A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Direct Measurement of KDM1A Target Engagement Using Chemoprobe-based Immunoassays
In This Article
Summary
Here, we present a protocol to measure KDM1A target engagement in a human or animal cell, tissue or blood samples treated with KDM1A inhibitors. The protocol employs chemoprobe tagging of the free KDM1A enzyme and direct quantification of the target occupation using chemoprobe-based immunoassays and can be used in preclinical and clinical studies.
Abstract
The assessment of the target engagement, defined as the interaction of a drug with the protein it was designed for, is a basic requirement for the interpretation of the biological activity of any compound in drug development or in basic research projects. In epigenetics, target engagement is most often assessed by the analysis of proxy markers instead of measuring the union of the compound to the target. Downstream biological readouts that have been analyzed include the histone mark modulation or gene expression changes. KDM1A is a lysine demethylase that removes methyl groups from mono- and dimethylated H3K4, a modification associated with the silencing of gene expression. Modulation of the proxy markers is dependent on the cell type and function of the genetic make-up of the cells investigated, which can make interpretation and cross-case comparison quite difficult. To circumvent these problems, a versatile protocol is presented to assess the dose effects and dynamics of the direct KDM1A target engagement. The assay described makes use of a KDM1A chemoprobe to capture and quantify uninhibited enzyme, can be broadly applied to cells or tissue samples without the need for genetic modification, has an excellent window of detection, and can be used both for basic research and analysis of clinical samples.
Introduction
Lysine specific demethylase 1 (KDM1A)1 is a demethylase involved in the control of gene transcription. This protein has emerged as a candidate pharmacological target2 in oncology; including Acute Myeloid Leukemia3 (AML), Myelodysplasia Syndrome (MDS)4, Myelofibrosis (MF)5,6, Small Cell Lung Cancer (SCLC)7; in Sickle Cell Disease (SCD)8,9, and in central nervous system diseases including Alzheimer's Disease (AD), Multiple Sclerosis (MS); and in aggression10.
Most of the KDM1A inhibiting compounds in clinical development are cyclopropylamine derivatives and inhibit the protein by covalent binding to its flavin adenine dinucleotide (FAD) cofactor11. Inhibition of KDM1A induces gene expression changes, but these changes vary enormously across tissues, cell types, or disease cases. Inhibition of KDM1A also changes histone marks12, yet these changes are generally produced locally at a specific site in the genome, and are again, highly tissue and cell specific.
The protocol was developed to directly measure KDM1A target engagement in biological samples and has been optimized for the use with cyclopropylamine derived inhibitors. The assay is based on ELISA technology and analyzes, in parallel, Total and Free (i.e. unbound by inhibitor) KDM1A in a native protein extract from a biological sample in a solid phase assay. As a first step, the biological sample is lysed in the presence of the biotinylated KDM1A selective chemoprobe OG-88113,14, derived from the selective KDM1A inhibitor ORY-1001 (iadademstat), a potent inhibitor of KDM1A in clinical development for the treatment of oncological disease. The chemoprobe has an IC50 for KDM1A of 120 nM and includes a FAD binding moiety linked to a biotinylated polyethylene glycol (PEG)-tail. The chemoprobe exclusively binds to the free KDM1A, but not to the inhibitor-bound KDM1A in the sample. After the chemoprobe binding, the KDM1A containing complexes in the sample are captured on microtiter plates with streptavidin coated surface to determine free KDM1A, or on plates coated with a monoclonal anti-KDM1A capture antibody to determine total KDM1A. After washing, both plates are incubated with an anti-KDM1A detection antibody, washed again, and incubated with a secondary HRP-conjugated donkey anti-rabbit IgG antibody for detection using a luminescent substrate and quantification by measuring relative light units (RLU) in a luminometer (Figure 1).
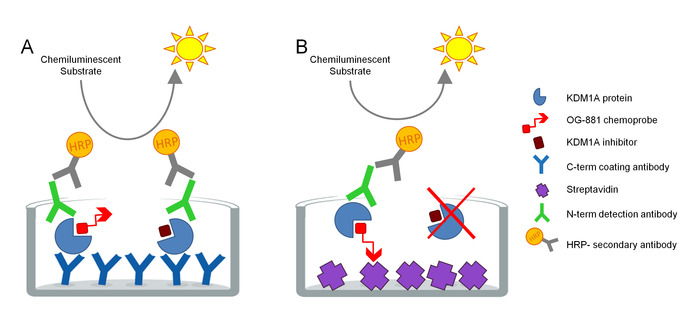
Figure 1. Schema of ELISA Enzyme linked chemoprobe immunoabsorbent assay for KDM1A target engagement: A) Determination of total KDM1A using sandwich ELISA and B) Determination of free KDM1A using chemoprobe ELISA. Please click here to view a larger version of this figure.
A standard curve is included in both ELISA plates to verify the linearity of each assay. The determination of KDM1A target engagement in each sample is then calculated as a relative value to the pre-dose or vehicle treated sample.
Protocol
Blood samples were obtained from the Instituto de Investigación Biomédica Sant Pau Biobank according to Spanish legislation (Real Decreto de Biobancos 1716/2011) and approval of the local ethics committees. Studies with animal tissues were performed in accordance with the institutional guidelines for the care and use of laboratory animals (European Communities Council Directive 86/609/EEC) established by the Ethical Committee for Animal Experimentation at the PRAAL-PCB.
1. Preparation of biological samples for the assay.
CAUTION: This protocol involves manipulation of biological samples which may be subjected to the Occupational Safety and Health Administration (OSHA) Blood Borne Pathogens standard (29 CFR 1910.1030), Directive 2000/54/EC of the European Parliament and of the Council of 18 September 2000 or equivalent regulations. In addition, the biological samples may contain traces of biologically active investigational chemical compounds and the protocol may involve further manipulation of such compounds. Review the safety data sheet (SDS) of the compounds used prior to the initiation of the experiment and strictly observe all applicable safety measures established in the research center, including the use of adequate personal protective equipment (PPE). Wear proper protective clothing and use proper shielding during the course of the experiment. Discard residues in the appropriate waste containers (biological/cytotoxic waste).
NOTE: This protocol starts with cells or samples of subjects treated with a KDM1A inhibitor and their untreated or vehicle/placebo treated controls3.
- Cells treated with vehicle or KDM1A inhibitor in vitro
- For the cells grown in suspension, as 10 mL cultures, transfer the suspensions into clean 15 mL conical tubes and proceed to 1.1.3.
- For the adherent cells (grown in 75 cm2 flasks), remove the medium from the flask and wash briefly using 4 mL PBS. Detach the cells from their vessels using 1.5 mL of 0.5% Trypsin-EDTA during 2 - 5 min (trypsinization conditions may vary, follow provider recommendations for the cell line), add 4 mL PBS and transfer the cells into clean 15 mL conical tubes.
- Insert the tubes in a bench top centrifuge and collect the cells by centrifugation for 5 min at 400 x g at 4 °C. Remove the supernatant, resuspend the pellet in 1 mL of PBS dispensed using a micropipette and transfer the suspension into a 1.5 mL microcentrifuge tube.
- Insert the samples in a microtube centrifuge and centrifuge them for 5 min at 400 x g at 4 °C. Remove the PBS by aspiration with a micropipette and either keep the pellets on the ice and proceed to Step 2; or freeze the pellets on dry ice and store them at -80 °C until Step 3.
- Samples from subjects or animals treated with vehicle/placebo or KDM1A inhibitor
- Tissues: Cut the tissue in small, ≈1 cm3 pieces using a scalpel. Freeze the tissues pieces in liquid N2 in a Dewar container and store them at -80 °C until Step 3.
- Polymorphic blood mononuclear cells (PBMCs): Dilute 10 mL of fresh blood (process maximum 2 h after blood withdrawal) collected in K2-EDTA tubes with 2 volumes of PBS in a 50 mL conical tube. Isolate the PBMCs from blood using commercially obtained PBMC separation tubes according to the manufacturer’s instructions. Keep pellets on the ice and proceed to Step 3.2; or freeze the pellets on dry ice and store them at -80 °C until Step 3.
NOTE: A wet cell pellet of 20 to 50 μL contains ≈ 1 x 107 cells, depending on the cell size. A wet PBMC pellet obtained from 10 mL of healthy human blood has a volume of ≈ 20 μL and contains ≈ 1 x 107 PBMCs. Tissues or cell pellets can be stored at -80 ºC for up to 6 months.
2. Solution preparation
- Prepare 2 μM OG-881 working solution: Take a 10 μL single-use aliquot of the 20 mM biotinylated probe OG-881 stock solution out of the 4 °C fridge and leave it at room temperature (RT) for 10 min. Prepare the 2 μM working solution by serial dilution of the OG-881 stock solution in PBS, using a micropipette with filter tips and changing the tip between the different dilution steps.
- Prepare 10x Protease inhibitor: dissolve 1 tablet in 1 mL PBS in a microcentrifuge tube.
- Prepare the desired volume of 1x Cell lysis buffer with 25 nM OG-881 chemoprobe. For each mL, mix 100 μL commercially obtained 10x Cell lysis buffer, 150 μL of 10x Protease Inhibitor, 12.5 μL of 2 μM OG-881, and 737.5 μL of Type 1 double distilled water.
- Optionally prepare the desired volume of 1x Cell lysis buffer but with 25 nM ORY-1001 instead of OG-881 as in Step 2.3. Less potent inhibitors may be used but may require higher concentrations, for use in the positive control with 100% inhibition (see Step 3.5).
NOTE: Take appropriate measures to avoid any unintended contamination of solutions or samples with the OG-881 or KDM1A inhibitor stock solutions. To calculate the desired volume of 1x Cell lysis buffer with 25 nM OG-881, assume that 400 µL is required per 40 mg of pulverized tissue, or 200 µL per wet pellet of 107 cells.
3. Native protein extraction
- From tissues:
- Pulverize and homogenize a ≈ 1 cm3 cube of frozen tissue with a mortar and pestle chilled on dry ice. Aliquot the samples in single-use vials containing ≈ 40 mg of tissue powder, avoid thawing at all times. Proceed to step 3.1.2. for immediate processing or store at -80 °C.
- Resuspend 40 mg of powdered tissue in 400 µL of 1x Cell Lysis buffer with 25 nM OG-881, vortex for 10 s, and force the sample at least five times through an 18-gauge blunt syringe needle until lysis of the tissue is achieved and a turbid light yellow to orange suspension is obtained. Avoid bubble formation.
- Continue to Step 3.3
- From cell pellets (PBMCs and cell lines):
- Resuspend a pellet of ≈ 1 x 107 cells in 200 µL of 1x cell lysis buffer containing 25 nM OG-881. Vortex the samples briefly and keep them on ice for 5 min.
- Sonicate the samples in a sonicator using 3 pulses of 20 s each at 45 kHz; place them on ice for 20 s between pulses.
NOTE: As soon as the biological samples have been resuspended in the 1x cell lysis buffer, keep them on the ice during the rest of the process.
- Keep the samples on ice for an additional 5 min, vortex briefly and centrifuge the samples for 10 min at 14 000 x g in a pre-cooled centrifuge at 4 °C.
- Using a 1 mL micropipette, transfer the supernatants into fresh 1.5 mL microcentrifuge tubes and leave them on the ice during 2 h. Continue to Step 4.
- Optionally, a positive control to simulate 100% target engagement may be prepared as follows:
- Resuspend the cell pellet or powdered tissue from a vehicle or untreated (predose) sample in the required volume of 1x Cell Lysis buffer with 25 nM ORY-1001 and process as outlined in Step 3.1 to 3.3.
- Transfer the supernatants of the positive control into fresh 1.5 mL microcentrifuge tubes and leave them on ice for 1 h. ORY-1001 willfully inhibit KDM1A and block chemoprobe binding.
- Add 5 μL of 2 μM OG-881 working solution to the positive control supernatant (volume for positive control generated from a 40 mg tissue sample) or 2.5 μL of 2 μM OG-881 working solution (volume for positive control generated from a 107-cell sample) to obtain the same OG-881 concentration as the other samples and leave on ice during 2 h. Continue to Step 4.
4. Quantification of native protein using Bradford assay
- Dilute the commercially sourced Bradford Protein Assay reagent 5 times with H2O Type 1 double distilled water. Calculate the volume of the reagent required for the total amount of samples and standards (1 mL per sample or standard + 5 mL excess volume).
- For the Bovine Serum Albumin (BSA) standard curve, prepare one microcentrifuge tube with 1 mL diluted Bradford Protein Assay solution (blank) and seven microcentrifuge tubes with 995 μL of the diluted Bradford Protein Assay solution. Add 5 μL of each of the BSA Standards (concentration ranging from 125 to 2,000 µg/mL) to each of the 7 microcentrifuge tubes and mix them by gently inverting the tubes several times. Incubate for 5 min at RT.
- Transfer the diluted Standards to cuvettes and read the OD of the blank and Bovine Serum Albumin Standard samples in a spectrophotometer at 280 nm.
- For the biological samples, prepare one microcentrifuge tube with 1 mL diluted Bradford Protein Assay solution (blank) and as many microcentrifuge tubes with 999 μL of the diluted Bradford Protein Assay reagent as samples that need to be quantified. Using an automatic P2 micropipette, add 1 μL of native protein extract prepared in Step 3 to each microcentrifuge tube and mix by gently inverting the tubes several times. Incubate the samples 5 min at RT.
- Transfer the volumes to cuvettes and read the OD of the samples in a spectrophotometer at 280 nm.
- Preferentially, proceed immediately to Step 5. Alternatively, store the native protein extracts at -80 °C until Step 5. Avoid freeze thaw cycles.
5. Luminescent ELISAs for Total and Free KDM1A determination
NOTE: Keep the lab temperature constant at 23-24 °C (RT).
- Coating of microtiter plates with capture KDM1A antibody or Streptavidin
- Total KDM1A ELISA: for each plate, prepare 10 mL of KDM1A capture antibody to a final concentration of 2 μg/mL in PBS. Transfer 100 µL into each well of the plate.
- Free KDM1A ELISA: for each plate, prepare 10 mL of streptavidin at 10 µg/mL in PBS. Transfer 100 µL into each well of the plate.
- Top-seal the Total and Free KDM1A ELISA plates with adhesive film and incubate the plates overnight at 4 °C in the refrigerator.
- Washing and Blocking the plates
- Take the plates out of the refrigerator and let them equilibrate for around 45 min at RT before use.
- Prepare 1,000 mL wash buffer (0.1% Tween in PBS) and 50 mL blocking buffer (1% BSA in PBS) per plate.
- Wash the plates 3 times with wash buffer. In this and subsequent steps, tap the plate on paper towels after every washing step to remove residual solution.
- Add 200 µL of blocking buffer per well to both plates, top-seal both plates with an adhesive film and incubate 2 h at RT.
- Biological sample preparation
- Dilute the native protein extracts obtained at the end of Step 3 to the appropriate concentration using PBS. The recommended concentration will vary in function of the level of KDM1A expression in the biological sample. Examples of appropriate ranges are (1) Cell pellets: 0.5 - 10 μg per well. (2) PBMCs: 5 - 30 μg per well. (3) Pulverized tissue (brain, lung, skin): 20 - 100 µg per well. Keep the samples on the ice during the preparation. When possible, run technical triplicate sample analyses.
- Prepare a Standard Curve using human rKDM1A:
- To prepare the KDM1A Standard working solution, pipet the appropriate volume of rKDM1A for a final concentration of 25 pg/µL, add 75 μL of 2 μM OG-881, and complete with 1 x PBS to a total volume of 6 mL in a 15 mL falcon tube. Keep the KDM1A Standard working solution on ice during 1 h and mix the solution gently by inverting the 15 mL falcon tube several times every 20 min.
- Prepare the KDM1A Standard Dilution Series in 1.5 mL microcentrifuge tubes according to Table 1 (Standard Preparation), in volume enough for the triplicate analysis of two 96 well microtiter plates.
| STANDARD SERIES | KDM1A Standard working solution (µL) | |
| (pg KDM1A/well) | PBS (µL) | |
| 2500 for C-* | 800 | - |
| 2500 | 800 | - |
| 1750 | 560 | 240 |
| 1250 | 400 | 400 |
| 750 | 240 | 560 |
| 250 | 80 | 720 |
| 25 | 8 | 792 |
| 0 | 0 | 800 |
| NOTE: | ||
| (1) The volume prepared of each dilution is enough to run in triplicate 2 plates of assay. | ||
| (2) The recommended range is between 2.5 and 5,000 pg / well | ||
| * For the negative control C-, without KDM1A detection antibody | ||
Table 1: Standard Preparation. To prepare the Standard Series of KDM1A protein, pipette the indicated volumes of KDM1A Standard working solution and PBS into eight properly labeled 1.5 mL microcentrifuge tubes.
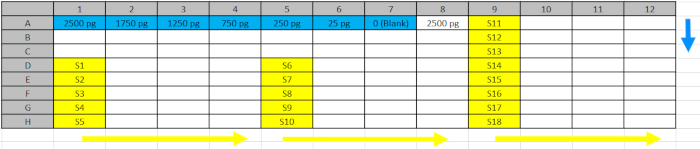
Table 2: Deep Well Plate Design.Standards (blue) and samples (yellow) from step 5.4.2. were pipetted into the reflected positions of the Deep Well Plate to facilitate loading into the ELISA plates following the direction of the blue (standard) and yellow (samples) arrows. Please click here to download this file.
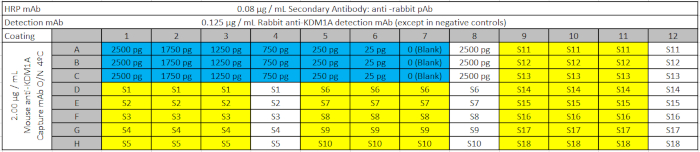
Table 3: ELISA Plate Design. The assay plate includes the standard curve with decreasing amounts of recombinant KDM1A target (in blue); the biological samples (S) in yellow; and corresponding negative controls (contain the samples but not the primary detection antibody) in white, to be loaded on the ELISA plates from the Deep Well Plate. The blank (0 in standard curve) contains all capture and detection reagents but no sample. Please click here to download this file.
- ELISA
- (Continued from Step 5.2.4.) After 2 h of incubation, discard the blocking buffer and wash the plates with wash buffer.
- Transfer the appropriately diluted samples (native protein extracts and standard curve from Step 5.3.) to a refrigerated 96 deep well storage block following the plate distribution shown in Table 2 (Deep Well Plate Design).
- Keep this block on ice until pipetting 100 μL sample / well in the Total and Free ELISA plates following the plate distribution shown in Table 3 (ELISA Plate Design).
- Incubate for 1 h at RT, discard the samples and wash the plates 5 times with wash buffer.
- Prepare 20 mL of rabbit anti-KDM1A detection antibody at 0.125 µg/mL in blocking buffer, add 100 µL per well in each plate of the assay, except in wells corresponding to the negative controls C-. Top-seal the plate and incubate 1 h at RT.
- Discard the detection antibody solution and wash the plates 6 times with wash buffer.
- Prepare 25 mL of secondary goat anti-rabbit antibody HRP to a dilution 1:5,000 in blocking buffer, add 100 µL per well to the microtiter plates; and incubate 1 h at RT.
- Chemiluminescent Detection
- 30 min before the end of Step 5.4.7. and under soft light conditions, mix equal parts of Luminol-Enhancer and Peroxide Solution (10.5 mL: 10.5 mL, for 2 plates) in an amber bottle and leave it at RT.
NOTE: Keep the Luminol Working Solution in an amber bottle and avoid prolonged exposure to any intense light. Short-term exposure to typical laboratory lighting will not harm the working solution. - At least 20 min before measuring the luminescence, switch on the microplate reader at 25 °C and set up readouts to 1,000 ms integration time and 150 ms settle time. Parameter settings may require optimization in function of the instrument.
- After 1 h of incubation in Step 5.4.7., discard the secondary antibody solution and wash the plates 6 times with wash buffer.
- Pipette 100 µL per well of the Luminol Working Solution (Chemiluminescent Substrate) prepared in Step 5.5.1. Pipette very slowly and avoid bubble formation. Use a timer to control the time between the addition of the addition of the solution and the luminescence measurement of the plates and keep this time constant to achieve a good inter assay reproducibility.
- Top-seal the plates and centrifuge to 500 x g at RT for 45 s in a plate centrifuge to eliminate any remaining bubbles. Incubate the plates for 1 min on a plate shaker at 100 rpm.
- Insert the plate inside the reader and leave it for 3 min to stabilize the temperature at 25 °C (without adhesive film). Always start with the Free ELISA plate.
- Read the relative luminescence units (RLU) of each ELISA plate assay (free and total KDM1A).
- Save and copy the Raw RLU values from the Raw Data excel files for further analysis of the results.
- 30 min before the end of Step 5.4.7. and under soft light conditions, mix equal parts of Luminol-Enhancer and Peroxide Solution (10.5 mL: 10.5 mL, for 2 plates) in an amber bottle and leave it at RT.
6. Calculation of the target engagement
- In a spreadsheet software, calculate the RLU Free and RLU Total values of samples SX and reference samples REF (untreated, vehicle or pre-dose sample) from their technical replicate Raw data as detailed below:
- Enter the individual Raw RLUi Total and Raw RLUi Free data from blanks, standard curve, negative controls C- and biological samples (SX and REF) into the analysis datasheet (e.g., Excel). Also, enter the amounts (in pg) of KDM1A from the Standard Curve in the datasheet.
- Calculate the Raw mean RLU, standard deviations σRLU, and coefficient of variation CVRLU from the individual Raw Total and Raw Free RLUi data for each technical replicate datapoint.
- Apply the outlier elimination (example for triplicates): for each individual Raw RLU Total and Raw RLU Free data point RLUi from a technical triplicate datapoint, apply Grubbs criteria when the CV for the triplicate > 0.15, and reject single suspect Raw RLU value when
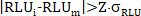 ,
,
whereby Z = 1.148 for n = 3 and 90 % confidence interval (CI). - If outlier elimination was applied, re-calculate the Raw mean RLU, standard deviation σRLU and CVRLU from the non-rejected (nr) Raw RLUi Total and Raw RLUi Free values for each datapoint.
- Apply the background correction: Calculate the mean RLU Free and RLU Total values for each standard sample, and each sample SX and reference sample REF as:


- Graphically represent the data as follows:
- Plot the RLUFree and RLUTotal values (Y-axis) relative to their sample identification (X-axis) in a bar graph.
- Also plot the RLU values (Y-axis) of the standards in a scatter plot relative to their amount of pg of rKDM1A protein (X-axis) for Free and Total measurements, as well as the corresponding lineal trendlines and calculate the r2 (the square of the linear correlation coefficient) values.
- Calculate the target engagement (TE); i.e. the percentage of KDM1A bound by the KDM1A inhibitor in each sample SX relative to a reference sample REF (untreated, vehicle or pre-dose sample) as follows:
- Calculate the ratio R of the mean RLU Free to Total values for the SX and REF samples as:

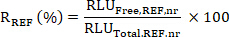
- Then calculate the target engagement (TE) of the sample SX as:
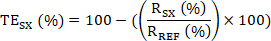
Optional: (1) If N biological replicate experiments were conducted, each with n technical replicates; first calculate the TESX for the technical replicate sets. Subsequently, calculate the mean TE, SD and the CV values for the biological replicate set.
- Calculate the ratio R of the mean RLU Free to Total values for the SX and REF samples as:
- Revise whether the assay acceptance criteria are met: Verify that (1) the assay background is acceptable and the mean Blank < 0.05 x 107 RLU; (2) the sample auto-luminescence is absent and the RLUs of the negative controls C- are below the Lower Limit of Quantification (LLOQ = Mean Blank + 10x SD); (3) the rKDM1A standard curve is linear and r2 ≥ 0.98; (4) the biological samples have RLU values that fall in the dynamic and linear range of the assay i.e. between LLOQ and 2,500 pg/well.
NOTE: Steps 6.1. to 6.4. can be readily automated in a calculus datasheet. - Export the TE data to an open source or commercially obtained statistics software of choice for the graphical representation of the TE values and additional statistical evaluations.
Results
The Linearity of Total and Free KDM1A determination.
A Standard Series was prepared as described in step 5.3.2., using 0 to 2500 pg of full-length human recombinant KDM1A enzyme. The RLU values of Total and Free rKDM1A were assessed to verify the linearity (Figure 2A and 2B). Data are represented as mean from 3 experiments with 3 technical replicates (n) ± SD. ...
Discussion
The protocol presented here was developed to directly measure KDM1A target engagement using a novel KDM1A chemoprobe capture based ELISA. The method has been validated on cultured human cell lines and ex vivo samples from human, rat and mouse and baboon (including PBMCs, lung, brain, skin, tumors), but can be readily applied to other species in which the KDM1A antibody target epitopes and catalytic center are conserved. As OG-881 is an activity based chemoprobe, the sample quality is important and proper manipulation and...
Disclosures
The author Tamara Maes is an executive director and shareholder, and the authors Cristina Mascaró and Raquel Ruiz Rodriguez are employees of Oryzon Genomics S.A. Oryzon Genomics S.A. develops KDM1A inhibitors and holds patents covering compounds and methods used in this Article.
Acknowledgements
This study was financed by Oryzon Genomics. S.A., Hoffman-La Roche, and partially supported by the CIIP-20152001 and RETOS collaboration program RTC-2015-3332-1.
Materials
| Name | Company | Catalog Number | Comments |
| 0,05% Trypsin-EDTA (1X) | Thermo Scientific | #25300-062 | |
| 10 X Protease Inhibitor Tablets | Roche | #11836153001 | |
| 96 deep well storage block | VWR | #734-1679 | |
| 96 well ELISA plates | Nunc | #436110 | |
| Adhesive black Film | Perkin Elmer | #6050173 | |
| Adhesive transparent Film | VWR | #60941-062 | |
| Biotinylated KDM1A probe OG-881 | Oryzon Genomics S.A. | NA | |
| Bovine Serum Albumin | Sigma | # 3117057001 | |
| Bovine Serum Albumin Standard | Thermo Scientific | #23208) | |
| Bradford Protein Assay | BioRad | #500-0001 | |
| Cell lysis buffer 10X | Cell Signaling | #9803 | |
| Centrifuge for 96- well plates | Hettich | Rotina 420R | |
| Flask | Thermo Scientific | #156499 | |
| Full length, enzymatically active human Recombinant LSD1 / KDM1A | Active Motif | #31426 | |
| Graphpad Prism 5 Project | GraphPad Software | NA | |
| Luminol-Enhacer and Peroxide Solution (Chemiluminescent Substrate) | Thermo Scientific | #37074 | |
| Micro Centrifuge | Eppendorf | 5415 R | |
| Microplate reader Infinite 200-Tecan | Tecan | Infinite 200 | |
| Mouse monoclonal capture antibody Anti-KDM1A (N-terminal epitope) | Abcam | #ab53269 | |
| Needle G18 gauge blunt | BD | #303129 | |
| ORY-1001 (iadademstat) | Oryzon Genomics S.A. | NA | |
| PBMC separation tubes 10 ml | Greiner bio-one | #163288 | |
| PBMC separation tubes 50 ml | Greiner bio-one | #227288 | |
| PBS 1x | Sigma | #D8537 | |
| Plate shaker | Heidolph Instruments | Rotamax 120 | |
| Polysorbate 20 | Sigma | #P7949 | |
| Rabbit monoclonal detection antibody Anti-KDM1A (C-terminal epitope) | Cell Signaling | #672184BF-100 | |
| Secondary antibody Peroxidase-conjugated Donkey Anti-rabbit IgG | Thermo Scientific | #31458 | |
| Spectrophotometer cuvette 1.5 | Deltalab | #302100 | |
| Spectrophotometer for cuvette | GE Healthcare | GeneQuant 1300 | |
| Streptavidin | Promega | #Z704A | |
| Syringe | BD | #303172 | |
| Type 1 ultrapure water | Millipore | Milli-Q Advantage A10 | |
| Ultrasonic cleaner | VWR | USC200T |
References
- Shi, Y. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 119 (7), 941-953 (2004).
- Maiques-Diaz, A., Somervaille, T. C. LSD1: biologic roles and therapeutic targeting. Epigenomics. 8 (8), 1103-1116 (2016).
- Maes, T. ORY-1001, a Potent and Selective Covalent KDM1A Inhibitor, for the Treatment of Acute Leukemia. Cancer Cell. 33 (3), 495-511 (2018).
- Sugino, N. A novel LSD1 inhibitor NCD38 ameliorates MDS-related leukemia with complex karyotype by attenuating leukemia programs via activating super-enhancers. Leukemia. 31 (11), 2303-2314 (2017).
- Kleppe, M., Shank, K., Efthymia, P., Riehnhoff, H., Levine, R. L. Lysine-Specific Histone Demethylase, LSD1, (KDM1A) As a Novel Therapeutic Target in Myeloproliferative Neoplasms. Blood. 126, 601 (2015).
- Jutzi, J. S., et al. LSD1 Inhibition Prolongs Survival in Mouse Models of MPN by Selectively Targeting the Disease Clone. HemaSphere. 2 (3), 54 (2018).
- Mohammad, H. P. DNA Hypomethylation Signature Predicts Antitumor Activity of LSD1 Inhibitors in SCLC. Cancer Cell. 28 (1), 57-69 (2015).
- Rivers, A., et al. RN-1, a potent and selective lysine-specific demethylase 1 inhibitor, increases γ-globin expression, F reticulocytes, and F cells in a sickle cell disease mouse model. Experimental Hematology. 43 (7), 546-553 (2015).
- Rivers, A. Oral administration of the LSD1 inhibitor ORY-3001 increases fetal hemoglobin in sickle cell mice and baboons. Experimental Hematology. 67, 60-64 (2018).
- Buesa, C., et al. The dual LSD1/MAO-B inhibitor ORY-2001 prevents the development of the memory deficit in samp8 mice through induction of neuronal plasticity and reduction of neuroinflammation. Alzheimer’s & Dementia. 11 (7), P905 (2015).
- Schmidt, D. M., McCafferty, D. G. Trans-2-Phenylcyclopropylamine is a mechanism-based inactivator of the histone demethylase LSD1. Biochemistry. 46 (14), 4408-4416 (2007).
- Forneris, F., Binda, C., Vanoni, M. A., Battaglioli, E., Mattevi, A. Human histone demethylase LSD1 reads the histone code. Journal of Biological Chemistry. 280 (50), 41360-41365 (2005).
- Gonz#225;lez, E. C., Maes, T., Crusat, C. M., Mu#241;oz, A. O. Oryzon Genomics, Methods to determine kdm1a target engagement and chemoprobes useful therefor. , (2016).
- Mascaró, C., Ortega, A., Carceller, E., Rruiz Rodriguez, R., Cicero, F., Lunardi, S., Yu, L., Hilbert, M., Maes, T. Chemoprobe-based assays of histone lysine demethylase 1A target occupation enable in vivo pharmacokinetics and -dynamics studies of KDM1A inhibitors. Journal of Biological Chemistry. , (2019).
- Rodriguez-Suarez, R. Development of Homogeneous Nonradioactive Methyltransferase and Demethylase Assays Targeting Histone H3 Lysine 4. Journal of Biomolecular Screening. 17 (1), 49-58 (2011).
- Lynch, J. T., Cockerill, M. J., Hitchin, J. R., Wiseman, D. H., Somervaille, T. C. CD86 expression as a surrogate cellular biomarker for pharmacological inhibition of the histone demethylase lysine-specific demethylase 1. Analytical Biochemistry. 442 (1), 104-106 (2013).
- Schulz-Fincke, J. Structure-activity studies on N-Substituted tranylcypromine derivatives lead to selective inhibitors of lysine specific demethylase 1 (LSD1) and potent inducers of leukemic cell differentiation. European Journal of Medicinal Chemistry. 144, 52-67 (2018).
- Ishii, T., et al. CETSA quantitatively verifies in vivo target engagement of novel RIPK1 inhibitors in various biospecimens. Scientific Report. 7, 13000 (2017).
- Maes, T. ORY-2001: An Epigenetic drug for the treatment of cognition defects in Alzheimer’s disease and other neurodegenerative disorders. Alzheimer’s & Dementia. 12 (7), P1192 (2017).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved