulations to study the neural circuits necessary for specific sensory modalities5.
The gustatory receptor GR66a in Drosophila is expressed in the labial palps at the distal end of the proboscis and in the legs, mediating bitter taste detection12,13. These neurons trigger avoidance behaviors in response to bitter substances. In approach 2, we used a single-fly red-light optogenetic positional preference assay to manipulate GR66a-expressing neurons. By expressing CsChrimson in these neurons, we activated them upon red-light exposure. Flies were placed in an arena with one half exposed to red light and the other half filtering red light. In the absence of red light, flies showed no preference. However, red-light activation of CsChrimson stimulated the bitter-sensing neurons, resulting in significant avoidance of the illuminated area, confirming successful optogenetic activation of GR66a neurons. Similar approaches have been used to identify the downstream circuits of heating cells sufficient for the avoidance behavior5.
We focused on optogenetic activation of appetitive behavior in approach 3. GR5a-expressing neurons, located in the taste sensilla on the labellum and legs, detect sugars and drive feeding behavior. Activation of these neurons triggers the proboscis extension response (PER)14. We used a red-light optogenetic proboscis extension response assay to activate GR5a neurons. By expressing CsChrimson in these neurons, we stimulated them with red light. Flies did not extend their proboscis under room-light conditions. However, red-light activation of CsChrimson led to proboscis extension without a sweet stimulus, demonstrating successful optogenetic activation of GR5a neurons. This approach has been used to investigate the neural circuit, including gustatory sensory neurons, taste projection neurons, and proboscis motor neurons15,16.
In approach 4, we investigated optogenetic activation of avoidance behaviors in groups of flies, using a red-light optogenetic fly maze assay targeting GR66a neurons. Flies were placed at the intersection of two tubes: one illuminated with red light and the other shaded. CsChrimson expression in GR66a neurons triggered avoidance. In the absence of red light, flies showed no preference, but red-light activation led GR66a-expressing flies to avoid red light, suggesting the successful activation of the pathway. Fly maze assays have been widely used to study various sensory modalities, including temperature, humidity, and olfaction. When combined with optogenetics, this approach is powerful for investigating both attractive and avoidance behaviors17,18,19.
These methods provide a reproducible framework for studying optogenetic activation and inhibition of Drosophila neural circuits. By utilizing a combination of different channelrhodopsins and accessible behavioral assays, this proof-of-concept study demonstrates the effectiveness of optogenetic manipulation, providing straightforward methods to manipulate neural circuit functions with potential broader applications in neuroscience research.
Protocol
1. Strains, fly rearing, and fly aspirator
- Strains and maintenance
- The strains used in the experiments include HC-Gal410, Gr5a-Gal420 (Bloomington Drosophila Stock Center (BDSC): 57592), Gr66a-Gal421 (BDSC: 57670), UAS-GtACR23, UAS-CsChrimson2 (BDSC: 55136). Rear flies at 25 °C on a standard cornmeal medium under a 12-h light:12-h dark cycle.
NOTE: 1 L cornmeal medium contains 1 L dH2O, 79 g of dextrose, 7.5 g of agar, 24 g of flaked yeast, 57 g of cornmeal, 2.1 g of methyl-4-hydroxybenzoate (dissolved in 11.1 mL of ethanol), 6 g of sodium potassium tartrate tetrahydrate, and 0.9 g of calcium chloride.
- The strains used in the experiments include HC-Gal410, Gr5a-Gal420 (Bloomington Drosophila Stock Center (BDSC): 57592), Gr66a-Gal421 (BDSC: 57670), UAS-GtACR23, UAS-CsChrimson2 (BDSC: 55136). Rear flies at 25 °C on a standard cornmeal medium under a 12-h light:12-h dark cycle.
- Fly rearing and experimental preparation
- Sort 0 to 3-day-old flies. By the time of experimentation, the flies will be 3 to 6 days old, ensuring age-related consistency (Figure 1A).
- Divide each genotype into four groups 3 days before conducting experiments: room-light ATR -, room-light ATR +, red or blue-light ATR -, and red or blue-light ATR +.
- Prepare 80 mM ATR stock solution by dissolving ATR in 100% ethanol. Supplement the food with 400 µM ATR to the ATR + groups.
- Supplement the food with the same concentration of ethanol but without ATR to the ATR - groups.
- Rear all groups in the dark22.
- Starve Gr5a>CsChrimson flies for 16-24 hours in vials containing only damp tissue. The ATR + vials contain 400 µM ATR mixed with water, while the ATR - vials contain dH2O mixed with the same concentration of ethanol.
- Fly aspirator
- Assemble the fly aspirator (Figure 1B) using plastic tubing, insect netting, and two 3 mL transfer pipettes.
- Cut the tips and bulbs of the transfer pipettes, ensuring that the fly suction tip allows for comfortable passage of a fly. This design enables gentle aspiration of flies through inhalation, minimizing harm and facilitating efficient collection and transfer of individual flies.
- Drape insect netting over the plastic tubing and secure the bulb ends of the pipette to the tubing using Parafilm.
2. Single-fly blue-light optogenetic thermotactic positional preference assay
- Drosophila strains and fly rearing
- Cross HC-Gal410 driver line to UAS-GtACR23, resulting in the experimental genotype HC>GtACR2. HC>GtACR2 flies express GtACR2 in heating cells.
NOTE: Ensure an equal number of male and female flies are used for each condition.
- Cross HC-Gal410 driver line to UAS-GtACR23, resulting in the experimental genotype HC>GtACR2. HC>GtACR2 flies express GtACR2 in heating cells.
- Experimental assay setup and execution
- Align two steel plates on separate hot plates so their edges meet. Place a plastic sheet protector on top, securing it with tape to minimize movement. Position a piece of white paper on the sheet protector to reduce background noise signals (Figure 1C).
NOTE: Replace the white paper at the start of each experiment or when soiled. - Place a clear plastic cover (2 mm height × 58 mm width × 83 mm length) on top of the white paper. This cover allows the fly to walk freely while preventing it from flying.
- Cut a hole on the bottom of a polystyrene foam box (27 cm height × 22 cm width × 16 cm length) to adhere to the camera and the blue light (1,000 mA) approximately 12 cm above the experimental surface.
- Position the camera and the blue light to minimize glare while enabling activation.
NOTE: A pretest may be required to determine the activation range of the blue light. - Set the camera to record with the following settings: 1 s timelapse, narrow field, 4000 × 3000 pixels.
- Adjust the hot plate settings to maintain a surface temperature of 25 ± 1 °C and 31 ± 1 °C on the respective steel plates.
NOTE: For conditions requiring temperatures below 25 °C, alternative cooling methods, such as Peltier devices or ice, are necessary. If the steel plates' temperatures exceed the desired levels, spray the surface with distilled water to achieve and maintain the required conditions. Then, use a napkin to absorb excess moisture from the plastic sheet protector. - Before and after each trial, monitor the temperatures using a surface temperature probe.
NOTE: Precise temperature control is crucial, as fluctuations can affect results. - Place the plastic cover on the 25 ˚C side. Using a fly aspirator, gently release a single fly under the cover. Position the box above the experimental area to create dim light (<10 lux) and allow the fly to acclimate for 1 min.
- After the acclimation period, lift the box and quickly adjust the plastic cover, positioning it with the center of the cover aligned with the steel plate boundary to ensure even coverage of the 25 °C and 31 °C sides.
NOTE: During this process, the air temperature and the temperature inside the cover on the 31 °C side increase to about 27 °C within 5 s (Supplementary Figure 1), sufficient to activate heating cells11. - Initiate the trial by turning on the camera and blue light (20 kLux).
NOTE: Do not turn on the blue light under room-light conditions. Turn on the camera and record a few frames before turning on the blue light in case of movement. - After more than 2 min, end the trial by turning off the camera and the light. Dispose of the flies using the aspirator.
NOTE: Record a few extra frames in case of movement when turning off the camera.
- Align two steel plates on separate hot plates so their edges meet. Place a plastic sheet protector on top, securing it with tape to minimize movement. Position a piece of white paper on the sheet protector to reduce background noise signals (Figure 1C).
- Behavioral and statistical analysis
- Only analyze data from flies that stay on the 25 °C side at the start of the trial.
NOTE: Discard data from flies starting trials on the 31 °C side, which often exhibit altered preferences23. Discard trials involving flies that do not approach the edge of the 31 °C side, ensuring they have made a choice between temperatures. Discard data from flies that remain stationary for over 30 s. - Analyze the results using the procedures described previously23. Calculate the preference index (PI) as the ratio of the difference in time that the fly spent in each temperature zone to the total time, as shown in the following formula:
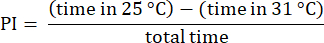
NOTE: Statistical details for the experiments are provided in the figure legends. Data analysis is performed using appropriate data analysis software.
- Only analyze data from flies that stay on the 25 °C side at the start of the trial.
3. Single-fly red-light optogenetic positional preference assay
- Drosophila strains and fly rearing
- Cross the Gr66a-Gal421 driver line to UAS-CsChrimson2, resulting in the experimental genotype Gr66a>CsChrimson. Gr66a>CsChrimson flies express CsChrimson in bitter taste neurons.
NOTE: Ensure an equal number of male and female flies are used for each condition.
- Cross the Gr66a-Gal421 driver line to UAS-CsChrimson2, resulting in the experimental genotype Gr66a>CsChrimson. Gr66a>CsChrimson flies express CsChrimson in bitter taste neurons.
- Experimental assay setup and execution
- Conduct the assay in a dark room or darkened environment to ensure controlled activation of CsChrimson (Figure 1D).
- Position a camcorder approximately 45 cm above the experimental surface.
- Use a camcorder without an infrared filter. If a camcorder has an internal short-pass infrared filter that blocks wavelengths longer than 700 nm, remove it.
- Adhere an 830 nm long-pass filter over the lens using super glue to capture the infrared wavelengths. This adjustment is essential for recording fly movement under infrared light conditions.
- Position a red-light source24 approximately 45 cm above the experimental surface, ensuring a light intensity of about 2.5 kLux (1,000 mA).
NOTE: Appropriate eye protection that can filter out red laser/light is recommended. Laser safety goggles help prevent potential eye damage from direct exposure to intense red light. - Use an infrared light to visualize the flies within the setup. Keep it on throughout the experiment.
NOTE: Avoid glare in the experimental area caused by the infrared light; glare can complicate data analysis. - Use glue to attach a 780 nm infrared long-pass filter to half of the plastic cover.
NOTE: Use any long-pass filter that blocks wavelengths under 780 nm. The unfiltered side permits red light and infrared to pass through, while the filtered side only allows infrared light to pass through. - Place a black matte material underneath the experimental area to minimize background noise.
- Place a single fly under a clear plastic cover (2 mm height × 58 mm width × 83 mm length) using a fly aspirator and allow it to acclimate for 1 min.
- Wait until the fly is visible on the unfiltered side. Start the trial by turning on the camcorder and red light.
NOTE: The camcorder is susceptible to motion immediately after starting. Secure the camera thoroughly while recording and turn on the camera several seconds before recording to avoid aberrant movement that may complicate analysis. Do not turn on the red light until the start of the trial. - Record the motion of the fly for 1 min.
NOTE: Discard trials in which the fly does not approach the midline or does not move for more than 15 s. - After 1 min, end the trial by turning off the camcorder and light. Dispose of the flies using the aspirator.
NOTE: Record a few extra frames to avoid the effects of movement when turning off the camera.
- Behavioral and statistical analysis
- Analyze the results using the procedures described previously23. Calculate the preference index (PI) as the ratio of the difference in the time the fly spent in the area without the filter and the area with the filter to the total time, as shown in the following formula:
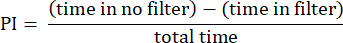
NOTE: Statistical details for the experiments are provided in the figure legends. Data analysis is performed using appropriate data analysis software.
- Analyze the results using the procedures described previously23. Calculate the preference index (PI) as the ratio of the difference in the time the fly spent in the area without the filter and the area with the filter to the total time, as shown in the following formula:
4. Red-light optogenetic proboscis extension response
- Drosophila strains and fly rearing
- Cross the Gr5a-Gal420 driver line to UAS-CsChrimson2, resulting in the experimental genotype referred to as Gr5a>CsChrimson. Gr5a>CsChrimson flies express CsChrimson in sweet taste receptor neurons.
NOTE: Ensure an equal number of male and female flies are used for each condition. - Starve flies for 16-24 h with ATR (ATR +) or without (ATR -) before testing.
NOTE: Longer starvation periods may be required to elicit a strong response.
- Cross the Gr5a-Gal420 driver line to UAS-CsChrimson2, resulting in the experimental genotype referred to as Gr5a>CsChrimson. Gr5a>CsChrimson flies express CsChrimson in sweet taste receptor neurons.
- Experimental assay setup and execution
- Anesthetize flies by cooling on ice (Figure 1E).
NOTE: Avoid the direct contact between flies and ice. Limit the cooling time to about 1 min; prolonged cooling can result in recovery issues and mortality. - Use the tip of a 1000 µL pipette tip to apply 7-10 small dots of glue on the glass slides.
NOTE: Small drops of glue on the slide ensure that the fly does not become submerged and help prevent the glue from sticking to its proboscis or legs. - Position one fly, ventral side up, on each glue dot. Ensure that the thorax and wings contact the glue to minimize movement. Fan the wings out to each side to increase the adhesive surface area.
NOTE: Avoid placing flies too close together, which may inadvertently activate multiple flies during light exposure. - Allow the glue to dry for about 10 min.
- Transfer slides to a humidity box (a plastic box with wet paper towels) and allow the flies to recover for 2 h.
- Place the slide under the microscope. Use a syringe to deliver a droplet of water to satiate the flies, preventing thirst-induced proboscis extensions.
- Manually hold a red laser pointer to shine red light at the proboscis/head of a single fly (700 lux) while observing the PER response through the microscope.
NOTE: Eye protection that can filter out red laser/light is essential when observing the red light through the microscope. - Observe and record proboscis extension within a 30-s window. Score each fly using the following recording system: 0 indicates no extension, 0.5 indicates 1-2 s extension, 1 indicates that extension lasts over 3 s.
- After testing light-induced proboscis extension, examine the response to 4% sucrose using a syringe. Expel a droplet of sucrose at the end of the needle and bring it close to the fly proboscis.
NOTE: Discard the data from flies that fail to respond to the sucrose droplet.
- Anesthetize flies by cooling on ice (Figure 1E).
- Behavioral and statistical analysis
- Perform statistical analysis using appropriate data analysis software.
NOTE: Statistical details for the experiments are provided in the figure legends.
- Perform statistical analysis using appropriate data analysis software.
5. Red-light optogenetic fly maze assay
- Drosophila strains and fly rearing
- Cross the Gr66a-Gal421 driver line to UAS-CsChrimson2, resulting in the experimental genotype Gr66a>CsChrimson. Gr66a>CsChrimson flies express CsChrimson in bitter taste neurons.
NOTE: Ensure each group contains an equal number of male and female flies.
- Cross the Gr66a-Gal421 driver line to UAS-CsChrimson2, resulting in the experimental genotype Gr66a>CsChrimson. Gr66a>CsChrimson flies express CsChrimson in bitter taste neurons.
- Fly-maze assembly
- Make the fly maze as shown in Figure 1F.
NOTE: Alternatively, 3D print the fly maze components, assembling as previously detailed25. - Cut three 5 mL plastic culture tubes to proper lengths to make a loading tube, an uncovered clear testing tube, and a foil-covered testing tube. Attach tubes to the fly maze as shown in Figure 1F.
- Depending on the maze material, cut the testing tubes to ensure equal distance for each testing condition. Based on the length of the holding chamber, shorten the uncovered testing tube in a clear maze, while shortening the foil-covered testing tube if the maze is opaque.
- Make the fly maze as shown in Figure 1F.
- Experimental assay setup and execution
- In dark or low-light conditions, place 10 males and 10 females into the loading tube. Connect the loading tube to the holding chamber. Tilt the elevator and gently tap the tube to move the flies into the holding chamber.
NOTE: Lower the elevator so that only half of the holding chamber is open. This manipulation facilitates the transfer of flies from the loading tube into the holding chamber. - After the flies are transferred to the holding chamber, use the elevator to lower the flies between the loading tube and testing tube holes. Then, remove the loading tube.
- Position the fly maze to approximately 13 cm away from the red-light source (1000 mA) without turning the light on.
- Lower the elevator until the holding chamber aligns with the testing tube holes, allowing the flies to move freely between the foil-wrapped and the uncovered testing tubes. Simultaneously, turn on the red light to activate CsChrimson. Ensure that the red-light intensity on the surface of the uncovered testing tube is about 40 kLux.
NOTE: Do not turn on the red light until the holding chamber is lowered and aligned with the testing tube holes. Appropriate eye protection that can filter out red laser/light is recommended due to the intense brightness of the red light. - Flies choose between the red-light exposed tube and the shaded tube for one minute.
- After 1 min, raise the elevator between the loading tube and testing tube holes. Count the flies in each tube.
NOTE: Only active and uninjured flies are counted. - Clean the fly elevator and fly maze using dH2O after each trial.
- In dark or low-light conditions, place 10 males and 10 females into the loading tube. Connect the loading tube to the holding chamber. Tilt the elevator and gently tap the tube to move the flies into the holding chamber.
- Data analysis and statistics
- Calculate the preference index (PI) as the ratio of the difference in fly numbers between uncovered and covered tubes to the total number of flies, as shown in the following formula:
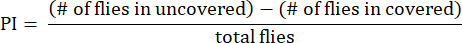
NOTE: Statistical details for the experiments are provided in the figure legends. Data analysis is performed using appropriate data analysis software
- Calculate the preference index (PI) as the ratio of the difference in fly numbers between uncovered and covered tubes to the total number of flies, as shown in the following formula:
Results
Single-fly blue-light optogenetic thermotactic positional preference assay
Four conditions were tested: room light with no ATR supplementation (room light, ATR -), room light with ATR supplementation (room light, ATR +), blue light with no ATR supplementation (blue, ATR -), and blue light with ATR supplementation (blue, ATR +). The first three conditions served as controls. In control experiments, flies avoided the 31 °C side. In blue light with ATR supplementation, flies exhibited no preferen...
Discussion
Optogenetic manipulation has transformed the field of neuroscience by enabling precise control of neural circuits with spatiotemporal accuracy27. A neural circuit includes populations of neurons interconnected by synapses, performing specific functions upon activation. The Drosophila whole-brain connectome has been completed, offering comprehensive insights into the synaptic pathways within the Drosophila brain28,29. Whil...
Disclosures
The authors declare that there are no conflicts of interest regarding the publication of this article. All authors have disclosed any potential conflicts and affirm that they have no financial or personal relationships that could influence the work presented in this study.
Acknowledgements
Schematic diagrams for all figures were created with Biorender.com. This work was supported by NIH R01GM140130 (https://www.nigms.nih.gov/) to L.N. The funders had no role in the study design, data collection and analysis, publication decision, or manuscript preparation.
Materials
| Name | Company | Catalog Number | Comments |
| 1000 mA LED driver | Luxeon Star | 3021-D-E-1000 | |
| 5 mL VWR Culture Tubes, Plastic, with Dual-Position Caps | VWR | 60818-664 | |
| 780 Longpass Filter / IR 780 nm 100 mm x 100 mm | Lee Filters | BH #LE8744 | Cut to approximately 47 x 100 fit the plastic cover |
| Agfabric 6.5 ft. x 15 ft. Insect Bug Netting Garden Net for Protecting Plants Vegetables Flowers Fruits | The Home Depot | EIBNW6515 | |
| All trans retinal | Sigma-Aldrich | 116-31-4 | |
| Aluminum Plate (30.5 cm x 30.5 cm x 0.6 cm) | Amazon | purchased from Amazon | |
| Black Plastic Box | LI-COR | 929-97101 | |
| CALCIUM CHLORIDE ANHYDRO 25GR | Thermo Fisher Scientific | AC297150250 | |
| CX405 Handycam with Exmor R CMOS sensor | SONY | HDR-CX405 | |
| Elmer’s “School Glue” | Elmer | ||
| Ethyl alcohol, Pure (200 Proof) | Sigma-Aldrich | E7023 | |
| Fisherbrand Isotemp Hot Plate Stirrer | Fisher Scientific | SP88850200 | |
| Fly line: Gr5a-Gal4 | Bloomington Drosophila Stock Center | 57592 | |
| Fly line: Gr66a-Gal4 | Bloomington Drosophila Stock Center | 57670 | |
| Fly line: HC-Gal4 (II) | Dr. Marco Gallio Lab | A kind gift | |
| Fly line: UAS-CsChrimson | Bloomington Drosophila Stock Center | 55136 | |
| Fly line: UAS-GtACR2/TM6B | Dr. Quentin Gaudry Lab | A kind gift | |
| Flystuff 62-101 Yellow Cornmeal (11.3 Kg), Yellow, 11.3 Kg/Unit | Genesee Scientific | 62-101 | |
| Flystuff 62-107 Inactive Dry Yeast, 10 Kg, Nutritional Flake, 10 Kg/Unit | Genesee Scientific | 62-107 | |
| Flystuff 66-103 Nutri-Fly Drosophila Agar, Gelidium, 100 Mesh, 5 Kg (11.02 lbs)/Unit | Genesee Scientific | 66-103 | |
| FreeMascot OD 8+ 190 nm–420 nm / 600 nm–1100 nm Wavelength Violet/Red/Infrared Laser Safety Glasses | FreeMascot | B08LGMQ65S | purchased from Amazon |
| GoPro Hero8 Black | GoPro | 6365359 | |
| LEE Filters 100×100 mm Infra Red #87 Infrared Polyester Filter | B&H Photo | LE8744 | |
| Longpass Filter, Colored Glass, 50.8 x 50.8 mm, 830 nm Cut-on, RG830 | Newport | FSQ-RG830 | |
| Methyl 4-hydroxybenzoate, 99%, Thermo Scientific Chemicals | Thermo Fisher Scientific | 126960025 | |
| MicroWell Mini Tray 60 Well, Low Profile NS PS | Thermal Scientific | NUNC 439225 | The lids are used as the "plastic cover" |
| Olympus Plastics 24-160RS, 1000 µL Olympus Ergonomic Pipet Tips Low Binding, Racked, Sterile, 8 Racks of 96 Tips/Unit | Eppendorf | 24-160RS | |
| Parafilm M Sealing Film | Heathrow Scientific | HS234526B | 4 in x 125 feet |
| Potassium chloride, ACS, 99.0-100.5%, | Thermo Fisher Scientific | AA1159530 | |
| Prism | GraphPad | Version 9 | data analysis software |
| Samco Graduated Transfer Pipettes | Thermo Fisher Scientific | 225 | 3 mL |
| Slides | Fisher Scientific | 12-544-2 | 5 mm x 75 mm x 1.0 mm |
| Stereo microscope | OLYMPUS | CZ61 | |
| Styrofoam box (27 cm height × 22 cm width × 16 cm length) | |||
| Sucrose | Fisher Scientific | 225911 | |
| Surface temperature probe | Fluke | 80PK-3A | |
| Syringe | BD Integra | 305270 | |
| Tate & Lyle 457 Dextrose, Tate & Lyle, Pow, Tate & Lyle 457 Dextrose, Tate & Lyle, Powder, 50 lbs/Unit | Genesee Scientific | 62-113 | |
| Traceable Calibrated Big-Digit Thermocouple Thermometer | Traceable by cple-parmer | UX-91210-07 | Fisherbrand Traceable BigDigit Type K Thermometer |
| Triple blue LED starboard | LEDSupply | 07007-PB000-D | 470 nm |
| Triple red LED starboard | LEDSupply | 07007-PD000-F | 627 nm |
| Tygon PVC Clear Tubing 1/4" ID, 3/8" OD, 5 ft. Length | McMaster Carr Supply Company | 6516T21 | |
| Univivi IR Illuminator, 850nm 12 LEDs Wide Angle IR Illuminator for Night Vision | Univivi | 4331910725 | |
| Wakefield Thermal 25.4 mm Round Heatsink Star LED Board - 882-100AB | Wakefield-Vette | 882-100AB | |
| Wireless Presenter | DinoFire Store | B01410YNAM | purchased from Amazon |
References
- Yizhar, O., Fenno, L. E., Davidson, T. J., Mogri, M., Deisseroth, K. Optogenetics in neural systems. Neuron. 71 (1), 9-34 (2011).
- Klapoetke, N. C., et al. Independent optical excitation of distinct neural populations. Nat Methods. 11 (3), 338-346 (2014).
- Mauss, A. S., Busch, C., Borst, A. Optogenetic neuronal silencing in drosophila during visual processing. Sci Rep. 7 (1), 13823 (2017).
- Govorunova, E. G., Sineshchekov, O. A., Janz, R., Liu, X., Spudich, J. L. Neuroscience. Natural light-gated anion channels: A family of microbial rhodopsins for advanced optogenetics. Science. 349 (6248), 647-650 (2015).
- Castaneda, A. N., et al. Functional labeling of individualized postsynaptic neurons using optogenetics and trans-tango in drosophila (flipsot). PLoS Genet. 20 (3), e1011190 (2024).
- Fischer, F. P., et al. Drosophila melanogaster as a versatile model organism to study genetic epilepsies: An overview. Front Mol Neurosci. 16, 1116000 (2023).
- Aragon, M. J., et al. Multiphoton imaging of neural structure and activity in drosophila through the intact cuticle. Elife. 11, e69094 (2022).
- Zhang, W., Ge, W., Wang, Z. A toolbox for light control of drosophila behaviors through channelrhodopsin 2-mediated photoactivation of targeted neurons. Eur J Neurosci. 26 (9), 2405-2416 (2007).
- Kono, M., Goletz, P. W., Crouch, R. K. 11-cis- and all-trans-retinols can activate rod opsin: Rational design of the visual cycle. Biochemistry. 47 (28), 7567-7571 (2008).
- Gallio, M., Ofstad, T. A., Macpherson, L. J., Wang, J. W., Zuker, C. S. The coding of temperature in the drosophila brain. Cell. 144 (4), 614-624 (2011).
- Ni, L., et al. A gustatory receptor paralogue controls rapid warmth avoidance in drosophila. Nature. 500 (7464), 580-584 (2013).
- Weiss, L. A., Dahanukar, A., Kwon, J. Y., Banerjee, D., Carlson, J. R. The molecular and cellular basis of bitter taste in drosophila. Neuron. 69 (2), 258-272 (2011).
- Ling, F., Dahanukar, A., Weiss, L. A., Kwon, J. Y., Carlson, J. R. The molecular and cellular basis of taste coding in the legs of drosophila. J Neurosci. 34 (21), 7148-7164 (2014).
- Dethier, V. G. . The Hungry Fly: A Physiological Study of the Behavior Associated with Feeding. , (1976).
- Shiu, P. K., Sterne, G. R., Engert, S., Dickson, B. J., Scott, K. Taste quality and hunger interactions in a feeding sensorimotor circuit. Elife. 11, e79887 (2022).
- Kim, H., Kirkhart, C., Scott, K. Long-range projection neurons in the taste circuit of drosophila. Elife. 6, e23386 (2017).
- Sayeed, O., Benzer, S. Behavioral genetics of thermosensation and hygrosensation in drosophila. Proc Natl Acad Sci U S A. 93 (12), 6079-6084 (1996).
- Coya, R., Martin, F., Calvin-Cejudo, L., Gomez-Diaz, C., Alcorta, E. Validation of an optogenetic approach to the study of olfactory behavior in the t-maze of drosophila melanogaster adults. Insects. 13 (8), 662 (2022).
- Chu, L. A., Tai, C. Y., Chiang, A. S. Thirst-driven hygrosensory suppression promotes water seeking in drosophila. Proc Natl Acad Sci U S A. 121 (34), e2404454121 (2024).
- Chyb, S., Dahanukar, A., Wickens, A., Carlson, J. R. Drosophila gr5a encodes a taste receptor tuned to trehalose. Proc Natl Acad Sci U S A. 100 (Suppl 2), 14526-14530 (2003).
- Dunipace, L., Meister, S., Mcnealy, C., Amrein, H. Spatially restricted expression of candidate taste receptors in the drosophila gustatory system. Curr Biol. 11 (11), 822-835 (2001).
- Ratnayake, K., et al. Blue light excited retinal intercepts cellular signaling. Sci Rep. 8 (1), 10207 (2018).
- Huda, A., Omelchenko, A. A., Vaden, T. J., Castaneda, A. N., Ni, L. Responses of different drosophila species to temperature changes. J Exp Biol. 225 (11), jeb243708 (2022).
- Tyrrell, J. J., Wilbourne, J. T., Omelchenko, A. A., Yoon, J., Ni, L. Ionotropic receptor-dependent cool cells control the transition of temperature preference in drosophila larvae. PLoS Genet. 17 (4), e1009499 (2021).
- Fu, Z., Huda, A., Kimbrough, I. F., Ni, L. Using drosophila two-choice assay to study optogenetics in hands-on neurobiology laboratory activities. J Undergrad Neurosci Educ. 22 (1), A45-A50 (2023).
- Joseph, R. M., Heberlein, U. Tissue-specific activation of a single gustatory receptor produces opposing behavioral responses in drosophila. Genetics. 192 (2), 521-532 (2012).
- Riemensperger, T., Kittel, R. J., Fiala, A. Optogenetics in drosophila neuroscience. Methods Mol Biol. 1408, 167-175 (2016).
- Dorkenwald, S., et al. Neuronal wiring diagram of an adult brain. Nature. 634 (8032), 124-138 (2024).
- Schlegel, P., et al. Whole-brain annotation and multi-connectome cell typing of drosophila. Nature. 634 (8032), 139-152 (2024).
- Talay, M., et al. Transsynaptic mapping of second-order taste neurons in flies by trans-tango. Neuron. 96 (4), 783-795.e4 (2017).
- Shiraiwa, T., Carlson, J. R. Proboscis extension response (per) assay in drosophila. J Vis Exp. (3), e193 (2007).
- Pulver, S. R., Hornstein, N. J., Land, B. L., Johnson, B. R. Optogenetics in the teaching laboratory: Using channelrhodopsin-2 to study the neural basis of behavior and synaptic physiology in drosophila. Adv Physiol Educ. 35 (1), 82-91 (2011).
- Titlow, J. S., Johnson, B. R., Pulver, S. R. Light activated escape circuits: A behavior and neurophysiology lab module using drosophila optogenetics. J Undergrad Neurosci Educ. 13 (3), A166-A173 (2015).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved