Method Article
Visualizing the Effects of Oxidative Damage on Drosophila Egg Chambers using Live Imaging
In This Article
Summary
The objective of this protocol is to use live imaging to visualize the effects of oxidative damage on the localization and dynamics of subcellular structures in Drosophila ovaries.
Abstract
Live imaging of Drosophila melanogaster ovaries has been instrumental in understanding a variety of basic cellular processes during development, including ribonucleoprotein particle movement, mRNA localization, organelle movement, and cytoskeletal dynamics. There are several methods for live imaging that have been developed. Due to the fact that each method involves dissecting individual ovarioles placed in media or halocarbon oil, cellular damage due to hypoxia and/or physical manipulation will inevitably occur over time. One downstream effect of hypoxia is to increase oxidative damage in the cells. The purpose of this protocol is to use live imaging to visualize the effects of oxidative damage on the localization and dynamics of subcellular structures in Drosophila ovaries after induction of controlled cellular damage. Here, we use hydrogen peroxide to induce cellular oxidative damage and give examples of the effects of such damage on two subcellular structures, mitochondria and Clu bliss particles. However, this method is applicable to any subcellular structure. The limitations are that hydrogen peroxide can only be added to aqueous media and would not work for imaging that uses halocarbon oil. The advantages are that hydrogen peroxide is readily available and inexpensive, acts quickly, its concentrations can be modulated, and oxidative damage is a good approximation of damage caused by hypoxia as well as general tissue damage due to manipulation.
Introduction
Multiple different cellular stressors may arise during the experimental culture and manipulation of tissues ex vivo, including heat shock, oxidative stress, osmotic stress, nutritional stress, and toxicity conditions. Live imaging is a powerful tool used to visualize real-time changes in ex vivo tissues after experimental treatment and manipulation. Fine tissue dissections and manipulation take practice, and the amount of time from dissection to imaging can vary depending on experience. The rationale for developing this method is based on the concern that preparing tissue for live imaging can cause cellular stress during dissection and imaging preparation. This could be particularly problematic for processes sensitive to changes in cellular metabolism and available oxygen levels, such as mitochondrial function. While having a parallel wild type sample is an important control, there is still the possibility that some or all observed changes in subcellular structures could be due to damage or cell stress from dissection and do not reflect normal physiology or the treatment or mutation being studied.
To address this potential problem, we use hydrogen peroxide addition during live imaging in order to induce cellular oxidative damage1. The purpose of this method is to induce damage to tissues in order to monitor the effect on subcellular structures. This protocol is useful for two purposes: 1) determining whether changes in subcellular localization of the structure of interest is due to the stress caused by inexperienced dissection and 2) once the researcher is confident with the dissection techniques described to monitor the effect of controlled stress on the structure of interest. Here we show two examples of how increased oxidative damage causes changes in two subcellular structures, mitochondria and Clu bliss particles. To do this, we use the Drosophila ovary which is a common model for live imaging studies. The first example examines mitochondrial localization. In our experience, normal mitochondrial localization in female germ cells is highly sensitive to perturbations and can act as a harbinger of cellular stress. Mitochondria in Drosophila female germ cells are normally evenly dispersed throughout the cytoplasm2. Hydrogen peroxide addition causes the organelles to quickly mislocalize and cluster in a similar manner to various mutations3,4,5. The second example are bliss particles formed by Clueless (Clu). Clu is a ribonucleoprotein that is diffuse throughout the cytoplasm; however, it also forms mitochondria-associated particles under optimal cellular conditions5. Because the presence of Clu particles is dependent upon healthy cellular conditions, we have termed them "bliss" particles3,5,6. Addition of hydrogen peroxide causes these particles to quickly disperse and become homogeneous in the cytoplasm5. In the course of our studies, we have observed changes in localization of both of these subcellular structures, but only after performing live imaging studies could we fully appreciate the effect of cellular stress and oxidative damage on localization and dynamics of mitochondria and bliss particles.
The utility of this protocol as an addition to already established or alternative methods depends on several factors. First, the imaging protocol must be amenable to drug-addition. If the sample is mounted under a coverslip and in halocarbon oil, this method would not be possible7. H2O2 addition causes a rapid rise in oxidative damage, therefore, this timescale may not be appropriate. Oxidative damage may be regarded as a proxy for hypoxia; however, it may be too harsh or too generalized to function as an appropriate control for damage for certain subcellular components. Finally, for imaging experiments that last hours such as those following a developmental process, H2O2 addition may be too strong (for example8). Testing a concentration curve may overcome this limitation.
Protocol
1. Preparation of dissection and imaging media
NOTE: The media best suited for this live imaging experiment contains Schneider's Drosophila media containing 15% heat inactivated fetal bovine serum, 0.6x Pen-Strep, and 200 µg/mL bovine insulin, hereafter referred to as Complete Schneider's media.
- Perform the media preparation under sterile conditions to ensure that it does not become contaminated. The media was developed to support Drosophila ovarioles for extended periods of time9.
- Add 15% heat inactivated fetal bovine serum, 0.6x Pen-Strep, and 200 µg/mL bovine insulin to the Schneider's media.
- Mix the contents well and store at 4 °C overnight.
NOTE: Insulin does not dissolve completely in the Complete Schneider's media, and you will notice a precipitate settle in the bottom of the tube. - Make aliquots of the media, being sure to leave the precipitate as it will interfere with imaging.
NOTE: This solution may be used within one month if stored in aliquots at 4 °C.
2. Collection of Drosophila for dissection
NOTE: Detailed Drosophila collection and dissection procedures may also be found in Weil et al.10 and Parker et al.11.
- For optimal female germ cell imaging, first prepare a vial containing standard cornmeal fly food and a dab of wet yeast paste that is the consistency of peanut butter. This ensures the female flies are well-fed and will produce all follicle developmental stages for imaging.
- For optimally healthy flies, collect 0-1 day old females and transfer with males into a fly food vial containing wet yeast paste.
NOTE: Make sure the sleeping flies do not contact the yeast paste as they can stick to it. - Feed the flies 3-7 days, changing the vial and the yeast paste daily.
NOTE: Make sure the yeast paste contacts the fly food so it does not dry out.
3. Drosophila ovary dissection
NOTE: It is important to prepare the media solutions fresh because hydrogen peroxide is susceptible to oxidation and TMRE degrades over time.
- Right before dissection, in Complete Schneider's media, prepare a fresh aliquot of 2 µM H2O2 solution, a fresh aliquot of 46 nM tetramethylrhodamine, ethyl ester (TMRE), and a fresh aliquot of a 46 nM TMRE solution containing 2 µM H2O2.
- For ovary dissection, use two pairs of fine forceps and a pair of electrolytically sharpened tungsten needles12. To dissect the ovaries, fill 2-3 wells of a glass bottom dissecting dish (watch glass) with Complete Schneider's that has been warmed to room temperature.
- Anaesthetize the vial of fattened flies with carbon dioxide and segregate the desired number of female flies to be dissected. Place a single fly in the media using forceps.
- Under a dissecting microscope, gently grasp the fly by the thorax using one pair of fine forceps. With the other pair of forceps, grasp the posterior, and gently pull to remove the ovaries.
NOTE: if the ovaries do not come out smoothly using this method, the entire abdomen may also be removed from the fly, and the ovaries can be gently squeezed out of either end of the abdomen using forceps. - Remove any extraneous cuticle or tissue, then transfer the ovaries to a new well containing fresh media. The ovaries should still be moving from the surrounding muscle sheath.
4. Preparing ovarioles for imaging
- Using sharpened tungsten needles, gently tease the ovarioles apart, taking care to remove the surrounding muscle sheath (Figure 1).
- Gently tease away any muscle sheath and nerve fibers attached to the isolated ovarioles (Figure 2).
NOTE: If the muscle sheath is not removed, the ovariole will twitch and move, causing problems with image acquisition (Video 1). - If the subcellular structures of interest are endogenously labeled, proceed to Step 4.4. If the structures of interest will be labeled with a fluorescent dye, proceed to Section 5.
- Once the ovarioles have been cleanly dissected, using a micropipette, transfer them in a 100 µL drop of Complete Schneider's imaging media into the glass depression of a glass bottom dish. The individual ovarioles will sink to the bottom of the droplet.
- Proceed to Section 6 for imaging.
NOTE: For imaging mitochondria and Clu bliss particles, no more than five-ten minutes should elapse from the start of dissection to imaging.
5. Staining mitochondria with TMRE
NOTE: Additional detailed procedures on live staining of mitochondria with fluorescent dyes may be found in Parker et al. 2017.
- After step 4.3, transfer the isolated ovarioles in a 100 µL drop of 46 nM TMRE media into the glass depression of a glass bottom dish. The individual ovarioles will sink to the bottom of the droplet.
- Incubate at room temperature for 20 minutes. Place the lid onto the glass-bottom dish and place the dish into a covered box for the duration of the experiment to protect from light.
NOTE: After incubation, samples can be imaged directly with no washes. - Repeat steps 5.1-5.2 to prepare at least two dishes of TMRE-labeled ovarioles, one to serve as an experimental group and one to serve as a control.
6. Live image acquisition
- Once the ovarioles are mounted, place one of the glass bottom dishes onto the microscope and configure the imaging parameters as necessary. The optimum excitation/emission wavelengths for the TMRE used here are 549 nm/574 nm.
- After locating the desired field of view, acquire still images or brief videos of one or more ovarioles as desired as a record of pre-treatment conditions.
7. Addition of hydrogen peroxide during imaging
- Pause live imaging, remove the lid from the glass bottom dish, and carefully add 100 µL of 2 µM H2O2 solution to the dish using a micropipette if imaging endogenously labeled structures. If imaging mitochondria, carefully add 100 µL of 46 nM TMRE with 2 µM H2O2 solution to the dish using a micropipette (Video 2).
- Avoid breaking the surface of the existing media droplet or adding the solution too quickly so as not to displace the ovarioles resting on the bottom of the dish (Video 2).
- Replace the dish lid (dish cover), relocate and refocus the desired field of view if necessary, and resume imaging (Video 3, Video 4, Figure 3, Figure 4).
- Take care to resume imaging as quickly as possible after H2O2 addition as the experimental treatment is time-sensitive (Video 5).
- Acquire still images or brief videos of one or more ovarioles as desired. This will serve as a record of post-treatment conditions.
- For the control, place the second glass bottom dish onto the microscope and repeat Section 6.
- Repeat step 7.1, this time adding 100 µL of TMRE-only media to the dish. If imaging endogenously labeled structures, add 100 µL of the Complete Schneider's media-only solution.
- Repeat steps 7.2 and 7.3 to acquire data for the control group.
NOTE: For imaging mitochondria and Clu bliss particles, no more than three-five minutes should elapse from the addition of hydrogen peroxide to imaging.
Results
The described protocols can be used to study the effects of hydrogen peroxide during live imaging of Drosophila ovaries. As shown in Figure 3, Figure 4, Video 3 and Video 4, this procedure provides an effective means to visualize tissue changes and dynamics after experimental treatment in real-time. Importantly, this protocol is specific for the addition of H2O2 to ovarioles while imaging; however, it may be adapted for the exogenous addition of other drugs or reagents of interest. In addition, follicles may be labeled with any fluorescent dyes of interest (e.g., tetramethylrhodamine (TMRE), LysoTracker) prior to imaging (Video 3). The most critical steps to obtaining clear imaging results are 1) the proper dissection and isolation of single ovarioles with all contractile elements removed (Figure 1 and Figure 2) and 2) the speed at which imaging is restarted after hydrogen peroxide addition. Video 4 is an illustration of a properly dissected follicle that remains steady throughout the imaging duration as compared to Video 1, which illustrates a poorly dissected follicle exiting the field of view during imaging. Video 5 is an illustration in which the H2O2 effects on TMRE-labeled mitochondria have already progressed prior to the start of imaging as a result of too much elapsed time. As compared to Video 3 in which imaging was restarted immediately after hydrogen peroxide addition (time 0) and intact, dispersed mitochondria are still visible, the mitochondria in Video 5 have already began to visibly clump and lose their membrane potential upon the restart of imaging. This issue is mostly attributed to disruption of the sample positions during H2O2 addition and can be alleviated by following the technique to keep the imaging media and sample position intact (Video 2). Of note, in control experiments performed without H2O2 added to the media, the mitochondria in follicles remains properly dispersed, and the TMRE dye remains sequestered in the mitochondria.
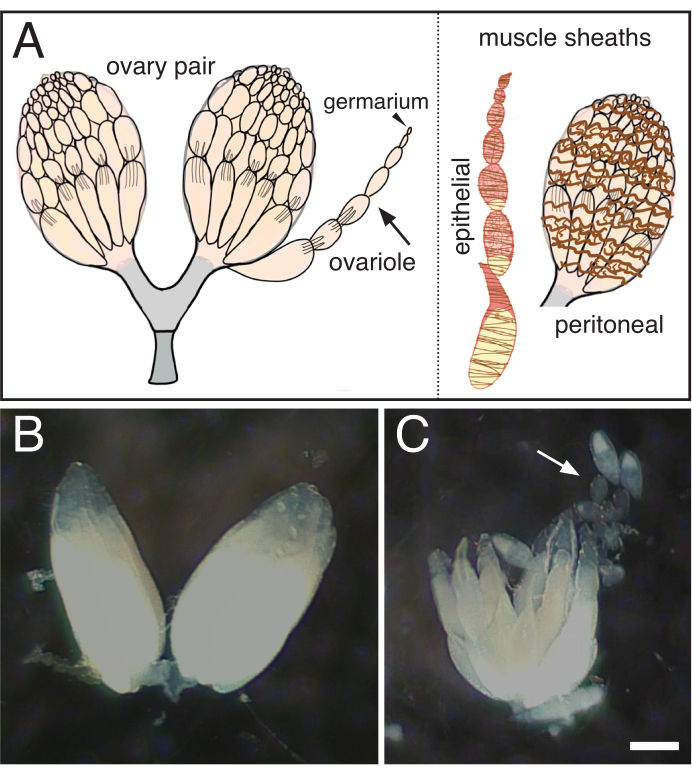
Figure 1: Isolation of single ovarioles from Drosophila ovaries. (A) Cartoon indicating a pair of ovaries, a single ovariole (arrow) with the germarium at the tip (arrowhead) and the two muscle sheaths that surround the ovariole (brown, epithelial) and the ovary (brown, peritoneal). (B) Dissected Drosophila ovary. (C) Subsequent separation of the teased ovary into individual ovarioles (arrow). Scale bar = 100 um. Please click here to view a larger version of this figure.
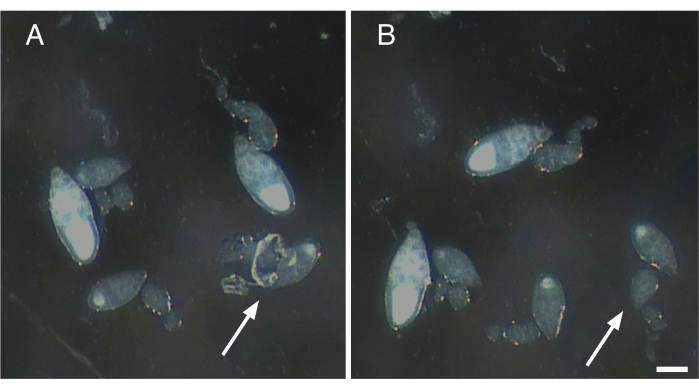
Figure 2: Removal of nerve fibers and contractile elements from single ovarioles. (A) Single ovariole with remnants of nerve tissue and the muscle sheath still attached (arrow). (B) Gentle removal of all remaining tissue attached to ovariole in A (arrow). Scale bar = 100 µm. Please click here to view a larger version of this figure.
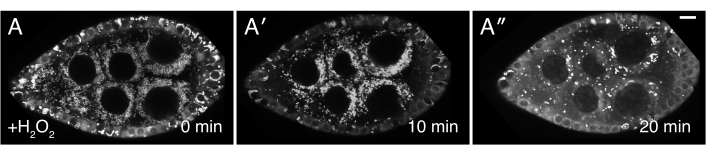
Figure 3: H2O2 causes mitochondrial mislocalization. (A-A″) Live-image stills of a well-fed cluCA06604 (Clu::GFP flies) follicle. Addition of H2O2 causes mitochondria to clump over the duration of imaging. TMRE labeling of mitochondria indicates that mitochondria are initially dispersed at time 0 (A), and that mitochondria start to clump after H2O2 addition (A′) At a later time-point, the TMRE labeling becomes spotty due to mitochondria losing their membrane potential and therefore their ability to sequester the dye (A″). White = TMRE (A-A″). Scale bars: 10 µm in (A) for A-A″5. Please click here to view a larger version of this figure.
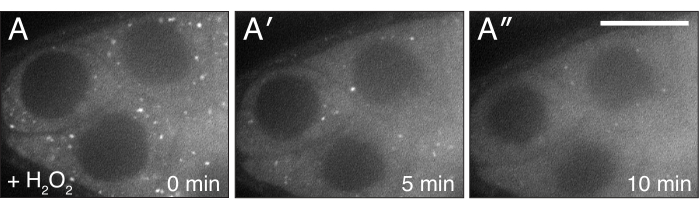
Figure 4: H2O2 disperses Clu. (A-A″) Live-image stills of a well-fed cluCA06604 follicle. Addition of H2O2 causes particles to disperse and become homogeneous in the cytoplasm over the duration of imaging. White = Clu::GFP (A-A″). Scale bar: 40 µm in (A) for A-A″5. Please click here to view a larger version of this figure.
Video 1: Follicle from a cluCA06604 female. As described in Figure 2, failure to remove nerve fiber contractile elements from single ovarioles will cause marked drifting and movement of the ovarioles during imaging and subsequent inability to analyze imaging data. White = Clu::GFP. Please click here to download this Video.
Video 2: Proper addition of H2O2 to sample dish. Hydrogen peroxide should be added to the sample dish using an appropriately sized micropipette. Dispensing H2O2 without breaking media surface. Care is taken to avoid breaking the surface of the imaging media to minimize sample drift during hydrogen peroxide addition. Please click here to download this Video.
Video 3: H2O2 addition during imaging of TMRE labeled follicles. Follicle from a cluCA06604 female stained with TMRE. H2O2 causes mitochondrial mislocalization in Drosophila follicles. White = TMRE dye. Imaged one frame per 15 s for 20 min, and the video is 10 frames per second5. Please click here to download this Video.
Video 4: H2O2 addition during imaging of Clu::GFP follicles. Follicle from a cluCA06604 female. H2O2 disperses Clu particles as described in Figure 4. White = Clu::GFP. Imaged one frame per 15 s for 15 min, and the video is 10 frames per second.5 Please click here to download this Video.
Video 5: Delayed imaging after H2O2 addition. Follicle from a cluCA06604 female. Addition of hydrogen-peroxide to follicles is a time-sensitive treatment. The restart of imaging was delayed in this video as a result of sample displacement during H2O2 addition. The mitochondria have already begun to visibly clump and lose their membrane potential upon the restart of imaging at time 0 (as compared to time 0 in Video 3). Failure to resume imaging quickly after treatment will result in inaccurate and unusable results as the early experimental effects will be missed prior to imaging. White = TMRE. Imaged one frame per 15 s for 20 min, and the video is 10 frames per second. Please click here to download this Video.
Discussion
This protocol could be a useful addition as a control for artifacts due to ovary dissection and tissue incubation for any live imaging experiment. The critical steps are similar to those found for other live imaging protocols. Learning how to dissect whole Drosophila ovaries takes practice; however, this skill can usually be learned fairly quickly with the appropriate dissection tools. More difficult to master is removing the muscle enveloping the ovaries and each ovariole13. This must be done to ensure muscle contractions do not interfere with image acquisition. If using sharpened tungsten needles to do this does not prove successful, the germarium at the tip of the ovariole can be grasped with forceps and the ovariole pulled from the muscle sheath. However, this technique is problematic if the earliest developmental stages are to be examined because they can become damaged. Another key step is to not dislodge the ovarioles resting on the bottom of the dish when adding H2O2. An additional important aspect is shared by all live imaging: the researcher should ensure that the structure of interest is fluorescently well-labeled before treatment. The dishes used here (Table of Materials) are commonly used for live imaging; however, any dish or slide with a glass coverslip on the bottom or even a large glass coverslip should work as long as the drop of media can be covered to prevent media evaporation. While we use a particular microscope, any inverted microscope with an objective of sufficient magnification to see the subcellular structure in question and an attached camera that has sufficient resolution and image capture rate should work.
While our laboratory is primarily interested in mitochondrial function, this method could be helpful examining the dynamics and localization of any subcellular structure or organelle, such as the nucleus, cytoskeleton or endoplasmic reticulum. However, this method has limitations. In order to add hydrogen peroxide, the tissue must be in an aqueous media. An alternative method for live imaging is to use halocarbon oil, which has been instrumental in describing many important processes in Drosophila ovarioles including the first example of dynamic movement of GFP in a model organism7,14. In addition, adding hydrogen peroxide to the media causes wide-spread oxidative damage which may be too general an insult to the tissue to be informative for the cellular process of interest, particularly for longer experiments examining development. Though it may not be feasible to perform experiments which require visualizing the cell over long periods of time due to this rapid, extensive, and likely irreversible oxidative damage, we have seen that the acute hydrogen peroxide treatment we have described is applicable to most stages of oogenesis as we are able to see the same effects in most stages within the imaging time period. Given the low cost and ease of the protocol, it may be a useful control for damage and can be used as a treatment before fixation and antibody labeling as well.
In our hands, H2O2 treatment mimics the changes in mitochondrial mislocalization and Clu bliss particle dispersion that we see in various Drosophila mutants. It also mimics results we see for new researchers in the lab learning dissection techniques. Therefore, this method clearly revealed that sample preparation and general cellular stress can lead to unexpected and previously unexplained changes to mitochondrial mislocalization and the presence of bliss particles. Moving this technique forward, hydrogen peroxide concentrations could be modulated using a higher or lower concentration. If a cellular effect is seen using a lower concentration, it is possible the stress phenotype may be reversible by replacing the media with Complete Schneider's. Different cell stressors such as carbonyl cyanide m-chlorophenyl hydrazone (CCCP), arsenite or simple heat shock might prove useful for general cellular stress for other subcellular structures. Since live imaging of ex vivo tissues requires manual manipulation and incubation in different media, this control should be a useful addition to ensure any observations are as close to normal physiology as possible.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We would like to thank Dr. Jeremy Smyth for imaging support and Ann C. Shenk for illustrations, production and videography. This work was supported by the National Institutes of Health (1R01GM127938 to R.T.C.).
Materials
| Name | Company | Catalog Number | Comments |
| Active dry yeast | Red Star® | ||
| CO2 gas | 99.9% purity | ||
| CO2 pad | |||
| Dissecting microscope, Nikon SMZ645 model | Nikon | ||
| Dissecting needles - PrecisionGlide needles | BD | 305165 | B-D 21G1 size |
| Dissecting needles - PrecisionGlide syringes | BD | 309657 | Luer-Lok tip, 3 mL size |
| Dissecting needles - tungsten wire | Electron Microscopy Sciences | 73800 | |
| Dumont #5 forceps (2 pairs) | Fine Science Tools | 11251-10 | |
| NI-150 High Intensity Illuminator | Nikon Instruments Inc. | ||
| Gibco Fetal Bovine Serum, Heat Inactivated | Fisher Scientific | 10082-147 | |
| Gibco Schneider's Drosophila Media | Sigma-Aldrich | 21720-024 | |
| Hydrogen peroxide solution, 30% (w/w) in H2O | Sigma-Aldrich | H1009 | |
| Insulin from bovine pancreas | Sigma-Aldrich | I6634 | |
| Spinning disk microscope | Nikon | Equivalent scopes may also be used | |
| Lonza BioWhittaker Antibiotics: Penicillin-Streptomycin mixtures | Fisher Scientific | 17-602E | |
| MatTek Corporation Glass Bottom Dishes, 35 mm | Fisher Scientific | NC9344527 | |
| Micropipettes and tips of appropiate size | Eppendorf | ||
| Microcentrifuge tubes, 1.7 mL | VWR | 87003-294 | |
| Tetramethylrhodamine, Ethyl Ester, Perchlorate (TMRE) | AnaSpec | AS-88061 | |
| w[1118] | Bloomington Drosophila Stock Center | 5905 | Wild-type flies |
| y w; clu[CA06604] | Available upon request. | Clu::GFP trap flies |
References
- Winterbourn, C. C. Toxicity of iron and hydrogen peroxide: the Fenton reaction. Toxicology Letters. 82-83, 969-974 (1995).
- Cox, R. T., Spradling, A. C. A Balbiani body and the fusome mediate mitochondrial inheritance during Drosophila oogenesis. Development. 130 (8), 1579-1590 (2003).
- Cox, R. T., Spradling, A. C. Clueless, a conserved Drosophila gene required for mitochondrial subcellular localization, interacts genetically with parkin. Disease Models & Mechanisms. 2 (9-10), 490-499 (2009).
- Sen, A., Kalvakuri, S., Bodmer, R., Cox, R. T. Clueless, a protein required for mitochondrial function, interacts with the PINK1-Parkin complex in Drosophila. Disease Models & Mechanisms. 8 (6), 577-589 (2015).
- Sheard, K. M., Thibault-Sennett, S. A., Sen, A., Shewmaker, F., Cox, R. T. Clueless forms dynamic, insulin-responsive bliss particles sensitive to stress. Developmental Biology. 459 (2), 149-160 (2020).
- Sen, A., Cox, R. T. Clueless is a conserved ribonucleoprotein that binds the ribosome at the mitochondrial outer membrane. Biology Open. 5 (2), 195-203 (2016).
- Parton, R. M., Valles, A. M., Dobbie, I. M., Davis, I. Isolation of Drosophila egg chambers for imaging. Cold Spring Harbor Protocols. 2010 (4), 5402(2010).
- Morris, L. X., Spradling, A. C. Long-term live imaging provides new insight into stem cell regulation and germline-soma coordination in the Drosophila ovary. Development. 138 (11), 2207-2215 (2011).
- Prasad, M., Montell, D. J. Cellular and molecular mechanisms of border cell migration analyzed using time-lapse live-cell imaging. Developmental Cell. 12 (6), 997-1005 (2007).
- Weil, T. T., Parton, R. M., Davis, I. Preparing individual Drosophila egg chambers for live imaging. Journal of Visualized Experiments. (60), e3679(2012).
- Parker, D. J., Moran, A., Mitra, K. Studying Mitochondrial Structure and Function in Drosophila Ovaries. Journal of Visualized Experiments. (119), e54989(2017).
- Brady, J. A simple technique for making fine, durable dissecting needles by sharpening tungsten wire electrolytically. Bulletin of the World Health Organization. 32 (1), 143-144 (1965).
- Hudson, A. M., Petrella, L. N., Tanaka, A. J., Cooley, L. Mononuclear muscle cells in Drosophila ovaries revealed by GFP protein traps. Developmental Biology. 314 (2), 329-340 (2008).
- Wang, S., Hazelrigg, T. Implications for bcd mRNA localization from spatial distribution of exu protein in Drosophila oogenesis. Nature. 369 (6479), 400-403 (1994).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved