A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
3D Cell-Printed Hypoxic Cancer-on-a-Chip for Recapitulating Pathologic Progression of Solid Cancer
* These authors contributed equally
In This Article
Summary
Hypoxia is a hallmark of tumor microenvironment and plays a crucial role in cancer progression. This article describes the fabrication process of a hypoxic cancer-on-a-chip based on 3D cell-printing technology to recapitulate a hypoxia-related pathology of cancer.
Abstract
Cancer microenvironment has a significant impact on the progression of the disease. In particular, hypoxia is the key driver of cancer survival, invasion, and chemoresistance. Although several in vitro models have been developed to study hypoxia-related cancer pathology, the complex interplay of the cancer microenvironment observed in vivo has not been reproduced yet owing to the lack of precise spatial control. Instead, 3D biofabrication approaches have been proposed to create microphysiological systems for better emulation of cancer ecology and accurate anticancer treatment evaluation. Herein, we propose a 3D cell-printing approach to fabricate a hypoxic cancer-on-a-chip. The hypoxia-inducing components in the chip were determined based on a computer simulation of the oxygen distribution. Cancer-stroma concentric rings were printed using bioinks containing glioblastoma cells and endothelial cells to recapitulate a type of solid cancer. The resulting chip realized central hypoxia and aggravated malignancy in cancer with the formation of representative pathophysiological markers. Overall, the proposed approach for creating a solid-cancer-mimetic microphysiological system is expected to bridge the gap between in vivo and in vitro models for cancer research.
Introduction
The cancer microenvironment is a critical factor driving cancer progression. Multiple components, including biochemical, biophysical, and cellular cues, determine the pathological features of cancer. Among these, hypoxia is strongly associated with cancer survival, proliferation, and invasion1. Due to the unlimited growth and division of cancer cells, nutrients and oxygen are continuously depleted, and a hypoxic gradient is generated. Under low-oxygen conditions, cells activate hypoxia-inducible transcription factor (HIF)-associated molecular cascade. This process induces a necrotic core, triggers metabolic changes, and initiates blood vessel hyperplasia and metastasis2,3. Subsequently, hypoxia in cancer cells causes the destruction of neighboring normal tissues. Furthermore, hypoxia is strongly associated with the therapeutic resistance of solid tumors in multifactorial manners. Hypoxia may severely impede radiotherapy, as radiosensitivity is limited owing to reactive oxygen species1,4. In addition, it decreases pH levels of cancer microenvironments, which decreases drug accumulation1. Therefore, reproducing pathological features related to hypoxia in vitro is a promising strategy for scientific and pre-clinical findings.
Modeling a specific microenvironment of cancer is essential for understanding cancer development and exploring appropriate treatments. Although animal models have been widely used because of their strong physiological relevance, issues related to species differences and ethical problems exist5. Furthermore, although conventional 2D and 3D models allow for the manipulation and real-time imaging of cancer cells for an in-depth analysis, their architectural and cellular complexity cannot be fully recapitulated. For example, cancer spheroid models have been widely used, as cancer cell aggregation in a spheroid can naturally generate hypoxia in the core. Moreover, large numbers of cellular spheroids of uniform size have been produced using plastic- or silicone-based multi-well systems6,7. However, the lower flexibility with regard to capturing the exact heterogeneous structure of cancerous tissues with conventional platforms has required the establishment of an advanced biofabrication technology to build a highly biomimetic platform to improve cancer research8.
3D microphysiological systems (MPSs) are useful tools to recapitulate the complex geometry and pathological progression of cancer cells9. As cancer cells sense the biochemical gradient of growth factors and chemokines and the mechanical heterogeneity reproduced on the system, important features of cancer development can be investigated in vitro. For instance, cancer viability, metastatic malignancy, and drug resistance depending on the varying oxygen concentrations has been studied using MPSs10,11. Despite recent advancements, generating hypoxic conditions of in vitro models relies on complex fabrication procedures, including connection with physical gas pumps. Therefore, simple, and flexible methods to build cancer-specific microenvironments are needed.
3D cell printing technology has gained considerable attention because of its precise control of the spatial arrangement of biomaterials to recapitulate native biological architectures12. In particular, this technology overcomes the existing limitations of 3D hypoxia models owing to its high controllability and feasibility for building the spatial features of the cancer microenvironment. 3D printing also facilitates computer-aided manufacturing through a layer-by-layer process, thereby providing a rapid, accurate, and reproducible construction of complex geometries to mimic actual tissue architectures. In addition to the advantages of existing manufacturing strategies for 3D MPSs, the pathophysiological features of cancer progression can be reproduced by patterning the biochemical, cellular, and biophysical components13,14.
Herein, we present a 3D cell-printing strategy for a hypoxic cancer-on-a-chip for recapitulating the heterogeneity of a solid cancer (Figure 1)15. The fabrication parameters were determined via a computational simulation of central hypoxia formation in the system. Cancer-stroma concentric rings were printed using collagen bioinks containing glioblastoma cells and endothelial cells to emulate the pathophysiology of glioblastoma, a type of solid cancer. The formation of a radial oxygen gradient aggravated cancer malignancy, indicating strengthened aggressiveness. Furthermore, we indicate future perspectives for the applications of the chip to patient-specific preclinical models. The proposed approach for creating a solid-cancer-mimetic microphysiological system is expected to bridge the gap between in vivo and in vitro models of cancer.
Access restricted. Please log in or start a trial to view this content.
Protocol
1. Computer simulation of oxygen gradient formation
- Generation of a 3D geometry model for hypoxic cancer-on-a-chip printing
- Run a 3D CAD software.
- Sketch the geometry model of hypoxic cancer-on-a-chip. Click on Sketch and select the desired plane to draw the geometry. Refer to the drawing (Figure 2A) for the detail scale of each part.
- Set the thickness of the geometry by clicking on Feature-Protrusion Boss/Base. Enter the desired thickness (refer to Figure 2A) in the empty box and select the green check icon to form the 3D geometry.
NOTE: The dimension of the cancer-on-a-chip is defined based on the desired volumes of media and hydrogel. In the present experiment, the desired volumes of media and hydrogel were approximately 1,500 µL and 500 µL, respectively, based on the previous practical experiences for resolution of extrusion-based bioprinter. - Save the geometry file as a 3D CAD file format (.prt or .stl).
- Determination of cellular density for induction of hypoxic core
- Run a physical diffusion simulation program.
- Click on LiveLink and select the CAD program used. Click on Synchronize to import the geometry of the hypoxic cancer-on-a-chip on the simulation program. As the inner space of the chamber will be filled with a culture medium in an actual experimental setting, oxygen will diffuse across the inner space of the chamber and the cellular construct, which will be composed of cell-laden hydrogels.
NOTE: Refer to previous study for details on the physical parameters15. - Define the imported 3D geometry as a control volume of the space wherein oxygen diffuses, and the cells consume oxygen (Figure 2B).
- Run a computer analysis for gas diffusion analysis following a user guide and previously established methods16,17.
- From the computer analysis results, export the estimated oxygen concentration data over cross-section A-A' at each time point following the user guide. The governing equation is based on Fick's first law, as expressed in Eq. (1) (Figure 2C).
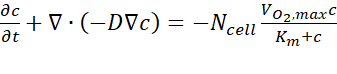
where c is the concentration, D is the oxygen diffusion coefficient, Ncell is the density of the cells,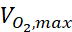 is the maximum up-take rate of oxygen, and Km is the Michaelis-Menten constant. The constants were applied as described in a previous publication15.
is the maximum up-take rate of oxygen, and Km is the Michaelis-Menten constant. The constants were applied as described in a previous publication15.
NOTE: Each time point means a step point to observe oxygen diffusion change over time. - Evaluate whether the minimal oxygen level reaches a threshold of hypoxia and repeat the computer analysis process with an increment or decrement of cellular density.
NOTE: Define that hypoxia gradient is formed in the construct if the oxygen level of 80% in the hydrogel area is less than 0.02 mM after 24 h. - Confirm the number of cells required to generate the oxygen gradient inducing hypoxia in the central region from Fick's first law in step 1.2.5 and the simulation results from step 1.2.6.
NOTE: In this protocol, cell number was 2 × 106 cells/each construct.
2. Cell culture of cancer cells and stromal cells
- Preparation of cell culture media to avoid physiological stress
- For U-87 MG cells (immortalized human glioblastoma cell line), place 12 mL of high-glucose Dulbecco's modified Eagle medium containing 10% fetal bovine serum, 100 U/mL penicillin, and 100 µg/mL streptomycin in a T-75 cell culture flask in a 37 °C, 5% CO2 humidified incubator for 30 min to minimize the thermal and alkaline effects of the medium on the cells.
NOTE: Glioblastoma was chosen as a type of solid cancer because it has aggressive characteristics in a hypoxic environment. Other various types of cancers can be applied to this model. - For human umbilical vein endothelial cells (HUVECs), place 12 mL of endothelial cell growth medium in a T-75 cell culture flask in a 5% CO2 humidified incubator at 37 °C for 30 min.
NOTE: HUVECs were chosen because it is one of the most representative endothelial cell lines. Various types of stromal cells can also be applied to this model.
- For U-87 MG cells (immortalized human glioblastoma cell line), place 12 mL of high-glucose Dulbecco's modified Eagle medium containing 10% fetal bovine serum, 100 U/mL penicillin, and 100 µg/mL streptomycin in a T-75 cell culture flask in a 37 °C, 5% CO2 humidified incubator for 30 min to minimize the thermal and alkaline effects of the medium on the cells.
- Rapid thawing of cryopreserved cancer cells and stromal cells and their maintenance
- Move cryovials containing 5 x 105 U-87 MG cells and HUVECs from the liquid nitrogen container to a laminar flow cabinet. Immediately loosen and retighten the cap to release the internal pressure.
- Gently place the cryopreserved cells in a water bath at 37 °C for 2 min, keeping the cap out of the water. Rinse the vials with 70% ethanol under laminar flow to prevent contamination.
- Transfer the thawed cells to the flasks containing the prepared cell culture media described in step 2.1 and place the cell-containing flasks in a 5% CO2 humidified incubator at 37 °C for cell recovery.
- Refresh the cell culture media every 2 days and maintain the cell growth.
- After 24 h of thawing, replace the cell culture media to avoid cytotoxicity of dimethyl sulfoxide (DMSO), which was used for cell freezing. Use HUVECs, which has undergone less than 6 passages.
3. Preparation of collagen pre-gel solution
- Solubilization of collagen sponge with 0.1 N hydrochloric acid (HCl)
- Prepare a solution of 0.1 N HCl and filter it with a 0.2 µm syringe filter.
- For 3 mL of a 1% (w/v) neutralized collagen pre-gel solution, prepare collagen sponges cut into 5 x 5 mm2 pieces and weighing 30 mg.
- Transfer the cut collagen pieces to a sterile 10 mL glass vial.
NOTE: Prepare 1.5 times volume of the required collagen hydrogel, considering the loss of the hydrogel due to the sticky characteristic of the collagen solution. - Add 2.4 mL of 0.1 N HCl into the collagen-containing glass vial and incubate it on the rocker at 15 rpm and 4 °C for 3 days.
NOTE: The volume of the 0.1 N HCl solution was four-fifths of the final volume of required collagen hydrogel. In this case, 3 mL of collagen was prepared. - After digestion, sieve the undigested collagen particles using a 40 µm cell strainer. Store the acidic collagen solution at 4 °C and use within 7 days.
- pH adjustment for 1% neutralized collagen pre-gel solution
- Centrifuge the acidic collagen solution at 1224 x g for 5 min at 4 °C.
- Add 30 µL of phenol red solution as a pH indicator to a final concentration of 1% (v/v) and 300 µL of 10x phosphate-buffered saline (PBS) buffer to a final concentration of 10% (v/v) in the collagen pre-gel solution.
- Neutralize the pH to 7 with 1 N sodium hydroxide (NaOH), verifying the color change.
NOTE: Based on the formula, moles H+ = molarity H+ x volume H+ = moles OH-= molarity OH- x volume OH-, add 240 µL of NaOH. - Add distilled water to obtain a total volume of 3 mL.
- After pH adjustment, store the 1% (w/v) neutralized collagen pre-gel solution at 4 °C and use within 3 days.
NOTE: To precheck the gelation of the neutralized collagen pre-gel solution, make 50 µL collagen droplets on a small dish using a positive displacement pipette and incubate them in a 37 °C incubator for 1 h. Refer to the following three methods to verify the cross-linking of collagen droplets. - Check whether the color of collagen has changed into opaque white from transparent color.
- Tilt the container and check whether the collagen is adhered to the bottom of the container.
- Pour 1x PBS on the droplets and check whether the collagen construct is not broken in the solution.
4. 3D printing of gas-permeable barrier
- 3D printing of a sacrificial poly (ethylene-vinyl acetate) (PEVA) mold
- Generate the 3D geometry of the sacrificial PEVA mold defined in step 1 using a 3D CAD software (Figure 3A).
NOTE: The 3D geometry and detailed model scale including dimension, units, and line types were shown in Figure 2A. - Convert the 3D CAD file into an STL file format by clicking on File | Save-File type as STL. Also, click on Option | Output form as ASCII for G-code generation.
- Click on File | Open STL file and select the saved STL file to import the generated STL file. Click on Slice model of STL-CAD exchanger to automatically generate the G-code of the sacrificial PEVA mold (Figure 3B, C).
NOTE: The printing path is generated with the connection of intersected points between the fundamental figure of the STL file and the slicing plane (i.e., layer). Basically, the fundamental figure of a fragment in an STL file is a triangle that contains the 3D coordinates. After the intersected points between the triangle and the layer are obtained, a G-code for printing is generated by connecting each point without an overlapped path on a layer18. Any G-code generation algorithm on board software can be used to generate printing paths for the chip fabrication. - Prepare a sterile adhesive and hydrophilic histology slide.
NOTE: The hydrophilic slide glass is critical for the permanent bonding of polydimethylsiloxane (PDMS) on the glass and the adhesion of the collagen constructs encapsulating cancer cells and stromal cells. - Print the sacrificial PEVA mold on the slide with a 50 G precision nozzle at a pneumatic pressure of 500 kPa at 110 °C.
NOTE: The line width is affected by the feed rate, nozzle gauge, and temperature of the material. The 50 G nozzle was used and a feed rate of 400 was applied to generate 500 µm line width for the sacrificial wall. The nozzle gauge, pneumatic pressure, and feed rate are defined with practical results19. The sacrificial wall needs to be sufficiently thick to hold the PDMS solution, which is the next fabrication step.
- Generate the 3D geometry of the sacrificial PEVA mold defined in step 1 using a 3D CAD software (Figure 3A).
- Casting of polydimethylsiloxane (PDMS) barrier
- Mix 6 mL PDMS base elastomer and 0.6 mL curing agent homogenously over 5 min in a plastic reservoir. This can fabricate 6 hypoxic cancer-on-chips, considering the loss due to the sticky characteristic of PDMS.
- Load the blended PDMS solution into a 10 mL disposable syringe and fit the syringe head with a 20 G plastic tapered dispense tip.
- Fill the sacrificial PEVA mold with the blended PDMS solution in the syringe. The blended PDMS will fill the sacrificial PEVA mold with a convex surface. The height of the PDMS barrier will be higher than that of the PEVA mold.
- Cure the PDMS barrier in an oven at 40 °C for over 36 h to avoid the melting of PEVA. Do not increase the temperature to over 88 °C, which is the melting temperature of PEVA.
- Detach the sacrificial PEVA mold with a pair of precision tweezers and sterilize the gas-permeable barrier at 120 °C in an autoclave.
5. Preparation of cell-encapsulated collagen bio-inks
- Detachment of the prepared cancer cells and stromal cells
NOTE: Considering cell viability, the entire printing process should be completed as soon as possible after detaching the cells.- Wash cancer and stromal cells with 10 mL of 1x PBS using a serological pipette; treat with 2 mL of 0.25% trypsin-ethylenediaminetetraacetic acid (EDTA) using a pipette and incubate them for 3 min at 37 °C.
- Neutralize the trypsinized cells with 3 mL of cell culture media; collect the suspensions of cells into 15 mL conical tubes and centrifuge at 516 x g for 5 min at 20 °C.
- Aspirate the supernatant slowly; resuspend the cell pellets in 5 mL cell culture media and count the number of cells using a hemocytometer.
- Transfer 5 x 106 cells of each cell type into new 15 mL conical tubes and centrifuge them at 516 x g for 5 min at 20 °C.
- Aspirate the supernatant off and place it on wet ice.
- Mixing of each cell type with the 1% neutralized collagen pre-gel solution
NOTE: To avoid thermal solidification of the 1% neutralized collagen pre-gel solution, this process should be performed on wet ice.- Resuspend each type of cell pellet collected in step 5.1.4 with 20 µL of cell culture media each.
- Add 1 mL of the 1% neutralized collagen pre-gel solution into each of the resuspended cell suspensions and mix them homogenously using a positive displacement pipette. The final concentration of each cell type will be 5 x 106 cells/mL.
- Transfer the cell-encapsulated collagen bioinks into 3 mL disposable syringes using a positive disposable pipette and store the syringes at 4 °C until 3D cell-printing.
6. 3D cell-printing of cancer-stroma concentric rings
- 3D cell-printing of collagen bioinks encapsulating cancer cells and stromal cells
- Generate the 3D geometry of the cancer-stroma concentric rings defined in step 1.2 using a 3D CAD software.
NOTE: The dimensions of the cancer stroma concentric rings are defined via simulated parameters. The final dimension parameter dimensions are shown in Figure 3A. - Convert the 3D CAD file into an STL file format and generate a G-code of the cancer-stroma concentric rings using a STL-CAD exchanger.
NOTE: Refer to the note in step 4.1.2 for the G-code generation algorithm. - Load the cell-encapsulated collagen bioinks contained in 3 mL disposable syringes to the head of the 3D printer and set the temperature of the head and plate to 15 °C.
NOTE: If the temperature of the head and plate of the printer reaches over 37 °C, the bioink gets cross-linked and no longer prints. - Load the generated printing path on the control software of the 3D printer.
- By clicking on the Start button, print the collagen bioinks encapsulating cancer cells and stromal cells on the gas-permeable barrier following the loaded G-code with an 18 G plastic needle at pneumatic pressure of approximately 20 kPa at 15 °C.
- At the end of every printing operation, manually place a sterilized 22 mm x 50 mm glass cover on top of the gas-permeable barrier to generate the hypoxic gradient.
NOTE: Compare two groups depending on the presence of glass cover (GR+) and absence (GR-) of that to verify the generation of the hypoxic gradient. - After generating three hypoxic cancer-on-chips, transfer the chips to an incubator at 37 °C for 1 h to cross-link the collagen bioinks.
- Generate the 3D geometry of the cancer-stroma concentric rings defined in step 1.2 using a 3D CAD software.
- Completion of the fabrication process and maintenance of the hypoxic cancer-on-a-chip
- After completion of all 3D cell-printing processes of the hypoxic cancer-on-a-chip, gently rub the cover glasses on top of the gas-permeable barriers with the cell-scrapper for tight bonding (Figure 4A,B).
NOTE: The cover glass and the gas-permeable barrier are assembled via hydrophobic bonding without chemical glues, simply scraping the bonded part between the cover glass and the PDMS barrier. - Introduce 1.5 mL of endothelial cell growth medium to each chip. To avoid detachment of the cancer construct, introduce cell culture medium from one side of the chip. Tilt the chip to allow the cell culture media to flow using a pipette.
- Refresh the cell culture media every day for a week. Use a pipette to aspirate the cell culture medium; do not use a pressure pump.
- After completion of all 3D cell-printing processes of the hypoxic cancer-on-a-chip, gently rub the cover glasses on top of the gas-permeable barriers with the cell-scrapper for tight bonding (Figure 4A,B).
7. Evaluation of post-printing cell viability
- Preparation of samples and treatment with calcein AM and EthD-1 solution
- Warm 1x PBS in a water bath at 37 °C.
- Prepare the assay solution by adding 0.75 µL of calcein acetoxymethyl (calcein AM) and 3 µL of ethidium homodimer (EthD-1) to 1.5 mL pre-warmed PBS.
- Carefully aspirate all media from the chip using a pipette.
- Wash the cancer construct with prewarmed PBS. Fill 1.5 mL PBS into the chip using a pipette and let it stand for 10 min at room temperature. To avoid deformation of the cancer construct, introduce 1x PBS from one side of the chips and tilt the chips to allow 1x PBS to flow.
- Aspirate the PBS from the chip; treat the 1.5 mL assay solution and incubate the chip at 37 °C for 20 min using a foil to protect from light. Use a pipette to aspirate 1x PBS; do not use a suction pump.
- Imaging of the cell viability using a fluorescence microscope
- View and capture the labeled cells using a fluorescence microscope (Figure 4C).
NOTE: Calcein AM marks live cells with green fluorescence (wavelength ~488 nm). EthD-1 represents the signal of dead cells with red fluorescence (wavelength ~594 nm). - Count the number of live and dead cells using imaging software, an open-source image-processing program, and calculate viability with the numbers .
- View and capture the labeled cells using a fluorescence microscope (Figure 4C).
8. Immunofluorescence to validate the formation of central hypoxia and its effect on cancer malignancy
- Fixation, permeabilization, and blocking of the cancer construct
- Prepare 1x PBS, 4% paraformaldehyde (PFA), 0.1% (v/v) Triton X-100, and 2% (w/v) bovine serum albumin (BSA) at room temperature.
- Carefully aspirate all the media from the chip using a pipette and rinse the chip three times with 1x PBS. To avoid deformation of the cancer construct, introduce 1x PBS from one side of the chips and tilt the chips to allow 1x PBS to flow. Between each washing step, let the chip stand with 1x PBS for 5 min to remove residual solutions.
NOTE: 1x PBS was aspirated using a pipette, not a pressure pump. - Add 500 µL of 4% PFA to the cancer construct on the chip using a pipette; leave it for 15 min and wash three times with 1x PBS to fix the cells in the cancer construct.
- Treat cancer construct with 500 µL of 0.1% Triton X-100 using a pipette at room temperature for 5 min and wash three times with 1x PBS to solubilize and permeabilize the cell membrane.
- Treat cancer construct with 500 µL of 2% BSA using a pipette at room temperature for 1 h to block reactive epitopes.
NOTE: Cover the chip with paraffin film to prevent evaporation. - After 1 h, wash the chip three times with 1x PBS.
- Treatment with primary antibody, secondary antibody, and DAPI and imaging of the structure using a confocal microscope.
- Prepare isotype control antibodies and the cocktail of primary antibodies by diluting the antibodies in 1x PBS to each desired working concentration.
NOTE: The specific details of the antibodies are listed in the Table of Materials. The same working concentrations of isotype control antibodies as the primary antibodies should be used. - Carefully aspirate all 1x PBS from the chip using a pipette and treat the chip with 200 µL primary antibody solution at 4 °C overnight. Cover the chips with paraffin film to prevent evaporation.
- Aspirate the primary antibody solution and wash the chip three times with 1x PBS.
- Dilute secondary antibodies and DAPI in 1x PBS to the desired working concentration.
NOTE: A Green fluorescence-conjugated secondary antibody is used in this case at a ratio of 1:200. DAPI was used at a ratio of 1:1000. - Carefully aspirate all 1x PBS from the chip using a pipette and treat the chip with 200 µL secondary antibody-DAPI solution at 4 °C for 3 h. Cover the chip with paraffin film to prevent evaporation and then wrap it with aluminum foil to prevent photobleaching.
- Aspirate the secondary antibody-DAPI solution and wash the chip three times with 1x PBS.
- After finishing the staining step, transfer the cancer construct to a confocal dish by gently gripping with forceps.
- Visualize and capture the labeled cells using a confocal microscope (Figure 5).
NOTE: The wavelength of the confocal microscope was adjusted, depending on the type of the fluorescent markers. The specific details of the antibodies are listed in the Table of Materials. To efficiently detect the cell position, it would be better to observe the DAPI stained nuclei of the construct at first. The detection excitation/emission wavelengths of the fluorescent signals were 358/461 nm (DAPI, Blue), 494/517 nm (Green), and 590/617 nm (Red). The magnifications were 4x, 10x, and 20x, adjusted from the lowest to the highest.
- Prepare isotype control antibodies and the cocktail of primary antibodies by diluting the antibodies in 1x PBS to each desired working concentration.
9. Statistical analysis
- Cell counting with image processing program
- Run an image processing program to count the number of live and dead cells.
- Open the fluorescent image files. Click on File | Open and import the TIFF images.
- Convert the images to 16-bit grayscale images. Click on Image | Type | 16-bit Grayscale.
- Adjust the threshold by clicking on Image | Adjust | Threshold and then select the color of the cells to be black.
- Cut merged cells apart by clicking on Process | Binary | Watershed for precise cell counting.
- Count the number of cells by clicking on Analyze and then on Analyze Particles three times; calculate the average and present the data as the mean ± standard error.
NOTE: Immunofluorescence markers were analyzed by comparing the fluorescence intensity.
Access restricted. Please log in or start a trial to view this content.
Results
The hypoxic cancer-on-a-chip was developed using computer-aided 3D cell-printing technology to recapitulate hypoxia and cancer-related pathology (Figure 1). Oxygen transportation and consumption were simulated using the 3D geometry model. The chip was designed in the form of concentric rings to mimic the radial oxygen diffusion and depletion, in cancer tissues (Figure 2A). After defining the control volume of a space where oxygen...
Access restricted. Please log in or start a trial to view this content.
Discussion
In this study, we describe the fabrication process of a hypoxic cancer-on-a-chip based on 3D cell-printing technology. The formation of the hypoxic gradient in the designed chip was predicted through computer simulations. The environment that can induce a heterogeneous hypoxic gradient was reproduced via a simple strategy combining the 3D-printed gas-permeable barrier and the glass cover. The hypoxia-related pathological features of glioblastoma, including pseudopalisade and a small population of cancer stem cells, were ...
Access restricted. Please log in or start a trial to view this content.
Disclosures
The authors have no disclosures.
Acknowledgements
This research was supported by the National Research Foundation of Korea (NRF) funded by the Ministry of Education (No. 2020R1A6A1A03047902 and NRF-2018H1A2A1062091) and the Korea government (MSIT) (No. NRF-2019R1C1C1009606 and NRF-2019R1A3A3005437).
Access restricted. Please log in or start a trial to view this content.
Materials
| Name | Company | Catalog Number | Comments |
| Cells | |||
| Human umbilical vein endothelial cells | Promocell | C-12200 | |
| U-87 MG cells | ATCC | ATCC HTB-14 | |
| Disposable | |||
| 0.2 μm syringe filter | Sartorius | 16534-K | |
| 10 mL disposable syringe | Jung Rim | 10ml 21G32 | |
| 10 mL glass vial | Hubena | A0039 | |
| 10 mL Serological pipette tip | SPL lifescience | 91010 | |
| 15 mL conical tube | SPL lifescience | 50015 | |
| 18G plastic needle | Musashi engineering | PN-18G-B | |
| 20G plastic tapered dispense tip | Musashi engineering | TPND-20G-U | |
| 22x50 glass cover | MARIENFIELD | 0101142 | |
| 25 mL Serological pipette tip | SPL lifescience | 90125 | |
| 3 mL disposable syringes | HENKE-JET | 4020-X00V0 | |
| 40 µm cell strainer | Falcon | 352360 | |
| 5 mL Serological pipette tip | SPL lifescience | 91005 | |
| 50 mL conical tube | SPL lifescience | 50050 | |
| 50 mL Serological pipette tip | SPL lifescience | 90150 | |
| 50N precision nozzle | Musashi engineering | HN-0.5ND | |
| Aluminum foil | SINKWANG | ||
| Capillary tips | Gilson | CP1000 | |
| Cell-scrapper | SPL lifescience | 90030 | |
| Confocal dish | SPL lifescience | 200350 | |
| Parafilm | Bemis | PM996 | |
| Pre-coated histology slide | MATSUNAMI | MAS-11 | |
| Reservoir | SPL lifescience | 23050 | |
| T-75 cell culture flask | SPL lifescience | 70075 | |
| Equipment | |||
| 3DX printer | T&R Biofab | ||
| Autoclave | JEIOTECH | AC-12 | |
| Centrifuger | Cyrozen | 1580MGR | |
| Confocal laser microscopy | Olympus Life Science | FV 1000 | |
| Fluorescence microscope | FISHER SCEINTIFIC | O221S366 | |
| Forcep | Korea Ace Scientific | HC.203-30 | |
| Hand tally counter | KTRIO | ||
| Hemocytometer | MARIENFIELD | 0650030 | |
| Incubator | Panasonic | MCO-170AIC | |
| Laminar flow cabinet | DAECHUNG SCIENCE | CB-BMMS C-001 | |
| Metal syringe | IWASHITA engineering | SUS BARREL 10CC | |
| Operating Scissors | Hirose | HC.13-122 | |
| Oven | JEIOTECH | OF-12, H070023 | |
| Positive displacement pipette | GILSON | NJ05652 | |
| Refrigerator | SAMSUNG | CRFD-1141 | |
| Voltex Mixer | DAIHAN scientific | VM-10 | |
| Water bath | DAIHAN SCIENTIFIC | WB-11 | |
| Water purifier | WASSER LAB | DI-GR | |
| Materials | |||
| 0.25 % Trypsin-EDTA | Gibco | 25200-072 | |
| 10x PBS | Intron | IBS-BP007a | |
| 4% Paraformaldehyde | Biosesang | ||
| 70% Ethanol | Daejung | 4018-4410 | |
| Anti-CD31 antibody | Abcam | ab28364 | |
| Anti-HIF-1 alpha antibody | Abcam | ab16066 | |
| Anti-SHMT2/SHMT antibody | Abcam | ab88664 | |
| Anti-SOX2 antibody | Abcam | ab75485 | |
| Bovine Serum Albumin | Thermo scientific | J10857-22 | |
| Collagen from porcine skin | Dalim tissen | PC-001-1g | |
| DAPI (4',6-Diamidino-2-Phenylindole, Dihydrochloride) | Thermofisher | D1306 | |
| Endothelial Cell Growth Medium-2 | Promocell | C22011 | |
| Fetal bovine serum | Gibco | 12483-020 | |
| Goat anti-Mouse IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 488 | Theromofisher | A-11001 | |
| Goat anti-Rabbit IgG (H+L) Cross-Adsorbed Secondary Antibody, Alexa Fluor 594 | Theromofisher | A-11012 | |
| High-glucose Dulbecco’s Modified Eagle Medium(DMEM) | Hyclone | SH30243-0 | |
| Hydrochloric acid | Sigma-Aldrich | 311413-100ML | |
| Live/dead assay kit | Invitrogen | L3224 | |
| Mouse IgG1, kappa monoclonal [15-6E10A7] - Isotype Control | Abcam | ab170190 | |
| Penicillin/streptomycin | Gibco | 15140-122 | |
| Phenol red solution | Sigma-Aldrich | P0290-100ML | |
| Poly(ethylene-vinyl acetate) | Poly science | 06108-500 | |
| Polydimethylsiloxane | Dowhitech | sylgard 184 | |
| Rabbit IgG, polyclonal - Isotype Control | Abcam | ab37415 | |
| Sodium hydroxide solution | Samchun | S0610 | |
| Triton X-100 | Biosesang | TRI020-500-50 | |
| Trypan Blue | Sigma-Aldrich | T8154 | |
| Software | |||
| COMSOL Multiphysics 3.5a | COMSOL AB | ||
| IMS beamer | in-house software | ||
| SolidWorks Package | Dassault Systems SolidWorks Corporation |
References
- Jing, X., et al. Role of hypoxia in cancer therapy by regulating the tumor microenvironment. Molecular Cancer. 18 (1), 157(2019).
- Al Tameemi, W., Dale, T. P., Al-Jumaily, R. M. K., Forsyth, N. R. Hypoxia-modified cancer cell metabolism. Frontiers in Cell and Developmental Biology. 7, 4(2019).
- Petrova, V., Annicchiarico-Petruzzelli, M., Melino, G., Amelio, I. The hypoxic tumour microenvironment. Oncogenesis. 7 (1), 1-13 (2018).
- Hockel, M., Vaupel, P. Tumor hypoxia: definitions and current clinical, biologic, and molecular aspects. Journal of the National Cancer Institute. 93 (4), 266-276 (2001).
- Kim, H., Lin, Q., Glazer, P. M., Yun, Z. The hypoxic tumor microenvironment in vivo selects the cancer stem cell fate of breast cancer cells. Breast Cancer Research. 20 (1), 16(2018).
- Jeong, G. S., Lee, J., Yoon, J., Chung, S., Lee, S. -H. Viscoelastic lithography for fabricating self-organizing soft micro-honeycomb structures with ultra-high aspect ratios. Nature Communications. 7 (1), 1-9 (2016).
- Razian, G., Yu, Y., Ungrin, M. Production of large numbers of size-controlled tumor spheroids using microwell plates. Journal of Visualized Experiments:JoVE. (81), e50665(2013).
- Nunes, A. S., Barros, A. S., Costa, E. C., Moreira, A. F., Correia, I. J. 3D tumor spheroids as in vitro models to mimic in vivo human solid tumors resistance to therapeutic drugs. Biotechnology and Bioengineering. 116 (1), 206-226 (2019).
- Wan, L., Neumann, C., LeDuc, P. Tumor-on-a-chip for integrating a 3D tumor microenvironment: chemical and mechanical factors. Lab on a Chip. 20 (5), 873-888 (2020).
- Nam, H., Funamoto, K., Jeon, J. S. Cancer cell migration and cancer drug screening in oxygen tension gradient chip. Biomicrofluidics. 14 (4), 044107(2020).
- Palacio-Castañeda, V., Kooijman, L., Venzac, B., Verdurmen, W. P., Le Gac, S. Metabolic switching of tumor cells under hypoxic conditions in a tumor-on-a-chip model. Micromachines. 11 (4), 382(2020).
- Ronaldson-Bouchard, K., Vunjak-Novakovic, G. Organs-on-a-chip: a fast track for engineered human tissues in drug development. Cell Stem Cell. 22 (3), 310-324 (2018).
- Mi, S., Du, Z., Xu, Y., Sun, W. The crossing and integration between microfluidic technology and 3D printing for organ-on-chips. Journal of Materials Chemistry B. 6 (39), 6191-6206 (2018).
- Yi, H. -G., Lee, H., Cho, D. -W. 3D printing of organs-on-chips. Bioengineering. 4 (1), 10(2017).
- Yi, H. -G., et al. A bioprinted human-glioblastoma-on-a-chip for the identification of patient-specific responses to chemoradiotherapy. Nature Biomedical Engineering. 3 (7), 509-519 (2019).
- Kang, T. -Y., Hong, J. M., Jung, J. W., Yoo, J. J., Cho, D. -W. Design and assessment of a microfluidic network system for oxygen transport in engineered tissue. Langmuir. 29 (2), 701-709 (2013).
- Woo Jung, J., et al. Evaluation of the effective diffusivity of a freeform fabricated scaffold using computational simulation. Journal of Biomechanical Engineering. 135 (8), (2013).
- Brown, A. C., De Beer, D. Development of a stereolithography (STL) slicing and G-code generation algorithm for an entry level 3-D printer. 2013 Africon (IEEE). , 1-5 (2013).
- Shim, J. -H., Lee, J. -S., Kim, J. Y., Cho, D. -W. Bioprinting of a mechanically enhanced three-dimensional dual cell-laden construct for osteochondral tissue engineering using a multi-head tissue/organ building system. Journal of Micromechanics and Microengineering. 22 (8), 085014(2012).
- Gillispie, G., et al. Assessment methodologies for extrusion-based bioink printability. Biofabrication. 12 (2), 022003(2020).
- Kim, B. S., Das, S., Jang, J., Cho, D. -W. Decellularized extracellular matrix-based bioinks for engineering tissue-and organ-specific microenvironments. Chemical Reviews. 120 (19), 10608-10661 (2020).
Access restricted. Please log in or start a trial to view this content.
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved