Method Article
In Situ Labeling of Mitochondrial DNA Replication in Drosophila Adult Ovaries by EdU Staining
In This Article
Summary
Drosophila oogenesis continues to be exceptionally useful in the study of mitochondrial proliferation and inheritance. This manuscript describes a detailed protocol used to label the replicating mitochondrial DNA (mtDNA) in Drosophila adult ovaries with 5-ethynyl-2´-deoxyuridine (EdU), which facilitates uncovering mechanisms associated with mitochondrial inheritance that were previously debatable.
Abstract
The mitochondrial genome is inherited exclusively through the maternal line. Understanding of how the mitochondrion and its genome are proliferated and transmitted from one generation to the next through the female oocyte is of fundamental importance. Because of the genetic tractability, and the elegant, ordered simplicity by which oocyte development proceeds, Drosophila oogenesis has become an invaluable system for mitochondrial study. An EdU (5-ethynyl-2´-deoxyuridine) labeling method was utilized to detect mitochondrial DNA (mtDNA) replication in Drosophila ovaries. This method is superior to the BrdU (5-bromo-2'-deoxyuridine) labeling method in that it allows for good structural preservation and efficient fluorescent dye penetration of whole-mount tissues.
Here we describe a detailed protocol for labeling replicating mitochondrial DNA in Drosophila adult ovaries with EdU. Some technical solutions are offered to improve the viability of the ovaries, maintain their health during preparation, and ensure high-quality imaging. Visualization of newly synthesized mtDNA in the ovaries not only reveals the striking temporal and spatial pattern of mtDNA replication through oogenesis, but also allows for simple quantification of mtDNA replication under various genetic and pharmacological perturbations.
Introduction
Besides the nuclear genome, each eukaryotic cell also contains thousands of copies of small circular DNA in the mitochondrial matrix. While mitochondrial DNA (mtDNA) encodes essential subunits of electron transport chain, the majority of the mitochondrial proteome, including all factors for replication and transcription of mtDNA are encoded by the nuclear genome. In contrast to the nuclear genome that follows Mendelian laws of inheritance, the animal mitochondrial genome is inherited exclusively through the maternal lineage. Therefore, understanding how mtDNA is proliferated and transmitted from one generation to the next through the female oocyte is fundamentally important. However, there is continuing debate over how mtDNA replication is regulated in the female germline. Additionally, the widely accepted mtDNA bottleneck theory for mtDNA transmission suggests that the population of mtDNA in primordial germ cells is subsampled to a relatively small number during development 12. It also implies that mtDNA replication is temporally and spatially regulated during oogenesis. Hence, the in situ detection of mtDNA replication during germline development will facilitate the understanding of the mechanism of mtDNA inheritance.
Drosophila oogenesis provides a genetically tractable system to study mtDNA replication and transmission. In each of the two Drosophila ovaries, there are 16-20 independent strings of egg chambers called ovarioles 3, which are the functional units of egg production (see Figure 1A). Each ovariole contains a progressive linear organization of oogenesis, where the anterior tip is composed of a structure called germarium. The germarium is further divided into four regions that contain germ cells at distinct developmental stages. In region 1, germline stem cells go through asymmetric division to produce daughter cells known as cystoblasts. Region 2a contains the cystoblasts that have completed their final division. Cystoblasts undergo four rounds of division producing 16-cell groups. The 16 cells remain connected to each other by cytoplasmic bridges called ring canals. Only one of the cells commits to differentiation as the oocyte, while the other 15 develop as polyploid nurse cells. An essential cytoplasmic structure, known as fusome, is proposed to facilitate with the formation of ring canals, determine the cyst polarity and the nurse cell-oocyte interactions 4,5. As the cysts move towards region 2b, the cyst structures span the entire width of the germarium, and become more associated with follicle cells. The germarium structures end with region 3 containing the first budding egg chamber. Subsequently, the egg chambers are assembled at the posterior end of the germarium, which progress through the ovariole in 14 morphologically distinct stages. The growth of oocyte depends on the nurse cells, which transport proteins, mRNA and endomembrane structures (e.g., Golgi) via the ring canals into the oocyte. During Drosophila oogenesis, it was found that a fraction of the mitochondria within each 16-cell cyst were associated with the fusome, moved through the ring canals and were delivered into the single oocyte in a large mass called Balbiani body 6. This phenomenon was proposed to contribute to the quality control of mitochondrial inheritance through female ovaries.
Detection of mtDNA synthesis relies on the incorporation of labeled DNA precursors into cellular DNA. Traditionally, a nucleoside analog of thymidine 5-bromo-2'-deoxyuridine (BrdU) was used to label the mtDNA replication in tissue culture cells 7,8. However, the antibody-based BrdU labeling exhibits several limitations, especially for whole-mount tissue staining. One major disadvantage of BrdU labeling is that it requires DNA denaturation to expose the BrdU epitope, so that it can be detected by the anti-BrdU antibody. The specimen has to be treated under harsh denaturing conditions such as chemicals (e.g., hydrochloric acid or mixtures of methanol and acetic acid), heat, or digestion with DNase, which could disrupt the structure of the specimen and complicate the following staining procedure 9,10.
Here, an alternative thymidine analog 5-ethynyl-2´-deoxyuridine (EdU) is used to label the replicating mitochondrial DNA in Drosophila adult ovaries. This method is faster and highly sensitive. EdU is readily incorporated into cellular DNA during DNA replication. The following detection is based on a "click" reaction, a Cu (I) -catalyzed covalent reaction between the terminal alkyne group and a fluorescent azide 10. Because the reaction does not require the denaturation of the specimen, it allows good structural preservation. Furthermore, the size of dye azide is only 1/500 of that of an antibody molecule 10, which enables fast and efficient penetration of whole-mount tissues. We have used this method to detect mtDNA replication during Drosophila oogenesis and found a striking spatial pattern in the germarium region of Drosophila ovary 11, which lead us to propose a replication-dependent mtDNA selective inheritance mechanism. We present here a detailed protocol on EdU labeling of mtDNA replication in Drosophila ovaries. To demonstrate the application of the protocol, we also tested the mtDNA replication in a mtDNA mutant Drosophila (mt:CoIT300I) 11, as well as with diverse treatment of mitochondrial uncouplers, which dissipate the mitochondrial membrane potential and potentially disrupt mtDNA replication.
Protocol
1. Tissue Collection and Dissection
- In each vial containing dry yeast, culture 10 adult female flies with 10 males for 2-3 days.
Note: Maintaining well-fed female flies will improve the overall quality and yield of ovary production and facilitate dissection. - Anesthetize the female flies on a carbon dioxide (CO2) fly pad.
- Under the stereoscope, place several drops of RT medium (Schneider's Drosophila medium supplemented with 10% fetal bovine serum (FBS)) on a dissecting pad. Conduct all the dissection procedures in the medium to keep the tissues alive and healthy.
- Grab a fattened female fly with sharp fine-nosed forceps at its lower thorax. Use another set of forceps to tug gently at the extreme posterior of the fly until the tissues in the abdomen are exposed. Detach the two ovaries from other tissues (e.g. guts).
Note: Each ovary displays an opaque structure that is composed of 16-20 attached ovarioles. - To enhance the penetration of reagents, open the ovaries by pulling them apart and passing the tips of the forceps in between each ovariole a couple of times. Keep the ovarioles connected within an ovary to minimize loss of tissues during the following procedures.
- Transfer the ovaries immediately to an 1.5 ml microcentrifuge tube containing 500 µl of Schneider's Drosophila medium with 10% FBS.
- Repeat the dissection and collect 10-15 ovaries in each microcentrifuge tube.
2. EdU Labeling
- Aspirate the medium in each tube, and replace with 500 µl of Schneider's Drosophila medium with 10% FBS containing 7 µM aphidicolin.
Note: Aphidicolin is used to block nuclear DNA synthesis by inhibiting DNA polymerase α without affecting mtDNA replication 7,12. It is possible to increase the aphidicolin concentration to up to 70 µM to achieve better inhibition. The stock solution of 3-30 mM aphidicolin in DMSO can be stored in the dark at -20 °C for up to 6 weeks. - To keep the ovaries healthy, ensure that they are immersed under solutions while changing the medium. Use Schneider's Drosophila medium with 10% FBS through step 2.6.
- Incubate ovaries for 3 hr at RT on a bench-top rocker with gentle rotation.
- For drug treatment, e.g., mitochondrial uncoupler carbonyl cyanide 4-trifluoromethoxy phenylhydrazone (FCCP), add the appropriate concentration of drug (e.g., 10 µM FCCP) into the medium after 2 hr of aphidicolin treatment. Continue incubating for another 1 hr.
- Remove the medium containing aphidicolin with or without drug. Briefly rinse the ovaries with medium (aphidicolin is not necessary) twice.
- Add 1 ml medium containing 10 µM EdU and 7 µM aphidicolin and continue incubating at RT for 2 hr. Store the 10 mM EdU stock solution (in DMSO) at -20 °C.
- Remove the medium containing EdU and aphidicolin. Wash with medium (without aphidicolin) twice for 3 min each.
3. Tissue Fixation and Permeabilization
- Prepare a 4% paraformaldehyde solution in phosphate buffered saline (PBS, pH 7.4).
Note: We used commercial available paraformaldehyde stored in pre-scored ampoules. Open a new ampoule and dilute to 4% before use.
CAUTION: Formaldehyde is toxic; it should be handled in a fume hood with skin and eye protection. - Fix ovaries with 4% paraformaldehyde for 20 min at RT with gentle rotation.
- Remove the fixative and wash ovaries twice in 1 ml of 3% BSA in PBS for 5 min each time.
- To permeabilize the tissues, remove the wash solution and add 1 ml of 0.5% Triton X-100 in PBS. Incubate at RT for 20 min.
- Remove the permeabilization solution and wash twice in 1 ml of 3% BSA in PBS.
4. EdU Detection
Note: EdU detection is based on "click" chemistry, a Cu(I)-catalyzed [3+2] cycloaddition 13, which adds fluorescent azides to the terminal alkyne group of EdU, and the fluorescent molecule is subjected to subsequent detection. Since Cu(I), which is easily oxidized to the non-catalytic Cu(II) species, is required to catalyze the reaction, it is recommended to reduce the Cu(II) sulfate in situ to obtain Cu(I). That is, Cu(II) sulfate is used in the presence of a reductant such as ascorbic acid (herein the "buffer additive") to generate copper (I).
- Before the experiment, prepare the working solution of the Alexa Fluor azide (Alexa Fluor 488 azide, Alexa Fluor 555 azide or others, depending on the preferred fluorophore, which are named as "dye azide" hereafter), EdU reaction buffer and EdU buffer additive according to manufacturer's instructions.
- Reconstitute the dye azide in 70 µl DMSO. Store the working solution at -20 °C for up to 1 year.
- Prepare fresh EdU reaction buffer by diluting the 10x EdU reaction buffer with deionized water.
Note: After use, store any remaining 1x solution at 4 °C. The 1x solution is stable for up to 6 months. - Make a 10x stock solution of the EdU buffer additive by fully dissolving the powder in 2 ml deionized water.
Note: This stock solution is stable for up to 1 year at -20 °C. If the solution develops a brown color, it has degraded and should be discarded.
- Prepare the EdU reaction cocktail fresh each time, and use within 15 min of preparation. To achieve reproducible results, make sure the reaction components are maintained at the same ratios.
- Make fresh 1x EdU buffer additive solution by diluting the 10x stock solution (prepared in step 4.1.3) 1:10 in deionized water. Use this solution on the same day.
- Prepare 1 ml of EdU reaction cocktail by combining the following ingredients in order: 860 µl 1x EdU reaction buffer, 40 µl CuSO4, 2.5 µl dye azide, 100 µl 1x EdU buffer additive.
Note: It is important that the ingredients are added in order to ensure optimal performance.
- Remove the PBS with 3% BSA and add 0.5 ml of the EdU reaction cocktail to each tube. Incubate at RT for 30 min with gentle rotation. Keep the samples protected from light.
- Remove the EdU reaction cocktail and wash once with 1 ml of 3% BSA in PBS.
- Wash the samples once with 1 ml of PBS.
5. Antibody Labeling
Note: Perform antibody labeling after EdU staining. If no additional staining is desired, one can proceed to mounting and imaging directly. It is important that the samples be protected from light in all the following procedures.
- Remove the wash solution. Block the ovaries in 1 ml of blocking solution containing 0.2% BSA and 0.1% Triton X-100 in PBS for 30 min.
- Remove the blocking solution and replace with primary antibody diluted in blocking solution (e.g., mouse ATP synthase subunit α antibody, 1:1,000 dilution). Incubate in the dark at 4 °C O/N.
- Wash the samples with 1 ml of blocking solution 3 times, 10 min each time. To minimize the background, wash two more times for 30 min each. Remove the blocking solution.
- Incubate with secondary antibody diluted in blocking solution (e.g., goat anti-mouse Alexa Fluor 568 secondary antibody, 1:200 dilution) for 2 hr at RT.
Note: Use a different color than the one coupled to EdU. - Repeat the wash steps performed in step 5.3.
- Rinse with 1 ml of PBS once to remove the detergent.
6. Mounting and Imaging
- Carefully remove all the PBS and immediately cover ovaries with 50 µl mounting medium (with or without DAPI).
Note: Use a mounting medium that does not harden while the ovarioles are being separated from each other. Ovaries can be stored in mounting medium at 4 °C for up to a week. - Cut the end of a pipette tip and use it to transfer ovaries carefully onto a microscope slide.
- Completely separate each ovariole with sharp fine-nosed forceps under the stereomicroscope. Remove the connecting tissues at the posterior end and stage 14 or mature egg chambers. The young egg chambers are at the anterior end and transparent.
- Align the egg chambers with forceps so that they do not overlap with each other.
- Slowly lower a # 1.5 glass coverslip on top of the samples. Allow the mounting medium to polymerize for several hours at RT. Seal with transparent nail polish.
- Image slides the next day, or store at 4 °C prior to imaging.
- Visualize under a confocal microscope with a 63X oil-immersion objective. Capture three-dimensional z-stacks images.
Note: There is strong EdU signal in later-stage egg chambers and background fluorescence in the epithelial sheath. When examining EdU labeling in a particular germarium, one should avoid germariums that are overlapped by or close to other tissues or later-stage egg chambers.
Results
The above protocol allows visualization of the punctate structures associated with mitochondria (Figure 1B-C), which indicate mitochondrial DNA replication during Drosophila oogenesis. The EdU puncta localized with mitochondria marked by staining for ATP synthase alpha subunit (Figure 2). The observed signals were absent in ovaries treated with ethidium bromide 11, an inhibitor for mtDNA replication 14, validating that these puncta indeed label replicating mtDNA.Aphidicolin was used to inhibit nuclear DNA staining without affecting mtDNA replication (Figure 2). Without aphidicolin treatment, intense EdU signals label the nuclei, and mtDNA puncta were barely detected (Figure 3). However, in the presence of aphidicolin, nuclear incorporation was dramatically reduced and many puncta associated with mitochondria were observed.
There is a high level of mtDNA replication in post-germarium egg chambers (Figure 1B). However, notably, mtDNA replication displayed a spatial pattern in the germarium. As indicated by the number of EdU puncta, there is a moderate level of mtDNA replication in region 1 of the germarium, but almost no EdU incorporation in region 2A (Figure 1C). As the cyst moves down to region 2B in the germarium, mtDNA replication resumed and the number of EdU puncta in the posterior cyst of region 2B was much higher than that in region 2A (Figure 1C). Specifically, intensive EdU incorporation was concentrated around the ring canals and the fusome structures, as stained by the hu li tai shao (Hts) protein. mtDNA kept replicating at a high level in region 3 of the germarium (Figure 1C)
To demonstrate the association of specific genes or treatment with mtDNA proliferation, Drosophila ovaries could be subjected to gene manipulation or drug treatment. We treated ovaries with different concentrations of FCCP, a classic mitochondrial protonophore, which dissipates mitochondrial membrane potential. As a control, DMSO had no effect on mtDNA replication (Figure 4A). High doses of FCCP (10 µM) almost depleted mtDNA replication entirely throughout the germarium (Figure 4D). Nonetheless, lower concentration of FCCP (2 or 5 µM) had a minor impact on mtDNA replication in region 1 but inhibited replication in regions 2B and 3 (Figure 4B-C), suggesting regions 2B and 3 are more sensitive to mitochondrial disruption, or they maintain relative slower replication kinetics. The above results indicated that mtDNA replication is associated with mitochondrial activity. Particularly, different regions of the germarium responded differently to mitochondrial impairment.
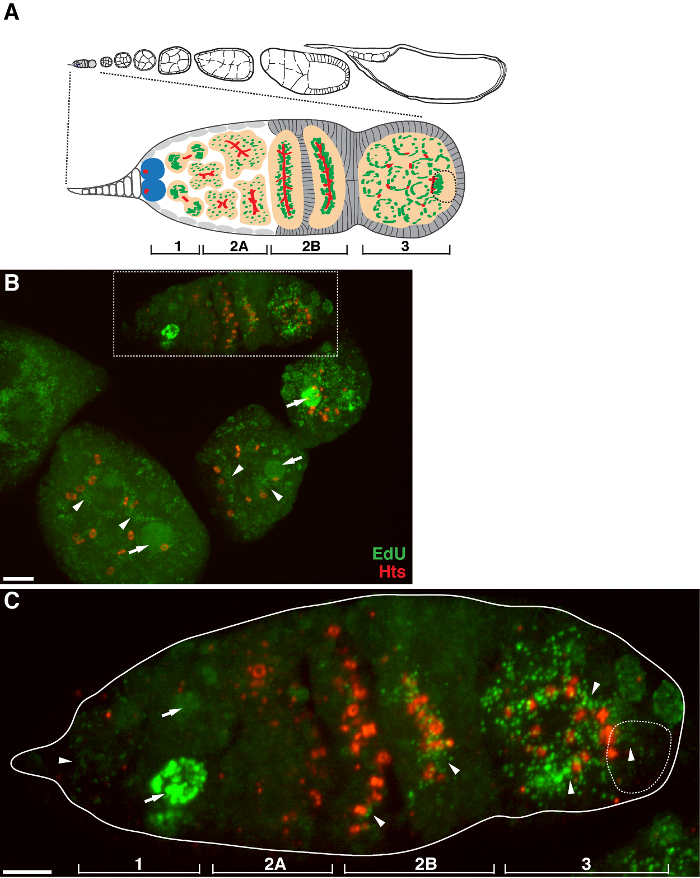
Figure 1. mtDNA replication during Drosophila oogenesis. (A) Diagram of a Drosophila ovariole and a magnified view of the germarium. The ovariole illustrates from left to right, anterior to posterior, successive developmental stages of egg chambers. In the germarium, the fusome (red), germline stem cells (blue), mitochondria (green), future oocyte (broken line, recognized by positioning, fusome structure and mitochondrial clusters), developing cysts (peach) and four developmental regions are shown. (B) Representative z-stack projection of a wild-type ovariole labeled by EdU and antibody against Hts-RC, a marker for ring canals and fusome. In the presence of aphidicolin, the EdU was incorporated into mtDNA (arrowheads) and nuclei (arrows). Scale bar, 10 µm. (C) Magnified view of germaria outlined in the boxed region in B. The four developmental regions are indicated. Scale bar, 5 µm 11 . Please click here to view a larger version of this figure.
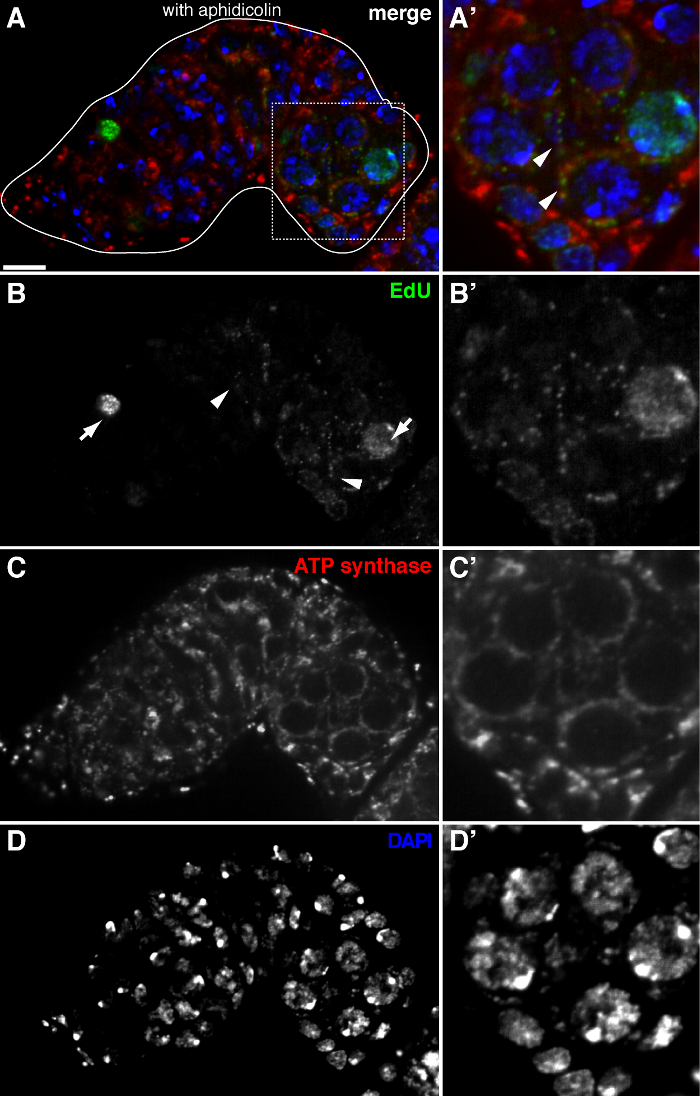
Figure 2. mtDNA replication in the Drosophila germarium visualized by EdU incorporation with aphidicolin treatment. (A)-(D) Representative confocal section of a wild type germarium showing EdU incorporation (green, b), mitochondria, marked by ATP Synthase alpha subunit staining (red, C), and nuclei, labeled with DAPI staining (blue, D) with pre-incubation with DNA polymerase-α inhibitor aphidicolin. (A'-D') A magnified image of the boxed area in (A) showing EdU incorporation (B'), mitochondria (C') and nuclei (D'). In the presence of aphidicolin, nuclear incorporation (arrows) is reduced and many puncta were localized within mitochondria (arrowheads). Scale bars, 10 µm. The figure has been modified from 11. Please click here to view a larger version of this figure.
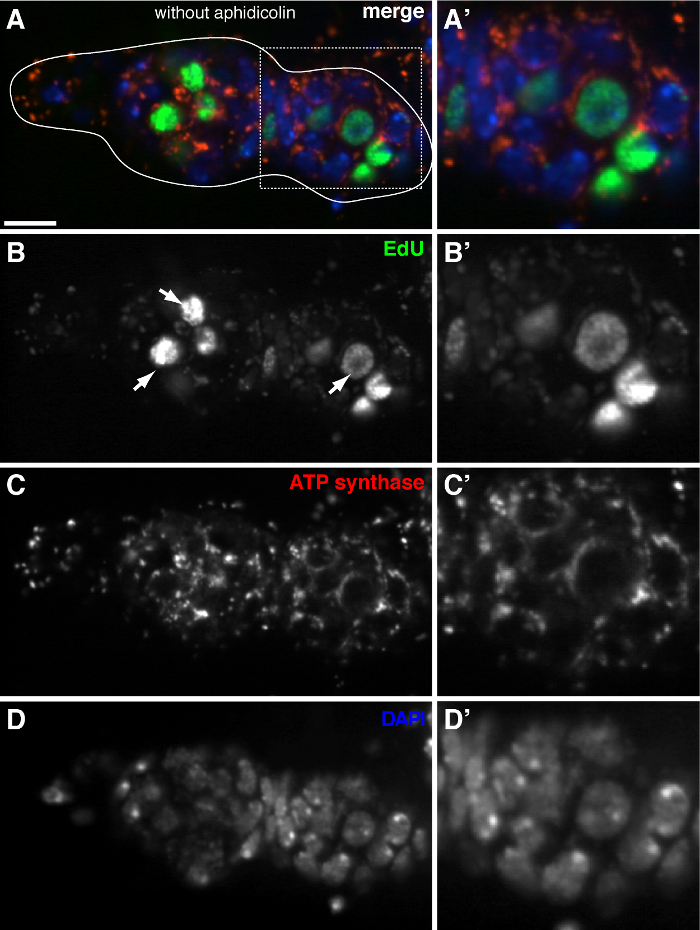
Figure 3. mtDNA replication is barely detected in the Drosophila germarium without aphidicolin treatment. (A)-(D) Representative confocal section of a wild type germarium showing EdU incorporation (green), mitochondria, marked by ATP Synthase alpha subunit staining (red), and nuclei, labeled with DAPI staining (blue) in the absence of the DNA polymerase-α inhibitor aphidicolin. (A'-D') A magnified image of the boxed area in (A) showing EdU incorporation (B'), mitochondria (C') and nuclei (D'). Without aphidicolin, intense EdU signals label nuclei (arrows), and mtDNA puncta were barely detected. Scale bars, 10 µm. The figure has been modified from 11. Please click here to view a larger version of this figure.
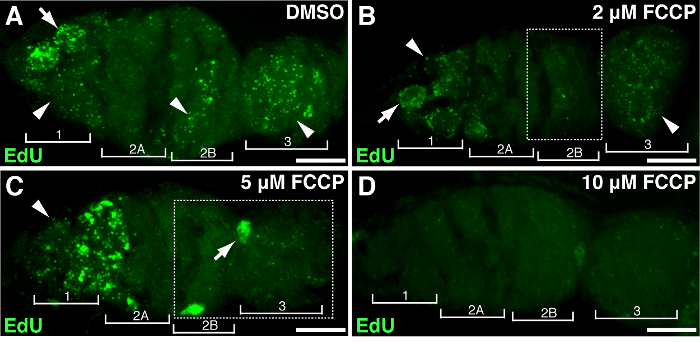
Figure 4. Mitochondrial uncoupler impairs the mtDNA replication. Representative z-stack projects showing wild-type germarium treated with DMSO (A) or the mitochondrial uncoupler FCCP at concentrations of 2 µM (B), 5 µM (C), 10 µM (D) during EdU incorporation. Note the impaired EdU labeling in region 2B treated with 2 µM FCCP (B), and in both region 2B and 3 treated with 5 µM FCCP (outlined in boxes). Four developmental regions are shown. Arrows, nuclear DNA; arrowheads, mtDNA. Scale bar, 10 µm. The figure has been modified from 11. Please click here to view a larger version of this figure.
Discussion
EdU labeling is a novel and efficient method to detect DNA synthesis in proliferating cells, which is based on the incorporation and staining of nucleoside analogues in newly synthesized DNA. This method is superior to the BrdU labeling method in that it is faster and highly sensitive. More importantly, it allows for good structural preservation and efficient EdU-dye penetration of whole-mount tissues 9, 10. Historically, as a superior alternative to BrdU labeling, EdU labeling was used for studying nuclear DNA replication during the S-phase of the cell cycle. Aphidicolin is an inhibitor of DNA polymerase α, which is the main polymerase for nuclear DNA replication at S-phase 1215. mtDNA replication is carried out by DNA polymerase γ, which is insensitive to aphidicolin treatment. Hence, treatment with aphidicolin prior to and during the EdU incubation significantly inhibited EdU incorporation into nuclear DNA. It should be noted that aphidicolin may be stable for several weeks if appropriately stored, and the efficacy of aphidicolin in inhibiting nuclear DNA replication was variable in our hands. The ovarioles or egg chambers with strong nuclear DNA labeling should be excluded from further data analyses.
mtDNA replication can be readily visualized as puncta in the cytoplasm, which also affords a straightforward way to quantify the level of mtDNA replication by counting the number of EdU puncta, normalized to the total volume of cytoplasm. Imaging software can be applied to automatically identify EdU puncta in microscopic images, which is especially useful for computational analyses of large data sets. However, precautions should be taken, because the incomplete inhibition of nuclear DNA replication can lead to the EdU incorporation in distinct loci on the chromosome and display as puncta inside the nucleus. Also in other cases, the intensity of EdU incorporation into replicating mtDNA might be weak, while the background and noise could be high. Therefore, the individual parameters for automatic image analyses should be carefully defined. It is also recommended that images should be examined with trained eyes to make sure that proper EdU puncta are identified.
Visualization of newly synthesized mtDNA during Drosophila oogenesis provides an opportunity to investigate how mtDNA replication is regulated under physiological or pathological conditions, by carrying out the experiment in flies subjected to a variety of pharmacological or genetic perturbations. In a previous study, EdU incorporation assay was performed in a mtDNA mutant Drosophila mt:CoIT300I11. Furthermore, to disrupt the mitochondrial membrane potential, ovaries were treated with various concentrations of mitochondrial uncoupler FCCP or 2,4-dinitrophenol (DNP) prior to EdU incorporation. Depending on the experimental purposes and drug characteristics, different methods might be adopted for effective delivery. For adult flies, drugs can be presented as a vapor (e.g., ethanol and cocaine) 16,17 or drugs can be injected into the abdomen, where it quickly diffuses throughout the body 18. The most common practice is that drugs are added to the fly food or a sucrose/drug-saturated filter paper. For example, the inhibitor of microtubule assembly, colchicine, was fed to flies for 2-3 days before ovary dissection 19. Hence, it is important to evaluate the drug delivery method and choose appropriate concentrations.
To ensure the successful imaging of mtDNA replication in Drosophila ovaries, several critical steps must be carefully executed. Foremost, preserving the viability and health of the ovaries during dissection and EdU incorporation is essential (steps 1 and 2). The Schneider's Drosophila medium with FBS needs to be warmed to RT before use. One should minimize direct contact between the egg chambers and dissecting tools or pipette tips. The ovaries should be immersed under solutions all times to avoid dehydration. Mishandling the tissues can readily lead to faint or no fluorescent signals. During the tissue mounting step, ovarioles should be separated from each other and spread out on the microscope slide. Make sure the egg chambers are not stacked on top of each other. We noticed that the egg chambers beyond stage 14 displayed little EdU incorporation. Additionally, because of the large size, the late stage egg chambers often cause the neighboring younger egg chambers to be out of focus. Thus it is recommended that the late stage egg chambers be discarded.
Here we provide a detailed protocol for labeling replicating mtDNA in Drosophila adult ovaries. This method allows for simple quantification of mtDNA replication under various genetic and pharmacological perturbations, and will be useful for dissecting mechanisms underlying developmental mitochondrial biogenesis and mtDNA inheritance.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank K. Delaney for comments on the manuscript. This work was supported by the National, Heart, Lung, and Blood Institute (NHLBI) Intramural Program.
Materials
| Name | Company | Catalog Number | Comments |
| Schneider’s Drosophila medium | Invitrogen | 21720-024 | |
| FBS | Invitrogen | 10100-147 | |
| Pair of Dumont #5 Forceps | Fine Science Tools | 11252-20 | |
| Aphidicolin | Sigma | A0781 | Aliquot after dissolving in DMSO. Avoid repetitive thawing and freezing. Protect from light. May be used within 6 weeks after dissolving. |
| FCCP | Sigma | C2920 | |
| DMSO | Sigma | D2650 | |
| Paraformaldehyde, 16% EM grade | Electron Microscopy Sciences | 15710 | Formaldehyde is toxic; it should be handled in a fume hood with skin and eye protection. |
| PBS | KD Medical | RGF-3190 | |
| BSA | Sigma | A7030 | |
| Triton X-100 | Sigma | T9284 | |
| Click-iT EdU Alexa Fluor 488 Imaging Kit | Invitrogen | C10337 | EdU, CuSO4, Alexa Fluor 488 azide, EdU reaction buffer and Edu buffer additive are included |
| Mouse ATP synthase subunit α antibody, (15H4C4) | MitoSciences | Ab14748 | 1:1,000 dilution |
| Mouse Hts antibody (clone RC) | Developmental Studies Hybridoma Bank (DSHB) | hts RC | 1:1,000 dilution |
| Goat anti-mouse Alexa Fluor 568 secondary antibody | Invitrogen | A-11004 | 1:200 dilution |
| Vectashield mounting medium with DAPI | Vector Laboratories | H-1500 | |
| Glass coverslips, #1.5 22 mm x 22 mm | Fisher Scientific | 12-541-B | |
| Microscope slide | Fisher Scientific | 22-038-103 | |
| Nail polish | Elf | Many of the pigments used in nail polishes are fluorescent and leach into specimens. Only clear nail polish should be used. |
References
- Taylor, R. W., Turnbull, D. M. Mitochondrial DNA mutations in human disease. Nat Rev Genet. 6 (5), 389-402 (2005).
- Wallace, D. C., Chalkia, D. Mitochondrial DNA genetics and the heteroplasmy conundrum in evolution and disease. Cold Spring Harb Perspect Biol. 5 (11), a021220 (2013).
- Spradling, A. C. . The development of Drosophila melanogaster. , (1993).
- Riechmann, V., Ephrussi, A. Axis formation during Drosophila oogenesis. Curr Opin Genet Dev. 11 (4), 374-383 (2001).
- Lin, H., Yue, L., Spradling, A. C. The Drosophila fusome, a germline-specific organelle, contains membrane skeletal proteins and functions in cyst formation. Development. 120 (4), 947-956 (1994).
- Cox, R. T., Spradling, A. C. A Balbiani body and the fusome mediate mitochondrial inheritance during Drosophila oogenesis. Development. 130 (8), 1579-1590 (2003).
- Davis, A. F., Clayton, D. A. In situ localization of mitochondrial DNA replication in intact mammalian cells. J Cell Biol. 135 (4), 883-893 (1996).
- Iborra, F. J., Kimura, H., Cook, P. R. The functional organization of mitochondrial genomes in human cells. BMC Biol. 2, (2004).
- Rakic, P. Neurogenesis in adult primate neocortex: an evaluation of the evidence. Nat Rev Neurosci. 3 (1), 65-71 (2002).
- Salic, A., Mitchison, T. J. A chemical method for fast and sensitive detection of DNA synthesis in vivo. Proc Natl Acad Sci U S A. 105 (7), 2415-2420 (2008).
- Hill, J. H., Chen, Z., Xu, H. Selective propagation of functional mitochondrial DNA during oogenesis restricts the transmission of a deleterious mitochondrial variant. Nat Genet. 46 (4), 389-392 (2014).
- Lentz, S. I., et al. Mitochondrial DNA (mtDNA) Biogenesis: Visualization and Duel Incorporation of BrdU and EdU Into Newly Synthesized mtDNA In Vitro. Journal of Histochemistry & Cytochemistry. 58 (2), 207-218 (2010).
- Tornoe, C. W., Christensen, C., Meldal, M. Peptidotriazoles on solid phase: [1,2,3]-triazoles by regiospecific copper(i)-catalyzed 1,3-dipolar cycloadditions of terminal alkynes to azides. J Org Chem. 67 (9), 3057-3064 (2002).
- Horwitz, H. B., Holt, C. E. Specific inhibition by ethidium bromide of mitochondrial DNA synthesis in physarum polycephalum. J. Cell Biol. 49, 546-553 (1971).
- Huberman, J. A. New views of the biochemistry of eucaryotic DNA replication revealed by aphidicolin, an unusual inhibitor of DNA polymerase alpha. Cell. 23 (3), 647-648 (1981).
- McClung, C., Hirsh, J. Stereotypic behavioral responses to free-base cocaine and the development of behavioral sensitization in Drosophila. Curr Biol. 8 (2), 109-112 (1998).
- Moore, M. S., et al. Ethanol intoxication in Drosophila: Genetic and pharmacological evidence for regulation by the cAMP signaling pathway. Cell. 93 (6), 997-1007 (1998).
- Dzitoyeva, S., Dimitrijevic, N., Manev, H. Gamma-aminobutyric acid B receptor 1 mediates behavior-impairing actions of alcohol in Drosophila: adult RNA interference and pharmacological evidence. Proc Natl Acad Sci U S A. 100 (9), 5485-5490 (2003).
- Koch, E. A., Spitzer, R. H. Multiple effects of colchicine on oogenesis in Drosophila: induced sterility and switch of potential oocyte to nurse-cell developmental pathway. Cell Tissue Res. 228 (1), 21-32 (1983).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved