Method Article
Detection of Protein Palmitoylation in Cultured Hippocampal Neurons by Immunoprecipitation and Acyl-Biotin Exchange (ABE)
In This Article
Summary
The reversible addition of palmitate to proteins is an important regulator of intracellular protein trafficking. This is of particular interest in neurons where many synaptic proteins are palmitoylated. We utilize a simple biochemical method to detect palmitoylated proteins in cultured neurons, which can be adapted for multiple cell types and tissues.
Abstract
Palmitoylation is a post-translational lipid modification involving the attachment of a 16-carbon saturated fatty acid, palmitate, to cysteine residues of substrate proteins through a labile thioester bond [reviewed in1]. Palmitoylation of a substrate protein increases its hydrophobicity, and typically facilitates its trafficking toward cellular membranes. Recent studies have shown palmitoylation to be one of the most common lipid modifications in neurons1, 2, suggesting that palmitate turnover is an important mechanism by which these cells regulate the targeting and trafficking of proteins. The identification and detection of palmitoylated substrates can therefore better our understanding of protein trafficking in neurons.
Detection of protein palmitoylation in the past has been technically hindered due to the lack of a consensus sequence among substrate proteins, and the reliance on metabolic labeling of palmitoyl-proteins with 3H-palmitate, a time-consuming biochemical assay with low sensitivity. Development of the Acyl-Biotin Exchange (ABE) assay enables more rapid and high sensitivity detection of palmitoylated proteins2-4, and is optimal for measuring the dynamic turnover of palmitate on neuronal proteins. The ABE assay is comprised of three biochemical steps (Figure 1): 1) irreversible blockade of unmodified cysteine thiol groups using N-ethylmaliemide (NEM), 2) specific cleavage and unmasking of the palmitoylated cysteine's thiol group by hydroxylamine (HAM), and 3) selective labeling of the palmitoylated cysteine using a thiol-reactive biotinylation reagent, biotin-BMCC. Purification of the thiol-biotinylated proteins following the ABE steps has differed, depending on the overall goal of the experiment.
Here, we describe a method to purify a palmitoylated protein of interest in primary hippocampal neurons by an initial immunoprecipitation (IP) step using an antibody directed against the protein, followed by the ABE assay and western blotting to directly measure palmitoylation levels of that protein, which is termed the IP-ABE assay. Low-density cultures of embryonic rat hippocampal neurons have been widely used to study the localization, function, and trafficking of neuronal proteins, making them ideally suited for studying neuronal protein palmitoylation using the IP-ABE assay. The IP-ABE assay mainly requires standard IP and western blotting reagents, and is only limited by the availability of antibodies against the target substrate. This assay can easily be adapted for the purification and detection of transfected palmitoylated proteins in heterologous cell cultures, primary neuronal cultures derived from various brain tissues of both mouse and rat, and even primary brain tissue itself.
Protocol
1. Extraction of Target Protein from Cultured Primary Hippocampal Neurons
- Before harvesting endogenous protein from primary hippocampal cells, prepare a Lysis Buffer (LB; Table 1) with protease inhibitors. To 10 ml of lysis buffer, add 0.1 ml phenylmethanesulfonyl fluoride solution (PMSF) to a final concentration of 10mM, and add 1 protease inhibitor cocktail tablet.
- Prepare N-ethylmaleimide (NEM) solution (Table 1) from lyophilized powder. Gently dissolve in 100% EtOH at room temperature (RT). Always prepare NEM solution fresh for the day of the experiment.
- Next combine the lysis buffer and NEM. Add 0.5 ml of 2M NEM solution to a fresh 50 ml conical tube. Add 20 ml LB over top, and quickly place LB +NEM (final concentration of 50mM) on a rocking platform at 4 °C to fully mix for 5 min. Always add NEM/EtOH solution first and LB second, so as to properly mix the two.
- Remove cultured neurons from incubator and place on ice*. Gently remove cellular media, and gently wash twice with cold phosphate buffered saline (PBS) 1x. Add 300 ml of LB + 50mM NEM to the first well of hippocampal neurons, and scrape cells using a disposable cell lifter. Collect the cell lysate, then discharge the lysate into the next well of that group. Scrape cells within the second well using the cell lifter, collect, and repeat using a third well, if needed to further concentrate the lysate.
- Collect cell lysate into a pre-cooled (on ice), 1.5 ml microcentrifuge tube. Repeat step 1.4 using 100 μl of LB + 50mM NEM to collect any remaining cell lysate. Pass total cell lysate through a 26½ gauge syringe 5-6 times to aid in mechanical lysis, being careful to not blow bubbles into the solution. One can instead sonicate for 15 sec on ice, if equipment is available. Nutate cell lysate for ≥30 min at 4 °C.
- Clarify the cell lysate by centrifugation at 16,000 x g/ 30 min, in order to pellet un-lysed cells and detergent-insoluble material.
- Collect cleared cell lysate (supernatant) in a new pre-cooled 1.5 ml tube on ice. Repeat harvest protocol for all groups of cells. Measure protein concentration in all lysate samples using BCA assay, and normalize the volume of all lysate samples from groups of cells to have exactly equal total protein, with a recommended minimum of 500 μg. Top up all samples with LB + 50mM NEM to 500 μl total volume.
- Add 1-5 μg of primary antibody directed against the target protein to each of lysate sample. Nutate the antibody-lysate mixed samples overnight at 4 °C.
* Primary rat hippocampal neurons should be cultured at a density of approximately 250,000 cells/well in a 6-well dish, in a serum-free media containing a B27 neuronal supplement, as previously described5, and grown for the desired length of time, typically 14-16 days-in-vitro (DIV) to achieve maturity. A minimum of 500 μg of total protein is recommended to successfully immunoprecipitate and biotinylate a target neuronal protein, which typically requires 2-3 wells of a 6-well dish.
2. Precipitation of Antibody-bound Target Protein
- Before precipitating and immobilizing a target protein, prepare a 50% slurry of protein A, or protein G-coated sepharose beads. Specifically, add ≥60 μl of sepharose beads per sample to 1.5 ml tubes, ensuring that all samples have equal amounts of beads. Magnetic beads are also suitable if the equipment is available.
- Add an equal volume of 50% slurry to each antibody-lysate sample, and nutate for ≥1 hr at 4 °C.
3. Acyl-Biotin Exchange: Hydroxylamine (HAM) Cleavage
- While performing step 2.2, prepare a number of tubes with lysis buffer (LB) of different pHs. The pH is very important for these steps and should always be adjusted using a pH meter. Prepare 2 ml of LB pH 7.2 per sample, and 0.5 ml of Stringent Buffer per sample (Table 1). Also prepare 0.5 ml LB + 10mM NEM per sample, as in steps 1.1-1.3. Add PMSF and protease inhibitor tablets to all lysis buffers, as in step 1.1.
-
- Hydroxylamine (HAM) is a powerful reducing agent, whose cleavage of palmitate from cysteines is required for biotinylation (Figure 1), and the omission of the HAM cleavage serves as a negative control. Split each sample of beads into two samples, one omitting the HAM cleavage step (-HAM), and one including the HAM step (+HAM). To normalize for protein degradation caused by HAM treatment, one should split each sample into thirds, with 1/3 of the beads used for the -HAM control, and the remaining 2/3 used for the +HAM treatment.
- Prepare additional 1.5 ml tubes on ice, labeled as the -HAM control for each sample. Following step 2.2, gently centrifuge all samples' beads at 0.5 x g/ 1 min at 4 °C (centrifuge at this speed, duration, and temperature for all remaining steps unless otherwise stated), place the tubes on ice, remove the supernatant, and re-suspend the beads in 600 μl of LB + 10mM NEM.
- After re-suspending the samples' beads in lysis buffer + NEM 10mM, immediately collect 200 μl of the sample's lysate-bead slurry, and discharge into that sample's -HAM tube on ice. Add an additional 300 μl of LB + 10mM NEM to the -HAM sample, and an additional 100 μl to the +HAM sample for a total of 500 μl in each sample. Incubate the tubes for 10 min on ice.
- Re-suspend (wash) the -HAM and +HAM samples gently with 0.5 ml stringent buffer, per sample. Perform this step quickly, as a longer SDS incubation will result in the removal of bound antibody from the beads.
- Gently wash all samples three times in 0.5 ml/sample of LB pH 7.2, and spin down between washes as in step 3.2b.
- While performing step 3.5, prepare a HAM buffer (Table 1). Add appropriate volume of hydroxylamine solution to a new tube, to achieve a final concentration of 1M in a volume of 0.5 ml LB pH 7.2, per +HAM sample. Always prepare the HAM buffer fresh for the day of the experiment. Add 0.5 ml/sample of LB pH 7.2 to all -HAM samples, and 0.5 ml/ sample of HAM buffer to all +HAM samples.
- Nutate all samples for 1 hr at room temperature (RT). Do not exceed 1 hr.
4. Acyl-Biotin Exchange: Biotin-BMCC Labeling
- While performing step 3.6, prepare 2 ml of LB pH 6.2 per sample (Table 1), as in step 3.1. Place the LB pH 6.2 on ice until needed. Also prepare a stock solution of Biotin-BMCC (Table 1) immediately before use. Weigh out exactly 2.1 mg of Biotin-BMCC, collect in a 1.5 ml tube, and gently dissolve in 0.5 ml Dimethyl sulfoxide (DMSO) by placing on a rocking platform at RT, to achieve 8mM concentration (do not use a pipette to break up the powder).
- Once step 3.6 is completed, gently wash the beads once in pH 6.2 lysis buffer, to remove residual HAM buffer. Remove the supernatant and place the samples on ice.
- Biotin-BMCC is a maliemide conjugated biotin molecule that has a high affinity for free thiol groups of cysteines (Figure 1). While performing step 4.2, prepare 0.5 ml of Biotin-BMCC Buffer (Table 1) per sample, at working concentration between 0.5 μM to 5 μM. Concentrations in excess of 5 μM will likely saturate the target protein6, and typically should not be exceeded, however this may differ depending on the desired target protein. Add 0.5 ml of Biotin-BMCC buffer to each sample, and nutate for exactly 1 hr at 4 °C. Do not exceed 1 hr.
5. Elution of Biotin-conjugated Target Protein and SDS-PAGE
- Once step 4.3 is completed, gently wash all samples once in LB pH 6.2. Keep all samples on ice in between washes. Remove 2x SDS Sample Buffer with no reducing agents (Table 1) from -20 °C storage and slowly thaw on ice.
- Gently wash samples three times in LB pH 7.5 + PMSF/Inhibitors, and keep samples on ice in between washes.
- Remove the supernatant of the third wash, leaving the pelleted beads in a slurry with remaining buffer in the bottom of the tube. Dip a pipette with a small-diameter tip (a gel loading tip or P2 pipette tip are sufficient) into the very bottom of the tube through the slurry, and quickly collect any remaining buffer, being careful not to take up any beads.
- Supplement the 2x SDS Sample Buffer with DL-Dithiothreitol (DTT) to achieve a final concentration of 5mM. Add 40-50 μl of 2x Sample Buffer + 5mM DTT to each sample. If necessary, vortex the beads to completely mix with the sample buffer, and immediately centrifuge at high speed to collect all beads in a pellet, submerged in sample buffer.
- Boil all samples for 10 min at 75-80 °C. Let samples cool to RT for ≥10 min on bench top, and centrifuge at 16,000 x g / 3 min to completely pellet the beads.
- Add complete volume of eluted protein (supernatant) from each sample to a single lane within a polyacrylamide gel suitable for western blotting, using SDS-PAGE7. Organize all samples so that the -HAM control eluent and +HAM eluent for a single sample are run immediately adjacent to one another during SDS-PAGE (Fig. 2). Following SDS-PAGE, transfer to a PVDF or nitrocellulose membrane.
6. Western Blotting and Detection of Palmitoylation of Target Protein
- Once step 5.6 is completed, wash the membrane twice in Tris-buffered Saline (TBS 1x) for 10 min on a rocking platform at RT.
- Prepare a Blocking Buffer to block non-specific labeling by weighing out Bovine Serum Albumin (BSA), to achieve 3% weight in a volume of TBS 1x + Tween-20 0.05% (TBS-T) that is sufficient to fully submerge the membrane. For streptavidin blotting, always block with BSA and not powdered milk. Block the membrane for 1 hr at RT on a rocking platform.
- Prepare an antibody solution of TBS-T + 0.3% BSA. Add Streptavidin-HRP antibody reconstituted at 1 mg/ml in distilled water, diluted at 1:5,000-1:10,000. Once step 6.2 is completed, wash the membrane once in TBS-T for 10 min, then incubate the membrane in Streptavidin antibody solution on a rocking platform for either for 1 hr at RT, or overnight at 4 °C.
- Wash the membrane three times in TBS-T for 10 min on a rocking platform. In order to detect the palmitoylation signal, expose the HRP luminescence using a chemiluminescent substrate kit.
- In order to normalize palmitoylation levels to the amount of the protein of interest that was immunoprecipitated, strip the membrane using western blot stripping buffer for 5-10 min on a rocking platform at RT. Repeat steps 6.1 - 6.4 using the primary antibody against the protein of interest that was originally used for the immunoprecipitation.
- Quantify palmitoylation of the protein of interest using image analysis software, and normalize palmitoylation levels to the amount of immunoprecipitated protein.
Results
The IP-ABE assay specifically detects thiol-palmitoylation of cysteine residues along substrate proteins, and can be used to detect palmitoylation of immunoprecipitated neuronal proteins in a manner depicted in Figure 1. Treatment of the immunoprecipitated neuronal protein with HAM (Figure 1) cleaves the thioester linkage between palmitate and a cysteine's thiol group, allowing for specific incorporation of biotin-BMCC onto the newly available thiol group, which can be subsequently detected using western blotting. The omission of HAM (minus-HAM) cleavage prevents the incorporation of biotin-BMCC and functions as a negative control for specific palmitoyl-biotinylation of the plus-HAM sample, and should therefore always be run adjacent to the plus-HAM sample for western blotting by SDS-PAGE (Figure 2).
In an optimized experiment (Figure 2A), the palmitoylation signal of the neuronal protein δ-catenin2 can be readily detected by blotting for biotin-BMCC at a concentration of 1 μM, using streptavidin conjugated with HRP, against parallel minus- and plus-HAM samples. The specific palmitoylation signal for δ-catenin appears at its predicted size of 160kD, only in the plus-HAM sample. The specificity of this signal was confirmed by reprobing for δ-catenin with the specific antibody that was initially used to immunoprecipitate it, resulting in a signal in both HAM treatments at the same predicted size. Optimization of the IP-ABE assay (Figure 2A) will result in a western blotting profile of a minus-HAM sample that is readily distinguishable from the plus-HAM sample, and exhibits minimal to no palmitoylation signal.
The concentration of biotin-BMCC used to label palmitoylated cysteines requires careful optimization, and if a super-saturating concentration of biotin-BMCC is used during the ABE chemistry steps (Figure 2B), excessive background and non-specific signals will be visible. A linear response curve for biotinylation of a palmitoylated neuronal protein, the α7 nicotinic acetylcholine receptor, using biotin-BMCC has previously been determined6, and shows near complete saturation at a concentration of 5-10 μM. Although the optimal concentration of biotin-BMCC will deviate slightly depending on the neuronal target protein, treatment with biotin-BMCC at a concentration in excess of 5 μM will likely super-saturate the target protein, and result in a western blot profile with visible background and non-specific signals. A concentration of 0.5 μM for biotin-BMCC was previously used to detect palmitoylation of other neuronal proteins including SNAP-25, huntingtin, AMPA receptor subunits GluA1 and GluA2, and paralemmin8, and determination of the optimal concentration for a novel target protein can be accomplished by trial and error in linear fashion, starting within the range of 0.5-5 μM that we describe here. The resulting streptavidin western blot profiles among the minus- and plus-HAM samples for δ-catenin palmitoylation appear similar when treated with 4 μM biotin-BMCC (Figure 2B), with additional background signals at different sizes, which indicates that inappropriate biotinylation occurred.
Hydroxylamine is a powerful reducing agent that results in limited degradation of the target protein in addition to its efficient cleavage of thioester linkage. Consequently, optimization of ABE chemistry necessitates one to normalize the amount of immunoprecipitated protein used for minus- and plus-HAM samples (steps 3.2 - 3.3). Normalization requires the use of double the amount of immobilized target protein in the plus- vs. the minus-HAM sample, which actually results in indistinguishable western blotting profiles for the target protein, as shown for δ-catenin (Figure 2A, B). However, if the amount of immobilized target protein is not normalized between minus- and plus-HAM samples, the resultant western blot signal for the target protein in the plus-HAM sample can be lost, as shown for δ-catenin when equal amounts of protein were used in both HAM treatments (Figure 2C).
The specificity of ABE chemistry for labeling palmitoylated proteins is very sensitive and requires optimization. The common pitfalls encountered during the IP-ABE assay include the sub-optimal results shown in Figures 2B and 2C, and degradation of the target protein, independent of HAM treatment, which are typically caused by older reagents and buffers, and incorrect pH. In order to avoid these pitfalls and correct for sub-optimal ABE chemistry, the freshness and concentration of the reagents required for the buffers listed in Table 1 should always be examined, and the pH adjustments of all the buffers should always be checked prior to running the experiment.
| Buffer | Working Concentration | Comments | |
| Lysis Buffer (LB) | In distilled H2O: 1% IGEPAL CA-630 50mM Tris-HCl pH7.5 150mM NaCl 10% Glycerol | N/A | Prepare before experiment and store at 4 °C |
| NEM Solution | In 100% EtOH: NEM lyophilized powder | 2 M | Prepare fresh, immediately before use. |
| Stringent Buffer | In LB: 10 mM MEM 0.1% SDS | N/A | Prepare shortly before use, keep on ice. |
| LB pH 7.2 | In LB: Adjust PH to exactly 7.2 | N/A | Use a pH meter to adjust pH immediately before use. |
| HAM Buffer | In LB pH 7.2: Stock HAM solution | 1 M (final HAM concentration) | Prepare immediately before use. |
| LB pH 6.2 | In LB: Adjust pH to exactly 6.2 | N/A | Same as for LB pH 7.2 |
| Stock Biotin-BMCC Solution | In DMSO: 2.1 mg Biotin-BMCC solution | 8 mM | Prepare immediately before use. |
| Biotin-BMCC Buffer | In LB pH 6.2: Add Biotin-BMCC solution | 1 - 5 μM (final biotin-BMCC concentration) | Prepare immediately before use. |
| 2x SDS Sample Buffer, no reducing agents | In distilled H2O: 5% SDS 5% Glycerol 125mM Tris-HCL pH 6.8 0.01% Bromophenol Blue | N/A | Prepare before experiment, and store at -20 °C until use. Supplement with 5 mM DTT before use. |
Table 1. Buffers and reagents required for the IP-ABE assay. Many of the lysis buffers can be prepared before beginning the experiment, however the pH should be checked and adjusted immediately before use with a pH meter, to ensure success of the ABE chemistry. The majority of stock solutions should be prepared fresh for every experiment, immediately before use. Many of the buffers require reagents common to most laboratories equipped for biochemistry, and specialty reagents with vendor information are listed in the table of reagents and equipment.
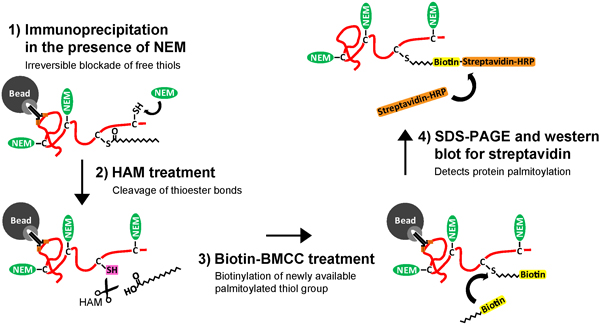
Figure 1. Schematic of the Immunoprecipitation and Acyl-Biotin Exchange (IP-ABE) assay to purify and detect palmitoylation of neuronal proteins. Cultured hippocampal neurons are lysed, (1) a target protein is then purified using a target-specific antibody, and immobilized on sepharose beads coated with protein G or A. The purified target protein is then (2) treated with N-ethylmaliemide (NEM) to irreversibly bind and block free thiol (-SH) groups along unmodified cysteines (C). The target protein is then (3) subjected to treatment with hydroxylamine (HAM), resulting in specific cleavage of thioester bonds at palmitoylated cysteines and the unmasking of a free palmitoylated thiol group (-SH). Next, the target protein is (4) treated with a thiol-reactive biotin molecule, biotin-BMCC, resulting in specific biotinylation of the palmitoylated cysteine. Finally, (5) the biotinylated target protein is eluted and removed from the antibody and bead. The target protein with its palmitoylated cysteine(s) tagged with biotin is now suitable for SDS poly-acrylamide gel electrophoresis (SDS-PAGE), and western blotting with streptavidin to detect for palmitoylation of the purified neuronal protein. Click here to view larger figure.
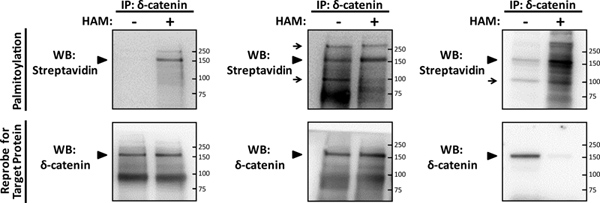
Figure 2. Detection of palmitoylation of the neuronal protein δ-catenin using the IP-ABE assay. Primary rat hippocampal neurons were grown as previously described5, at a density of 130 cells/mm2 until maturity at 14DIV, were then lysed, and subjected to the IP-ABE assay to detect palmitoylation of a recently identified palmitoylated neuronal substrate, δ-catenin2. Purified immunoprecipitates (IPs) of δ-catenin (target protein) were isolated using 5 μg of δ-catenin antibody per sample (BD Transduction Laboratories), and were then subjected to SDS-PAGE in parallel minus- and plus-hydroxylamine (HAM) samples. Palmitoylation was detected by western blotting (WB) with streptavidin-HRP (palmitoylation), and the membrane was stripped and subjected to a reprobe WB for δ-catenin. (A) An optimized representative IP-ABE result for palmitoylated δ-catenin (arrowhead) treated with 1 μM biotin-BMCC, illustrating the specificity of ABE chemistry for detecting thiol-palmitoylated proteins from IP samples. (B) A sub-optimal representative IP-ABE result for δ-catenin, where an excess concentration of 4 μM biotin-BMCC was used, resulting in non-specific signals and background in both minus- and plus-HAM samples (arrows). (C) Another sub-optimal representative IP-ABE WB for δ-catenin, where an excessive 4 μM concentration of biotin-BMCC was used, and the amount of protein used for the plus-HAM IP sample was not normalized for HAM-mediated protein degradation, resulting in a lost signal in the reprobe WB for δ-catenin. Click here to view larger figure.
Discussion
The IP-ABE assay presented here is a simple and highly sensitive method for detecting palmitoylated proteins in cultured primary hippocampal neurons, using mostly standard biochemical techniques. This makes the assay easily adaptable for labs already equipped with western blotting materials. The IP-ABE assay combines a standard immunoprecipitation protocol to isolate and immobilize one's target protein, followed by previously described ABE chemistry2-4 to rapidly detect levels of palmitate on a substrate protein. The ABE technique has a wide range of applications for examining palmitoylated proteins in various tissues and cell lines, including large-scale palmitoyl-proteomic screening of global palmitoylation profiles, and rapid detection of palmitoylation levels of a single target protein.
Previous descriptions of ABE chemistry have utilized different methods cell lysis and detergent extraction of neuronal proteins2, 9, free thiol-blockade10, 11, biotinylation, and purification of the target protein4, depending on the application of the experiment. The addition of palmitate to neuronal proteins results in their targeting toward cellular membranes, and detection of the palmitoylated fraction of a target protein will require its extraction from these membranes. Previously described ABE methodologies have utilized a variety of ionic9 and non-ionic detergents8 to extract desired target proteins, and the use of a particular detergent depends on the target protein and its known membrane trafficking. Here, we utilize the non-denaturing and non-ionic detergent IGEPAL CA-630 to extract palmitoylated proteins, which we have validated for extraction and detection of the palmitoylated neuronal protein δ-catenin (Figure 2). The structure of IGEPAL CA-630 includes a bulky and rigid non-polar head group that does not disrupt native protein conformation or interactions, but which allows its association with the hydrophobic regions of membrane-associated proteins, thereby conferring miscibility to them and permitting their extraction. Many neuronal proteins localized to the synaptic membrane or post-synaptic density of synapses should be extracted using IGEPAL CA-630, however some may not completely solubilize, and may require use of a different non-denaturing detergent like Triton X-100, which has been previously utilized for ABE chemistry2, 8.
Several descriptions of ABE chemistry have also utilized a different thiol-reactive biotinylation reagent from the molecule we describe here, called Biotin-HPDP (Thermo Scientific), which also necessitates a different means of protein purification4. Biotin-HPDP is another thiol-reactive, maliemide-conjugated biotin molecule that contains a disulfide bond in the maliemide linker arm, structurally differentiating it from Biotin-BMCC, which does not contain this disulfide bond. Following thiol-biotinylation of a protein's free or palmitoylated cysteine by Biotin-HPDP, the resulting disulfide bond between the target protein and the biotin group can be cleaved by reducing reagents to release the biotin group and regenerate the protein in its unmodified form, making biotinylation by Biotin-HPDP transient and reversible. Therefore, use of this biotinylation reagent requires purification of a target protein by immobilized avidin reagents, such as streptavidin-coated beads. However, thiol-biotinylation of a target protein by Biotin-BMCC, as we describe here, is stable and relatively permanent, and requires purification of the target protein by an antibody directed against the target, and immobilized on sepharose beads. Both ABE techniques have been previously utilized for large-scale screens, and detection of palmitoylation of single proteins2, 8-10, 12, and the IP-ABE assay we describe here is optimal for rapid detection of palmitoylation of single neuronal target proteins.
An overwhelming majority of cellular palmitoylation involves the reversible addition of palmitate to the thiol group of cysteines (termed S-palmitoylation, which we generalize as 'palmitoylation' in this protocol), however a very limited group of proteins are subjected to irreversible palmitoylation at glycine and cysteine residues through the formation of stable amide bonds, termed N-palmitoylation13. One limitation of ABE chemistry is its inability to detect N-palmitoylation owing to the stability of the amide bond, and so screening of a novel target protein for palmitoylation should be confirmed using 3H-palmitate metabolic labeling1.
As previously mentioned, the IP-ABE assay can be easily adapted for examining palmitoylation in protein extracts from multiple sources, including transfected heterologous cell lines, primary tissue, and primary neuronal cultures from multiple brain regions. The IP-ABE assay has been utilized for examining palmitoylation sufficiency of mutated cysteine-to-serine proteins to identify the location of a palmitoylated cysteine along a target protein8, for examining dynamic palmitate turnover on synaptic proteins in cultured neurons under basal conditions10, and changes in palmitoylation levels of neuronal proteins following induction of neuronal activity9. The sensitivity and adaptability of the IP-ABE assay make it ideally suited for examining palmitoylation profiles of neuronal proteins, and optimal for detecting dynamic changes in palmitoylation.
Disclosures
No conflicts of interest declared.
Acknowledgements
This work was supported by grants from Canadian Institutes of Health Research MOP-81158.
Materials
| Name | Company | Catalog Number | Comments |
| IGEPAL CA-630 | Sigma | I8896 | |
| PMSF | Fluka | 93482 | |
| Protease Inhibitor Cocktail | Roche Complete Mini | 11 836 153 001 | |
| NEM | Sigma | E3876 | |
| HAM solution | Sigma | 46780-4 | |
| BCA Protein Assay Kit | Thermo Scientific | 23225 | Ensure compatibility with chosen detergent |
| Protein G/A-coated sepharose beads | GE Healthcare | 17-6002-35 | |
| Biotin-BMCC | Thermo Scientific | 21900 | |
| Streptavidin-HRP | Thermo Scientific | 21126 | Reconstitute at 1 mg/ml in water |
| Western Blot Stripping Buffer | Thermo Scientific | 21059 |
References
- Fukata, Y., Fukata, M. Protein palmitoylation in neuronal development and synaptic plasticity. Nat. Rev. Neurosci. 11, 161-175 (2010).
- Kang, R., et al. Neural palmitoyl-proteomics reveals dynamic synaptic palmitoylation. Nature. 456, 904-909 (2008).
- Drisdel, R. C., Green, W. N. Labeling and quantifying sites of protein palmitoylation. Biotechniques. 36, 276-285 (2004).
- Wan, J., Roth, A. F., Bailey, A. O., Davis, N. G. Palmitoylated proteins: purification and identification. Nat. Protoc. 2, 1573-1584 (2007).
- Xie, C., Markesbery, W. R., Lovell, M. A. Survival of hippocampal and cortical neurons in a mixture of MEM+ and B27-supplemented neurobasal medium. Free Radic. Biol. Med. 28, 665-672 (2000).
- Drisdel, R. C., Alexander, J. K., Sayeed, A., Green, W. N. Assays of protein palmitoylation. Methods. 40, 127-134 (2006).
- Eslami, A., Lujan, J. W. e. s. t. e. r. n. Blotting: Sample Preparation to Detection. J. Vis. Exp. (44), e2359 (2010).
- Huang, K., et al. Neuronal palmitoyl acyl transferases exhibit distinct substrate specificity. FASEB J. 23, 2605-2615 (2009).
- Noritake, J., et al. Mobile DHHC palmitoylating enzyme mediates activity-sensitive synaptic targeting of PSD-95. J. Cell Biol. 186, 147-160 (2009).
- Thomas, G. M., Hayashi, T., Chiu, S. L., Chen, C. M., Huganir, R. L. Palmitoylation by DHHC5/8 Targets GRIP1 to Dendritic Endosomes to Regulate AMPA-R Trafficking. Neuron. 73, 482-496 (2012).
- Hayashi, T., Thomas, G. M., Huganir, R. L. Dual palmitoylation of NR2 subunits regulates NMDA receptor trafficking. Neuron. 64, 213-226 (2009).
- Ho, G. P., et al. S-Nitrosylation and S-Palmitoylation Reciprocally Regulate Synaptic Targeting of PSD-95. Neuron. 71, 131-141 (2011).
- Iwanaga, T., Tsutsumi, R., Noritake, J., Fukata, Y., Fukata, M. Dynamic protein palmitoylation in cellular signaling. Prog. Lipid Res. 48, 117-127 (2009).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved