A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Oligomerization Dynamics of Cell Surface Receptors in Living Cells by Total Internal Reflection Fluorescence Microscopy Combined with Number and Brightness Analysis
In This Article
Summary
We describe an imaging approach for the determination of the average oligomeric state of mEGFP-tagged-receptor oligomers induced by ligand binding in the plasma membrane of living cells. The protocol is based on Total Internal Reflection Fluorescence (TIRF) microscopy combined with Number and Brightness (N&B) analysis.
Abstract
Despite the importance and ubiquity of receptor oligomerization, few methods are applicable for detecting clustering events and measuring the degree of clustering. Here, we describe an imaging approach to determine the average oligomeric state of mEGFP-tagged-receptor homocomplexes in the membrane of living cells. The protocol is based on Total Internal Reflection Fluorescence (TIRF) microscopy combined with Number and Brightness (N&B) analysis. N&B is a method similar to fluorescence-correlation spectroscopy (FCS) and photon counting histogram (PCH), which are based on the statistical analysis of the fluctuations of the fluorescence intensity of fluorophores diffusing in and out of an illumination volume during an observation time. In particular, N&B is a simplification of PCH to obtain information on the average number of proteins in oligomeric mixtures. The intensity fluctuation amplitudes are described by the molecular brightness of the fluorophore and the average number of fluorophores within the illumination volume. Thus, N&B considers only the first and second moments of the amplitude distribution, namely, the mean intensity and the variance. This is, at the same time, the strength and the weakness of the method. Because only two moments are considered, N&B cannot determine the molar fraction of unknown oligomers in a mixture, but it only estimates the average oligomerization state of the mixture. Nevertheless, it can be applied to relatively small time series (compared to other moment methods) of images of live cells on a pixel-by-pixel basis, simply by monitoring the time fluctuations of the fluorescence intensity. It reduces the effective time-per-pixel to a few microseconds, allowing acquisition in the time range of seconds to milliseconds, which is necessary for fast oligomerization kinetics. Finally, large cell areas as well as sub-cellular compartments can be explored.
Introduction
We describe a Total Internal Reflection Fluorescence-Number and Brightness (TIRF-N&B) imaging approach for determining the average oligomeric state of receptor molecules at the plasma membrane of live cells, aiming at linking the receptor assembly dynamics to the biological function of the proteins (Figure 1).
Upon extracellular ligand binding, receptors initiate the intracellular signal transduction depending on their conformation, oligomerization, potential co-receptors and membrane composition. Despite the importance and ubiquity of receptor oligomerization, recognized as a key event in cellular signaling1,2,3,4,5,6,7, few methods can detect clustering events and measure the degree of clustering experimentally8,9. The confocal volume (x,y ≈ 300 nm, z ≈ 900 nm) is insufficiently resolved for proving molecular interaction and stoichiometry, even after optimization by image restoration algorithms10. The sub-unit composition of protein oligomers cannot be resolved on a purely spatial basis even by super-resolution methods at x,y resolution of 20-70 nm such as PALM11, STORM12, and STED13. Moreover, their temporal resolution (in the order of minutes per image) cannot follow kinetics in the range of seconds. Single molecule step-bleaching resolves the stoichiometry of protein oligomers only if they are immobile14.
One of the most versatile methods to measure density and oligomerization of fluorescently tagged proteins within single images is the spatial intensity distribution analysis (SpIDA), which relies on spatial sampling. It is applicable to both chemically fixed and live cells, and allows the analysis of several regions of interest of the cell simultaneously using standard fluorescence microscopy15. Alternatively, moment methods, such as fluorescence-correlation spectroscopy (FCS)16, photon counting histogram (PCH)17, and Number and Brightness (N&B)18,19, are suitable for quantitative oligomer measurements. These methods analyze the fluorescence intensity fluctuations that can be observed in time when the fluorophores diffuse in and out of an illumination volume. The amplitudes of the intensity fluctuations can be uniquely described by the molecular brightness of the fluorophore (ε) and the average number of fluorophores (n) within the illumination volume17 (Figure 2). Typically, the diffusion coefficient of the fluorophores and the average number of molecules (inversely related to the G(0) value) within the illumination volume can be obtained by FCS20. However, since the diffusion time only scales with the cubic root of the mass, FCS is not sufficiently sensitive to detect changes in molecular mass21. In practice, single color FCS cannot detect dimerization of membrane receptors. PCH resolves mixtures of different oligomers accurately. Using more than two moments of the amplitude distribution, it detects molecules of different brightness that occupy the same illumination volume. Scanning FCS22 and developments, such as the interesting pair-correlation of molecular brightness (pCOMB) approach23, introduced to extend the range of applicability of fluorescence correlation methods in biological systems24, remain single point methods lacking the capability of fast measurements in a large area of a cell, requiring many consecutive observations at each pixel and data acquisition in the order of seconds.
N&B is a simplified version of PCH that considers only the first and second moments of the amplitude of the fluorescence distribution, namely the mean intensity, <I>, and the variance, σ2 (Figure 2)18,19 and, because of that, it cannot determine the molar fraction of unknown oligomers in a mixture, but only estimates the average oligomerization state of the mixture. Nevertheless, N&B has the advantage of working with relatively smaller time series of images of live cells than PCH on a pixel-by-pixel basis, simply by monitoring the fluctuations on time of the fluorescence intensity. Because N&B reduces the time-per-pixel to a few microseconds, it can follow fast oligomerization kinetics over large cell areas, allowing image acquisition on a time scale of seconds in raster scanning microscopy (e.g., confocal, 2-photon) and milliseconds in camera-based microscopy (e.g., TIRFM).
Several reports have demonstrated the capability of N&B to quantify the number of subunits in protein clusters by imaging extended cell regions. Paxillin-EGFP clusters were detected at the adhesion sites in CHO-K1 cells25, and the intracellular aggregation of the pathogenic Httex1p peptide was described in COS-7 cells26. N&B was applied for following the ligand-driven oligomerization of the ErbB receptor27, and the effect of the ligand FGF21 on Klothob (KLB) and FGFR1c in HeLa cells28. The combination of TIRF imaging and N&B analysis was used to show that dynamin-2 is primarily tetrameric throughout the entire cell membrane29. We applied N&B to both raster scanning and TIRF images to prove ligand-driven dimerization of uPAR and FGFR1 cell membrane receptors30,31.
Fluorescence correlation methods, such as N&B, FCS and PCH, are based on the notion that in an open volume the occupation number of particles follows a Poisson distribution. Because only the photons that the fluorophores emit can be detected, the mean value for a measured fluorescence intensity versus time in a pixel of the image, 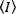 , is the product of the average number of fluorophores in the illumination volume, n, and their molecular brightness, ε17:
, is the product of the average number of fluorophores in the illumination volume, n, and their molecular brightness, ε17:
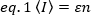
where ε is expressed as the number of photons emitted per unit of time (conventionally per second) per molecule when the molecule is at the center of the illumination volume.
Brightness is a property of each fluorophore in a given acquisition set up, while intensity is the sum of all contributions from all fluorophores. In biological contests, brightness will increase with the increase of the number of fluorophores that fluctuate together, giving information on the oligomerization state of the fluorescently-tagged protein. The fluctuation amplitudes at a given pixel is measured from the variance of the fluorescence signal, σ2:
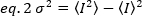
Where the mean of the square of intensity, 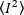 , and the square of the mean of intensity,
, and the square of the mean of intensity, 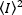 , are computed from the individual intensity values in each pixel of each frame:
, are computed from the individual intensity values in each pixel of each frame:
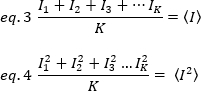
where K is the number of total frames in the time series. Experimentally, it is necessary to compute for the entire image series the variance that describes the scatter of the individual intensity values at each pixel of a single image around the mean intensity value. The variance includes all fluctuations of different origins. In a first approximation, the variance due the diffusing particles in the illumination volume, σ20, can be separated from the variance due to the detector shot noise, σ2d. The two variances are independent; thus, the total variance is given by their sum:
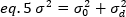
The variance, due to molecular fluctuations in and out of the detection volume, is linearly dependent on the molecular brightness and intensity:
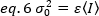
Rearranging eq. 6 according to eq. 1:
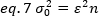
According to the typical concept in fluorescence correlation spectroscopy, equation 7 states that the variance due to the number of fluctuations depends on the square of the particle brightness.
Then, the variance due to detector fluctuations is a linear function of the detected intensity, under the assumption that the detector is operated below its saturation limit19:
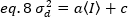
In the case of photon counting detectors a=1 and c=0, thus the detector variance is equal to the average intensity:
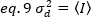
To apply these concepts to real measurements in live cells, Gratton and colleagues18 define the apparent brightness, B, for each pixel as the ratio of the variance over the average intensity:
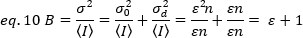
B is the parameter that is measured experimentally. In this work, time series images of FGFR1 receptors at the plasma membrane of HeLa cells are captured by TIRF microscopy and the average apparent brightness, B, is determined by the N&B analysis. Then, after addition of FGF2, consecutive time series are captured to follow the changes in the self-assembly of the receptor molecules in the membrane surface after stimulation of the receptor with the canonical ligand.
However, since the detector of the TIRF microscope is a EMCCD camera, the expression for the apparent brightness needs to be modified as19:
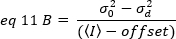
where offset is the intensity offset of the detection electronics that is a characteristic of the detector settings. The variance and average intensity for an analog detector are respectively given by:
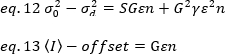
where G is the analog gain in digital levels (DL/photons), S, the digital levels per photon19, is given by the slope of an intensity versus variance plot for a light source with constant intensity (no temporal fluctuations). The γ factor is related to the shape of the pixel detection volume. According to Hassler et al.32, the γ factor is equal to 0.3 for TIRF imaging working at the maximum gain of the detection camera19. The offset, S and G parameters are characteristics of the camera and the microscope. The apparent brightness, B, is obtained by rearranging eq. 11 according to eq. 12 and 13:
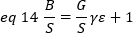
Experimentally, ε is a complex function of laser intensity and the detection efficiency of the system. Nevertheless, since B/S is linearly dependent on ε, it is only important to determine the relative value of ε for a given detection mode:
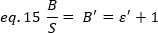
where ε' is proportional to ε. Still, a calibration is performed using an internal reference.
Protocol
1. Sample Preparation
- Day 1. Seed HeLa cells in complete medium at a concentration of 100,000-200,000 cell/mL in glass-bottom dishes. Seed 6-8 replicate dishes.
NOTE: In this example, the medium is supplemented with 10% heat inactivated Fetal Bovine Serum (FBS), 1 mM sodium pyruvate, 100 U/100 µg penicillin/streptomycin. Several replicate dishes are prepared. - Day 2-3. When cells are at sub-confluency, transfect half of the dishes with the protein plasmid and the second half with reference plasmids (monomer and dimer), in serum-free medium.
NOTE: Transfection is made in serum free medium supplemented with antibiotics, 0.1% Bovine Serum Albumin and 25 mM HEPES buffer, without Phenol Red. - Day 3-4. Check that the transfected cells are adherent to the bottom of the dishes and the cell membrane is fluorescent. Discard dishes with overgrown cells or with very low fluorescence.
NOTE: Do not let cells overgrow. Cells must be well distributed and be adhered to the glass area of the dish (Figure 1A). Precoated glass bottom dishes can be used for favoring cell adhesion. The cell culture is tested for mycoplasma contamination before any experiment. In this example, cells are transfected with a (A207K)mEGFP-FGFR1 plasmid and the reference cells are transfected with a GPI-(A207K)mEGFP and a GPI-(A207K)mEGFP-(A207K)mEGFP plasmids using standard protocols. For live cell microscopy, an indicator-free medium is recommended; 25 mM HEPES buffer is added to prevent pH changes during imaging.
2. TIRF Imaging — Alignment of the Laser Line and Optimization of TIRF Illumination
- Four hours before experiment, activate the temperature incubator of the microscope at 37 °C.
- Turn on the microscope, computers and cameras and wait for the cameras to reach the proper working temperature.
NOTE: The working temperature of the camera used in this study is -75 °C. - Place a little drop of oil on the objective. Put a sample dish in place. Close the doors of the incubator and let the temperature of the dish equilibrate (~10 min).
- Turn on the epifluorescence lamp and the 488 nm laser.
- Select the epifluorescence contrast mode to explore the sample, searching a cell to focus from the ocular.
NOTE: The use of a fluorescent lamp for searching cells trough the ocular is not mandatory. A suitable laser line can be used instead. - Select the proper filter for collecting the green emission through the microscope camera (Band Pass Ex 490/20 (500) Band Pass Em 525/50, or similar.
- Switch from the ocular to the camera port (camera #1 in Figure 1) in epifluorescence mode, refine the focus and change to TIRF mode. Epifluorescence and TIRF modes might be named with a different nomenclature depending on the brand of the microscope.
NOTE: There may be issues focusing or aligning the laser if there are no fluorescent markers at the coverslip interface. To align the laser properly (essential for good TIRF), focus on the coverslip. It is often very difficult to determine whether the coverslip is in focus. As a suggestion, focus on the edges of the cells. - Activate the auto-alignment following the instructions of the TIRF microscope.
NOTE: Briefly, for steps from 2.4 to 2.8, first find the cells through the ocular and focus on them, then send the emission to the camera port of the TIRF microscope, re-focus the cells on the microscope computer screen and activate the procedure for laser alignment. The alignment consists in finding the critical angle at which illumination becomes evanescent (Figure 3). Commercial microscopes might have slightly different alignment protocols and also be fully automated; others might have a small camera for facilitating the visualization of the critical angle conditions. - Choose a suitable illumination depth and optimize the direction of the evanescent field (Figure 3).
NOTE: The penetration depth is kept constant for all controls and samples.
3. TIRF Imaging: Capture of the Time Series
- Define a region of interest (ROI) of at least 256 x 256 pixels.
NOTE: In this set up, the capture is done with camera #2 under software that directly controls only the camera (See Figure 1 legend). - Set the exposure to 1 ms and the EM gain to 1,000 (this is the G factor in eq. 12 and 13). At such a speed, it might be necessary to adjust or increase the laser power. Here laser power is 0.5 mW.
NOTE: Depending on the type of the camera and the limits imposed by the diffusion coefficient of the protein, fluorescence intensity and background, the general criteria for setting the laser power are not to saturate the detector, minimize photobleaching, and capture as fast as possible at a reasonable S/N. The EM gain is always set at the maximum of the camera (see Introduction). - Run a first trial sequence under initial conditions and roughly estimate the S/N value. The conditions are acceptable at S/N = 2-3 or higher, as measured in the first frame of the first time series.
- Use the slider of the emission splitting system connecting camera #2 to the microscope for masking a side of the image (Figure 1B, Figure 4A-B)
NOTE: In this set up a multichannel imaging connector is installed on camera #2 to enable the acquisition of two spatially identical images simultaneously. The system is equipped with slides for mounting different emission filters. One of the sliders mounts a black mask to cover a side of the image. The masked area is used for the internal calibration of each time series, to determine the camera parameters (eq. 12 and eq. 13). In this way there is no need for an independent calibration step and, importantly, calibration is performed in parallel to the capture of each time series. In the absence of this system, the camera can be calibrated applying published protocols33. - Select the camera file autosave option.
- Start the acquisition of the image series. Acquire a minimum of 700 frames at a minimum S/N ratio of 2.
NOTE: The number of frames that are necessary for analysis depends on the sample stability to photobleaching and on the dispersion of the data. Therefore, the quality of each time series is assessed during N&B analysis. - Without taking the dish out of the microscope, add the ligand.
- Select a cell with a bright fluorescence membrane and quickly start the first time series of the kinetic run.
NOTE: If the addition of the ligand is done quickly, this first capture sets the point = 0 time of the ligand kinetics. The software registers the exact time of the capturing. - Search a second cell and acquire the second time point of the kinetics.
NOTE: Point-visiting routines are available in some microscopes equipped with x,y,z motorized stages. These allow the memorization of multiple positions on the cell dish, and can help in keeping a more constant interval of time between image-series on different cells. - Capture a new cell for each time point of the kinetic run.
NOTE: After capturing, a cell is partially photobleached and it cannot be re-imaged. Because of that, the kinetics is obtained by combining time series of many cells, each captured at a different time point. - For each new dish, repeat the protocol from step 2.3 to 3.9.
NOTE: For reference dishes, add a volume of the vehicle (PBS supplemented with 0.01% bovine serum albumin) equivalent to that used for the ligand.
4. Number & Brightness (N&B): Quality Check of the Time Series
- Convert and save as .TIFF the files acquired with the camera software (.sif files in this example).
- Import .TIFF files in the analysis software routine by activating the N&B graphical user interface (GUI) MATLAB.
NOTE: A customized Matlab executable N&B routine is used here (N&B analysis at https://www.cnic.es/en/investigacion/2/1187/tecnologia). By opening an imported .TIFF file, the routine generates the average intensity image, the average intensity profile and it allows inspecting the series frame-by-frame (Supplemental Figure 1). Other software are available for N&B analysis (e.g., SimFCS software). - Discard series for which the average intensity profile shows more than 10% photobleaching, and series in which there has been an evident cell membrane distortion or translation during acquisition.
- Crop frames that are evidently out-of-focus.
NOTE: A cropping tool is implemented in the routine to discards single or multiple frames within the image series. This operation is allowed because frame-to-frame time is not critical whereas the pixel dwell time (exposure time) is (see Discussion). - Keep for the analysis only series with at least 500 time frames.
5. Number & Brightness (N&B): Determination of the Camera Parameters (Offset, σ and S)
- Activate the routine Calibrate Camera.
- Select an area of at least 20 x 50 pixels in the detector noise region (Figure 4).
NOTE: The routine originates a histogram of the values (also defined Digital Level, DL) and it returns a logarithm plot of the Frequency versus Digital Levels. - In the log Frequency versus Digital level plot, move the linear red cursor to delimit the Gaussian and the linear part of the curve.
NOTE: The red cursor divides the two sections of the curve, and activates the routine returning the offset, which is the center of the Gaussian function of the camera response, the σ of the Gaussian fit, and the S factor, which is the slope of the linear part of the camera response (Figure 4C-D).
6. Number & Brightness (N&B): Computation of the B-values in Selected Region-of-interest (ROI)
- Activate the B key.
NOTE: This action generates the average intensity image (Figure 5, first column) and the B-image in which each individual B-value is associated to the related pixel in the image (Supplemental Figure 1). - Apply a minimum binning (2 2) to reduce the dispersion of the data and to generate the B-I histogram (Figure 5, second column).
NOTE: The B-I histogram represents the distribution of the B-values of all pixels of the image versus the pixel intensity. Y = B/S; X = (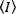 - offset)/S (Supplemental Figure 1 and eq. 11 and 15).
- offset)/S (Supplemental Figure 1 and eq. 11 and 15). - Inspect the B-I histogram using the interactive square cursor.
- Select a square ROI for the analysis (Figure 5, third column).
NOTE: The cursor synchronizes a mobile mask on the average intensity image, highlighting the pixels that are selected inside the square cursor area (Supplemental Figure 1). By this inspection, it is possible to exclude from the analysis the background and areas with very low intensity. - Generate the B-map of the selected ROI (Figure 5, fourth column).
- Save the ASCII file of the B-values associated to the selection.
- Import the ASCII file in a graphic software to compute the frequency distribution of the data and obtain the average B-value ± S.E (Figure 5, fifth column).
NOTE: If data are homogeneous, the frequency distribution of the B-values approximates a Gaussian distribution. - Apply eq. 15 to derive the average brightness
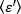 =
= 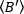 - 1 [(counts/molecule) per dwell time] for each cell at each time point of the kinetic run. Normalize the data according to:
- 1 [(counts/molecule) per dwell time] for each cell at each time point of the kinetic run. Normalize the data according to:
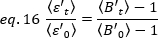
where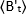 is the average B-value measured at time "t" after ligand addition, and
is the average B-value measured at time "t" after ligand addition, and 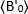 is the average B-value measured at the time t=0 (10-20 s after ligand addition).
is the average B-value measured at the time t=0 (10-20 s after ligand addition).
NOTE: The normalization of the results allows the direct comparison of experiments that are carried out in different days. It compensates for differences in the measured brightness due to laser power and technical fluctuations. - Plot the Normalized Average Brightness versus acquisition time to build the kinetic run (Figure 6).
Results
The results for two representative HeLa-mEGFP-FGFR1 cells seeded in the same culture dish are shown in Figure 5 and Supplemental Table 1. The two cells were captured at time 0 min (Figure 5A, top) and 7 min (Figure 5A, bottom) after addition of the FGF2 ligand.
Figure 5...
Discussion
N&B requires several precautions in the choice of the cell model and labelling strategy. It can be applied only to live cells that remain stably adhered during the image capture time. Extra fluctuations due to the whole cell rigid displacement might be handled with appropriate image restoration approaches38. However, generally when a cell moves, the cell membrane also deforms, and structure deformation, producing large extra variance, introduces serious limitation to the analysis of membrane p...
Disclosures
The authors have nothing to disclose.
Acknowledgements
The CNIC is supported by the Ministry of Ciencia, Innovacion y Universidades and the Pro CNIC Foundation, and is a Severo Ochoa Center of Excellence (SEV-2015-0505). We are also supported by European Regional Development Fund (FEDER) "Una manera de hacer Europa". UC acknowledges the support from the Associazione Italiana Ricerca sul Cancro, the Association for International Cancer Research (now known as Worldwide Cancer Research), and the Italian Ministry of Health. A.T. acknowledge the "Fondazione Banca del Monte di Lombardia" for partly supporting his work with the PV Fellowship "Progetto Professionalità Ivano Becchi" 2011-2012.
Materials
| Name | Company | Catalog Number | Comments |
| 3-Colour Fast TIRF Leica AM TIRF MC inverted microscope, with smi-automatic TIRF alignment. The microscope is equipped with a diode 488 nm laser, a 100x1.46 oil TIRF objective, Ex/Em Bandpass filters at 490/20 and 525/50, temperature/CO2 incubator and Andor DU 8285 VP EMCCD camera. The microscope is operated by Leica LIF software. | Leica Microsystems, Wetzlar, Germany | ||
| Albumin from Bovine Serum 98% minimun | Sigma-Aldrich, St. Louis, MI, USA | A7906-100G | |
| DMEM without Phenol Red with 25 mM HEPES | GIBCO Thermo Fisher Scientific,Waltham, MA, USA | 21063029 | Used serum free for microscopy |
| DMEM high-glucose GlutaMAX I | GIBCO Thermo Fisher Scientific,Waltham, MA, USA | 10566-016 | Used for complete medium |
| Dulbecco's Phosphate Buffered Saline 10x (PBS) | Biowest, Nuaillé, France | X0515-500 | |
| Emission splitting system Photometrics DV2 | TeledynePhotometrics, Tucson, AZ, USA | ||
| Fetal Bovine Serum, qualified, Brazil | GIBCO Thermo Fisher Scientific,Waltham, MA, USA | 10270106 | 10% inactivated supplement for complete medium |
| Glass bottom 35-mm sterile 1.5 dishes | MatTek, Ashland, MA, USA | P35G-0.170-14-C | uncoated, glass thickness 0.17 microns |
| GraphPad Prism | GraphPad Software Inc., San Diego, CA, USA | ||
| Human cervical carcinoma (HeLa), serum-free animal component (AC) cells | Millipore-Sigma ECACC, Darmstadt, Germany | CB_08011102 | |
| iXonEM+ 897 EMCCD (back-illuminated) ANDOR camera controlled by ANDOR Solis software | Oxford Instruments, Andor TM Technology, Abingdon-on-Thames, UK | This camera, installed in an additional port of the microscope, is used for acquiring the N&B time series | |
| Matlab Executable N&B routine | Unit of Microscopy and Dynamic Imaging, CNIC, Madrid, Spain | download at https://www.cnic.es/en/investigacion/2/1187/tecnologia | |
| MatLab v.2018b | The MathWorks, Inc. Natick, MA, USA | download at https://www.mathworks.com/products/matlab.html | |
| Penicillin:Streptomycin for tissue culture 100x | Biowhittaker Inc. Walkersville, MD, USA | LONZA 17-602E | supplement for medium at Penicillin/Streptomycin 100U/100µg. |
| pN1-mEGFP-FGFR1 expression vector | Unit of Gynecological Oncology Research, European Institute of Oncology IRCCS, Milan, Italy | Zamai et al., 2019 | |
| pN1-N-Gly-mEGFP-GPI expression vector | Unit of Microscopy and Dynamic Imaging, CNIC, Madrid, Spain | Hellriegel et al., 2011 | |
| pN1-N-Gly-mEGFP-mEGFP-GPI expression vector | Unit of Microscopy and Dynamic Imaging, CNIC, Madrid, Spain | Hellriegel et al., 2011 | |
| Recombinant FGF2 | PeproTech EC, Ltd., London, UK | Ligand solution: 20ng/mL of FGF2 in PBS supplemented with 0.01%BSA. | |
| Sodium pyruvate GIBCO | ThermoFisher Scientific | 11360070 | 1mM supplement for medium |
| TransIt-LT1 Transfection Reagent | MirusBio LLC, Madison, WI, USA | MIR 2300 | |
| Trypsin-EDTA (0.25%), phenol red | GIBCO Thermo Fisher Scientific,Waltham, MA, USA | 25200056 | |
| Type F Immersion liquid 10 mL | Leica Microsystems, Wetzlar, Germany | 11513 859 | |
| UltraPure BSA (50 mg/mL) | ThermoFisher Scientific | AM2618 | 0.1% supplement for medium without phenol red used for transfections |
References
- Agwuegbo, U. C., Jonas, K. C. Molecular and functional insights into gonadotropin hormone receptor dimerization and oligomerization. Minerva Ginecologica. 70 (5), 539-548 (2018).
- Ferre, S., et al. G protein-coupled receptor oligomerization revisited: functional and pharmacological perspectives. Pharmacological Reviews. 66 (2), 413-434 (2014).
- Marsango, S., Ward, R. J., Alvarez-Curto, E., Milligan, G. Muscarinic receptor oligomerization. Neuropharmacology. 136 (Pt C), 401-410 (2018).
- Oishi, A., Cecon, E., Jockers, R. Melatonin Receptor Signaling: Impact of Receptor Oligomerization on Receptor Function. International Review of Cell and Molecular Biology. 338, 59-77 (2018).
- Thelen, M., Munoz, L. M., Rodriguez-Frade, J. M., Mellado, M. Chemokine receptor oligomerization: functional considerations. Current Opinion in Pharmacology. 10 (1), 38-43 (2010).
- Van Craenenbroeck, K. GPCR oligomerization: contribution to receptor biogenesis. Subcellular Biochemistry. 63, 43-65 (2012).
- Wnorowski, A., Jozwiak, K. Homo- and hetero-oligomerization of beta2-adrenergic receptor in receptor trafficking, signaling pathways and receptor pharmacology. Cell Signaling Technology. 26 (10), 2259-2265 (2014).
- Fricke, F., Dietz, M. S., Heilemann, M. Single-molecule methods to study membrane receptor oligomerization. Chemphyschem. 16 (4), 713-721 (2015).
- Vidi, P. A., Ejendal, K. F., Przybyla, J. A., Watts, V. J. Fluorescent protein complementation assays: new tools to study G protein-coupled receptor oligomerization and GPCR-mediated signaling. Molecular and Cellular Endocrinology. 331 (2), 185-193 (2011).
- Trussell, H. J., et al., Trussell, J., et al. . Academic Press Library in Signal Processing. 4, 3-9 (2014).
- Betzig, E., et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science. 313 (5793), 1642-1645 (2006).
- Rust, M. J., Bates, M., Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nature Methods. 3 (10), 793-795 (2006).
- Nagerl, U. V., Willig, K. I., Hein, B., Hell, S. W., Bonhoeffer, T. Live-cell imaging of dendritic spines by STED microscopy. Proceedings of the National Academy of Sciences of the United States of America. 105 (48), 18982-18987 (2008).
- Tsekouras, K., Custer, T. C., Jashnsaz, H., Walter, N. G., Presse, S. A novel method to accurately locate and count large numbers of steps by photobleaching. Molecular Biology of the Cell. 27 (22), 3601-3615 (2016).
- Godin, A. G., et al. Revealing protein oligomerization and densities in situ using spatial intensity distribution analysis. Proceedings of the National Academy of Sciences of the United States of America. 108 (17), 7010-7015 (2011).
- Qian, H., Elson, E. L. Distribution of molecular aggregation by analysis of fluctuation moments. Proceedings of the National Academy of Sciences of the United States of America. 87 (14), 5479-5483 (1990).
- Chen, Y., Muller, J. D., So, P. T., Gratton, E. The photon counting histogram in fluorescence fluctuation spectroscopy. Biophysical Journal. 77 (1), 553-567 (1999).
- Dalal, R. B., Digman, M. A., Horwitz, A. F., Vetri, V., Gratton, E. Determination of particle number and brightness using a laser scanning confocal microscope operating in the analog mode. Microscopy Research and Technique. 71 (1), 69-81 (2008).
- Unruh, J. R., Gratton, E. Analysis of molecular concentration and brightness from fluorescence fluctuation data with an electron multiplied CCD camera. Biophysical Journal. 95 (11), 5385-5398 (2008).
- Hess, S. T., Huang, S., Heikal, A. A., Webb, W. W. Biological and chemical applications of fluorescence correlation spectroscopy: a review. Biochemistry. 41 (3), 697-705 (2002).
- Muller, J. D., Chen, Y., Gratton, E. Fluorescence correlation spectroscopy. Methods in Enzymology. 361, 69-92 (2003).
- Levi, V., Ruan, Q., Kis-Petikova, K., Gratton, E. Scanning FCS, a novel method for three-dimensional particle tracking. Biochemical Society Transactions. 31 (Pt 5), 997-1000 (2003).
- Hinde, E., et al. Quantifying the dynamics of the oligomeric transcription factor STAT3 by pair correlation of molecular brightness. Nature Communications. 7, 11047 (2016).
- Waithe, D., et al. Optimized processing and analysis of conventional confocal microscopy generated scanning FCS data. Methods. 140-141, 62-73 (2018).
- Digman, M. A., Dalal, R., Horwitz, A. F., Gratton, E. Mapping the number of molecules and brightness in the laser scanning microscope. Biophysical Journal. 94 (6), 2320-2332 (2008).
- Ossato, G., et al. A two-step path to inclusion formation of huntingtin peptides revealed by number and brightness analysis. Biophysical Journal. 98 (12), 3078-3085 (2010).
- Nagy, P., Claus, J., Jovin, T. M., Arndt-Jovin, D. J. Distribution of resting and ligand-bound ErbB1 and ErbB2 receptor tyrosine kinases in living cells using number and brightness analysis. Proceedings of the National Academy of Sciences of the United States of America. 107 (38), 16524-16529 (2010).
- Ming, A. Y., et al. Dynamics and Distribution of Klothobeta (KLB) and fibroblast growth factor receptor-1 (FGFR1) in living cells reveal the fibroblast growth factor-21 (FGF21)-induced receptor complex. Journal of Biological Chemistry. 287 (24), 19997-20006 (2012).
- Ross, J. A., et al. Oligomerization state of dynamin 2 in cell membranes using TIRF and number and brightness analysis. Biophysical Journal. 100 (3), L15-L17 (2011).
- Hellriegel, C., Caiolfa, V. R., Corti, V., Sidenius, N., Zamai, M. Number and brightness image analysis reveals ATF-induced dimerization kinetics of uPAR in the cell membrane. FASEB J. 25 (9), 2883-2897 (2011).
- Zamai, M., et al. Number and brightness analysis reveals that NCAM and FGF2 elicit different assembly and dynamics of FGFR1 in live cells. Journal of Cell Science. 132 (1), (2019).
- Hassler, K., et al. Total internal reflection fluorescence correlation spectroscopy (TIR-FCS) with low background and high count-rate per molecule. Optics Express. 13 (19), 7415-7423 (2005).
- Di Rienzo, C., Gratton, E., Beltram, F., Cardarelli, F. From fast fluorescence imaging to molecular diffusion law on live cell membranes in a commercial microscope. Journal of Visualized Experiments. (92), e51994 (2014).
- Beenken, A., Mohammadi, M. The FGF family: biology, pathophysiology and therapy. Nature Reviews Drug Discovery. 8 (3), 235-253 (2009).
- Joubert, J., Sharma, D. Light microscopy digital imaging. Current Protocols in Cytometry. , (2011).
- Gell, C., Berndt, M., Enderlein, J., Diez, S. TIRF microscopy evanescent field calibration using tilted fluorescent microtubules. Journal of Microscopy. 234 (1), 38-46 (2009).
- Burghardt, T. P. Measuring incidence angle for through-the-objective total internal reflection fluorescence microscopy. Journal of Biomedical Optics. 17 (12), 126007 (2012).
- Trullo, A., Corti, V., Arza, E., Caiolfa, V. R., Zamai, M. Application limits and data correction in number of molecules and brightness analysis. Microscopy Research and Technique. 76 (11), 1135-1146 (2013).
- Caiolfa, V. R., et al. Monomer-dimer dynamics and distribution of GPI-anchored uPAR are determined by cell surface protein assemblies. Journal of Cell Biology. 179 (5), 1067-1082 (2007).
- Campbell, R. E., et al. A monomeric red fluorescent protein. Proceedings of the National Academy of Sciences of the United States of America. 99 (12), 7877-7882 (2002).
- Cutrale, F., et al. Using enhanced number and brightness to measure protein oligomerization dynamics in live cells. Nature Protocols. 14 (2), 616-638 (2019).
- Dunsing, V., Chiantia, S. A Fluorescence Fluctuation Spectroscopy Assay of Protein-Protein Interactions at Cell-Cell Contacts. Journal of Visualized Experiments. (142), (2018).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved