Method Article
Fluorespirometria de alta resolução para avaliar mudanças dinâmicas no potencial da membrana mitocondrial em células imunes humanas
Neste Artigo
Resumo
Os métodos para estudar a bioenergética mitocondrial sob concentrações de substrato fisiologicamente relevantes em células imunes são limitados. Fornecemos um protocolo detalhado que usa fluorespirometria de alta resolução para avaliar as mudanças na resposta do potencial da membrana mitocondrial à demanda de energia em células T humanas, monócitos e células mononucleares periféricas.
Resumo
As células mononucleares periféricas (PBMCs) exibem alterações robustas na capacidade respiratória mitocondrial em resposta à saúde e à doença. Embora essas mudanças nem sempre reflitam o que ocorre em outros tecidos, como o músculo esquelético, essas células são uma fonte acessível e valiosa de mitocôndrias viáveis de seres humanos. Os PBMCs são expostos a sinais sistêmicos que afetam seu estado bioenergético. Assim, expandir nossas ferramentas para interrogar o metabolismo mitocondrial nessa população elucidará os mecanismos relacionados à progressão da doença. Os ensaios funcionais das mitocôndrias geralmente se limitam ao uso de saídas respiratórias após concentrações máximas de substrato, inibidor e desacoplador para determinar toda a gama de capacidade respiratória, o que pode não ser alcançável in vivo. A conversão de difosfato de adenosina (ADP) em trifosfato de adenosina (ATP) pela ATP-sintase resulta em uma diminuição no potencial de membrana mitocondrial (mMP) e um aumento no consumo de oxigênio. Para fornecer uma análise mais integrada da dinâmica mitocondrial, este artigo descreve o uso de fluorespirometria de alta resolução para medir a resposta simultânea do consumo de oxigênio e do potencial de membrana mitocondrial (mMP) a concentrações fisiologicamente relevantes de ADP. Esta técnica usa éster metílico de tetrametilrodamina (TMRM) para medir a polarização de mMP em resposta a titulações de ADP após hiperpolarização máxima com substratos complexos I e II. Essa técnica pode ser usada para quantificar como as mudanças no estado de saúde, como envelhecimento e doenças metabólicas, afetam a sensibilidade da resposta mitocondrial à demanda de energia em PBMCs, células T e monócitos de seres humanos.
Introdução
A capacidade de uma célula de funcionar e sobreviver em um período de estresse fisiológico depende em grande parte de sua capacidade de atender ao requisito energético para restaurar a homeostase 1,2. A demanda de energia aumenta em resposta a uma variedade de estímulos. Por exemplo, o aumento da contração muscular durante o exercício aumenta a utilização de ATP e glicose pelo músculo esquelético, e um aumento na síntese de proteínas após a infecção aumenta a utilização de ATP pelas células imunes para a produção e proliferação de citocinas 3,4,5,6. Um aumento na demanda de energia desencadeia uma série de processos bioenergéticos para restaurar a relação ATP / ADP. À medida que o ATP é consumido, os níveis de ADP aumentam e estimulam a ATP-sintase F1F0 (complexo V), que requer uma força protonmotriz para conduzir sua rotação mecânica e conversão catalítica de ADP em ATP dentro da mitocôndria7. A força protonmotriz é um gradiente eletroquímico criado pelo bombeamento de prótons durante a transferência de elétrons de substratos para oxigênio através do sistema de transporte de elétrons (ETS) dentro da membrana mitocondrial interna. A diferença resultante na concentração de prótons (delta pH) e no potencial elétrico (potencial de membrana) cria a força protonmotriz que impulsiona a síntese de ATP e o consumo de oxigênio em resposta à demanda de energia, reduzindo a relação ATP / ADP ou aumentando os níveis de ADP. A afinidade das mitocôndrias ao ADP pode ser determinada pelo cálculo do Km ou EC50 da respiração estimulada por ADP de mitocôndrias isoladas ou células permeabilizadas 8,9. Este método mostrou que as fibras musculares permeabilizadas de humanos mais velhos requerem uma concentração maior de ADP para estimular 50% de sua capacidade máxima de fosforilação oxidativa do que as de indivíduos mais jovens9. Da mesma forma, o envelhecimento do músculo esquelético de camundongos requer mais ADP para diminuir a produção de espécies reativas de oxigênio mitocondriais (ROS) 10 , 11 . Além disso, a sensibilidade ao ADP é reduzida em fibras musculares permeabilizadas de camundongos com obesidade induzida por dieta em relação aos controles e é aumentada na presença de insulina e após o consumo de nitrato12,13. Assim, a capacidade das mitocôndrias de responder à demanda de energia varia sob diferentes condições fisiológicas, mas isso não foi explorado anteriormente no contexto das células imunes.
As células mononucleares do sangue periférico (PBMCs) são comumente usadas para investigar a bioenergética celular em seres humanos 14,15,16,17,18,19,20. Isso se deve em grande parte ao fato de as células serem facilmente obtidas a partir de amostras de sangue não coagulado em estudos clínicos, à capacidade de resposta das células a perturbações metabólicas e aos métodos desenvolvidos por vários grupos para interrogar o metabolismo mitocondrial usando inibidores e desacopladores para determinar a capacidade máxima e mínima da respiração mitocondrial21,22. Esses métodos levaram a uma apreciação dos papéis da bioenergética no envelhecimento, doenças metabólicas e função imunológica 14,20,23,24. A capacidade respiratória mitocondrial é frequentemente reduzida no músculo esquelético e PBMCs em condições de insuficiência cardíaca 18,25. A bioenergética da PBMC também está correlacionada com fatores de risco cardiometabólicos em adultos saudáveis17 e responde a tratamentos como o ribosídeo de nicotinamida18. As PBMCs incluem neutrófilos, linfócitos (células B e células T), monócitos, células natural killer e células dendríticas, que contribuem para a capacidade mitocondrial da PBMC26 , 27 , 28 . Além disso, a bioenergética celular desempenha um papel crucial na ativação, proliferação e renovação das células imunes23. No entanto, uma limitação desses métodos é que as células não estão funcionando sob uma faixa fisiológica de substratos. Métodos adicionais são, portanto, necessários para interrogar a função mitocondrial em concentrações de substrato que são mais relevantes para o que as células experimentam in vivo.
O potencial de membrana mitocondrial (mMP) é o principal componente de uma força protonmotriz e é essencial para uma variedade de processos mitocondriais além da produção de ATP, como regulação do fluxo respiratório, produção de espécies reativas de oxigênio, importação de proteínas e íons, autofagia e apoptose. O mMP pode ser avaliado com sondas eletroquímicas ou corantes fluorescentes sensíveis a mudanças na polarização da membrana como JC-1, Rhod123, DiOC6, tetrametil rodamina (TMRE) ou éster metílico (TMRM) e safranina. Os dois últimos são corantes catiônicos lipofílicos que têm sido usados com sucesso na fluorespirometria de alta resolução de homogeneizados teciduais, mitocôndrias isoladas e tecido permeabilizado 11,29,30,31,32,33. Nesta técnica, o TMRM é usado no modo de têmpera, onde as células são expostas a uma alta concentração de TMRM que se acumula na matriz mitocondrial quando polarizada (alta mMP e força protonmotriz), resultando na extinção da fluorescência citosólica de TMRM. Quando as mitocôndrias se despolarizam em resposta ao ADP ou desacopladores, o corante é liberado da matriz, aumentando o sinal fluorescente TMRM34,35. O objetivo deste método é medir simultaneamente as mudanças na respiração mitocondrial e mMP em resposta a titulações de ADP em PBMCs derivados de humanos, monócitos circulantes e células T, e também pode ser aplicado a células T esplênicas de camundongos.
Protocolo
A coleta de amostras de sangue para desenvolvimento de dados e métodos aqui apresentados foi aprovada pelo Comitê de Revisão Interna da Universidade de Washington. Os resultados representativos também incluem dados de camundongos C57BL / 6J machos (5-7 meses de idade) adquiridos da Jackson Laboratories. Todos os procedimentos com animais foram aprovados pelo Escritório de Bem-Estar Animal da Universidade de Washington. A visão geral do protocolo é mostrada na Figura 1. A preparação do reagente para este protocolo pode ser encontrada no Arquivo Suplementar 1.
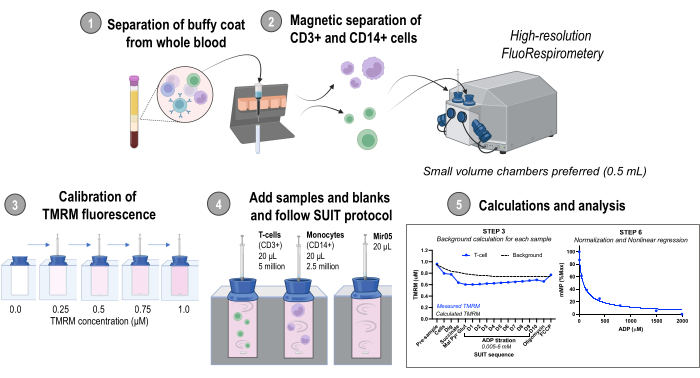
Figura 1: Visão geral do protocolo. Fluxo de trabalho usando fluorespirometria de alta resolução para avaliar alterações no potencial da membrana mitocondrial em monócitos isolados (CD14+) e células T (CD3+) de amostras frescas de sangue humano. Abreviaturas: TMRM, éster metílico de tetrametilrodamina; SUIT, titulações de substrato-desacoplador-inibidor; ADP, difosfato de adenosina; Dig, digitonina; Mal, malato; Piruvato, piruvato; Glut, glutamato; D1-10, 10 titulações consecutivas de ADP. Clique aqui para ver uma versão maior desta figura.
1. Separação da pelagem amarelada do sangue total
NOTA: O isolamento celular é modificado de Kramer et al.27.
- Permita que RPMI, gradiente de densidade e centrífuga atinjam a temperatura ambiente. Esterilize o gabinete de biossegurança e os materiais antes de começar.
- Colete sangue venoso em três tubos de 10 mL K2EDTA. Inverta os tubos pelo menos 3 vezes.
- Centrifugar os tubos a 500 x g 10 min (22 °C, 9 acelerações [acc], 2 desacelerações [dec]).
- Remover 1 ml de plasma de cada tubo e conservá-lo a -80 °C para análises futuras.
- Transfira o plasma e metade da camada de glóbulos vermelhos de cada tubo para um único tubo cônico de 50 mL. Adicione RPMI até a marca de 40 mL. Inverta pelo menos 3 vezes.
- Camada lenta de 10 mL da solução de plasma em quatro tubos cônicos de 15 mL contendo 3 mL do gradiente de densidade.
- Centrifugue a 700 x g por 30 min (22 °C, 5 acc, 2 dec).
- Colete todo o plasma e o revestimento leucocitário contendo as células mononucleares periféricas (PBMCs) sem interromper os glóbulos vermelhos.
- Centrifugue a 500 x g por 10 min (22 ° C, 5 acc, 5 dec) e aspire o sobrenadante.
- Lave o pellet PBMC 1x-2x ressuspendendo-o em 10 mL de RPMI e centrifugando a 500 x g por 10 min (22 ° C, 5 acc, 5 dec).
2. Separação magnética de células CD14+ e CD3+
- Coloque uma coluna no campo magnético de um separador de célula magnética (consulte a Tabela de Materiais). Lave a coluna com 3 mL de RP-5.
- Ressuspenda o pellet PBMC em 80 μL de RP-5 e 20 μL de microesferas anti-CD14 (consulte a Tabela de Materiais). Incubar durante 15 min a 4 °C.
- Ressuspenda as células com 1 mL de RP-5 e carregue a suspensão na coluna. Colete células não marcadas que fluem para um tubo cônico de 15 mL rotulado como "Flow-through 1". Aguarde até que toda a suspensão celular tenha passado pela coluna e, em seguida, continue lavando com 3 mL de RP-5 3x, coletando todo o fluxo.
- Remova cuidadosamente a coluna do campo magnético e coloque-a em um novo tubo cônico de 15 mL. Adicione 5 mL de RP-5 e use imediatamente o êmbolo para purgar o conteúdo da coluna em um tubo de coleta rotulado como "CD14+".
- Centrifugar "Flow-through 1" a 500 x g durante 10 min (22 °C, 5 acc, 5 dec) e aspirar o sobrenadante.
- Usando células de Flow-through 1, repita as etapas 2.2-2.5 usando microesferas anti-CD3 (consulte a Tabela de Materiais) para isolar as células T.
- Tubos de centrífuga contendo células T (CD3+) e monócitos (CD14+) a 300 x g por 5 min. Aspire o sobrenadante e ressuspenda o pellet em 1 mL de RP-5.
- Determine a concentração de células usando um hemocitômetro ou contador automático de células.
NOTA: As células podem ser contadas adicionando 10 μL de uma diluição celular de 1:10 ou 1:20 a um hemocitômetro. Pode-se referir a protocolos publicados anteriormente para contagem de células usando um hemocitômetro36. - Pipete 2,5 milhões de monócitos ou 5 milhões de células T em um novo tubo de centrífuga. Centrifugar durante 30 s a 2000 x g, aspirar o sobrenadante e ressuspender as células em MiR05 para um volume total de 20 μL e concentração final de 125 milhões de monócitos ou 250 milhões de células T por ml.
NOTA: A concentração final foi selecionada para injetar 2,5 milhões de monócitos ou 5 milhões de células T em um volume de 20 μL. Células T de camundongos isoladas do baço também foram testadas usando este método. O procedimento encontra-se no Processo Suplementar 1.
3. Fluorespirometria de alta resolução - Calibração de fluorescência de oxigênio e TMRM
NOTA: Este método foi adaptado de trabalhos anteriores realizados em fibras permeabilizadas por Pharaoh et al.11. Uma concentração alta e não inibitória de TMRM é usada para o modo de têmpera, onde a relação da concentração de mMP e TMRM na matriz é invertida. Assim, uma diminuição na mMP leva à liberação do corante TMRM da matriz e a um aumento na fluorescência32.
- Instale câmaras de 0.5 mL no respirômetro O2K de acordo com as instruções do fabricante (consulte a Tabela de Materiais). Ligue o instrumento e conecte-o ao software fornecido pelo fabricante para aquisição de dados.
- Ajustar a temperatura para 37 °C e agitar para 750 rpm.
- Lave as câmaras com água destilada 3x. Substitua a água por 0,54 mL de Mir05, feche totalmente as rolhas e remova o excesso de tampão com o sistema de sucção integrado (ISS). Levante as rolhas para permitir que o oxigênio ambiente se equilibre com o oxigênio da câmara usando um espaçador de rolha.
- Quando o fluxo de oxigênio estiver estável, execute a calibração de oxigênio do ar (R1) de acordo com as instruções do fabricante.
NOTA: Pode levar >30 minutos para que o fluxo de oxigênio se estabilize. Os sensores de oxigênio requerem a determinação do fluxo de oxigênio de ponto zero (R0) e oxigênio de fundo de 50-200 μM de experimentos separados usando titulações de ditionita. Métodos específicos podem ser encontrados no manual do fabricante. - Feche a câmara fechando as rolhas.
NOTA: Ao contrário dos experimentos que usam fibras permeabilizadas, as câmaras não requerem hiperoxigenação para PBMCs. A vedação da câmara após a calibração R1 fornece oxigênio suficiente para o experimento. Os níveis de oxigênio devem ser mantidos entre 50-250 μM. Se a concentração de oxigênio cair abaixo do limite, a câmara pode ser parcialmente aberta para que o oxigênio da câmara possa se equilibrar com o oxigênio do ar ambiente. - Calibração TMRM
- Use os sensores fluo de LED verde (ex. 525 nm) com o conjunto de filtros AmR (consulte a Tabela de Materiais). Defina o ganho do fluorômetro para 1000 e a intensidade para 1000. Ligue os sensores fluo e comece a gravar a linha de base.
- Injete 2,5 μL de 0,05 mM TMRM e deixe o sinal estabilizar (~ 2 min) antes da próxima injeção de 2,5 μL até que um total de 4 injeções sejam realizadas para uma concentração total de TMRM de 1 μM TMRM na câmara. Use uma seringa Hamilton para todas as injeções.
- Calibre o sensor fluo selecionando o sinal fluorescente (voltage) para cada injeção representando 0, 0.25, 0.5, 0.75 e 1.0 μM de TMRM para uma calibração de cinco pontos.
4. Protocolo de titulação substrato-desacoplador-inibidor (SUIT)
NOTA: Execute experimentos em branco onde 20 μL de Mir05 são injetados na câmara em vez de 20 μL de suspensão celular, pois o sinal TMRM mudará em resposta apenas às injeções (discutido em resultados representativos). Deixe o sinal de fluxo de oxigênio se estabilizar (cerca de 2-3 min) antes da próxima injeção para experimentos em branco e amostra. O seguinte protocolo de titulação e as observações esperadas estão na Tabela 1.
- Quando o fluxo de oxigênio estiver estável, selecione e rotule o fluxo de oxigênio e o sinal TMRM como "pré-célula".
- Injete a suspensão celular contendo 5 milhões de células T ou 2,5 milhões de monócitos em ~ 20 μL e meça por cerca de 10 min. Selecione e rotule o fluxo de oxigênio e o sinal TMRM como "Célula".
- Permeabilize as células injetando 2 μL de digitonina 1 mg/mL (concentração final: 4 μg/mL). Aguarde 20 min. Selecione e rotule o fluxo de oxigênio e o sinal TMRM como "Dig".
NOTA: Sugere-se otimizar a concentração de digitonina em experimentos separados. - Adicionar 2,5 μL de succinato 1 M (concentração final: 5 mM). Selecione e rotule o fluxo de oxigênio e o sinal TMRM como "SUCC".
- Quando o fluxo de oxigênio estiver estável, adicione 5 μL de malato 100 mM (concentração final: 1,0 mM), 5 μL de glutamato 1 M (concentração final: 10 mM) e 5 μL de piruvato 500 mM (concentração final: 5 mM). Selecione e rotule o fluxo de oxigênio e o sinal TMRM como "MPG".
- Quando o fluxo de oxigênio estiver estável, titule o ADP. Selecione as taxas para cada titulação e rotule-as como "D" sequencialmente de 1 a 10, dependendo do número de titulações. Use o esquema de titulação na Tabela 2.
- Quando o fluxo de oxigênio estiver estável, execute uma série de titulações de 1 μL de cianeto de carbonila p-(tri-fluromethoxy) fenil-hidrazona (FCCP) 0,25 mM até que o sinal fluorescente atinja seu máximo. Selecione e rotule o fluxo de oxigênio e o sinal TMRM que representam o potencial mínimo de membrana e rotule-o como "FCCP".
NOTA: A concentração de FCCP de 0,5-1,0 μM é geralmente necessária para esgotar o potencial da membrana mitocondrial.
CUIDADO: O FCCP é tóxico. Consulte a ficha de dados de segurança (SDS) para manuseio adequado. - OPCIONAL: Quando o fluxo de oxigênio estiver estável, injete 1 μL de rotenona 0,25 mM para inibir o complexo I e determinar a capacidade respiratória através do complexo II.
NOTA: As alterações no potencial de membrana não são mais relevantes após a titulação com o desacoplador.
CUIDADO: A rotenona é tóxica. Consulte o SDS para obter o manuseio adequado. - Quando o fluxo de oxigênio estiver estável, injete 1 μL de 1,25 mM (concentração final: 2,5 μM) de antimicina A para inibir a respiração mitocondrial.
CUIDADO: A antimicina A é tóxica. Consulte o SDS para obter o manuseio adequado.
5. Cálculo do potencial de membrana mitocondrial e análise
- Utilizando experiências em branco, registar os valores de TMRM calibrados (TMRM micromolar) antes da injecção da amostra em branco ("pré-amostra") e para cada uma das injecções. Veja a Figura 2.
- Para cada experimento em branco, calcule a proporção de fundo definindo a concentração de TMRM "pré-amostra" para 1,0. Calcule a diminuição proporcional subsequente no TMRM. Calcule a proporção média de fundo de todos os experimentos em branco.
NOTA: O número de experimentos em branco a serem incluídos pode depender da precisão do instrumento. Veja o exemplo de cálculo na Tabela 3 de cinco experimentos em branco diferentes, onde o desvio padrão da razão média de fundo caiu entre 0 e 0,016 para cada titulação. - Cálculo de fundo: Calcule o fundo para cada experimento de amostra multiplicando o TMRM "pré-amostra" do experimento de amostra pela proporção média de fundo para cada injeção. Veja o exemplo de cálculo na Tabela 3.
- Correção de fundo: Subtraia o fundo do experimento para os valores de TMRM medidos da amostra. Veja o exemplo de cálculo na Tabela 4.
- Correção FCCP: Subtraia o mMP corrigido em segundo plano do FCCP de cada injeção. Veja o exemplo de cálculo na Tabela 4.
- Curva de sensibilidade do ADP: Normalize a diminuição do mMP impulsionada pelo ADP, definindo o potencial de membrana mais alto e mais baixo como 100% e 0%, respectivamente, usando os valores de mMP coletados durante a titulação do ADP. Ajuste os dados em um modelo de regressão de ajuste não linear usando o software estatístico preferido para calcular a concentração inibitória semimáxima (IC50) de ADP em mMP.
NOTA: A curva se ajusta à resposta [Inibidor] vs. normalizada - Inclinação variável no prisma.
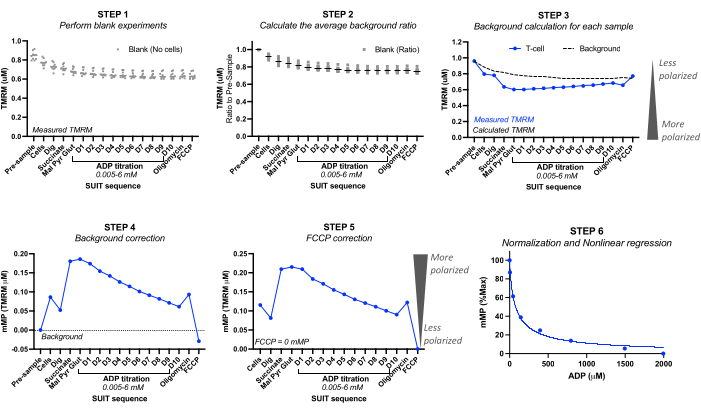
Figura 2: Cálculo do potencial de membrana mitocondrial (mMP) e sensibilidade ao ADP a partir da fluorescência TMRM. Etapas para calcular o potencial de membrana mitocondrial (mMP) e a sensibilidade ao ADP a partir de medições de fluorescência TMRM por fluorespirometria de alta resolução de uma amostra de células T (n = 1). Etapa 1: A fluorescência TMRM é medida em amostras em branco, conforme feito na amostra biológica. Etapa 2: Determine a proporção no sinal TMRM com cada titulação em relação ao sinal anterior à amostra para cada experimento em branco. Calcule a média para cada titulação de todos os experimentos em branco. Etapa 3: Calcule o fundo para cada experimento de amostra multiplicando a fluorescência "pré-amostra" pela proporção média de fundo para cada titulação. Etapa 4: Calcule a diferença entre a fluorescência TMRM de fundo e da amostra para cada titulação para expressar dados como mMP ou captação mitocondrial de TMRM. Etapa 5: corrija o mMP para que o desacoplamento total com o FCCP reflita zero mMP. Etapa 6: Realize a regressão não linear para representar graficamente as alterações no mMP com o aumento das concentrações de ADP. As medições foram realizadas em câmaras de 0,5 mL, uma contendo 5 milhões de células T de um voluntário saudável. Os dados médios são expressos como média ± SEM. Os pontos de dados únicos de uma única réplica são expressos sem barras de erro. Clique aqui para ver uma versão maior desta figura.
Resultados
Para ilustrar as diferenças na concentração celular ideal para o ensaio, 5 milhões de células T foram carregadas em uma câmara de 0,5 mL (10 milhões de células/mL) e 1,25 milhão de células foram carregadas em outra câmara (2,5 milhões de células/mL) contendo 1 μM TMRM (Figura 3A-G). Três experimentos em branco também foram incluídos para calcular o fundo TMRM. Descobrimos que uma concentração mais alta de células T resultou em uma mudança mais distinguível na fluorescência do TMRM em relação ao fundo ( Figura 3B , D ). Além disso, uma maior concentração celular nos permitiu detectar o aumento esperado no consumo de oxigênio e a depleção simultânea do mMP em resposta à adição de FCCP (Figura 3E,F). O uso de uma baixa concentração de células produziu uma mudança fraca na fluorescência paralela ao fundo. Como o cálculo de mMP subtrai o fundo do sinal, uma baixa concentração de células não permite a determinação de mudanças no mMP em resposta a substratos e desacopladores. Além de usar as concentrações mais altas de células neste ensaio, recomendamos manter a concentração de células constante para cada tipo de célula entre os experimentos.
Para validar a influência da ATP-sintase na dissipação de mMP com titulações de ADP, realizamos experimentos paralelos em PBMCs e células T onde uma câmara recebeu oligomicina antes da titulação de ADP (Figura 4). Não encontramos dissipação de mMP em resposta ao ADP em células tratadas com oligomicina, sugerindo que a diminuição gradual do mMP com ADP é resultado do fluxo de prótons através da ATP-sintase (Figura 4A-F). Também comparamos a sensibilidade do ADP entre células T e PBMCs do mesmo participante e descobrimos que a sensibilidade do ADP é menor (maior EC50) na fração de células T (Figura 4G, H).
Realizamos uma série de experimentos em branco para determinar a influência do tempo ou do protocolo SUIT na fluorescência TMRM. Descobrimos que o sinal TMRM em experimentos em branco é influenciado principalmente pelas titulações SUIT (Figura 5A) em oposição ao tempo das titulações (Figura 5B).
Comparamos as mudanças impulsionadas pelo ADP nas taxas de consumo de oxigênio (OCR) e no mMP em células T e monócitos de 11 voluntários saudáveis que vivem na comunidade (Figura 6A-H). Semelhante aos resultados de experimentos publicados anteriormente usando fluxo extracelular e ensaios enzimáticos, os monócitos exibiram maior capacidade respiratória mitocondrial do que os linfócitos26,27 (Figura 6A,H). No entanto, não detectamos um aumento típico de dose-resposta no OCR com ADP em nenhum dos tipos de células ( Figura 6C, D ), ao contrário do que este método mostra ao usar tecidos altamente metabólicos como o fígado de camundongo ( Figura 7A-H ). Por outro lado, o uso de TMRM nos permitiu detectar um declínio gradual na mMP com ADP em células imunes humanas (Figura 6E-G) e em células T esplênicas de camundongos (Figura 7E-H). Embora não tenhamos comparado diretamente as células T humanas e de camundongos usando o mesmo protocolo de titulação, descobrimos que o IC50 das células T de camundongos era menor por um fator de 10 em comparação com o das células T circulantes de seres humanos.
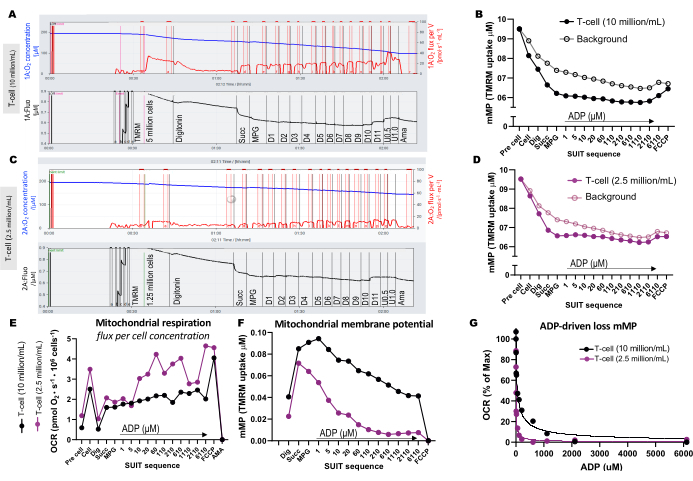
Figura 3: Experimentos de fluorespirometria de alta resolução. (A-D) Traço de experimentos de fluorespirometria de alta resolução usando concentrações de células T de 10 milhões de células/mL e 2,5 milhões de células/mL em câmaras de 0,5 mL. (A) 10 milhões de células/mL em câmaras de 0,5 mL. (C) 2,5 milhões de células/mL em câmaras de 0,5 mL. O fluxo de oxigênio (pmol/s/mL) é mostrado no painel superior (vermelho) e o sinal TMRM calibrado é mostrado no painel inferior (preto). As mudanças no TMRM ao longo do SUIT para a amostra e seu fundo calculado foram plotadas para as câmaras contendo (B) 10 milhões de células/mL e (D) 2,5 milhões de células/mL. (E) Para cada concentração celular, foram calculados o fluxo de oxigênio (pmol/s/milhão de células) e (F) o potencial de membrana mitocondrial. (G) A curva de sensibilidade do ADP foi plotada e ajustada a um modelo de regressão não linear (linhas sólidas). Abreviaturas: mMP, potencial de membrana mitocondrial; TMRM, éster metílico de tetrametilrodamina; SUIT, titulações de substrato-desacoplador-inibidor; ADP, difosfato de adenosina; Dig, digitonina; Mal, malato; Piruvato, piruvato; Glut, glutamato; D1-11, 11 titulações consecutivas de ADP; U, FCCP desacoplador de 0,5 e 1,0 μM; AMA, antimicina A. Clique aqui para ver uma versão maior desta figura.
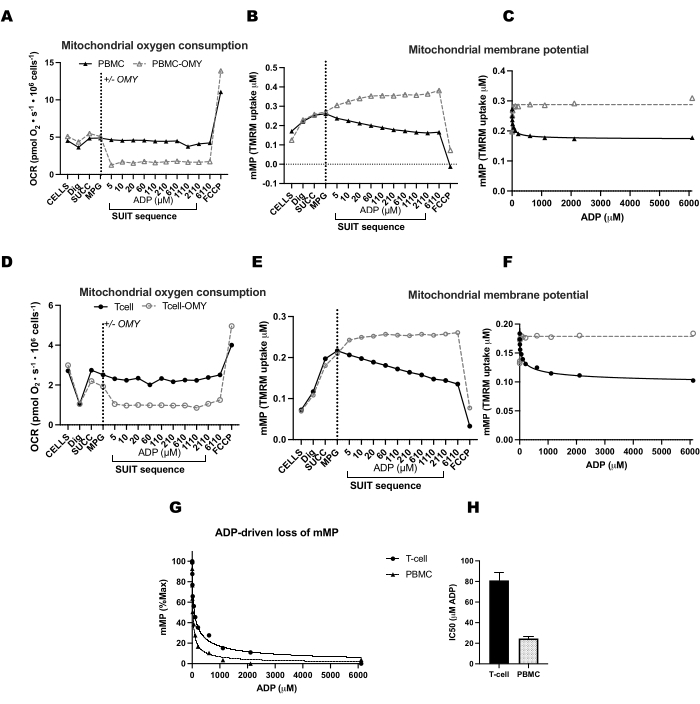
Figura 4: A ATP-sintase impulsiona a diminuição do potencial de membrana impulsionada pelo ADP em células T e PBMCs. (AH) O protocolo descrito aqui foi testado em PBMCs e células T. Duas câmaras de O2K foram injetadas com PBMCs e duas câmaras de um O2K adicional foram injetadas com células T do mesmo participante. Após a injeção dos substratos malato, piruvato e glutamato em todas as câmaras, uma câmara de PBMCs e células T recebeu oligomicina. A oligomicina impediu qualquer aumento da respiração impulsionado por ADP em (A) PBMCs e (D) células T ou declínio no potencial de membrana mitocondrial em (B, C) PBMCs e (E, F) células T. (G, H) A sensibilidade do ADP foi maior nas PBMCs em comparação com as células T. Clique aqui para ver uma versão maior desta figura.
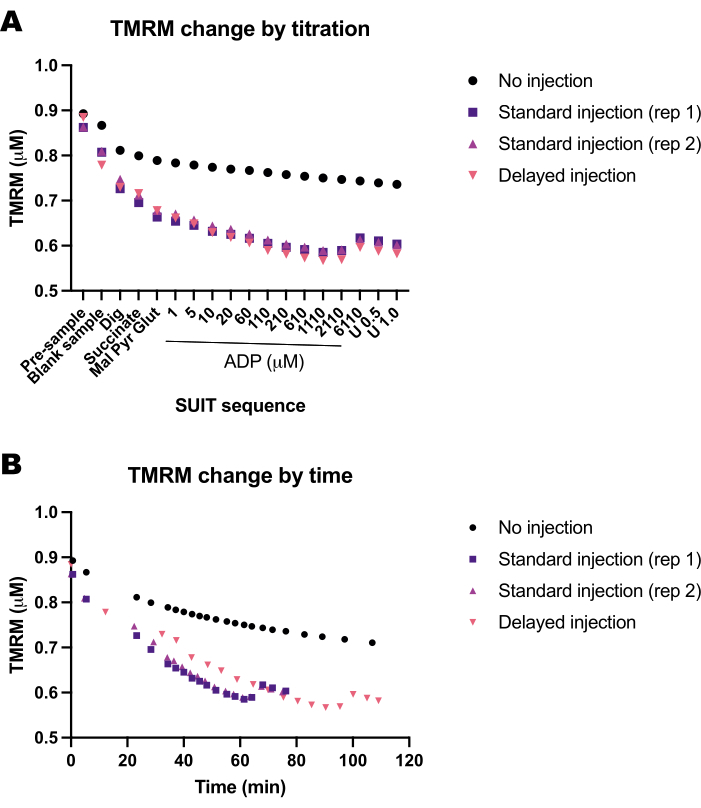
Figura 5: Experimentos em branco mostram a mudança na fluorescência TMRM em resposta ao tempo e titulações de substratos, desacopladores e inibidores (SUIT). (A) Mudança na fluorescência TMRM em resposta à titulação. (B) Mudança na fluorescência TMRM em resposta ao tempo. Os experimentos foram conduzidos em câmaras de 0,5 mL preenchidas com Mir05 contendo 1 μM de TMRM. Uma câmara não recebeu nenhuma titulação SUIT (sem injeção); duas câmaras em dois instrumentos diferentes receberam um protocolo de traje padrão (injeção padrão); uma câmara recebeu as mesmas titulações SUIT, mas com um atraso entre cada injeção (injeção retardada). Clique aqui para ver uma versão maior desta figura.
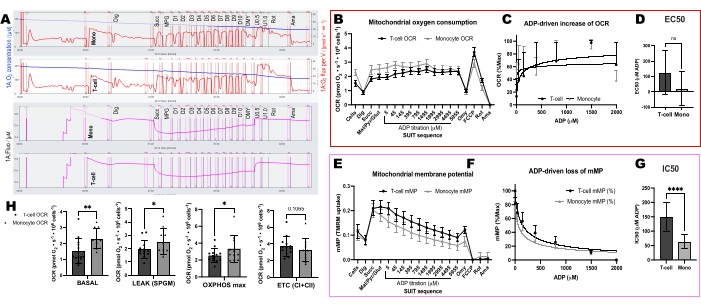
Figura 6: Diferenças na sensibilidade do ADP entre células T e monócitos usando OCR e mMP. (A) Traço do experimento de fluorespirometria de alta resolução da amostra de monócitos e células T de um sujeito. (B) Consumo de oxigênio em monócitos (n = 11) e células T (n = 13) do sangue de voluntários saudáveis. (C, D) Ajuste de regressão não linear do aumento plotado na respiração com titulações de ADP para calcular um EC50. (E) Medição simultânea do potencial de membrana mitocondrial. (F, G) Ajuste de regressão não linear do declínio plotado no potencial de membrana mitocondrial com titulações de ADP para calcular um IC50. (H) Parâmetros de capacidade respiratória de monócitos e células T. Os dados são expressos como média ± SEM para gráficos de linhas e média ± DP para gráficos de barras. As diferenças estatisticamente significativas após os testes t são expressas como *p < 0,05. **p < 0,01 e ****p < para 0,0001. Clique aqui para ver uma versão maior desta figura.
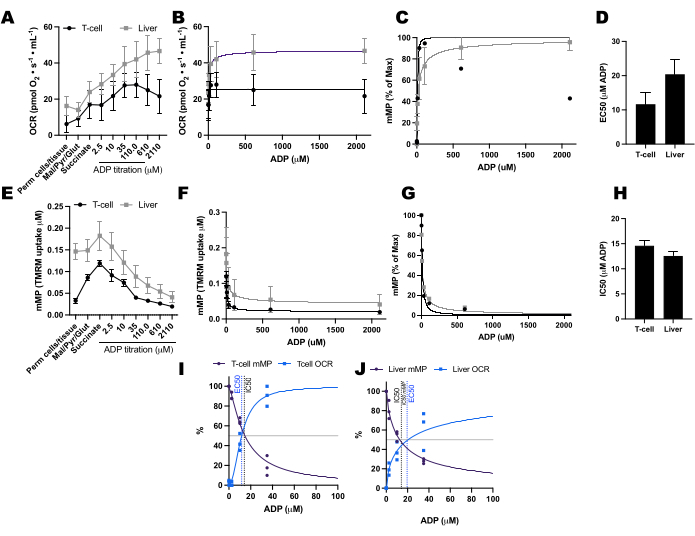
Figura 7: Comparando a resposta do ADP na respiração e o potencial de membrana mitocondrial (mMP) em células T esplênicas de camundongo permeabilizadas e fígado. (AD) Resposta na respiração em células T esplênicas de camundongo permeabilizadas e fígado. (E-H) Resposta em mMP em células T esplênicas de camundongo permeabilizadas e fígado. Fígado e baço frescos foram dissecados de três camundongos após luxação cervical. As células T Pan esplênicas foram isoladas usando separação de esferas magnéticas conjugadas com anticorpos. Ambas as amostras foram submetidas ao mesmo protocolo SUIT na presença de 1 μM de TMRM. (I, J) Comparação de EC50 calculada a partir do aumento do consumo de oxigênio (OCR) e IC50 a partir da diminuição da mMP em resposta ao ADP. N = 3 por grupo. Os dados são expressos como média ± SEM. Clique aqui para ver uma versão maior desta figura.
Tabela 1: Exemplo de protocolo SUIT para avaliar o potencial de membrana mitocondrial em células T e monócitos recém-isolados usando as câmaras de 0,5 mL. Clique aqui para baixar esta tabela.
Tabela 2: Titulação de ADP recomendada para câmara de 0,5 mL. Clique aqui para baixar esta tabela.
Tabela 3: Calculando a razão média de fundo usando cinco experimentos independentes em branco. Clique aqui para baixar esta tabela.
Tabela 4: Cálculo do potencial de membrana mitocondrial (mMP) do experimento de amostra. Clique aqui para baixar esta tabela.
Figura suplementar 1: Efeito de Mir05 e DMSO na respiração mitocondrial e no potencial de membrana. Clique aqui para baixar este arquivo.
Arquivo Suplementar 1: Preparação de reagentes e protocolo para isolamento de células T do baço de camundongo. Clique aqui para baixar este arquivo.
Discussão
Este protocolo usa fluorespirometria de alta resolução para medir a sensibilidade da resposta mitocondrial à demanda de energia, medindo a dissipação de mMP em resposta ao aumento dos níveis de ADP em PBMCs, monócitos e células T. Isso é feito adicionando substratos complexos I e II para maximizar o potencial da membrana mitocondrial e titulando o ADP para estimular gradualmente a ATP-sintase a usar o gradiente de prótons para a geração de ATP.
As etapas críticas do protocolo incluem definir o ganho e a intensidade do fluoróforo para 1000 e garantir que um sinal fluorescente TMRM seja adquirido durante a titulação do TMRM. Como a fluorescência TMRM diminui após cada titulação (uma limitação desse método), é imperativo executar experimentos em segundo plano usando amostras em branco. Também descobrimos que o DMSO tem um efeito inibitório na respiração mitocondrial e no potencial de membrana e, portanto, recomendamos diluir a solução de trabalho de TMRM em Mir05 (Figura Suplementar 1).
Algumas modificações que podem ser usadas ao tentar este protocolo são ajustar as concentrações celulares e usar a câmara padrão de 2 mL. No entanto, a câmara de 0,5 mL é preferida para células T e monócitos devido à alta concentração de células necessárias para uma resposta ideal no potencial de membrana e no fluxo de oxigênio. Uma concentração mais baixa de células pode ser ideal ao testar células com maior capacidade respiratória, como macrófagos.
Limitações adicionais do método apresentado aqui incluem a exigência de pelo menos 5 milhões de células T e 2,5 milhões de monócitos. Muitas vezes podemos obter células suficientes de ~ 20 mL de sangue de participantes saudáveis, mas esses números podem variar de acordo com o estado de saúde, idade e sexo26. Além disso, como na maioria dos métodos que avaliam a capacidade mitocondrial, as células precisam ser recém-isoladas. No entanto, este método pode ser tentado em células criopreservadas no futuro. Em comparação com o rendimento do sangue humano, o rendimento de células T do baço de camundongos saudáveis é alto o suficiente para realizar este ensaio.
As células T circulantes, particularmente as células de memória de longa duração (TM) e as células reguladoras (Treg), dependem da fosforilação oxidativa para obter energia37. Embora sua demanda de energia e consumo de oxigênio sejam baixos (por exemplo, em comparação com o músculo em repouso), sua sobrevivência é essencial para uma resposta imune eficaz à reinfecção e ao câncer 38,39,40. A redução da fosforilação oxidativa das células T resulta em comprometimento da capacidade proliferativa e promove exaustão e senescência das células T 5,41. Além disso, a hiperpolarização mitocondrial promove uma produção sustentada de citocinas (IL-4 e IL-21) pelas células T CD4 efetoras durante a ativação42. Após a infecção, a necessidade de energia para ativação e proliferação de células imunes pode chegar a 25% a 30% da taxa metabólica basal43. Portanto, as células imunes funcionam em uma ampla e extrema gama de demandas de energia, e este protocolo pode testar as respostas mitocondriais dentro dessa faixa.
A inflamação crônica é uma característica comum da obesidade, diabetes e envelhecimento. Níveis desregulados de hormônios, lipídios e glicose circulantes têm impactos sistêmicos e, portanto, podem afetar a forma como as mitocôndrias respondem a um desafio energético. Aqui, apresentamos um método para avaliar a sensibilidade do ADP mitocondrial em PBMCs circulantes. Mais estudos são necessários para determinar como a sensibilidade ao ADP pode ser modulada na doença metabólica e como isso afeta o estado de saúde.
Divulgações
Os autores declaram não haver conflitos de interesse.
Agradecimentos
Gostaríamos de agradecer aos gentis voluntários que doaram sangue para este projeto. Também estendemos nosso sincero agradecimento à Dra. Ellen Schur e sua equipe por nos fornecer amostras adicionais de seu estudo. Também gostaríamos de agradecer a Andrew Kirsh por revisar o manuscrito e editá-lo para legibilidade. Este trabalho foi apoiado pelas seguintes fontes de financiamento: P01AG001751, R01AG078279, P30AR074990, P30DK035816, P30DK017047, R01DK089036, K01HL154761, T32AG066574.
Materiais
| Name | Company | Catalog Number | Comments |
| Adenosine Diphosphate | Sigma-Aldrich | A5285 | Fluorespirometry |
| Antimycin A | Sigma-Aldrich | A8674 | Fluorespirometry |
| Bovine Serum Albumin (BSA) | Sigma-Aldrich | A6003 | Mir05 buffer |
| Bovine Serum Albumin | Sigma-Aldrich | A6003 | Cell isolation |
| Carbonyl cyanide 4-(trifluoromethoxy)phenylhydrazone | Sigma-Aldrich | C2920 | Fluorespirometry |
| Cell strainers | Fisher Scientific | 22-363-548 | Isolation of T-cells from mouse spleen protocol |
| CD14 Microbeads, human | Miltenyi Biotec | 130-050-201 | Cell isolation |
| CD3 Microbeads, human | Miltenyi Biotec | 130-050-101 | Cell isolation |
| DatLab | Oroboros | Version 8 | |
| Digitonin | Sigma-Aldrich | D141 | Fluorespirometry |
| D-Sucrose | Sigma-Aldrich | 84097 | Mir05 buffer |
| Ethylene glycol-bis(β-aminoethyl ether)-N,N,N′,N′-tetraacetic acid (EGTA) | Sigma-Aldrich | E4378 | Mir05 buffer |
| Filter Set AmR | Oroboros | 44321-01 | |
| HBSS (10x) | Gibco | 12060-040 | |
| HEPES sodium salt | Sigma-Aldrich | H7523 | Mir05 buffer |
| Histopaque 1077 | Sigma-Aldrich | 10771 | Cell isolation |
| K2EDTA blood collection tubes | BD Vacutainer | 366643 | Cell isolation |
| Lactobionic acid | Sigma-Aldrich | 153516 | Mir05 buffer |
| L-Glutamic acid | Sigma-Aldrich | G1626 | Fluorespirometry |
| L-Malic Acid | Sigma-Aldrich | M1000 | Fluorespirometry |
| LS Columns | Miltenyi Biotec | 130-042-401 | Cell isolation |
| Magnesium Chloride (MgCl2) | Sigma-Aldrich | M9272 | Mir05 buffer |
| Multi-MACS stand and MidiMACS Separator | Miltenyi Biotec | 130-042-301 | Cell isolation |
| O2k-Fluo Smart-Module | Oroboros | 12100-03 | |
| O2k-FluoRespirometer series J | Oroboros | 10201-03 | |
| O2k-sV-Module (0.5 chamber) | Oroboros | 11200-01 | |
| Oligomycin | Sigma-Aldrich | 04876 | Fluorespirometry |
| Pan T Cell Isolation Kit II, mouse | Miltenyi | 130095130 | Isolation of T-cells from mouse spleen protocol |
| Potassium dihydrogen phosphate (KH2PO4) | Sigma-Aldrich | P0662 | Mir05 buffer |
| Potassium Hydroxide (KOH) | Sigma-Aldrich | 221473 | Mir05 buffer |
| Prism | GraphPad | Version 10 | |
| Rotenone | Sigma-Aldrich | R8875 | Fluorespirometry |
| RPMI Buffer | Corning | 17-105-CV | Cell isolation |
| Sodium Pyruvate | Sigma-Aldrich | P2256 | Fluorespirometry |
| Succinate disodium salt | Sigma-Aldrich | S2378 | Fluorespirometry |
| Taurine | Sigma-Aldrich | T0625 | Mir05 buffer |
| Tetramethyrhodamine methyl ester perchlorite | Sigma-Aldrich | T5428 | Fluorespirometry |
Referências
- Eisner, V., Picard, M., Hajnóczky, G. Mitochondrial dynamics in adaptive and maladaptive cellular stress responses. Nat Cell Biol. 20 (7), 755-765 (2018).
- Sokolova, I. Bioenergetics in environmental adaptation and stress tolerance of aquatic ectotherms: linking physiology and ecology in a multi-stressor landscape. J Exp Biol. 224, Pt Suppl 1 236802(2021).
- Buttgereit, F., Brand, M. D. A hierarchy of ATP-consuming processes in mammalian cells). Biochem J. 312, Pt 1 163-167 (1995).
- Schmid, D., Burmester, G. R., Tripmacher, R., Kuhnke, A., Buttgereit, F. Bioenergetics of human peripheral blood mononuclear cell metabolism in quiescent, activated, and glucocorticoid-treated states. Biosci Rep. 20 (4), 289-302 (2000).
- Vardhana, S. A., et al. Impaired mitochondrial oxidative phosphorylation limits the self-renewal of T cells exposed to persistent antigen. Nat Immunol. 21 (9), 1022-1033 (2020).
- Nelson, S. R., Li, A., Beck-Previs, S., Kennedy, G. G., Warshaw, D. M. Imaging ATP consumption in resting skeletal muscle: One molecule at a time. Biophys J. 119 (6), 1050-1055 (2020).
- Meyrat, A., von Ballmoos, C. ATP synthesis at physiological nucleotide concentrations. Sci Rep. 9 (1), 3070(2019).
- Gouspillou, G., et al. Accurate determination of the oxidative phosphorylation affinity for ADP in isolated mitochondria. PLoS One. 6 (6), e20709(2011).
- Holloway, G. P., et al. Age-associated impairments in mitochondrial ADP sensitivity contribute to redox stress in senescent human skeletal muscle. Cell Rep. 22 (11), 2837-2848 (2018).
- Pharaoh, G., Brown, J., Ranjit, R., Ungvari, Z., Van Remmen, H. Reduced adenosine diphosphate sensitivity in skeletal muscle mitochondria increases reactive oxygen species production in mouse models of aging and oxidative stress but not denervation. JCSM Rapid Commun. 4 (1), 75-89 (2021).
- Pharaoh, G., et al. The mitochondrially targeted peptide elamipretide (SS-31) improves ADP sensitivity in aged mitochondria by increasing uptake through the adenine nucleotide translocator (ANT). Geroscience. 45 (6), 3529-3548 (2023).
- Brunetta, H. S., Petrick, H. L., Vachon, B., Nunes, E. A., Holloway, G. P. Insulin rapidly increases skeletal muscle mitochondrial ADP sensitivity in the absence of a high lipid environment. Biochem J. 478 (13), 2539-2553 (2021).
- Brunetta, H. S., et al. Nitrate consumption preserves HFD-induced skeletal muscle mitochondrial ADP sensitivity and lysine acetylation: A potential role for SIRT1. Redox Biol. 52, 102307(2022).
- Tyrrell, D. J., et al. Blood-cell bioenergetics are associated with physical function and inflammation in overweight/obese older adults. Exp Gerontol. 70, 84-91 (2015).
- Liepinsh, E., et al. Low-intensity exercise stimulates bioenergetics and increases fat oxidation in mitochondria of blood mononuclear cells from sedentary adults. Physiol Rep. 8 (12), e14489(2020).
- Hedges, C. P., et al. Peripheral blood mononuclear cells do not reflect skeletal muscle mitochondrial function or adaptation to high-intensity interval training in healthy young men. J Appl Physiol. 126 (2), 454-461 (2019).
- DeConne, T. M., Muñoz, E. R., Sanjana, F., Hobson, J. C., Martens, C. R. Cardiometabolic risk factors are associated with immune cell mitochondrial respiration in humans. Am J Physiol Heart Circ Physiol. 319 (2), H481-H487 (2020).
- Zhou, B., et al. Boosting NAD level suppresses inflammatory activation of PBMCs in heart failure. J Clin Invest. 130 (11), 6054-6063 (2020).
- Altintas, M. M., DiBartolo, S., Tadros, L., Samelko, B., Wasse, H. Metabolic changes in peripheral blood mononuclear cells isolated from patients with end stage renal disease. Front Endocrinol (Lausanne). 12, 629239(2021).
- Pence, B. D., Yarbro, J. R. Aging impairs mitochondrial respiratory capacity in classical monocytes). Exp Gerontol. 108, 112-117 (2018).
- vander Windt, G. J. W., Chang, C. H., Pearce, E. L. Measuring bioenergetics in T Cells using a Seahorse extracellular flux analyzer. Curr Protoc Immunol. 113, 11-14 (2016).
- Chacko, B. K., et al. Methods for defining distinct bioenergetic profiles in platelets, lymphocytes, monocytes, and neutrophils, and the oxidative burst from human blood. Lab Invest. 93 (6), 690-700 (2013).
- Buck, M. D., et al. Mitochondrial dynamics controls T Cell fate through metabolic programming. Cell. 166 (1), 63-76 (2016).
- Quinn, K. M., et al. Metabolic characteristics of CD8. Nat Commun. 11 (1), 2857(2020).
- Scandalis, L., et al. Skeletal muscle mitochondrial respiration and exercise intolerance in patients with heart failure with preserved ejection fraction. JAMA Cardiol. 8 (6), 575-584 (2023).
- Rausser, S., et al. Mitochondrial phenotypes in purified human immune cell subtypes and cell mixtures. Elife. 10, e70899(2021).
- Kramer, P. A., et al. Bioenergetics and the oxidative burst: protocols for the isolation and evaluation of human leukocytes and platelets. J Vis Exp. (85), e51301(2014).
- Kramer, P. A., Ravi, S., Chacko, B., Johnson, M. S., Darley-Usmar, V. M. A review of the mitochondrial and glycolytic metabolism in human platelets and leukocytes: implications for their use as bioenergetic biomarkers. Redox Biol. 2, 206-210 (2014).
- Teodoro, J. S., Machado, I. F., Castela, A. C., Rolo, A. P., Palmeira, C. M. The evaluation of mitochondrial membrane potential using fluorescent dyes or a membrane-permeable cation (TPP+) electrode in isolated mitochondria and intact cells. Methods Mol Biol. 2184, 197-213 (2020).
- Hassan, H., Zakaria, F., Makpol, S., Karim, N. A. A link between mitochondrial dysregulation and idiopathic autism spectrum disorder (ASD): Alterations in mitochondrial respiratory capacity and membrane potential. Curr Issues Mol Biol. 43 (3), 2238-2252 (2021).
- Williams, A. S., et al. Disruption of Acetyl-Lysine turnover in muscle mitochondria promotes insulin resistance and redox stress without overt respiratory dysfunction. Cell Metab. 31 (1), 131-147 (2020).
- Krumschnabel, G., Eigentler, A., Fasching, M., Gnaiger, E. Use of safranin for the assessment of mitochondrial membrane potential by high-resolution respirometry and fluorometry. Methods Enzymol. 542, 163-181 (2014).
- Chowdhury, S. R., Djordjevic, J., Albensi, B. C., Fernyhough, P. Simultaneous evaluation of substrate-dependent oxygen consumption rates and mitochondrial membrane potential by TMRM and safranin in cortical mitochondria. Biosci Rep. 36 (1), 00286(2015).
- Vianello, C., et al. High-throughput microscopy analysis of mitochondrial membrane potential in 2D and 3D models. Cells. 12 (7), (2023).
- Perry, S. W., Norman, J. P., Barbieri, J., Brown, E. B., Gelbard, H. A. Mitochondrial membrane potential probes and the proton gradient: a practical usage guide. Biotechniques. 50 (2), 98-115 (2011).
- JoVE, JoVE. Science Education Database. Basic Methods in Cellular and Molecular Biology. Using a Hemacytometer to Count Cells. JoVE. , Cambridge, MA. (2023).
- Geltink, R. I. K., Kyle, R. L., Pearce, E. L. Unraveling the complex interplay between T cell metabolism and function. Annu Rev Immunol. 36, 461-488 (2018).
- Sukumar, M., et al. Inhibiting glycolytic metabolism enhances CD8+ T cell memory and antitumor function. J Clin Invest. 123 (10), 4479-4488 (2013).
- Gerriets, V. A., et al. Foxp3 and Toll-like receptor signaling balance T. Nat Immunol. 17 (12), 1459-1466 (2016).
- MacIver, N. J., Michalek, R. D., Rathmell, J. C. Metabolic regulation of T lymphocytes. Annu Rev Immunol. 31, 259-283 (2013).
- Desdín-Micó, G., et al. T cells with dysfunctional mitochondria induce multimorbidity and premature senescence. Science. 368 (6497), 1371-1376 (2020).
- Yang, R., et al. Mitochondrial Ca2+ and membrane potential, an alternative pathway for Interleukin 6 to regulate CD4 cell effector function. Elife. 4, 06376(2015).
- Straub, R. H., Cutolo, M., Buttgereit, F., Pongratz, G. Energy regulation and neuroendocrine-immune control in chronic inflammatory diseases. J Intern Med. 267 (6), 543-560 (2010).
Reimpressões e Permissões
Solicitar permissão para reutilizar o texto ou figuras deste artigo JoVE
Solicitar PermissãoThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. Todos os direitos reservados