A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Pulmonary Structural MRI using Free-Breathing, Self-Gated Ultra-short Echo Time Imaging
In This Article
Summary
A protocol is described for generating high-resolution structural images of the lungs using ultra-short-echo time (UTE) Magnetic Resonance Imaging (MRI). This protocol allows for images to be acquired using a simple MRI pulse sequence during free-breathing.
Abstract
High quality MRI of the lungs is challenged by low tissue density, fast MRI signal relaxation, and respiratory and cardiac motion. For these reasons, structural imaging of the lungs is performed almost exclusively using Computed Tomography (CT). However, CT imaging delivers ionizing radiation, and thus is less well suited for certain vulnerable populations (e.g., pediatrics) or for research applications. As an alternative, MRI using ultra-short echo times (UTE) is attracting interest. This technique can be performed during free-breathing over the course of a ~5-10 min scan. Respiratory motion information is encoded alongside images; this information can be used to "self-gate" images. Self-gating thus removes the requirement of advanced MRI pulse sequence programming or the use of respiratory bellows, which simplifies image acquisition. In this protocol, simple, robust, and computationally efficient acquisition and reconstruction methods for acquiring high quality UTE MRI of the lungs are presented. This protocol was developed for use on a 3T MRI scanner, but the same principles can be implemented at lower magnetic field strength. The protocol includes recommended parameter settings for 3D radial UTE image acquisition as well as directions for self-gated image reconstruction to generate images at distinct respiratory phases. Through the implementation of this protocol, users can generate high-resolution UTE images of the lungs with minimal to minimal-to-no motion artifacts. These images can be used to evaluate pulmonary structure, which can be implemented for research use in a variety of pulmonary conditions.
Introduction
High-resolution imaging of the pulmonary structure is an essential part of diagnostic work-ups for many pulmonary conditions. Typically, this is performed using Computed Tomography (CT) imaging, which is ideally suited to generate high-resolution images of the lungs1. However, CT imaging delivers a non-trivial dose of ionizing radiation, making it ill-suited for regular repeat imaging, imaging at multiple different respiratory phases, or imaging certain populations (e.g., pediatrics). Magnetic resonance imaging (MRI) does not carry the same risk of ionizing radiation, and thus is amenable to such imaging tasks. However, it is challenging to image the lungs using MRI owing to low tissue density, respiratory and cardiac motion, and very fast signal relaxation2,3,4.
One MRI technique that is able to mitigate these challenges is ultra-short echo time (UTE) MRI4,5,6. In UTE MRI, the MRI signal is sampled immediately following signal excitation, which reduces the impact of fast signal relaxation. Moreover, this technique samples k-space from the center outward, which leads to significant oversampling at the center of k-space. This oversampling at the center of the k-space makes this imaging technique robust to motion. In addition to this inherent robustness to motion, repeated sampling of the center of k-space encodes information about respiratory motion, which enables the self-gating of images7,8,9. This self-gating can be used to generate images at a variety of respiratory phases. Because humans spend the majority of the respiratory phase at expiration, it is common to generate an image for end-expiration, as this phase has the most imaging data acquired.
There are a variety of strategies for respiratory self-gating in pulmonary MRI. The first distinction to be made is image-based vs. k-space-based gating10 (Figure 1). In image-based gating, a set of images with high temporal resolution is generated by reconstructing small temporal subsets of the imaging data. Subsequently, the position of the diaphragm in these images is used to identify the respiratory phase for a given set of image projections10,11. In k-space-based gating, data from the center of k-space ("k0") is examined8,9,12. The signal intensity of the image is encoded in k0, and thus, the intensity of the k0 point varies with respiration. Projections can thus be binned into different respiratory phases based on the intensity of k0. In both image-based and k-space-based gating, projections with like-respiratory phases are grouped for image reconstruction. It has been suggested that image-based gating provides improved fidelity in estimating the respiratory phase, thereby providing images with reduced blurring10,13.
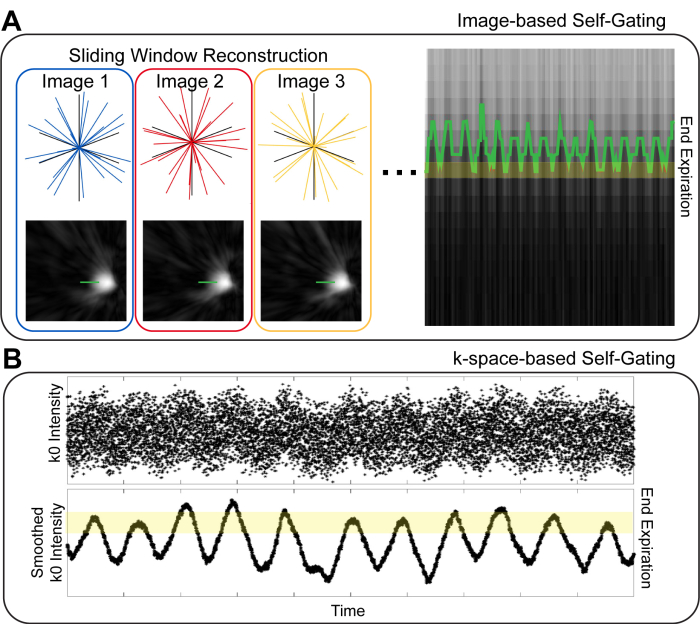
Figure 1: Image-based and k-space based self gating techniques. (A) In image-based gating, low spatial resolution, high temporal resolution images showing the diaphragm are generated from temporal subsets of the overall data. Using a line over the diaphragm, respiratory motion can be visualized and binned for image reconstruction. (B) In k-space-based gating, the first point on a center-out k-space projection ("k0") is used to visualize respiratory motion. After smoothing k0, signal intensity differences based on the respiratory cycle are clearly visible and can be used to identify different respiratory phases. Please click here to view a larger version of this figure.
Both image and k-space-based gating can be performed using either hard gating or soft gating11,14. In hard gating, only the projections corresponding to the desired respiratory phase are reconstructed. However, this discarding of unwanted projections can lead to reduced image signal-to-noise ratio (SNR) and increased undersampling artifacts. These undesired effects can be mitigated by using soft gating. In soft gating, all projections are used for image reconstruction, but projections from an unwanted respiratory phase are weighted such that they have a lesser impact on the final image. In doing so, images can be reconstructed with minimal artifacts and high SNR while still suppressing the impact of respiratory motion.
Through the combination of UTE MRI acquisition with post-acquisition self-gating, high-quality images can be generated that, while not equivalent to CT, have a contrast and resolution that is approaching that of CT imaging6,15,16,17,18,19. Herein, a simple protocol is provided for collecting and reconstructing UTE MRI images to generate high quality images of pulmonary structure.
This protocol is written primarily for 3T MRI scanners; 3T is the most common field strength used for research MRI. Lower magnetic field strengths such as 1.5T or the recently available 0.55 T20 can provide improved image quality and signal intensity within the lungs, as signal relaxation within the lungs is slower at these field strengths.
While every attempt has been made to provide clarity and simplicity in this protocol and the provided image reconstruction code, the protocol will likely require a dedicated MRI physicist (or similar MRI expert) to establish an appropriate UTE MRI sequence on the MRI scanner. The MRI sequence should implement a 3D non-Cartesian encoding strategy with Center-out k-space trajectories. Examples include 3D radial or 3D spiral (e.g., "FLORET")21,22 imaging sequences. Importantly, the order of projections should have good temporal stability: Over any given subset of time, the projections should cover the full range of k-space23. Examples of projection ordering strategies with good temporal stability are golden means or Halton-randomized Archimedean spiral. If a projection ordering with poor temporal stability is used, post-acquisition self-gating will omit large regions of k-space, leading to image artifacts. Finally, the sequence should be capable of achieving an echo time (TE) of <100 µs. The T2* relaxation time in the lungs at 3T is <1 ms24, so using a very short TE is essential to generating high-quality images.
Protocol
All human subject imaging was performed with approval from the KUMC IRB. Written informed consent was obtained from all participants. Images in this study were obtained under a generic technical development protocol, and the inclusion/exclusion criteria were deliberately broad. Inclusion Criteria: Age ≥ 18. Exclusion Criteria: MRI contraindicated based on responses to the MRI screening questionnaire, and pregnancy. The accessories and the equipment used for this study are listed in the Table of Materials.
1. UTE image acquisition
- Prepare imaging sequence. Prepare the imaging sequence one time and use this same sequence for all participants.
- Set parameters according to Table 1.
- Place an MRI phantom at the center of the MRI and run the imaging sequence.
NOTE: Because this sequence requires fast gradient performance and many RF pulses, it is important to verify that the protocol setup can be run prior to testing in a human.
- Prepare the participant for MRI. Use institutional-standard MRI safety screening to ensure the participant can safely enter the MRI.
- Position the participant on the MRI bed and place a chest coil over the participant's torso. Position the coil close to the participant's chin in order to ensure full coverage of the lung apices.
- Move the participant into the MRI scanner. Place the positioning landmark just below the sternum of the participant.
- Collect a localizer scan to ensure that the participant's lungs are within the field of view for the UTE scan. Do not move the geometry of the UTE scan. If the participant's lungs are not within the field of view, move the participant and collect additional localizer scans until the lungs are fully within the field of view.
- Run the UTE sequence. During this sequence, the participant can breathe normally.
- Export the raw data from the scanner. Depending on the imaging sequence used, the scanner may or may not reconstruct images on the scanner. For the proposed retrospective gating reconstruction, raw imaging data is required to determine whether or not images are generated on the scanner. Note the raw data will be large (>10 GB).
- Export or calculate k-space trajectories (i.e., the location in k-space of every raw data point).
NOTE: For some imaging sequences, k-space trajectories may be stored alongside raw data on the MRI scanner and can be directly exported. For other imaging sequences, the k-space trajectories will need to be calculated based on imaging parameters.
| Parameter | Generic Recommended Settings | Settings Implemented Herein |
| Imaging Sequence | 3D Non-Cartesian with Center-out k-space trajectories | 3D Radial with Golden Means Projection ordering |
| Field-of-View | 400 x 400 x 400 mm3 | 400 x 400 x 400 mm3 |
| Matrix Size | As desired for target resolution | 320 x 320 x 320 (1.25 mm isotropic resolution) |
| Bandwidth | As needed for readout duration < 1.0 ms | 888 Hz/Pixel |
| TE | < 0.1 ms | 0.07 ms |
| TR | Minimum (Target 3 – 4 ms) | 3.5 ms |
| Flip Angle | Approximately 5° | 4.8° |
| Number of Projections | Minimum 100,000 | 1,35,386 |
| Image Duration | Minimum 5 min | 7 min, 54 s |
Table 1: Recommended settings for UTE imaging. Generic recommended settings are provided that can be used to guide protocol setup. Specific recommended settings that were used for the data are also provided, as shown as representative results. Parameter specifications are generic across vendors, except for bandwidth. Some major MRI vendors specify bandwidth as Hz/Pixel. Other major MRI vendors specify absolute bandwidth. The recommended bandwidth (888 Hz/Pixel) corresponds to an absolute bandwidth of 284,160 Hz.
2. UTE image reconstruction using image-based respiratory soft-gating
NOTE: MATLAB code to complete the following steps is provided at https://github.com/pniedbalski3/UTE_Reconstruction.
- Import data and k-space trajectories into MATLAB. Code for importing raw MRI data is available for all of the major MRI vendors.
- Discard the first 1000 projections to ensure that data is at steady-state magnetization.
NOTE: If the imaging sequence used includes dummy scans prior to data acquisition, this step can be skipped. - Reconstruct a low-resolution image using a very small subset of data.
- Reconstruct the image using a non-uniform fast Fourier transform to a matrix size of 96 x 96 x 96.
- Use approximately 200 projections, corresponding to 0.6 s to 0.8 s worth of data.
- Reconstruct and store images from all coil elements as well as a final, coil-combined image.
- In the resulting coil-combined image, select a coronal slice that clearly shows the diaphragm.
NOTE: The provided code will prompt the user to select a slice containing the diaphragm. - Once this slice has been selected, view the individual coil images for this slice and select one or two coil elements that best show the diaphragm (Figure 2).
NOTE: The provided code will prompt the user to select coil elements. - Reconstruct images using a sliding window to generate images with ~0.5 s temporal resolution (Figure 2).
- Reconstruct only the data from the coil elements selected in step 2.4.
NOTE: While all coil elements can be reconstructed, only the elements closest to the diaphragm are needed to visualize the diaphragm for the purposes of respiratory self-gating. By reconstructing only the coil elements closest to the diagram, the reconstruction time and computational burden is drastically reduced. - Use the first 200 projections to reconstruct an image using a non-uniform fast Fourier transform (Figure 2). Store only the slice showing the diaphragm (as found in step 2.4).
NOTE: Ultimately, up to 1500 images will be generated; only a 2D slice is needed to visualize the diaphragm position, and storing 3D images for each of the sliding window steps would be prohibitive. - Shift by 100 projections (i.e., the first image is reconstructed using projections 1-200. The second is reconstructed using projections 101 - 300) and reconstruct an additional image, storing the slice selected in step 2.4.
- Continue until all projections have been used to generate images.
- Reconstruct only the data from the coil elements selected in step 2.4.
- Select a line over the diaphragm in the first of the sliding window images. Ensure that the line is long enough to extend into the lungs by 5-10 voxels and into the diaphragm by 5-10 voxels.
- Visualize respiratory motion by viewing this respiratory navigator for all projections.
- Determine the location of the diaphragm for all respiratory navigators. There are a variety of ways to do this, but a straightforward method is to use Otsu's method25 to divide the darker side (lung) from the brighter side (diaphragm).
- Use the diaphragm location to label projections as belonging to a given respiratory bin. If a given respiratory navigator shows the diaphragm at "position 1", then the 200 projections used to generate the image for that navigator would belong to "bin 1".
NOTE: Because images were generated using a sliding window with a 100-projection overlap, some projections may be labeled as belonging to multiple bins. The coarse spatial resolution of sliding window images leads to a total of ~4-6 bins that cover the full range of inspiration to expiration. - Select the bin to reconstruct by determining which bin has the greatest number of projections, which should correspond to end expiration.
- Alternatively, reconstruct images for the desired respiratory phases based on visual inspection of the respiratory navigator.
- Generate weights for soft-gating14.
- Use an exponential filter to provide a weight of 1 to projections within the primary bin and a sharply reducing weight to projections within different respiratory bins.
- Use the Berkely Advanced Reconstruction Toolbox (BART; https://mrirecon.github.io/bart/)26,27 to reconstruct a high-resolution image at the desired respiratory bin.
NOTE: BART is a freely available toolbox for MRI image reconstruction.- Calculate density compensation weights using iterative density combination.
- Scale the density compensation weights by the soft-gating weights.
- Scale data based on density compensation and soft-gating weights
- Perform a basic non-uniform fast Fourier transform (NUFFT) to facilitate coil combination.
- Convert the NUFFT image into gridded k-space to be used for coil combination.
- Generate a coil combination matrix and use it to combine coils for both the raw data and the gridded k-space.
- Estimate coil sensitivities.
- Perform parallel imaging compressed sense reconstruction using the weighted density compensation, coil combined data, and coil sensitivity maps.
- Save the final image. The NIFTI format is easily implemented. If the image is to be uploaded to a PACs system, a DICOM format may be required.
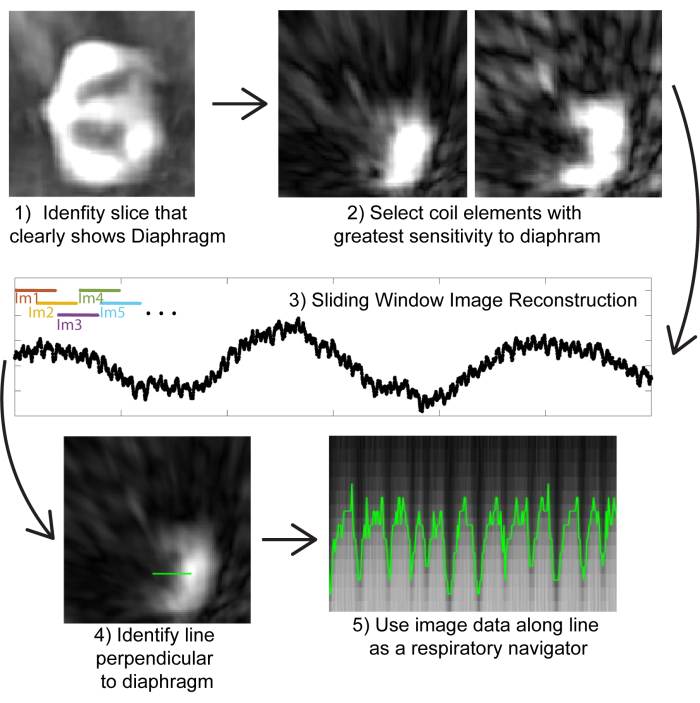
Figure 2: Image-based self gating. (1) Using a low-resolution image reconstructed from a small number of projections (for computational efficiency), identify a coronal slice that clearly shows the diaphragm. (2) By examining images from individual coil elements, select the coil elements that are closest to the diaphragm. (3) Performing a sliding window reconstruction only of the coil elements closest to the diaphragm (for computational efficiency). Images can be generated from subsets of 200 projections (corresponding to ~0.8 s); by overlapping projections, a pseudo-temporal resolution of ~0.5 s can be achieved in images. (4) Identifying a line that is perpendicular to the diaphragm to be used as a respiratory navigator. (5) Visualizing the image data on this line shows respiratory motion, which can be used to bin images. Please click here to view a larger version of this figure.
3. UTE image reconstruction using k-space-based respiratory soft-gating
- Complete steps 2.1-2.4 so that the coil element closest to the diaphragm can be identified.
- Generate a k0 time series trace by using the absolute value of the first point on the projection for all projections for the selected coil element. This will provide a visualization of a respiratory waveform.
- In steps of 5000 projections, normalize k0 by the mean signal intensity of those same k0 points28. This mitigates signal intensity drift over time and provides an improved ability to quantitatively bin projections.
- Label each k0 point as occurring during inspiration or expiration.
- Smooth the k0 time series and take the derivative to assess the slope for every point on the gating trace.
- Label inspiration points based on the sign of the slope. A positive slope corresponds to expiration, while a negative slope corresponds to inspiration.
- Bin projections based on signal intensity. Because the depth of breathing can be variable, bin projections are based on signal amplitude rather than location in the respiratory phase.
NOTE: A simple and rapid method by which to accomplish this is to implement k-means clustering to identify different signal intensity levels. - For bins intermediate between end-inspiration and end-expiration, identify projections as occurring during inspiration and expiration based on step 3.4.
- Complete image reconstruction following the steps provided in step 2.10 through step 2.13.
- If desired, reconstruct images for all respiratory bins rather than only at end-expiration.
Results
Representative results (Figure 3) were generated using the settings shown in Table 1. The imaging duration used provides high-quality images that are tolerable by most participants.
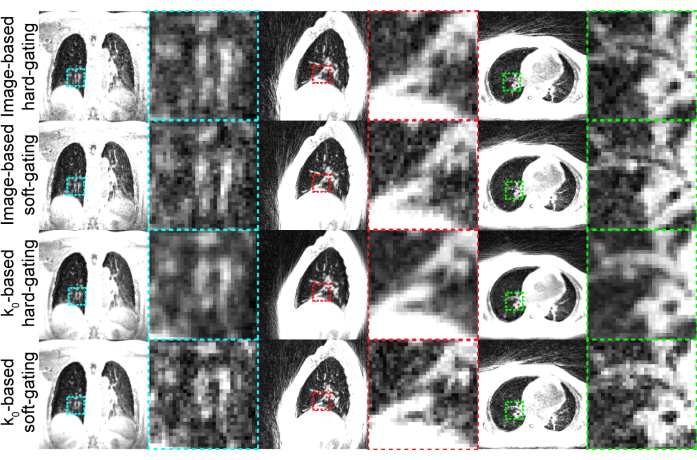
Figure 3: Representative UTE images generated. Coronal, sagittal, and axial slices of ima...
Discussion
When performing UTE imaging of the lungs, many variations of both acquisition and reconstruction can be used to generate images of the lungs. This protocol focuses on ease of implementation and computational efficiency. Imaging using 3D radial UTE is relatively simple, with imaging sequences generally available from the major MRI vendors. MATLAB-based tools are provided for data handling and self-gating. Because most academic institutions have access to MATLAB licenses, this code should be broadly useable and easily impl...
Disclosures
Peter Niedbalski receives research funding from the National Scleroderma Foundation, the American Heart Association, and the NIH. He is a consultant for Polarean Imaging Plc., a company that develops hyperpolarized 129Xe MRI technology.
Acknowledgements
The development of this protocol and the images shown as representative results were supported by the National Scleroderma Foundation.
Materials
| Name | Company | Catalog Number | Comments |
| Chest MRI Coil | Siemens, GE, Philips,, Other Clinical MRI Imaging Coil Vendor | N/A | A 26 - 32 channel Chest coil should be used |
| High Performance Workstation | HP, Apple, or other Computer Hardware company | N/A | A computer with a minimum of 64 GB of Memory is needed for image reconstruction |
| Matlab | Mathworks | R2016A or newer | A Matlab license is needed to run the provided computer code |
| MRI Phantom | Siemens, GE, Philips, or Other MRI Phantom Vendor | N/A | Any Phantom can be used to test the MRI sequence prior to its use in human subjects. |
| MRI Scanner | Siemens, GE, Philips, or Other Clinical MRI Scanner Vendor | N/A | The protocol was developed on a 3T scanner, but 1.5T or 0.55T would also work with minimal adaptation |
References
- Raju, S., Ghosh, S., Mehta, A. C. Chest ct signs in pulmonary disease: A pictorial review. Chest. 151 (6), 1356-1374 (2017).
- Biederer, J., et al. MRI of the lung (2/3). Why, when, how. Insights Imaging. 3 (4), 355-371 (2012).
- Biederer, J., et al. MRI of the lung (3/3)-current applications and future perspectives. Insights Imaging. 3 (4), 373-386 (2012).
- Johnson, K. M., Fain, S. B., Schiebler, M. L., Nagle, S. Optimized 3D ultrashort echo time pulmonary MRI. Magn Reson Med. 70 (5), 1241-1250 (2013).
- Ma, W., et al. Ultra-short echo-time pulmonary MRI: Evaluation and reproducibility in COPD subjects with and without bronchiectasis. J Magn Reson Imaging. 41 (5), 1465-1474 (2015).
- Roach, D. J., et al. Ultrashort echo-time magnetic resonance imaging is a sensitive method for the evaluation of early cystic fibrosis lung disease. Ann Am Thorac Soc. 13 (11), 1923-1931 (2016).
- Tibiletti, M., et al. Multistage three-dimensional UTE lung imaging by image-based self-gating. Magn Reson Med. 75 (3), 1324-1332 (2016).
- Weick, S., et al. Dc-gated high-resolution three-dimensional lung imaging during free-breathing. J Magn Reson Imaging. 37 (3), 727-732 (2013).
- Fischer, A., et al. Self-gated non-contrast-enhanced functional lung imaging (SENCEFUL) using a quasi-random fast low-angle shot (FLASH) sequence and proton MRI. NMR Biomed. 27 (8), 907-917 (2014).
- Tibiletti, M., et al. Respiratory self-gated 3D UTE for lung imaging in small animal MRI. Magn Reson Med. 78 (2), 739-745 (2017).
- Jiang, W., et al. Motion robust high resolution 3D free-breathing pulmonary MRI using dynamic 3D image self-navigator. Magn Reson Med. 79 (6), 2954-2967 (2018).
- Higano, N. S., et al. Retrospective respiratory self-gating and removal of bulk motion in pulmonary UTE MRI of neonates and adults. Magn Reson Med. 77 (3), 1284-1295 (2016).
- Metze, P., et al. Non-uniform self-gating in 2D lung imaging. Front Phys. 10, (2022).
- Gandhi, D. B., et al. Comparison of weighting algorithms to mitigate respiratory motion in free-breathing neonatal pulmonary radial UTE-MRI. Biomed Phys Eng Express. 10 (3), 035030 (2024).
- Fauveau, V., et al. Performance of spiral UTE-MRI of the lung in post-covid patients. Magn Reson Imaging. 96, 135-143 (2023).
- Metz, C., et al. Comparison of diagnostic quality of 3D ultrashort-echo-time techniques for pulmonary magnetic resonance imaging in free-breathing. Acta Radiologica. 64 (5), 1851-1858 (2023).
- Periaswamy, G., et al. Comparison of ultrashort TE lung MRI and HRCT lungs for detection of pulmonary nodules in oncology patients. Indian J Radiol Imaging. 32 (04), 497-504 (2022).
- Darçot, E., et al. Comparison between magnetic resonance imaging and computed tomography in the detection and volumetric assessment of lung nodules: A prospective study. Frontiers in Medicine. 9, 858731 (2022).
- Dournes, G., et al. 3D ultrashort echo time MRI of the lung using stack-of-spirals and spherical k-space coverages: Evaluation in healthy volunteers and parenchymal diseases. J Magn Reson Imaging. 48 (6), 1489-1497 (2018).
- Campbell-Washburn, A. E. 2019 American Thoracic Society Bear Cage winning proposal: Lung imaging using high-performance low-field magnetic resonance imaging. Am J Respir Crit Care Med. 201 (11), 1333-1336 (2020).
- Robison, R. K., Anderson, A. G., Pipe, J. G. Three-dimensional ultrashort echo-time imaging using a FLORET trajectory. Magn Reson Med. 78 (3), 1038-1049 (2017).
- Willmering, M. M., Robison, R. K., Wang, H., Pipe, J. G., Woods, J. C. Implementation of the FLORET sequence for lung imaging. Magn Reson Med. 82 (3), 1091-1100 (2019).
- Chan, R. W., Ramsay, E. A., Cunningham, C. H., Plewes, D. B. Temporal stability of adaptive 3D radial MRI using multidimensional golden means. Magn Reson Med. 61 (2), 354-363 (2009).
- Yu, J., Xue, Y., Song, H. K. Comparison of lung T2* during free-breathing at 1.5 T and 3.0 T with ultrashort echo time imaging. Magn Reson Med. 66 (1), 248-254 (2011).
- Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern. 9 (1), 62-66 (1979).
- Martin Uecker, F. O., et al. Berkely advanced reconstruction toolbox. Proc Intl Soc Magn Reson Med. 23, 2486 (2015).
- . . Bart Toolbox for Computational Magnetic Resonance Imaging. , (2024).
- Munidasa, S. . Treatment monitoring of pediatric cystic fibrosis lung disease using free breathing lung MRI. , (2024).
- Zhu, X., Chan, M., Lustig, M., Johnson, K. M., Larson, P. E. Z. Iterative motion-compensation reconstruction ultra-short TE (IMOCO UTE) for high-resolution free-breathing pulmonary MRI. Magn Reson Med. 83 (4), 1208-1221 (2020).
- Tan, F., et al. Motion-compensated low-rank reconstruction for simultaneous structural and functional UTE lung MRI. Magn Reson Med. 90 (3), 1101-1113 (2023).
- Bhattacharya, I., et al. Oxygen-enhanced functional lung imaging using a contemporary 0.55 T MRI system. NMR Biomed. 34 (8), e4562 (2021).
- Kim, M., et al. Feasibility of dynamic T2*-based oxygen-enhanced lung MRI at 3T. Magn Reson Med. 91 (3), 972-986 (2024).
- Klimeš, F., et al. 3D phase-resolved functional lung ventilation MR imaging in healthy volunteers and patients with chronic pulmonary disease. Magn Reson Med. 85 (2), 912-925 (2021).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved