A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Atomic Force Microscopy to Study the Physical Properties of Epidermal Cells of Live Arabidopsis Roots
In This Article
Summary
The atomic force microscopy indentation protocol offers the possibility to dissect the role of the physical properties of the cell wall of a particular cell of a tissue or organ during normal or constrained growth (i.e., under water deficit).
Abstract
A method is described here to characterize the physical properties of the cell wall of epidermal cells of living Arabidopsis roots through nanoindentations with an atomic force microscope (AFM) coupled with an optical inverted fluorescence microscope. The method consists of applying controlled forces to the sample while measuring its deformation, allowing quantifying parameters such as the apparent Young's modulus of cell walls at subcellular resolutions. It requires a careful mechanical immobilization of the sample and correct selection of indenters and indentation depths. Although it can be used only in external tissues, this method allows characterizing mechanical changes in plant cell walls during development and enables the correlation of these microscopic changes with the growth of an entire organ.
Introduction
Plant cells are surrounded by a cell wall that is a complex structure composed of interacting networks of polysaccharides, proteins, metabolites, and water that varies in thickness from 0.1 to several µm depending on the cell type and the phase of growth1,2. Cell wall mechanical properties play an essential role in the growth of plants. Low stiffness values of the cell wall have been proposed as a precondition for cell growth and cell-wall expansion, and there is increasing evidence that all cells sense mechanical forces to perform their functions. However, it is still debated whether changes in the physical properties of the cell wall determines cell fate2,3,4. Because plant cells do not move during development, the final shape of an organ depends on how far and in what direction a cell expands. Thus, Arabidopsis root is a good model to study the impact of cell wall physical properties in cell expansion because different types of expansion occur in different regions of the root. For example, anisotropic expansion is evident in the elongation zone and particularly noticeably in the epidermal cells5.
The method described here was used to characterize the physical properties of the cell wall of epidermal cells at the nanoscale of living Arabidopsis roots using an Atomic Force Microscope (AFM) coupled with an inverted fluorescence phase microscope6. For an extensive revision of the AFM technique, read7,8,9.
This protocol outlines a basic sample preparation method and a general method for AFM-based elasticity measurements of plant cell walls.

Figure 1: Schematic overview of force-indentation experiment in Arabidopsis roots using atomic force microscopy (AFM). The scheme gives an overview of the steps of a Force-Indentation experiment from the preparation of the substrate to immobilize the root sample firmly (1-2), root viability confirmation through propidium iodide staining (3), cantilever positioning on the surface of an elongated epidermal cell of the primary root (4-5), force curves measurement (6), and force curve processing to calculate the apparent Young's modulus (7-8). EZ: elongation zone. Please click here to view a larger version of this figure.
Protocol
1. Preparation of the plant material and growth conditions
- To generate the needed plant material, sterilize the seeds of Arabidopsis wild type and mutant lines of interest.
NOTE: In this protocol, we used the following: ttl1: T-DNA insertion lines Salk_063943 (for TTL1; AT1G53300) - Columbia-0 (Col-0) wild-type; the Procuste1 (prc1-1) mutant, which consists of a knock-out mutation (Q720stop) in the CESA6 gene (AT5G64740), as previously described in10,11; and ttl1 x prc1-1 double mutant.- Put an approximate volume of 50 μL of seeds in a microtube. Add 500 μL of 70% ethanol + 0.01% Tween solution at room temperature. Mix and incubate for 7 min.
- Pour the ethanol solution and add 500 μL of 20% sodium hypochlorite. Mix and incubate for 7 min.
- Pour the sodium hypochlorite solution and wash the seeds three to four times with sterile water.
- Plating
- Prepare in advance basal Murashige and Skoog (MS) medium12 + 1.5% sucrose + 1% agar (pH 5.7) (see Table of Materials). Autoclave the medium. Prepare a laminar flow hood to work in a sterile environment and label the square Petri dishes needed for the experiment. Pour the medium into the Petri dishes and allow the medium to solidify.
- Plate the sterilized seeds using sterile pipette tips onto square Petri dishes containing the medium. Distribute the seeds evenly in two rows. When the seeds are dry, close the lids and seal the plates. Stratify the seeds at 4 °C in the dark for 4 days.
NOTE: Stratification is needed to synchronize germination. - Place the plates vertically in the growth chamber in a long-day regime (16 h of light/8 h of dark) with 25 μmol m2 s−1 light intensity and 23 °C (day/night). Grow the seedlings for 7 days.
2. Osmotic stress treatment (optional)
NOTE: This section provides details on the growth of Arabidopsis roots in osmotic potential of -1.2 MPa as estimated by cryoscopic osmometer (Table of Materials). This part can be omitted or changed depending on the experimental question at hand.
- Prepare basal MS medium + 1.5% sucrose + 400 mM mannitol + 1% agar (pH 5.7). Autoclave the medium. Pour the medium in square Petri dishes in the laminar flow hood and allow the medium to solidify.
- With tweezers (Table of Materials), place 5-day old seedlings pre-grown in basal MS + 1.5% sucrose medium in square Petri dishes containing the medium supplemented with 400 mM mannitol. Place the plates vertically in the growth chamber in the same conditions as in step 1.2.3. Grow the seedlings under this osmotic stress condition for 7 days.
NOTE: To evaluate root growth adaptation of the primary root during severe osmotic stress, it is suggested to use seedlings 5 days after germination grown in controlled conditions to ensure that the root meristem size is already established13. Alternatively, the analysis can be performed at every stage of development to determine the developmental effect of osmotic stress action.
3. Atomic force microscopy (AFM) nanoindentation experiments
- Preparation of the sample for nanoindentation experiments
NOTE: This is a crucial step for applying AFM to the study of live biological samples. The sample must be firmly attached to the substrate to be mechanically stable during indentation measurements. At the same time, sample structural qualities should be preserved. An effective strategy to stabilize a big sample for AFM is to use adhesives. The adhesive must dry quickly and must not be toxic or reactive with the surrounding medium. In this protocol, polystyrene Petri dishes were used as the substrate and non-acidic silicone glue (Table of Materials) was used to bind the Arabidopsis seedlings.- Spread a thin layer of silicone glue into the Petri dish with a coverslip. Leave the glue in the air for 45 s.
- With tweezers, place the seedling on the glue, orienting it in a direction that avoids contact between the seedling's protruding parts and the cantilever. Then, press the root gently to the silicone glue layer to bind it firmly. Leave the seedling in contact with the glue for 45 s before continuing with the next steps of the protocol.
- Incubate the attached seedlings with 2 μg/mL propidium iodide (PI; Table of Materials) for 5 min in dark and carefully wash them thrice with 1x phosphate-buffered saline (PBS) solution (Table of Materials).
- Place the PI-incubated seedlings into an inverted fluorescence microscope coupled with the AFM. To observe fluorescence, set the excitation and emission wavelengths at 572 nm and 617 nm, respectively. The absence of fluorescence inside the epidermal cells confirms the viability of the roots.
- Cover the seedling with 1x phosphate-buffered saline (PBS) solution.
- Indentation protocol
NOTE: Perform all AFM measurements within 1 h after the immobilization of the sample on its support at room temperature. The room where the AFM is located should be maintained at 25 °C with an air conditioner. The PBS solution should be at the same temperature.- Mount a standard silicon nitride cantilever with a pyramidal tip (see Table of Materials) into the AFM probe holder for fluid (Table of Materials).
- Align the laser on the cantilever close to the position of the tip. Next, move the photodiode to place the laser spot at the center of the detector.
- To calibrate the deflection sensitivity, position a polystyrene Petri dish with 1x PBS under the AFM.
NOTE: This step is needed to calculate the indentation. When a force measurement is made on a hard surface, there is no indentation into the surface, so any change in the z scanner displacement corresponds to the cantilever deflection. Therefore, it is very important to calibrate the deflection sensitivity on a hard surface. Using a soft surface will overestimate the deflection sensitivity, which will result in spring constants that are too low. - Adjust the photodetector such that the non-contact deflection is near 0 V. Calibrate the deflection sensitivity by performing an indentation (force curve), with a ramp size of 3 μm, an indentation-and-retraction rate of 0.6 μm/s, and a trigger threshold of 0.5 V.
- To calibrate the spring constant of the cantilever, use the Thermal Tune utility of the AFM software (Table of Materials) recommended for probes with spring constants less than approximately 5 N/m or use the method described in reference14. Before activating Thermal Tune in the software, ensure that the probe does not interact with the sample.
- Enter the cantilever temperature. Click on Calibrate > Thermal Tune or on the Thermal Tune icon in the NanoScope toolbar. Select a frequency range (1-100 Hz).
- Click on Acquire Data in the Thermal Tune panel. Wait for the AFM to acquire data, and then click on the Simple Harmonic Oscillator (Fluid) button.
- Adjust Median Filter Width to 3. Adjust PSD Bin Width to reduce the noise in the acquired data by averaging. Set fit boundaries around the first resonance peak.
- Click on Calculate Spring Constant k, and then, click on Yes in the pop-up window asking if the user wants to use this value.
- Repeat the steps described above three times and manually take an average of spring constant values. Enter this average value in the Spring Constant box in the Ramp parameter list in the PicoForce view. The calibration ends at this point.
- Using the inverted optical microscope at 10x, 20x, and 40x eyepiece magnification, position the AFM probe on the surface of the fourth elongated epidermal cell of the primary root, ensuring to position it in the center of the cell.
- With the calculated spring constant value (step 3.2.5.5), obtain force curves with a ramp size of 3 µm, a trigger threshold of 11 nN, and an indentation-and-retraction rate of 0.6 μm/s at selected points.
NOTE: Previous work15,16 found that low-frequency of indentation-and-retraction helps to minimize hysteresis and drag force. - Obtain force curves from three cells per root for each treatment (use three biological replicates per treatment). Capture at least 150 force curves for each root.
4. Measure the apparent Young's modulus
- Fit each force curve to the following model for pyramidal indenters17:
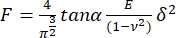 , where E is the apparent Young's modulus of the sample,
, where E is the apparent Young's modulus of the sample, 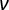 is the Poisson modulus of the sample, and α is the semi-angle to the vertex. Discard the fits with a correlation coefficient (r2) < 0.99. Consider the Poisson's ratio for totally incompressible material:
is the Poisson modulus of the sample, and α is the semi-angle to the vertex. Discard the fits with a correlation coefficient (r2) < 0.99. Consider the Poisson's ratio for totally incompressible material: 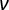 = 0.5 and the geometry of the indenter given by the manufacturers.
= 0.5 and the geometry of the indenter given by the manufacturers.
NOTE: Use the first 100 nm from the contact point of the loading curve for the fitting to be as sensitive as possible to cell wall mechanics and to reduce the impact of turgor pressure on the apparent Young's modulus values8. The fitting has been done using a custom MATLAB program (available upon personal request by writing to J. C. B) that allows setting a specific indentation depth from the contact point of the loading curve. - Create a normalized histogram with the apparent Young's modulus data and fit it with a Gaussian distribution. Discard data points outside the 95% confidence interval and recalculate both the histogram and the Gaussian fit. Calculate and report the mean and standard deviation of the apparent Young's modulus.
- Determine the significance of the comparisons between groups by multiple comparison tests like ANOVA.
Results
Force-Indentation experiments
The following text presents some results expected when a force-indentation experiment is conducted to show the typical output to expect when the protocol is well executed.
Force-displacement curves
Representative force indentation plots that were obtained indenting live samples at a position placed in the center of the cell of the root elongation zone are presented in Figure 2. When the AF...
Discussion
Cell and cell-wall mechanics are increasingly becoming relevant to gain insight into how mechanics affects growth processes. As physical forces propagate over considerable distances in solid tissues, the study of changes in the physical properties of the cell wall and how they are sensed, controlled, tuned, and impact the plant's growth are becoming an important field of study2,3,8.
A method is pr...
Disclosures
The authors have no conflicts of interest. The MATLAB script used for fitting the data is available upon personal request by writing to J. C. B.
Acknowledgements
This research was funded by CSIC I+D 2018, grant No. 95 (Mariana Sotelo Silveira).; CSIC Grupos (Omar Borsani) and PEDECIBA.
Materials
| Name | Company | Catalog Number | Comments |
| 1 x Phosphate-Buffered Saline (PBS) | Include sodium chloride and phosphate buffer and is formulated to prevent osmotic shock and maintain water balance in living cells. | ||
| AFM software | Bruker, Billerica, MA, USA | ||
| Atomic force microscopy (AFM) | BioScope Catalyst, Bruker, Billerica, MA, USA | ||
| Catalyst Probe holder-fluid | Bruker, Billerica, MA, USA | CAT-FCH | A probe holder for the Bioscope Catalyst, designed for fluid operation in contact or Tapping Mode. Also compatible with air operation. |
| Cryoscopic osmometer; model OSMOMAT 030 | Gonotech, Berlin, Germany | ||
| Murashige & Skoog Medium | Duchess Biochemie | M0221 | Original concentration, (1962) |
| Optical inverted microscope coupled to the AFM | Olympus IX81, Miami, FL, USA | ||
| PEGAMIL | ANAEROBICOS S.R.L., Buenos Aires, Argentina | 100429 | Neutral, non acidic silicone glue |
| Petri dishes | Deltalab | 200201.B | Polystyrene, 55 x 14 mm, radiation sterile. |
| Propidium iodide | Sigma | P4170 | For root viability test. |
| Silicon nitride probe, DNP-10, cantilever A | Bruker, Billerica, MA, USA | DNP-10/A | For force modulation microscopy in liquid operation. Probe tip radius of 20-60 nm. 175-μm-long triangular cantilever, with a spring constant of 0.35 N/m. |
| Tweezers | Sigma | T4537 |
References
- Anderson, C. T., Kieber, J. J. Dynamic construction, perception, and remodeling of plant cell walls. Annual Review of Plant Biology. 71, 39-69 (2020).
- Roeder, A. H. K., et al. Fifteen compelling open questions in plant cell biology. The Plant Cell. 34 (1), 72-102 (2022).
- Zhang, B., Gao, Y., Zhang, L., Zhou, Y. The plant cell wall: Biosynthesis, construction, and functions. Journal of Integrative Plant Biology. 63 (1), 251-272 (2021).
- Hamant, O., Haswell, E. S. Life behind the wall: Sensing mechanical cues in plants. BMC Biology. 15 (59), 1-9 (2017).
- Scheres, B., Benfey, P., Dolan, L. Root development. The Arabidopsis Book. 1, 0101 (2002).
- Cuadrado-Pedetti, M. B., et al. The arabidopsis tetratricopeptide thioredoxin-like 1 gene is involved in anisotropic root growth during osmotic stress adaptation. Genes. 12 (2), 236 (2021).
- Milani, P., Braybrook, S. A., Boudaoud, A. Shrinking the hammer: micromechanical approaches to morphogenesis. Journal of Experimental Botany. 64 (15), 4651-4662 (2013).
- Braybrook, S. A. Measuring the elasticity of plant cells with atomic force microscopy. Methods in Cell Biology. 125, 237-254 (2015).
- Bidhendi, A. J., Geitmann, A. Methods to quantify primary plant cell wall mechanics. Journal of Experimental Botany. 70 (14), 3615-3648 (2019).
- Desnos, T., et al. Procuste1 mutants identify two distinct genetic pathways controlling hypocotyl cell elongation, respectively in dark- and light-grown Arabidopsis seedlings. Development. 122 (2), 683-693 (1996).
- Fagard, M., et al. Procuste1 encodes a cellulose synthase required for normal cell elongation specifically in roots and dark-grown hypocotyls of arabidopsis. The Plant Cell. 12 (12), 2409-2423 (2000).
- Murashige, T., Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiologia Plantarum. 15 (3), 473-497 (1962).
- Perilli, S., Sabatini, S. Analysis of root meristem size development. Methods in Molecular Biology. 655, 177-187 (2010).
- Sader, J. E., et al. A virtual instrument to standardise the calibration of atomic force microscope cantilevers. Review of Scientific Instruments. 87 (9), 093711 (2016).
- Collinsworth, A. M., Zhang, S., Kraus, W. E., Truskey, G. A. Apparent elastic modulus and hysteresis of skeletal muscle cells throughout differentiation. American Journal of Physiology - Cell Physiology. 283 (4), 1219-1227 (2002).
- Mathur, A. B., Collinsworth, A. M., Reichert, W. M., Kraus, W. E., Truskey, G. A. Endothelial, cardiac muscle and skeletal muscle exhibit different viscous and elastic properties as determined by atomic force microscopy. Journal of Biomechanics. 34 (12), 1545-1553 (2001).
- Sirghi, L., Ponti, J., Broggi, F., Rossi, F. Probing elasticity and adhesion of live cells by atomic force microscopy indentation. European Biophysics Journal. 37 (6), 935-945 (2008).
- Peaucelle, A. AFM-based mapping of the elastic properties of cell walls: At tissue, cellular, and subcellular resolutions. Journal of Visualized Experiments: JoVE. (89), e51317 (2014).
- Peaucelle, A., et al. Pectin-induced changes in cell wall mechanics underlie organ initiation in Arabidopsis. Current Biology. 21 (20), 1720-1726 (2011).
- Fernandes, A. N., et al. Mechanical properties of epidermal cells of whole living roots of Arabidopsis thaliana: An atomic force microscopy study. Physical Review E. 85 (2), 21916 (2012).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved