A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Reusable Single Cell for Iterative Epigenomic Analyses
In This Article
Summary
The present protocol describes a single-cell method for iterative epigenomic analyses using a reusable single cell. The reusable single cell allows analyses of multiple epigenetic marks in the same single cell and statistical validation of the results.
Abstract
Current single-cell epigenome analyses are designed for single use. The cell is discarded after a single use, preventing analysis of multiple epigenetic marks in a single cell and requiring data from other cells to distinguish signal from experimental background noise in a single cell. This paper describes a method to reuse the same single cell for iterative epigenomic analyses.
In this experimental method, cellular proteins are first anchored to a polyacrylamide polymer instead of crosslinking them to protein and DNA, alleviating structural bias. This critical step allows repeated experiments with the same single cell. Next, a random primer with a scaffold sequence for proximity ligation is annealed to the genomic DNA, and the genomic sequence is added to the primer by extension using a DNA polymerase. Subsequently, an antibody against an epigenetic marker and control IgG, each labeled with different DNA probes, are bound to the respective targets in the same single cell.
Proximity ligation is induced between the random primer and the antibody by adding a connector DNA with complementary sequences to the scaffold sequence of the random primer and the antibody-DNA probe. This approach integrates antibody information and nearby genome sequences in a single DNA product of proximity ligation. By enabling repeated experiments with the same single cell, this method allows an increase in data density from a rare cell and statistical analysis using only IgG and antibody data from the same cell. The reusable single cells prepared by this method can be stored for at least a few months and reused later to broaden epigenetic characterization and increase data density. This method provides flexibility to researchers and their projects.
Introduction
Single-cell technology is entering the era of single-cell multiomics, which integrates individual single-cell omics technologies1. Recently, single-cell transcriptomics has been combined with methods for detecting chromatin accessibility (scNMT-seq2 and SHARE-seq3) or histone modifications (Paired-seq4 and Paired-Tag5). More recently, single-cell transcriptomics and proteomics were integrated with chromatin accessibility (DOGMA-seq6). These methods use transposase-based tagging for detecting chromatin accessibility or histone modifications.
Transposase-based approaches cleave genomic DNA and add a DNA barcode at the end of the genomic DNA fragment. Each cleaved genomic fragment can only accept up to two DNA barcodes (= one epigenetic mark per cleavage site), and the genomic DNA at the cleavage site is lost. Therefore, cleavage-based approaches have a trade-off between the number of epigenetic marks tested and the signal density. This hampers the analysis of multiple epigenetic marks in the same single cell. A single-cell epigenomic method that does not cleave the genomic DNA was developed to overcome this issue7,8.
In addition to the cleavage-derived issue mentioned above, transposase-based approaches have other limitations. In single-cell epigenome analysis, it is critical to know the location of histones and DNA-associated proteins on the genome. In current approaches, this is accomplished by using unfixed single cells and retention of protein-DNA and protein-protein interactions. However, this generates strong bias to accessible chromatin regions, even in the analysis of histone modifications 9. The location of histones and genome-associated proteins on the genome can be preserved without crosslinking protein-DNA and protein-protein, using a polyacrylamide scaffold7,8. This approach reduces the structural bias observed in current approaches that depend upon protein-DNA and protein-protein interactions.
Transposase-based approaches can acquire signals only once from a single cell. Therefore, it is difficult to delineate the complete epigenome of a single cell due to the drop-off of the signals. Reusable single cells have been developed to overcome current limitations by allowing iterative epigenomic analysis in the same single cell.
Protocol
NOTE: A schematic representation of the method is shown in Figure 1.
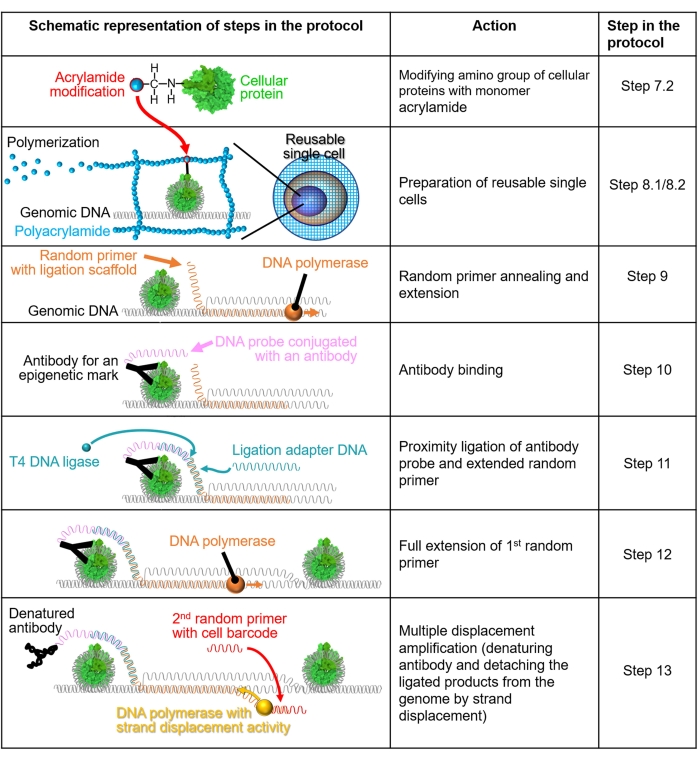
Figure 1: Schematic representation of the protocol workflow. Steps 7.2-13 are explained through schematic representations. Each row indicates a step in the protocol. A cellular protein colored in green is a human nucleosome generated based on a crystal structure (PDB: 6M4G). Please click here to view a larger version of this figure.
1. Equilibration of desalting columns
NOTE: Desalting spin columns are equilibrated as described in the following steps. The equilibrated desalting columns are used in steps 2.1, 3.4, and 4.6.
- Remove the bottom closure of a desalting column (7 kDa cut-off, 0.5 mL resin bead volume, see the Table of Materials) and loosen the cap at the top of the desalting column.
- Place the column in a 1.5 mL protein low-binding tube (a collection tube, see the Table of Materials) and centrifuge at 1,500 × g for 1 min at room temperature to remove the storage solution in the column.
NOTE: Use a swing rotor centrifuge to flatten the top of the bead bed. - Remove the flowthrough in the 1.5 mL tube and add 300 µL of 150 mM NaCl/100 mM phosphate buffer, pH 8.0 (see Table 1), on top of the resin bed.
- Centrifuge at 1,500 × g for 1 min at room temperature and remove the flowthrough in the collection tube.
- Repeat steps 1.3-1.4 three additional times.
- Discard the buffer from the collection tube and place the column in a new collection tube.
- Use the equilibrated desalting column in steps 2.1, 3.4, and 4.4.
2. Buffer exchange of antibodies
NOTE: Remove glycerol, arginine, and sodium azide from anti-H3K27ac10, anti-H3K27me310, anti-Med111, and anti-Pol II10 (see buffer composition shown in Table 1). All following procedures are performed under a clean hood to avoid DNase contamination. Time: 1 h
- Apply the antibody solution (100 µL, see Table 2) to an equilibrated desalting column prepared following step 1.
- Centrifuge at 1,500 × g for 2 min at 4 °C and transfer the flowthrough from the collection tube to a 1.5 mL protein low-binding tube.
- Measure the IgG concentration using absorbance at 280 nm12 (use a microvolume spectrophotometer).
- Transfer the antibody solution to an ultrafiltration cassette (molecular weight cut-off 100 kDa, 0.5 mL, see the Table of Materials) and centrifuge at 12,000 × g for 5 min at 4 °C.
- Measure the IgG concentration using the absorbance at 280 nm.
- Repeat steps 2.4-2.5 until the IgG concentration reaches 1 mg/mL.
3. Antibody activation
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 2.5 h
- Dissolve 1 mg of succinimidyl 6-hydrazinonicotinate acetone hydrazone (S-HyNic, see the Table of Materials) with 100 µL of anhydrous N,N-dimethylformamide (DMF, see the Table of Materials).
NOTE: DMF is a flammable organic solvent and a potent liver toxin absorbed through the skin. Wear gloves, safety goggles, and a lab coat. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines. - Add 0.6 µL of S-HyNic/DMF to 100 µL of the antibody solution (1 mg/mL in 150 mM NaCl/100 mM Sodium phosphate, pH8.0. See Table 2 for the used antibodies and control IgG.
- Incubate at room temperature for 2 h (protecting from light).
- Apply 100 µL of S-HyNic-reacted antibody to the top of the equilibrated desalting column (see step 1) and centrifuge at 1,500 × g for 2 min at 4 °C to collect the sample.
- Discard the desalting column after use.
NOTE: The stability of HyNic groups on proteins and other biomolecules varies. It is recommended to conjugate HyNic-modified biomolecules immediately.
4. Activation of DNA probe
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 2.5 h
- Dissolve a pellet of an amine-modified DNA probe for antibody or control IgG (Antibody Probe, Table 3) with 20 µL of 150 mM NaCl/100 mM Sodium Phosphate buffer, pH 8.0.
NOTE: Look for a thin, transparent pellet/film at the bottom of the tube. Apply the buffer directly to the pellet. If the pellet is not visible, the pellet may have detached and adhered to the wall or the lid. In this case, centrifuge the tube and look for a pellet at the bottom of the tube. - Dissolve 1 mg of succinimidyl 4-formylbenzoate (S-4FB, see the Table of Materials) with 50 µL of anhydrous DMF, and add 10 µL of DMF to the dissolved Antibody Probe.
- Add 4 µL of S-4FB/DMF prepared in step 4.2, mix, and incubate at room temperature for 2 h (protecting from light).
- Apply 34 µL of S-4FB-reacted Antibody Probe to the top of the equilibrated desalting column (see step 1).
- Apply 15 µL of 150 mM NaCl/100 mM Sodium phosphate buffer, pH 8.0, to the top of the gel bed after the sample has been fully absorbed, and centrifuge at 1,500 × g for 2 min at 4 °C to collect the S-4FB-modified Antibody Probe.
- Use the flowthrough for the subsequent conjugation; measure the absorbance at 260 nm; and calculate the recovery rate of the Antibody Probe.
5. Conjugation of S-HyNic-modified antibody and S-4FB-modified Antibody Probe
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 2 h
- Mix the S-HyNic-modified antibody and the S-4FB-modified Antibody Probe, and pipette the solution up and down to mix.
- Incubate for 2 h at room temperature (protecting from light).
- To quench the reaction, add 478.8 µL of Quenching & Storage solution (see Table 1).
- Transfer the Antibody Probe-conjugated antibody solution to an ultrafiltration cassette (molecular weight cut-off 100 kDa).
- Centrifuge for 5 min at 12,000 × g, 4 °C.
- Check the volume of the solution inside the cassette by pipetting.
- Repeat steps 5.5-5.6 until the volume reaches 100 µL and store at -20 °C.
NOTE: IgG concentration is measured by sandwich ELISA13,14,15 using an IgG standard.
6. Preparation of core magnetic beads
NOTE: In this method, a single cell is embedded into a bilayered acrylamide bead (see Figure 2). The core is a magnetic polyacrylamide bead. The outer layer is polyacrylamide alone. The core magnetic beads are generated in this section. This section is not essential for the experiment. Time: 3 h
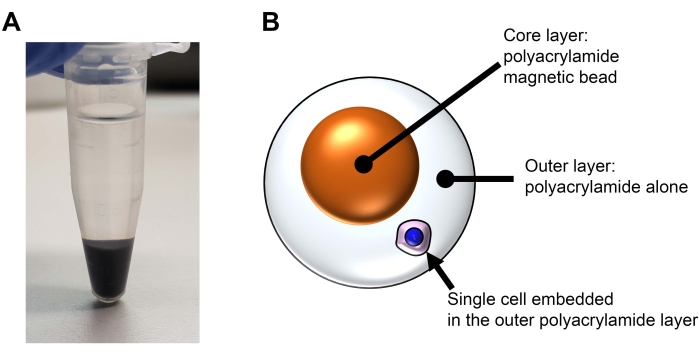
Figure 2: Structure of a bilayered polyacrylamide bead for visibility and easy handling in REpi-seq experiments. (A) Magnetic nanoparticles from step 6.6 after centrifugation. The magnetic nanoparticles are modified with monomeric acrylamide and integrated into the polyacrylamide magnetic bead shown in B.(B) Schematic representation of a reusable single cell with a polyacrylamide magnetic bead. Please click here to view a larger version of this figure.
- Mix 50 µL of 1 M sodium bicarbonate buffer, pH 8.5, and 450 µL of 40% acrylamide solution.
NOTE: Acrylamide is a neurotoxin. Wear gloves, an eye protector, and a lab coat. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines. - Suspend 1 g of NHS ester-functionalized 30 nm of iron oxide powder (see the Table of Materials) in the acrylamide/sodium bicarbonate buffer and incubate at 4 °C overnight.
NOTE: Because acrylamide beads are transparent, it may be difficult to see and manipulate the position of the beads. The inclusion of a core bead made of iron oxide improves visibility and facilitates manipulation because the position of the beads can be controlled using a magnet. However, if users are familiar with the REpi-seq experiments, use of the core magnetic polyacrylamide beads can be skipped. - Transfer the nanobead suspension into 2 tubes (1.5 mL), centrifuge at 21,300 × g for 1 h (use an angled rotor) at 4 °C, and remove the supernatant.
- Suspend the bottom slurry with 1 mL of 40% Acrylamide/Bis-acrylamide (19:1, see the Table of Materials).
NOTE: Bis-acrylamide is a neurotoxin. Wear gloves and a lab coat. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines. - Centrifuge at 21,300 × g for 1 h (use an angled rotor) at 4 °C.
- Centrifuge at 5,000 × g for 30 min (use a swing rotor without a brake) at 4 °C.
- Remove the supernatant using a P1000 pipette with slow-speed aspiration.
- Adjust the volume to 400 µL of 40% Acrylamide/Bis-acrylamide (19:1, see the Table of Materials).
- Add 25 µL of 10% ammonium persulfate solution (see Table 1).
NOTE: Ammonium persulfate is a strong oxidizing agent. Airborne dust containing ammonium persulfate may irritate the eye, nose, throat, lung, and skin upon contact. Wear gloves, safety goggles, and a lab coat. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines. - To generate core magnetic polyacrylamide beads, transfer 0.5 µL of the acrylamide-modified iron oxide suspension to a PCR tube.
- Add 50 µL of 4% N,N,N',N'-tetramethylethylenediamine/mineral oil (TEMED, see Table 1) and incubate overnight at room temperature.
NOTE: TEMED is a flammable solvent. Work under a hood. Do not inhale. Keep away from open flames, hot surfaces, and sources of ignition. Wear gloves, safety goggles, and a lab coat. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines.
7. Modifying the amino group of cellular proteins with monomer acrylamide
NOTE: REpi-seq was designed to analyze the epigenome of mouse and human cells at the single-cell level. Each step must be optimized when using this method on cells of species other than mouse or human.
- Harvesting the cells
NOTE: Time: 30 min- Measure the cell concentration and viability using a cell counter (see the Table of Materials).
NOTE: The viability of the cells at this step affects how many cells are live cells in the data analysis. - Adjust the cell concentration to 1 × 105 cells/mL with culture medium (*) containing 10% fetal bovine serum.
NOTE: *Culture medium is an optimal culture medium for the cells of interest. - Transfer 1 mL of the cell suspension to a 1.5 mL tube.
- Centrifuge the cell suspension at 240 × g for 5 min at 4 °C and remove the supernatant.
- Add 1 mL of phosphate-buffered saline (PBS), mix the cells by gentle pipetting, and centrifuge the cell suspension at 240 × g for 5 min at 4 °C.
- Remove the supernatant.
NOTE: All the above steps should be performed in a laminar flow clean hood to prevent contamination.
- Measure the cell concentration and viability using a cell counter (see the Table of Materials).
- Modifying the amino group of cellular proteins with monomeric acrylamide
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 1.5 h- Add 1 mL of Amino Group Modification solution (see Table 1) to the cell pellet and suspend the cell pellet by gentle pipetting.
- Incubate the tube on ice for 1 h and centrifuge the cell suspension at 240 × g for 5 min at 4 °C.
- Remove the supernatant and resuspend the cells with 100 mL of 4% acrylamide/1 mM EDTA/PBS containing an intercalator dye for DNA (see Table 1, 1 cell/µL).
8. Preparation of reusable single cells
- Preparation of reusable single cells (manual version)
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 9 h/96 cells- Mix 1 mL of the cell suspension (1 cell/µL) and 199 mL of 1 mM EDTA/PBS containing an intercalator dye for DNA (see Table 1).
- Transfer 200 µL of the cell suspension to each well of flat-bottom 96-well plates (total 10 plates, see the Table of Materials).
- Put the cover onto the 96-well plate and scan the 10 plates using a scanning microscope (see the Table of Materials) to identify wells containing a single cell.
- Transfer the contents of the well containing a single cell to a PCR tube.
- Tilt the plate (toward the operator), and wait for a few minutes until the single cell has sunk to the lower edge of the well.
- Place the pipette tip on the bottom corner and transfer the single cell and the buffer (aspirate 210 µL/well = 200 µL of buffer + 10 µL of air) to a PCR tube.
NOTE: Be sure to use a P200 low-retention tip. - Check the well using a fluorescence microscope to confirm the transfer.
- If a single cell is still in the well, add 200 µL/well of 1 mM EDTA/PBS containing an intercalator dye for DNA.
- Then, repeat the transfer.
- Centrifuge the PCR tube at 240 × g for 5 min at 4 °C using a swing rotor without brake.
NOTE: Braking causes a swirling flow of water in the tube, which disrupts the precipitation of the single cell. When centrifuging with a swinging rotor without brake, the single cell will always sink to the bottom of the tube. However, if an angled rotor is used, the single cell attaches to the sidewall of the PCR tube and could be lost. - Remove 195 µL of the supernatant with a very slow pipetting speed.
- Add 195 µL/tube of Acrylamide/Bis-acrylamide/APS solution (see Table 1).
- Centrifuge the PCR tube at 240 × g for 5 min at 4 °C using a swing rotor without brake.
- Remove 195 µL of the supernatant with very slow pipetting speed.
- Transfer the bottom 3 µL to the PCR tube containing 4% TEMED/mineral oil and a magnetic polyacrylamide bead (generated in step 6).
NOTE: Be sure to use a P10 low-retention tip. Dispense the cell near the magnetic polyacrylamide bead. In the mineral oil, the liquid containing the single cell adheres to the surface of the magnetic polyacrylamide bead by surface tension. - Incubate overnight at room temperature.
NOTE: This process generates a bilayer acrylamide gel bead. The core is a magnetic gel bead. The outer layer is polyacrylamide gel containing a single cell. The single cell is embedded in the outer layer of the gel. Safe stopping point: After washing the reusable single cells with 50% glycerol/1 mM EDTA/0.05% Tween 20/0.5% BSA/TBS buffer, the reusable single cells can be stored at -20 °C for up to 6 months.
- Preparation of reusable single cells (semiautomated version)
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 3 h/96 cells- Mix 1 mL of cell suspension (1 cell/µL) and 199 mL of 1 mM EDTA/PBS containing an intercalator dye for DNA (see Table 1).
- Transfer 200 µL of the cell suspension to each well of a 4 nL nanowell plate (see Figure 3 and the Table of Materials) and place the 4 nL nanowell plate on an automated single-cell-picking robot (see the Table of Materials).
- Place a 96-well PCR plate as a destination plate on the automated single-cell-picking robot. Ensure that each well contains 200 µL/well Acrylamide/Bis-acrylamide/APS solution (see Table 1).
- Transfer a single cell from a 4 nL nanowell to a well of the 96-well PCR plate (see Supplemental Video S1).
- Put a cover on top of the 96-well PCR plate and centrifuge the 96-well plate at 240 × g for 5 min at 4 °C using a swing rotor without a brake.
- Place the 96-well PCR plate on the deck of an automatic liquid-handling robot (see the Table of Materials, Figure 4, and Supplemental Code 1).
- Drag and drop the Supplemental Code 1 to the software window (see the Table of Materials) of the liquid-handling robot.
- Run the program on the automatic liquid-handling robot (see Supplemental Video S2 and Supplemental Video S3).
- Remove 195 µL of the supernatant with a very slow pipetting speed.
- Add 50 µL of 4% TEMED/mineral oil.
NOTE: Steps 8.2.8.1-8.2.8.2 can be performed by manual pipetting.
- Incubate overnight at room temperature (Figure 5). Safe stopping point: After washing the reusable single cells with 50% glycerol/1 mM EDTA/0.05% Tween 20/0.5% BSA/TBS buffer, store the reusable single cells at -20 °C for up to 6 months.
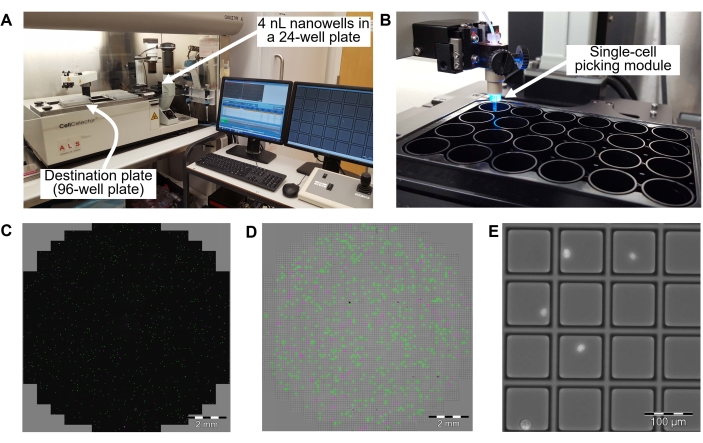
Figure 3: Automatic single-cell picking and transfer into a 96-well PCR plate in step 8.2. (A) Overview of a single cell-picking system. A single cell-picking robot is in a laminar flow clean hood to avoid contamination. (B) A 24-well plate with 4 nL nanowells inside the well. (C) Cell distribution in a well from the 24-well plate. Green dots are cells identified as a single cell in each 4 nL nanowell. Magenta dots are cells identified as doublets or multiplets of cells. (D) Brightfield image of the well in the 24-well plate. A green square is a 4 nL nanowell containing a single cell. A magenta square is a 4 nL nanowell containing multiple cells. (E) Magnified field of some 4 nL nanowells. Bright dots are single cells in 4 nL nanowells. The single cell-picking system identifies nanowells containing a single cell by acquiring brightfield and fluorescence images of cells with DAPI staining. Identified single cells are transferred from the 4 nL nanowell to a well of a 96-well PCR plate. Scale bars = 2 mm (C, D), 100 µm (E). Abbreviation = DAPI = 4',6-diamidino-2-phenylindole. Please click here to view a larger version of this figure.
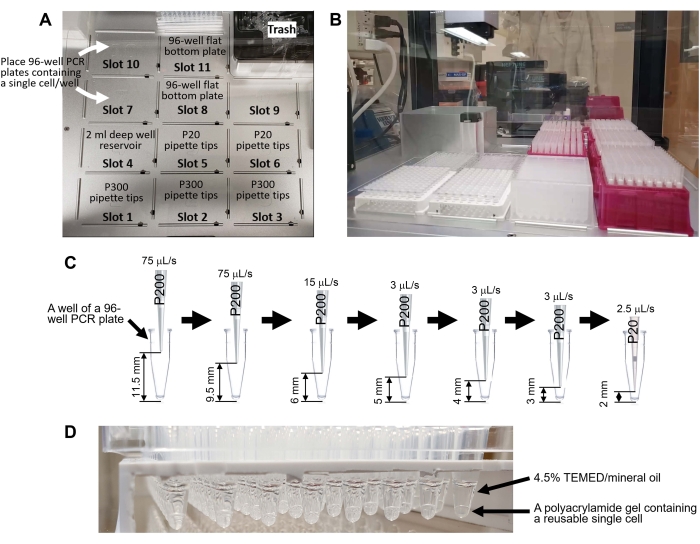
Figure 4: Generating reusable single cells using a liquid-handling robot. (A) A deck of the liquid-handling robot. The deck has 11 slots for pipette tip racks (P300 tip: Slots 1-3, P20 tip: Slots 5-6), 2 mL deep-well 96-well plate (Slot 4), two 96-well PCR plates containing a single cell per well (Slots 7 and 10), and two flat-bottom 96-well plates for liquid waste (Slots 8 and 11). (B) The deck after placing the labware. (C) Schematic representation of robotic pipetting in step 8.8.1. The program removes the supernatant without aspirating a single cell from the bottom of the 96-well PCR plate. (D) Reusable single cells generated using the Supplemental Code 1. Please click here to view a larger version of this figure.
9. Random primer annealing and extension
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Asterisks (*) at the following steps indicate that a magnetic separator can be used to control the position of the polyacrylamide beads containing a reusable single cell in the tube. However, the use of the magnetic separator is not essential. By moving down the pipette tip slowly along the wall of the tube, the beads are pushed up for washing or buffer exchange. Time: 9 h
- Remove the mineral oil by pipetting(*) and wash (*) 5 times with 200 µL of 1x TP Mg(-) buffer (dilute 10x TP Mg(-) buffer to 1x buffer with ultrapure water, see Table 1).
- Remove the supernatant (*) and add 15 µL of Annealing buffer (see Table 1).
- Incubate for 1 h on ice.
NOTE: The purpose of this incubation is to permeabilize the plasma and nuclear membranes and deliver the random primer to the cell nucleus. - Place the PCR tubes on a thermal cycler and heat at 94 °C for 3 min.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 20 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is 105 °C. - Transfer the tubes to an ice-cooled metal block and incubate for 2 min.
- Add 4 µL of MgSO4/NaCl/dNTP mix (see Table 1) and mix with a vortex mixer at a medium speed.
- Add 1 µL of Bst polymerase, large fragment (see the Table of Materials) and mix by using a vortex mixer at a medium speed.
- Incubate for 4 h on a shaker (600 rpm at 4 °C).
NOTE: The purpose of this incubation is to deliver the Bst polymerase to the cell nucleus. - Place the PCR tube on a thermal cycler and run one of the following programs.
- Run a 4 h program: 10 °C for 30 min, 20 °C for 30 min, and 25 °C for 180 min.
- Alternatively, run an overnight program: 4 °C for 4 h, 10 °C for 2 h, 20 °C for 2 h, 25 °C for 4 h, and hold at 4 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 25 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is 105 °C. Safe stopping point: Stop the experiment here for up to 1 day by storing the reusable single cells at 4 °C.
10. Antibody binding
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 1.5 h
- Add 1.625 µL of NaCl/EDTA/BSA solution (see Table 1) and mix by vortexing at low speed.
NOTE: This step aims to 1) facilitate chelation of Mg2+ by EDTA, 2) stabilize the binding of the extended 1st random primer by 300 mM NaCl, and 3) block nonspecific binding of antibodies using bovine serum albumin (BSA) in the subsequent reaction. - Incubate for 1 h on ice and add 0.1 µg/mL each of antibody and control IgG conjugated with Antibody Probe (prepared in step 5).
- Incubate overnight on ice with gentle shaking on a shaker.
11. Proximity ligation of antibody probe and proximally extended random primer
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Asterisks (*) in the following steps indicate where a magnetic separator can be used to control the position of polyacrylamide beads containing a reusable single cell in the tube. However, the use of the magnetic separator is not essential. By moving down the pipette tip slowly along the wall of the tube, the beads can be pushed up for washing or buffer exchange. Time: 6 h
- Wash (*) twice with 200 µL of 1x TPM-T buffer (20 min incubation in each wash) on ice. Dilute 10x TPM-T buffer (Tris-HCl/Potassium chloride/Magnesium sulfate/Triton X-100) to 1x with ultrapure water ).
- Remove (*) the supernatant and wash (*) once with 1x T4 DNA ligase buffer (see the Table of Materials).
- Remove (*) the supernatant and add 19 µL of Ligation Adapter Solution (see Table 1).
- Incubate for 1 h at 25 °C.
- Add 1 µL T4 DNA ligase (see the Table of Materials) and mix the tube on a vortex mixer at medium speed.
- Place the tube on a thermal cycler and run the following program: proximity ligation program, 4 h at 16 °C and 30 min at 25 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 25 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is room temperature. - Wash (*) twice briefly with 200 µL of 1x Bst Mg(-) EDTA(+) buffer (see Table 1; prepare 1x buffer from 10x stock buffer) and store the cell at 4 °C, overnight. Safe stopping point: Stop the experiment here for up to 1 day by storing the reusable single cells at 4 °C.
12. Full extension of the 1st random primer
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 4.5 h
- Wash twice with 200 µL 1x Bst Mg(-) EDTA(-) buffer (see Table 1, prepare 1x buffer from 10x stock buffer), remove the supernatant, and add 20 µL of Bst/dNTPs/MgSO4 mixture (see Table 1).
- Mix with a vortex mixer at medium speed and incubate for 4 h on an orbital shaker (at 6 °C and 500 rpm).
- Place the tube on a thermal cycler and run the following program: full-extension program: 10 °C for 1 h, 20 °C for 1 h, 30 °C for 1 h, 40 °C for 1 h, 50 °C for 1 h, 65 °C for 1 h, 94 °C for 10 min, and hold at 4 °C.
NOTE: Safe stopping point: The experiment can be stopped here for up to 1 day by storing the reusable single cells at 4 °C.
13. Multiple displacement amplification
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 2.5 h (steps 13.1-13.2) + 15 min (steps 13.3-13.4) + 1 day (steps 13.5-13.10)
- Add 0.4 µL of 100 µM 2nd random primer (see Table 3) and mix with a vortex mixer at medium speed.
- Incubate for 2 h at 6 °C and 500 rpm on an orbital shaker, and place the tube on a thermal cycler and heat at 94 °C for 3 min.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 25.4 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is 105 °C. - Place the tubes in an ice-cooled metal block and add 1 µL/tube of Bst DNA polymerase.
- Vortex at slow speed, place the tubes on a thermal cycler, and run the following program:
4 °C for 4 h, 10 °C for 30 min, 20 °C for 30 min, 30 °C for 30 min, 40 °C for 30 min, 50 °C for 30 min, 65 °C for 60 min, 94 °C for 3 min, and hold at 4 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 26.4 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is 105 °C. Safe stopping point: Stop the experiment here for up to 1 day by storing the reusable single cells at 4 °C. - Collect the supernatant (approximately 20 µL), and transfer it to a PCR tube.
- Store the collected supernatant at -80 °C.
- Add 20 µL/tube of 0.05% Tween 20/0.1x TE buffer to a PCR tube containing the reusable single cell (see Table 1) and incubate the reusable single cell at 4 °C, overnight. Safe stopping point: Stop the experiment here for a few days by extending the incubation time.
- Collect the supernatant and combine the supernatant with the sample collected at step 13.5.
- Repeat steps 13.7-13.8 once more. Soak the reusable single cell in 50% glycerol/5 mM EDTA/0.5% BSA/0.05% Tween20/TBS buffer and incubate it for 30 min on an orbital shaker (4 °C, 600 rpm). Store the reusable single cell at -20 °C until the next experiment.
- Add 40 µL of Exo- master mix (see Table 1) and place the tube on a thermal cycler and run the following program: i) 95 °C for 5 min, ii) 95 °C for 30 s, iii) 60 °C 30 s, iv) 72 °C for 30 s, v) repeat steps ii-iv 19 times, vi) 72 °C for 5 min, vii) hold at 4 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is approximately 45 µL, including the volume of the polyacrylamide bead. The temperature of the thermal cycler lid is 105 °C. Safe stopping point: The experiment can be stopped here for at least a few days by storing the sample at -80 °C.
14. Phenol-chloroform purification and polyethylene glycol precipitation
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 1.5 h
- Transfer the product to a 0.5 mL DNA low-binding tube and add 100 µL of Phenol:Chloroform:Isoamyl Alcohol.
NOTE: Phenol:Chloroform:Isoamyl Alcohol causes irritation and possibly burns by contact. Wear gloves, safety goggles, and a lab coat. Use only with adequate ventilation, or wear an appropriate respirator. Follow the institutional safety guidelines. Discard the used labware according to the institutional guidelines. - Shake for 30 s by hand and centrifuge at 12,000 × g for 10 min at 4 °C.
- Collect the aqueous phase (80 µL) into a 0.5 mL DNA low-binding tube, and add 40.84 µL/tube of Linear acrylamide/MgCl2 mixture (see Table 1).
- Add 47.06 µL/tube of 50% (w/v) PEG8000 (RNase-free) and mix by pipetting.
- Incubate for 20 min at room temperature and centrifuge at 240 × g for 10 min at room temperature.
- Remove the supernatant and add 400 µL/tube of 80% ethanol (EtOH).
- Wash with 80% EtOH, remove the supernatant using an aspirator, and air-dry the pellet.
- Suspend the pellet with 20 µL of 1 mM EDTA/10 mM Tris-HCl, pH 7.4 buffer, and store the solution at -80 °C.
NOTE: Measure the DNA concentration using a double-stranded DNA-specific intercalator dye (see the Table of Materials). Safe stopping point: The experiment can safely be stopped here for at least a week.
15. In vitro transcription
NOTE: The following procedure is performed under a clean hood to avoid DNase and RNase contamination. Time: 5 h
- Thaw DNA of single cell-derived products (from step 14) and prepare a mixed library of single cell-derived products by mixing 2 µL/cell of the DNA products (total volume 20 µL).
- Add 26 µL of In vitro transcription master mix (see the Table of Materials and the manufacturer's protocol) and mix by pipetting.
- Place the PCR tube on a thermal cycler and incubate for 4 h at 37 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is 46 µL. The temperature of the thermal cycler lid is 105 °C. - Add 5 µL of 10x DNase I buffer (see the Table of Materials) and add 4 µL of DNase I (RNase-free, 4 U, see the Table of Materials).
- Mix and incubate for 15 min at 37 °C, and transfer the sample to a 0.5 mL tube.
- Add 300 µL/tube of Guanidinium thiocyanate-phenol-chloroform (see the Table of Materials) and mix by gentle vortexing.
NOTE: Guanidinium thiocyanate-phenol-chloroform can lead to serious chemical burns by contact. Wear gloves, safety goggles, and a lab coat. Use only with adequate ventilation or wear an appropriate respirator. Follow institutional safety guidelines. Discard the used labware according to the institutional guidelines. - Incubate for 5 min at R.T. on a shaker and then store samples at -80 °C for up to 3 days.
NOTE: Safe stopping point: The experiment can safely be stopped here for up to 3 days.
16. RNA purification
NOTE: The following procedure is performed under a clean hood to avoid DNase and RNase contamination. Time: 2 h
- Add 80 µL/tube of chloroform to the sample from step 15.7 and shake the tube vigorously by hand for 15 s.
- Incubate for 2-15 min at room temperature and centrifuge the sample at 12,000 × g for 15 min at 4 °C.
- Collect and transfer the aqueous phase of the sample (~200 µL) to a new 1.5 mL tube.
- Add 600 µL/tube of Guanidinium thiocyanate-phenol-chloroform and transfer the sample to a 1.5 mL tube.
- Add 180 µL/tube of Chloroform and shake the tube vigorously by hand for 15 s.
- Centrifuge the sample at 12,000 × g for 10 min at 4 °C.
- Collect and transfer the aqueous phase of the sample (~450 µL) to a 1.5 mL tube.
- Add 60 µL/tube of Linear Acrylamide (5 µg/µL) and add 400 µL/tube of 100% isopropanol to the aqueous phase.
- Incubate the tube at room temperature for 10 min and centrifuge it at 12,000 × g for 10 min at 4 °C.
- Remove the supernatant carefully.
- Wash the RNA pellet 3 times with 200 µL of 75% ethanol (EtOH/RNase-free water), remove the supernatant, and air-dry the RNA pellet.
NOTE: Do not allow the RNA to dry completely because the pellet can lose solubility. - Resuspend the RNA pellet in 20 µL of RNase-free water containing an RNAse inhibitor (1 µL/20 µL) and dissolve the pellet by pipetting.
- Measure the absorbance at 260 nm.
17. Reverse transcription
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 1 h
- Add 7 µL/tube of Reverse Transcription primer + dNTP mix (see Table 1 and Table 3) and place the tubes on a thermal cycler.
NOTE: The tube size is 0.2 mL. The volume of the solution is 27 µL. The temperature of the thermal cycler lid is 105 °C. - Heat at 65 °C for 5 min and place the tubes on ice for at least 1 min.
- Add 14 µL/tube of Reverse transcriptase master mix (see Table 1) and mix by pipetting.
- Place the tubes on a thermal cycler, and incubate them at 55 °C for 10 min and then at 80 °C for 10 min.
NOTE: The tube size is 0.2 mL. The volume of the solution is 60 µL. The temperature of the thermal cycler lid is 105 °C.
18. Second strand synthesis
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 2.5 h
- Add 40 µL/tube of ultrapure water and split the 100 µL solution into two 0.2 mL PCR tubes (50 µL/tube).
- Add 60 µL/tube of Second Strand Synthesis mix (see Table 1) and place the tubes on a thermal cycler and run the following program: i) 95 °C for 5 min, ii) 95 °C for 30 s, iii) 60 °C for 30 s, iv) 72 °C for 30 s, v) repeat steps ii-iv 20 times, and vi) hold at 4 °C.
NOTE: The tube size is 0.2 mL. The volume of the solution is 110 µL. The temperature of the thermal cycler lid is 105 °C. - Add 4.4 µL/tube of 0.5 M EDTA (final 20 mM) and store at -80 °C, overnight.
NOTE: Safe stopping point: The experiment can safely be stopped here for up to a few days. - Purify DNA by phenol-chloroform purification and polyethylene glycol precipitation (described in step 14).
NOTE: Safe stopping point: The experiment can safely be stopped here for at least one week.
19. Restriction enzyme digestion and size selection
NOTE: The following procedure is performed under a clean hood to avoid DNase contamination. Time: 3 h (steps 19.1-19.7)
- Measure the DNA concentration of the purified DNA (from step 18.4) by measuring the absorbance at 260 nm.
- Transfer 6 µg of DNA to a PCR tube, add 30 µL of 10x Digestion buffer, and adjust the volume to 294 µL with ultrapure water.
- Add 6 µL of BciVI restriction enzyme and incubate at 37 °C for 1 h.
- Perform EtOH precipitation with Linear polyacrylamide.
- Add 60 µL/tube of 3 M sodium acetate (pH 5.2) and then add 40 µL/tube of Linear acrylamide (5 mg/mL, see the Table of Materials).
- Add 400 µL/tube of EtOH and incubate overnight at -20 °C.
NOTE: Safe stopping point: The experiment can safely be stopped here for one day. - Centrifuge 12,000 × g for 10 min at 4 °C and remove the supernatant.
- Wash the pellet twice with 80% EtOH and dry the pellet.
- Dissolve the pellet with 20 µL of 1xTE buffer and add 4 µL/tube of fresh 6x Gel Loading buffer.
- Load the sample in a 5% agarose gel/0.5x TAE buffer and perform electrophoresis (50 V for 40 min).
- Cut and collect the gel over 50 bp (see Figure 6) and extract the DNA using a Gel Extraction kit.
NOTE: Safe stopping point: The experiment can safely be stopped here for at least one week. - Construct the sequencing library using a DNA library preparation kit (see the Table of Materials)16,17.
Results
K562 single cells were generated using the protocol described in step 8 (see Figure 5). Single cells were embedded in the outer layer of the polyacrylamide bead. Cell DNA was stained and visualized using an intercalator dye for DNA staining.
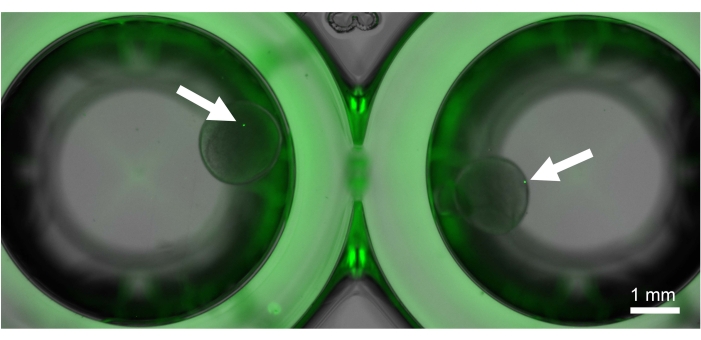

Figure 5: Generated reusable single cells. Cells are sta...
Discussion
This article describes the step-by-step protocol for the recently reported single-cell multiepigenomic analysis using reusable single cells7. In the subsequent paragraphs, we discuss critical points, emphasizing potential limitations in the protocol.
One of the critical points throughout the protocol (from steps 7.2-13) is avoiding DNase contamination. A single cell only has two copies of genomic DNA. Therefore, damaging genomic DNA critically reduces the signal number...
Disclosures
Drs. Ohnuki and Tosato are co-inventors on a patent entitled "Methods for preparing a reusable single cell and methods for analyzing the epigenome, transcriptome and genome of a single cell" (EP3619307 and US20200102604). The patent application was filed in part based on preliminary results related to the technology described in the current manuscript. The invention or inventions described and claimed in this patent application were made while the inventors were full-time employees of the U.S. Government. Therefore, under 45 Code of Federal Regulations Part 7, all rights, title, and interest to this patent application have been or should by law be assigned to the U.S. Government. The U.S. Government conveys a portion of the royalties it receives to its employee inventors under 15 U.S. Code § 3710c.
Acknowledgements
We thank Drs. David Sanchez-Martin and Christopher B. Buck for comments during the conceptualization stage of the project. We also thank the Genomics Core, Center for Cancer Research, National Cancer Institute, National Institutes of Health for help in preliminary experiments, and the Collaborative Bioinformatics Resource, CCR, NCI, NIH for advice in computational analysis. We thank Ms. Anna Word for helping with the optimization of DNA polymerases used in the method. This work utilized the computational resources of the NIHHPC Biowulf cluster (http://hpc.nih.gov). This project is supported by the Intramural Program of the Center for Cancer Research, National Cancer Institute, National Institutes of Health, the NCI Director's Innovation Award (#397172), and Federal funds from the National Cancer Institute under Contract No. HHSN261200800001E. We thank Drs. Tom Misteli, Carol Thiele, Douglas R. Lowy, and all members of Laboratory of Cellular Oncology for productive comments.
Materials
| Name | Company | Catalog Number | Comments |
| 10x CutSmart buffer | New England BioLabs | B6004 | 10x Digestion buffer |
| 200 proof ethanol | Warner-Graham Company | 200 proof | Ethanol |
| 5-Hydroxymethylcytosine (5-hmC) Monoclonal Antibody [HMC/4D9] | Epigentek | A-1018-100 | Anti-5hmC |
| Acridine Orange/Propidium Iodide Stain | Logos Biosystems | F23001 | Cell counter |
| Acrylamide solution, 40% in H2O, for molecular biology | MilliporeSigma | 01697-500ML | 40% acrylamide solution |
| All-in-One Fluorescence Microscope BZ-X710 | Keyence | BZ-X710 | Scanning microscope |
| Amicon Ultra-0.5 Centrifugal Filter Unit | MilliporeSigma | UFC510024 | Ultrafiltration cassette |
| Ammonium persulfate for molecular biology | MilliporeSigma | A3678-100G | Ammonium persulfate powder |
| Anhydrous DMF | Vector laboratories | S-4001-005 | Anhydrous N,N-dimethylformamide (DMF) |
| Anti-RNA polymerase II CTD repeat YSPTSPS (phospho S5) antibody [4H8] | Abcam | ab5408 | Anti-Pol II |
| Anti-TRAP220/MED1 (phospho T1457) antibody | Abcam | ab60950 | Anti-Med1 |
| BciVI | New England BioLabs | R0596L | BciVI |
| Bovine Serum Albumin solution, 20 mg/mL in H2O, low bioburden, protease-free, for molecular biology | MilliporeSigma | B8667-5ML | 20% BSA (Table 7) |
| Bst DNA Polymerase, Large Fragment | New England BioLabs | M0275L | Bst DNA polymerase |
| BT10 Series 10 µl Barrier Tip | NEPTUNE | BT10 | P10 low-retention tip |
| CellCelector | Automated Lab Solutions | N/A | Automated single cell picking robot |
| CellCelector 4 nl nanowell plates for single cell cloning, Plate S200-100 100K, 24 well,ULA | Automated Lab Solutions | CC0079 | 4 nL nanowell plate |
| Chloroform | MilliporeSigma | Chloroform | |
| Corning Costar 96-Well, Cell Culture-Treated, Flat-Bottom Microplate | Corning | 3596 | Flat-bottom 96-well plates |
| Deep Vent (exo-) DNA Polymerase | New England BioLabs | M0259L | Exo- DNA polymerase |
| DNA LoBind Tubes, 0.5 mL | Eppendorf | 30108035 | 0.5 mL DNA low-binding tube |
| DNA Oligo, 1st random primer | Integrated DNA Technologies | N/A, see Table 3 | 1st random primer |
| DNA Oligo, 2nd random primer Cell#01 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#02 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#03 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#04 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#05 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#06 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#07 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#08 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#09 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#10 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#11 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd random primer Cell#12 | Integrated DNA Technologies | N/A, see Table 3 | 2nd random primer |
| DNA Oligo, 2nd synthesis primer | Integrated DNA Technologies | N/A, see Table 3 | 2nd synthesis primer |
| DNA Oligo, Ligation Adaptor | Integrated DNA Technologies | N/A, see Table 3 | Ligation Adaptor |
| DNA Oligo, Reverse Transcription primer | Integrated DNA Technologies | N/A, see Table 3 | Reverse Transcription primer |
| DNase I (RNase-free) | New England BioLabs | M0303L | DNase I (RNase-free, 4 U). |
| DNase I Reaction Buffer | New England BioLabs | B0303S | 10x DNase I buffer (NEB) |
| dNTP Mix (10 mM each) | Thermo Fisher | R0192 | 10 mM dNTPs |
| Fetal Bovine Serum, USA origin, Heat-inactivated | MilliporeSigma | F4135-500ML | Fetal bovine serum |
| HiScribe T7 High Yield RNA Synthesis Kit | New England BioLabs | E2040S | In-vitro-transcription master mix |
| Histone H3K27ac antibody | Active motif | 39133 | Anti-H3K27ac |
| Histone H3K27me3 antibody | Active motif | 39155 | Anti-H3K27me3 |
| IgG from rabbit serum | Millipore Sigma | I5006-10MG | Control IgG |
| Iron oxide(II,III) magnetic nanopowder, 30 nm avg. part. size (TEM), NHS ester functionalized | MilliporeSigma | 747467-1G | NHS ester functionalized 30 nm iron oxide powder |
| K-562 | American Type Culture Collection (ATCC) | CCL-243 | cells |
| Linear Acrylamide (5 mg/mL) | Thermo Fisher | AM9520 | Linear Acrylamide |
| LUNA-FL Dual Fluorescence Cell Counter | Logos Biosystems | L20001 | Cell counter |
| LUNA Cell Counting Slides, 50 Slides | Logos Biosystems | L12001 | Cell counter |
| Mineral oil, BioReagent, for molecular biology, light oil | MilliporeSigma | M5904-500ML | Mineral oil |
| N,N,N′,N′-Tetramethylethylenediamine for molecular biology | MilliporeSigma | T7024-100ML | N,N,N′,N′-Tetramethylethylenediamine |
| NaCl (5 M), RNase-free | Thermo Fisher | AM9760G | 5M NaCl |
| NanoDrop Lite | Thermo Fisher | 2516 | Microvolume spectrophotometer |
| NEST 2 mL 96-Well Deep Well Plate, V Bottom | Opentrons | N/A | 2 mL deep well 96-well plate |
| Non-skirted 96-well PCR plate | Genesee Scientific | 27-405 | 96-well PCR plate |
| NuSive GTG Agarose | Lonza | 50081 | Agarose |
| OmniPur Acrylamide: Bis-acrylamide 19:1, 40% Solution | MilliporeSigma | 1300-500ML | 40%Acrylamide/Bis-acrylamide |
| OT-2 lab robot | Opentrons | OT2 | Automated liquid handling robot |
| Paraformaldehyde, EM Grade, Purified, 20% Aqueous Solution | Electron Microscopy Sciences | 15713 | 20% Pararmaldehyde |
| PBS (10x), pH 7.4 | Thermo Fisher | 70011044 | 10x PBS |
| PIPETMAN Classic P1000 | GILSON | F123602 | A P1000 pipette |
| Protein LoBind Tubes, 1.5 mL | Eppendorf | 925000090 | 1.5 mL Protein low-binding tube |
| QIAgen Gel Extraction kit | Qiagen | 28706 | A P1000 pipette |
| Quant-iT PicoGreen dsDNA Assay | Thermo Fisher | P11495 | dsDNA specific intercalator dye |
| Quick Ligation kit | New England BioLabs | M2200L | T4 DNA ligase (NEB) |
| RNaseOUT Recombinant Ribonuclease Inhibitor | Thermo Fisher | 10777019 | RNAse inhibitor |
| S-4FB Crosslinker (DMF-soluble) | Vector laboratories | S-1004-105 | Succinimidyl 4-formylbenzoate (S-4FB) |
| S-HyNic | Vector laboratories | S-1002-105 | Succinimidyl 6-hydrazinonicotinate acetone hydrazone (S-HyNic) |
| Sodium Acetate, 3 M, pH 5.2, Molecular Biology Grade | MilliporeSigma | 567422-100ML | 3M Sodium acetate (pH 5.2) |
| Sodium bicarbonate, 1M buffer soln., pH 8.5 | Alfa Aesar | J60408 | 1M sodium bicarbonate buffer, pH 8.5 |
| Sodium phosphate dibasic for molecular biology | MilliporeSigma | S3264-250G | Na2HPO4 |
| Sodium phosphate monobasic for molecular biology | MilliporeSigma | S3139-250G | NaH2PO4 |
| SuperScript IV reverse transcriptase | Thermo Fisher | 18090050 | Reverse transcriptase |
| SYBR Gold Nucleic Acid Gel Stain (10,000x Concentrate in DMSO) | Thermo Fisher | S11494 | An intercalator dye for DNA |
| T4 DNA Ligase Reaction Buffer | New England BioLabs | B0202S | 10x T4 DNA ligase reaction buffer |
| ThermoPol Reaction Buffer Pack | New England BioLabs | B9004S | 10x TPM-T buffer (Tris-HCl/Pottasium chloride/Magnesium sulfate/Triton X-100) |
| TRIzol LS reagent | Thermo Fisher | 10296-028 | Guanidinium thiocyanate-phenol-chloroform extraction |
| TruSeq Nano DNA library prep kit | Illumina | 20015965 | A DNA library preparation kit (see also the manufacturer's instruction) |
| Ultramer DNA Oligo, Anti-5hmC_Ab#005 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for antibody |
| Ultramer DNA Oligo, Anti-H3K27ac_Ab#002 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for antibody |
| Ultramer DNA Oligo, Anti-H3K27me3_Ab#003 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for antibody |
| Ultramer DNA Oligo, Anti-Med1_Ab#004 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for antibody |
| Ultramer DNA Oligo, Anti-Pol II_Ab#006 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for antibody |
| Ultramer DNA Oligo, Control IgG_Ab#001 | Integrated DNA Technologies | N/A, see Table 3 | An amine-modified DNA probe for control IgG |
| UltraPure 0.5 M EDTA, pH 8.0 | Thermo Fisher | 15575020 | 0.5M EDTA, pH 8.0 |
| UltraPure DNase/RNase-Free Distilled Water | Thermo Fisher | 10977023 | Ultrapure water |
| Zeba Splin Desalting Columns, 7K MWCO, 0.5 mL | Thermo Fisher | 89882 | Desalting column |
References
- Perkel, J. M. Single-cell analysis enters the multiomics age. Nature. 595 (7868), 614-616 (2021).
- Clark, S. J., et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nature Communications. 9 (1), 781 (2018).
- Ma, S., et al. Chromatin potential identified by shared single-cell profiling of RNA and Chromatin. Cell. 183 (4), 1103-1116 (2020).
- Zhu, C., et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nature Structural & Molecular Biology. 26 (11), 1063-1070 (2019).
- Zhu, C., et al. Joint profiling of histone modifications and transcriptome in single cells from mouse brain. Nature Methods. 18 (3), 283-292 (2021).
- Mimitou, E. P., et al. Scalable, multimodal profiling of chromatin accessibility, gene expression and protein levels in single cells. Nature Biotechnology. 39 (10), 1246-1258 (2021).
- Ohnuki, H., Venzon, D. J., Lobanov, A., Tosato, G. Iterative epigenomic analyses in the same single cell. Genome Research. 31 (10), 1819-1830 (2021).
- Tosato, G., Ohnuki, H. . Methods of preparing a re-usable single cell and methods fro analyzing the epigenome, transcriptome, and genome of a single cell. , (2021).
- Harada, A., et al. A chromatin integration labelling method enables epigenomic profiling with lower input. Nature Cell Biology. 21 (2), 287-296 (2019).
- Egelhofer, T. A., et al. An assessment of histone-modification antibody quality. Nature Structural & Molecular Biology. 18 (1), 91-93 (2011).
- Marina, R. J., et al. TET-catalyzed oxidation of intragenic 5-methylcytosine regulates CTCF-dependent alternative splicing. EMBO Journal. 35 (3), 335-355 (2016).
- Weast, R. C., Weast, R. C., Astle, M. J., Beyer, W. H., et al. . Handbook of chemistry and physics: a ready-reference book of chemical and physical data. 56th edn. , (1975).
- Aydin, S. A short history, principles, and types of ELISA, and our laboratory experience with peptide/protein analyses using ELISA. Peptides. 72, 4-15 (2015).
- Gan, S. D., Patel, K. R. Enzyme immunoassay and enzyme-linked immunosorbent assay. Journal of Investigative Dermatology. 133 (9), 12 (2013).
- Porstmann, T., Kiessig, S. T. Enzyme immunoassay techniques. An overview. Journal of Immunological Methods. 150 (1-2), 5-21 (1992).
- Chao, H. P., et al. Systematic evaluation of RNA-Seq preparation protocol performance. BMC Genomics. 20 (1), 571 (2019).
- Song, Y., et al. A comparative analysis of library prep approaches for sequencing low input translatome samples. BMC Genomics. 19 (1), 696 (2018).
- Rotem, A., et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nature Biotechnology. 33 (11), 1165-1172 (2015).
- Ku, W. L., et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nature Methods. 16 (4), 323-325 (2019).
- Carter, B., et al. Mapping histone modifications in low cell number and single cells using antibody-guided chromatin tagmentation (ACT-seq). Nature Communications. 10 (1), 3747 (2019).
- Maskell, D. P., et al. Structural basis for retroviral integration into nucleosomes. Nature. 523 (7560), 366-369 (2015).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved