Method Article
Optogenetic Signaling Activation in Zebrafish Embryos
* These authors contributed equally
In This Article
Summary
Optogenetic manipulation of signaling pathways can be a powerful strategy to investigate how signaling is decoded in development, regeneration, homeostasis, and disease. This protocol provides practical guidelines for using light-oxygen-voltage sensing domain-based Nodal and bone morphogenic protein (BMP) signaling activators in the early zebrafish embryo.
Abstract
Signaling pathways orchestrate fundamental biological processes, including development, regeneration, homeostasis, and disease. Methods to experimentally manipulate signaling are required to understand how signaling is interpreted in these wide-ranging contexts. Molecular optogenetic tools can provide reversible, tunable manipulations of signaling pathway activity with a high degree of spatiotemporal control and have been applied in vitro, ex vivo, and in vivo. These tools couple light-responsive protein domains, such as the blue light homodimerizing light-oxygen-voltage sensing (LOV) domain, with signaling effectors to confer light-dependent experimental control over signaling. This protocol provides practical guidelines for using the LOV-based bone morphogenetic protein (BMP) and Nodal signaling activators bOpto-BMP and bOpto-Nodal in the optically accessible early zebrafish embryo. It describes two control experiments: A quick phenotype assay to determine appropriate experimental conditions, and an immunofluorescence assay to directly assess signaling. Together, these control experiments can help establish a pipeline for using optogenetic tools in early zebrafish embryos. These strategies provide a powerful platform to investigate the roles of signaling in development, health, and physiology.
Introduction
Signaling pathways allow cells to respond to their environment and coordinate activities at tissue- and organism-wide scales. Signals crucial for embryonic development include the TGF-beta superfamily members bone morphogenetic protein (BMP) and Nodal1,2,3. During embryogenesis, the pathways regulated by these signals and others pattern the body plan by controlling gene expression and additional processes to ensure that diverse tissues and organs develop and interface properly. Pathologies, including birth defects and cancer, can occur when signaling or responses to signaling are perturbed4,5,6,7. Despite rigorous investigation into signaling, much more remains to be discovered about how levels and dynamics are decoded in a variety of contexts8,9,10,11, especially during development12,13,14,15,16,17,18,19.
To understand how signaling is decoded, an ideal experiment would be to manipulate signaling levels, timing, and/or dynamics-with a high degree of spatial and temporal control-and assess outcomes. For example, precise spatial signaling gradients are proposed to pattern developing tissues20,21. Altering signaling gradient spatial distributions would help test this hypothesis22. Additionally, the importance of signaling dynamics in generating diverse cellular responses is becoming clearer: The same signaling pathway can instruct cells to differentiate or proliferate depending on signaling frequency, for example9,23. Experimental paradigms in which signaling dynamics can be easily manipulated will be valuable to explore the relationship between dynamics and cell fate decisions8,12,13,14,15.
Historically, multiple methods have been used to manipulate signaling in developmental contexts, leading to fundamental discoveries1,2,3. Signaling can be blocked using pathway loss-of-function mutants, ectopic inhibitor expression, or antagonist drugs. Methods to activate signaling include agonist drugs, recombinant ligands, ectopic expression of ligands or constitutively active receptors, and pathway inhibitor loss-of-function mutants. These methods range along a continuum of experimental control. For example, mutants and ectopic expression may fall on the sledgehammer side of the continuum: With these approaches, dramatic, systemic changes in pathway activity may cause early death and preclude investigations at later stages, or over time may result in pleiotropic effects that are difficult to disentangle. Additionally, it is often challenging to independently manipulate one signaling feature at a time, such as level or duration. Toward the other end of the continuum, some methods offer more precise experimental control, such as microfluidic devices that expose samples to drugs or recombinant proteins with temporal and sometimes spatial control18,24,25, or genetic methods, including heat shock-inducible and tissue-specific promoters that can offer similar benefits16,26,27. However, these methods can be difficult to execute, may not be reversible, may have relatively slow kinetics or poor resolution, and may be unavailable in some model systems.
Molecular optogenetic approaches are a powerful addition to this toolkit. These approaches use proteins that respond to different light wavelengths to manipulate biological processes, including signaling8,12,13,14,15, and have been developed over decades for use in a variety of systems from cell culture to whole animals12,13,28. Compared to historical approaches, molecular optogenetics can often offer a higher degree of spatiotemporal control over biological processes: The controller in optogenetic systems is light, and control of light wavelength, intensity, duration, and exposure frequency is relatively straightforward. With sophisticated systems such as confocal and two-photon microscopes, spatial control in the subcellular range is possible29,30,31. Tools to optogenetically manipulate signaling have been developed and applied in several systems, including those described in Johnson et al.22, Čapek et al.32, Krishnamurthy et al.33, and Huang et al.34. For example, exploiting the spatial control afforded by optogenetics, this strategy was recently used to modify a signaling gradient in Drosophila embryos, demonstrating that fly embryogenesis is surprisingly robust to changes in this gradient22. The reversibility and fast on/off kinetics of optogenetic signaling activators have also made them attractive tools for investigating the decoding of signaling dynamics8,12,13,14,15,34,35,36.
The early zebrafish embryo is an in vivo system well-suited for optogenetic studies because it is externally fertilized, transparent, microscopy-friendly, and genetically tractable. Light exposure is easier to deliver to embryos that develop outside of the mother, light can penetrate and access their non-opaque tissues, live zebrafish embryos tolerate imaging well (in addition to being transparent), and existing genetic methods provide straightforward opportunities for knockdown and overexpression experiments, in addition to the development of useful transgenics37.
Recently, optogenetic tools were developed to activate BMP38 and Nodal39 signaling in zebrafish embryos with blue light exposure (Figure 1). We refer to these tools as bOpto-BMP and bOpto-Nodal (b for blue light-activated and Opto for optogenetic). bOpto-BMP/Nodal are based on similar pathway activation mechanisms. The binding of BMP or Nodal ligands to their respective receptor serine-threonine kinases drives receptor kinase domain interactions that lead to the phosphorylation of signaling effectors (Smad1/5/9 for BMP and Smad2/3 for Nodal). Phosphorylated signaling effectors then translocate to the nucleus and regulate target gene expression3 (Figure 1A,D). These receptor kinase interactions can be made light-responsive by coupling receptor kinases to light-responsive dimerizing proteins: With light exposure, these chimeric proteins should dimerize, causing the receptor kinase domains to interact and activate signaling (Figure 1B,C,E,F). Importantly, in contrast to endogenous receptors, bOpto-BMP/Nodal do not contain extracellular ligand-binding domains, ensuring ligand-independent activity (Figure 1C,F). This optogenetic activation strategy was first achieved with receptor tyrosine kinases40,41,42 and then applied to receptor serine-threonine kinases.
bOpto-BMP/Nodal use the blue light-responsive (~450 nm) homodimerizing light-oxygen-voltage sensing (LOV) domain from the algae Vaucheria fridiga AUREO1 protein (VfLOV)43,44. These constructs consist of a membrane-targeting myristoylation motif followed by either BMP or Nodal receptor kinase domains, fused to a LOV domain (Figure 1B,E). Blue light exposure should cause LOV homodimerization, resulting in receptor kinase domain interactions that lead to respective Smad phosphorylation and pathway activation (Figure 1C,F). For bOpto-BMP, a combination of constructs with the type I receptor kinase domains from Acvr1l (also known as Alk8) and BMPR1aa (also known as Alk3) and the type II receptor kinase domain from BMPR2a was found to optimally activate signaling38 (Addgene #207614, #207615, and #207616). For bOpto-Nodal, a combination of constructs with the type I receptor kinase domain from Acvr1ba and the type II receptor kinase domain from Acvr2ba is used39.
bOpto-BMP/Nodal have been introduced into early zebrafish embryos by injecting mRNA at the one-cell stage, and used to investigate the role of signaling duration in Nodal interpretation39, to determine why zebrafish lose the ability to respond to Nodal45, and to examine how BMP target genes respond to different BMP signaling levels38. It is likely that these tools will continue to be useful in a diverse range of future investigations. However, the strength of optogenetic signaling activators is also their weakness: light-sensitive samples must be treated with care to avoid inadvertent ectopic signaling activity. Exposure to room light or sunlight can activate bOpto-BMP/Nodal.
This protocol provides practical suggestions for using mRNA-encoded LOV-based BMP and Nodal activators in early zebrafish embryos. It begins by detailing one strategy to build a light box to control uniform light exposure and temperature (Figure 2, Supplementary File 1, Supplementary File 2, Supplementary File 3, Supplementary File 4, Supplementary File 5, Supplementary File 6, Supplementary File 7, Supplementary File 8). It then describes two key control experiments that determine whether an optogenetic signaling activator is behaving as expected-i.e., activating pathway activity only when exposed to light (Figure 3). The first control assay involves examining phenotypes at one day post-fertilization in light-exposed and unexposed embryos (Figure 3A). mRNA-injected light-exposed embryos, but not unexposed embryos, should phenocopy BMP or Nodal overexpression (Figure 4A,B; BMP phenotypes in particular are clearly distinguishable at this time point46). This assay provides a fast activity readout. In the second control assay, to determine whether phenotypes are caused specifically by excess BMP or Nodal signaling and to directly observe the change in signaling levels, immunofluorescence staining is used to detect phosphorylated signaling effectors (pSmad1/5/9 or pSmad2/3, respectively) after a 20 min light exposure around late blastula / early gastrulation stage, when signaling activity has been well described12,16,17,47,48,49,50 (Figure 3B and Figure 4C). (Note that, although spatially localized activation has been demonstrated for both bOpto-BMP38 and bOpto-Nodal39, this protocol only describes uniform light exposure and signaling activation strategies.) It is advisable to execute these control experiments prior to applying bOpto-BMP/Nodal to specific research questions in order to determine ideal local experimental conditions.
Protocol
Zebrafish research protocols were reviewed and approved by the NICHD Animal Care and Use Committee at the National Institutes of Health (ASP 21-008). All zebrafish studies were carried out in compliance with the Guide for the Care and use of Laboratory Animals.
1. Building a light box
- To control light exposure and temperature, construct a light box that uses a light-emitting diode (LED) microplate illuminator as a light source (Figure 2A, Table of Materials, Supplementary File 1 , Supplementary File 2). This customizable illuminator provides dynamic, programmable control over multiple wavelengths.
NOTE: There are many possible strategies to build a light box, and an alternative approach may be more appropriate (see e.g., Gerhardt et al.51, Bugaj et al.52, Kumar and Khammash53, and more at https://www.optobase.org/materials/). - Include the following features in the light box: Temperature control (28 °C is ideal for zebrafish embryos), exclusion of unwanted light (e.g., room light and sunlight), uniform delivery of blue light that covers the target area (e.g., 6-well plate), and control over light intensity and exposure dynamics.
NOTE: bOpto-BMP/Nodal are activated by blue light, but some optogenetic tools respond to other wavelengths. Use the appropriate wavelength for the optogenetic tool. - Drill a hole through the top of an incubator (Figure 2B) that is slightly wider than the LED output lens.
- Ensure there are no electrical components in the top of the incubator that will be destroyed by drilling (Table of Materials). This can be ascertained by contacting the incubator manufacturer and asking directly.
- If there is an existing hole on the incubator's top panel, use a step drill to increase the hole size to 1.25 inches. Otherwise use a 1.25 inch hole saw with arbor.
- If there are internal panels that block the light source, drill the appropriate hole size to ensure the light cone is not blocked (Figure 2A). Panels are typically made of thin sheet metal, so use slow speeds and metal drill bits to prevent damage (cobalt drill bit recommended).
- Construct an LED holder to secure the LED microplate illuminator to the top of the incubator (Figure 2C, Supplementary File 3, Supplementary File 4, Supplementary File 5, Supplementary File 6, Supplementary File 7, Supplementary File 8).
- Mount four M3 cube standoffs to the vertical bracket piece using M3 screws.
- Attach the two side brackets to the left and right of the vertical bracket's cube standoffs. Mount another cube standoff to the leftover hole on the side brackets.
- Mount the vertical bracket piece to LED microplate illuminator assembly (Table of Materials) using M6 screws. Place the LED lens gasket over the LED system lens.
- Mount the assembled pieces to the incubator panel mount, then to the M3 standoffs on the vertical and side brackets. Place the incubator gasket over the drilled hole on top of the incubator.
- Place the incubator panel mount on top of the incubator gasket, ensuring the gasket and panel openings are concentric with the incubator's hole. Mount panel to incubator top using no. 8 screws. Ensure this seal is light-tight.
- Place a 6-well plate on the top shelf of the incubator. Determine whether the light beam uniformly covers the plate (Figure 2A). Use a sheet of paper to visualize light coverage.
- If the entire plate is not covered by the beam, increase the distance between the LED and plate by moving the shelf down. For the system described here, ~14 inches between the LED and shelf is sufficient.
- Use a light meter to determine the irradiance level and spatial uniformity (Table of Materials).
- To avoid inadvertent sunlight and room light exposure, use weather stripping to ensure that the incubator door is light-tight (Figure 2A).
- Zebrafish embryos develop robustly at 28 °C54. Use a memory card thermometer to ensure that the light box holds 28 °C.
2. Generating mRNA for injection
NOTE: pCS2+ is the vector backbone for bOpto-BMP constructs38 and bOpto-Nodal constructs39. This vector is ampicillin resistant. bOpto-BMP is composed of three constructs (Figure 1B): BMPR1aa-LOV (Addgene # 207614): Putative kinase domain of the type I BMPR1aa receptor (also known as Alk3) fused to LOV; Acvr1l-LOV (Addgene # 207615): Putative kinase domain of the type I Acvr1l receptor (also known as Alk8) fused to LOV; and BMPR2a-LOV (Addgene # 207616): Putative kinase domain of the type II BMPR2a receptor and following C-terminal domain fused to LOV. bOpto-Nodal is composed of two constructs (Figure 1E): Acvr1ba-LOV: Putative kinase domain of the type I Acvr1ba receptor (also known as Acvr1b) fused to LOV; Acvr2ba-LOV: Putative kinase domain of the type II Acvr2ba receptor (also known as Acvr2b) fused to LOV.
- To linearize plasmids, digest between 2-5 µg plasmid DNA using NotI restriction enzyme at 37 °C for 1-3 h (Table of Materials).
NOTE: It is also possible to generate linearized DNA using the plasmid as PCR template. - Purify DNA using a standard column-based purification kit (Table of Materials).
- Use an in vitro SP6 transcription kit such as a mMessage mMachine kit to transcribe RNA from the linearized template (Table of Materials). Set up two reactions according to manufacturer's recommendations to ensure higher yield.
- Purify RNA using a standard column-based RNA cleanup kit (Table of Materials). It is also possible to purify via precipitation.
3. Injecting mRNA
- At least 1 day in advance of injections, make injection dishes and agarose-coated 6-well plates.
- Prepare 200 mL of 1% agarose in zebrafish embryo medium and microwave until the agarose has dissolved completely. Any standard zebrafish embryo medium should be acceptable; however, exclude methylene blue from embryo medium as it can affect downstream imaging in other applications.
- Carefully pour molten agarose into 100 mm x 15 mm plastic petri dishes. Fill the dishes halfway.
- Rinse an injection dish mold with embryo medium and place gently onto molten agarose, ensuring no bubbles are trapped between the mold and agarose. Use tape to make a tab on the back of the mold for easy placement and retrieval.
- The mold should float in the molten agarose. If the mold sinks, carefully remove from the molten agarose and repeat step 3.1.3.
- After agarose has solidified, use the tab to gently remove the mold. This can be accelerated by placing the dish at 4 °C.
- Make agarose-coated 6-well plates if working with dechorionated embryos for immunofluorescence experiments.
- Use a disposable 10 mL plastic pipette to transfer enough molten agarose to cover the bottom of each well of a 6-well plate.
- Store injection dishes and 6-well plates at 4 °C. They can be used immediately or stored until the agarose is dried out or contaminated (usually 2-3 weeks).
- Prepare flamed glass pipette tips if working with dechorionated embryos for immunofluorescence experiments.
- Insert the end of a glass pipette into a Bunsen burner flame and rotate continuously until the edges are smooth. Dechorionated embryos must comfortably fit through the end of the pipette. Do not allow the opening to shrink below the diameter of an embryo.
- Either purchase or pull microinjection needles (Table of Materials). It is advisable to have extra needles available on the day of injections in case a replacement is necessary.
- The day before injections, set up zebrafish breeders according to the institute's standard operating procedures (SOPs). Keep males and females separated.
- Switch on the light box temperature regulator to maintain 28 °C. To ensure the light box temperature stays at 28 °C, monitor temperature using a memory card thermometer (Table of Materials).
- Prepare mRNA injection mix(es). Inject equimolar amounts of each construct. It is necessary to empirically determine what amount to inject.
- bOpto-BMP transcript sizes are as follows:
Acvr1l-LOV = 2007 nucleotides (nt)
BMPR1aa-LOV = 1983 nt
BMPR2a-LOV = 3409 nt - To inject equimolar amounts, inject 1.01x more of the Acvr1l construct than BMPR1aa; inject 1.72x more of the BMPR2a construct than BMPR1aa.
- bOpto-Nodal transcript sizes are as follows:
Acvr1ba-LOV and Acvr2ba-LOV are both 1962 nt. - To inject equimolar amounts, inject same amount of each construct.
- Prepare equimolar injection mixes, combining all mRNAs targeting one pathway into one injection mix. Include phenol red injection tracer if desired. For the data shown in Figure 4, 15 pg of each bOpto-Nodal construct (Acvr1ba-LOV and Acvr2ba-LOV) was used, and for bOpto-BMP 7.8 pg Acvr1l-LOV and BMPR1aa-LOV, and 13.4 pg BMPR2a-LOV was used.
- Store injection mixes at -20 °C. Once the optimal concentration of injection mix has been determined, make 5-10 µL aliquots and store at -20 °C or -80 °C.
- Injection day
- If performing a phenotyping assay (Figure 3A), inject through the chorion directly into the center of the cell at the one-cell stage according to your lab's SOP. The chorion protects embryos from environmental stressors and keeps lysed embryos contained. This is helpful during scoring for accurate quantification of lysed embryos (Figure 4A,B).
- If performing an immunofluorescence assay (Figure 3B), embryos will eventually need to be dechorionated for imaging (Figure 4C). Therefore, inject directly into the center of the cell of dechorionated embryos at the one-cell stage (see Rogers et al.55 for dechorionating protocol). Alternatively, embryos can be manually dechorionated after fixation, but this is more cumbersome than dechorionating with pronase.
- Use flamed glass pipettes to handle dechorionated embryos (step 3.1.10). Take care to not expose dechorionated embryos to air or plastic, which will cause embryos to lyse. Handle dechorionated embryos gently.
- Prepare an extra dish of non-injected embryos as a proxy to evaluate stage progression (see step 4.3.5). Ensure proxy embryos are from the same set of experimental embryos, such that all embryos were fertilized at the same time from the same parents. This is useful for the immunofluorescence assay where stage is relevant but is unnecessary for the phenotyping assay.
- Use the following conditions for both the phenotyping and immunofluorescence assays: 1) non-injected, unexposed, 2) non-injected, exposed to light, 3) injected, unexposed, 4) injected, exposed to light. Select at least 30 embryos per condition. For optimal embryo health, do not incubate more than 30 embryos per well in a 6-well plate.
- After injection, transfer embryos to labeled Petri dishes or agarose-coated 6-well plates (for dechorionated embryos) and incubate at 28 °C. Embryos are not yet light sensitive. Treat injected embryos as though they are light sensitive after 1.5 hours post-fertilization (hpf).
4. Light exposure experiment
NOTE: Exposure to ~450 nm light with an irradiance of 45 W/m2 robustly activates bOpto-BMP/Nodal without obvious phototoxicity (for light meter information, see Table of Materials). The level of optogenetically activated signaling can be tuned by changing irradiance values38. However, phototoxicity will need to be assessed at higher irradiances.
- Time-sensitive step. At the 4- to 16-cell stage, around 1.5 hpf, remove unfertilized and unhealthy embryos. Redistribute if necessary to ensure the same number of embryos in each well (no more than 30).
NOTE: It is important to perform this step around the 4- to 16-cell stage because embryos will become light sensitive as the injected mRNA is translated to protein. We have not observed evidence that embryos are significantly light sensitive prior to 1.5 hpf. To minimize inadvertent photoactivation if assessing embryos after 1.5 hpf, use red lights, or cover light sources-including microscope stages-with red gel filter paper that blocks LOV-dimerizing blue wavelengths (Table of Materials).- Evaluate embryos consistently to ensure unbiased distributions between conditions and experiments.
- For the phenotyping assay, use the following 1-day light exposure protocol starting at 1.5 hpf (Figure 3A).
- Wrap the unexposed control plate in aluminum foil. This plate should include both non-injected and injected embryos. Be sure the plate is entirely covered and take care not to introduce tears into the foil. Place this plate on the lower shelf of the 28 °C light box (Figure 2A).
- Place the exposed plate on the top shelf of the 28 °C light box (Figure 2A). Ensure the cover is on the dish to avoid embryo medium evaporation. Turn on the blue light (an irradiance of 45 W/m2 robustly activates signaling).
- Close the light box door to avoid inadvertent room light exposure. If desired, include a memory card thermometer inside the light box before closing the door. Do not open the door until phenotype scoring at 1-day post-fertilization (dpf; see step 5.1).
- For the immunofluorescence assay, expose embryos to blue light for 20 min starting around 40% epiboly56 (~6 hpf) and fix immediately after light exposure, together with the unexposed controls (Figure 3B). A 20 min blue light exposure at 40% epiboly reproducibly activates signaling.
- Around 1.5 hpf, separately wrap both the exposed and unexposed dishes in aluminum foil. Be sure the dishes are entirely covered and take care not to introduce tears into the foil. Leave the proxy dish unwrapped. Place the wrapped dishes and the unwrapped proxy dish in the 28 °C light box (Figure 2A). Do not turn on the LED yet.
- If testing more than one mRNA amount, for the unexposed condition, sort embryos injected with different amounts into individual agarose-coated 6-well plates, which will help minimize inadvertent light exposure during subsequent fixation.
- Close the light box door to avoid inadvertent room light exposure.
- Dilute formaldehyde stock to 4% in 1x phosphate-buffered saline (PBS) and aliquot 1 mL into pre-labeled 2 mL round bottom microcentrifuge tubes, one tube per condition. Store at 4 °C.
- Around 5 hpf, remove the proxy dish containing non-injected embryos and assess the developmental stage using a dissecting scope. The stage of the proxy embryos should reflect the stage of the light-sensitive embryos in the wrapped dishes.
- Remove the wrapped dishes as well, so all dishes experience the same temperature. Repeat until proxy embryos have reached 40% epiboly (~6 hpf). Embryogenesis progression is temperature-sensitive54; therefore, the longer the dishes are outside of the incubator, the longer it will take for the embryos to reach 40% epiboly.
- Once the proxy embryos have reached 40% epiboly, unwrap the exposed dish and place it on the top shelf of the light box (Figure 2A). Leave the unexposed dish(es) wrapped and place on the lower shelf. Immediately turn on the blue light, close the door, and set a timer for 20 min (an irradiance of 45 W/m2 robustly activates signaling).
- To prepare for fixation, eliminate as much room light as possible (close window blinds, turn off the overhead lights, turn off screens, etc.). Ensure the formaldehyde-containing tubes are appropriately labeled. Immediately before fixation, remove formaldehyde-containing tubes from 4 °C and place next to the light box.
- Time- and light-sensitive step. Be prepared to move quickly at the end of the 20 min light exposure. After 20 min, open the light box door and remove the unexposed dish.
- Use a flamed glass pipette tip to transfer light-sensitive embryos quickly but gently into the prepared corresponding tube of 4% formaldehyde.
- Minimize the transfer time (<45 s) of light-sensitive embryos to avoid inadvertent light exposure. Ensure there are no air bubbles in the pipette. Air exposure will destroy dechorionated embryos.
- To eject embryos into formaldehyde, submerge the glass pipette tip in the formaldehyde and allow the embryos to sink into the liquid. Minimize the amount of embryo medium that is transferred to the formaldehyde by keeping the embryos at the end of the tip.
- After embryos are transferred, return the pipette to the same well and pipette up and down to dislodge any stuck embryos. This prevents embryos from multiple conditions accidentally ending up in one tube.
- Immediately repeat steps 4.3.11-4.3.13 for the unexposed, non-injected embryos, followed by the exposed embryos (injected and non-injected).
- Store fixed embryos at 4 °C overnight.
5. Experiment evaluation
- Phenotype scoring and imaging
- At 1 dpf, ideally between 24-32 hpf, remove embryos from the light box to assess phenotypes using a dissecting scope and create a scoring rubric. This is the experimental endpoint; inadvertent photoactivation is no longer a concern.
- Score embryos while still in the chorion for ease. Use a pipette or probe to move embryos in order to view from multiple angles.
- Embryos experiencing excess BMP signaling will be ventralized with varying degrees of severity as described in detail in Kishimoto et al. 199746 (Figure 4A, left panel). Embryos experiencing excess Nodal signaling will have a range of developmental defects related to excess mesendoderm (Figure 4A, right panel)1,3,47,57,58,59,60. They will often have lysed by 1 dpf.
- Score each embryo in all conditions (Figure 4B). Acquire an overview image of all embryos in each well. If desired, dechorionate and image individual representative embryos in methylcellulose (Figure 4A).
- Immunofluorescence staining and imaging
- After incubating embryos in 4% formaldehyde at 4 °C overnight, remove formaldehyde and wash 3-5x with 1x phosphate-buffered saline with Tween20 (PBST). Remove PBST and add 100% methanol.
- Close tubes and gently invert them to mix residual PBST and methanol. Wash 2x with methanol and store at -20 °C for at least 2 h up to years.
- For pSmad1/5/9 (BMP) immunofluorescence protocol see Rogers et al.38. For pSmad2/3 (Nodal) immunofluorescence protocol see van Boxtel et al.17 and Rogers et al.47.
- Image immunostained embryos, using a microscope capable of optical sectioning (e.g., a confocal or light sheet microscope). Avoid saturation and maintain identical imaging conditions between all samples stained with the same antibody.
- Image within 5 days of completing the immunofluorescence staining because fluorescence may fade over time.
Results
The goal of the two control experiments described here is to determine whether bOpto-BMP/Nodal activate their respective pathways in response to blue light exposure without affecting signaling in the absence of light, as expected. Use these controls to establish the appropriate experimental workflow in your lab before applying bOpto-BMP/Nodal to your research questions of interest.
The phenotyping assay can be completed in only 2 days and provides a useful indication of signaling activity and phototoxicity (Figure 3A). Injected, blue light-exposed embryos should phenocopy excess BMP signaling (ventralization46; Figure 4A, left panel) or Nodal signaling (developmental defects related to extra mesendoderm1,3,47,57,58,59,60 (Figure 4A, right panel)). If injected, light-exposed embryos are aphenotypic, test the quality of the mRNA and consider injecting more, and double-check the light exposure strategy to ensure constant exposure to bright light (~450 nm light with an irradiance of 45 W/m2 should strongly activate signaling). In contrast, injected, unexposed embryos should look identical to non-injected siblings. If injected, unexposed embryos exhibit phenotypes, reduce the amount of mRNA injected and reassess the experimental set up to ensure that unexposed embryos are protected from light exposure. The data shown in Figure 4B shows the results from typical phenotyping experiments with appropriate mRNA amounts and exposure conditions: strong signaling activity is evident in injected, light-exposed embryos, with only a small fraction of injected, unexposed embryos exhibiting phenotypes.
The phenotyping assay also provides an opportunity to assess phototoxicity. If phototoxicity is negligible, non-injected, light-exposed embryos should appear wild type, similar to non-injected, unexposed embryos. If non-injected, light-exposed embryos have defects, but not non-injected, unexposed embryos, consider decreasing the light irradiance. An irradiance of 45 W/m2 robustly activates signaling without obvious phototoxicity. The data shown in Figure 4B shows no concerning differences between non-injected, light-exposed and non-injected, unexposed embryos, indicating negligible phototoxicity.
Although the immunofluorescence assays require more time and effort (~1 week) compared to the phenotyping assay (2 days), immunofluorescence staining provides a direct readout of signaling pathway activity and may reveal subtle signaling changes that might not be reflected by gross morphology. Immunofluorescence is especially important to assess responses to bOpto-Nodal, because excess Nodal signaling often results in embryos lysing by 1 dpf-which can have many causes-in contrast to the specific ventralization phenotypes characteristic of excess BMP signaling46 (Figure 4A). Injected, blue light-exposed embryos should exhibit a uniform increase in Smad1/5/9 or Smad2/3 phosphorylation compared to non-injected, light-exposed embryos. If levels are not increased, or only weakly increased, test the quality of the mRNA and consider injecting more, and double-check the light exposure strategy. A 20 min exposure to blue light with an irradiance of 45 W/m2 around 40% epiboly should strongly activate signaling. If pSmad staining is non-uniform, try injecting mRNA into the center of the cell (rather than the yolk), which may result in more uniform mRNA distribution.
Injected, unexposed embryos should have pSmad levels comparable to non-injected embryos. Anecdotally, we have observed leakier Smad phosphorylation with bOpto-Nodal than bOpto-BMP. If pSmad levels are increased in injected, unexposed embryos, reduce the amount of mRNA injected. In addition, re-assess the experimental set up to ensure 1) unexposed embryos are not inadvertently exposed to light, and 2) light exposure during fixation is minimal. During the fixation step, it is critical to allow no more than 45 s to elapse between removal from the light box and immersion in formaldehyde. In addition, during this step minimize exposure to room light and sunlight by closing window blinds, turning off white light sources, using red lights, or covering white light sources with blue light-blocking gel filter paper (Table of Materials).
The data in Figure 4C shows the results from typical immunofluorescence staining experiments with appropriate mRNA amounts and light exposure conditions: pSmad levels are similar in non-injected and unexposed embryos, whereas injected, light-exposed embryos exhibit higher levels of Smad phosphorylation.
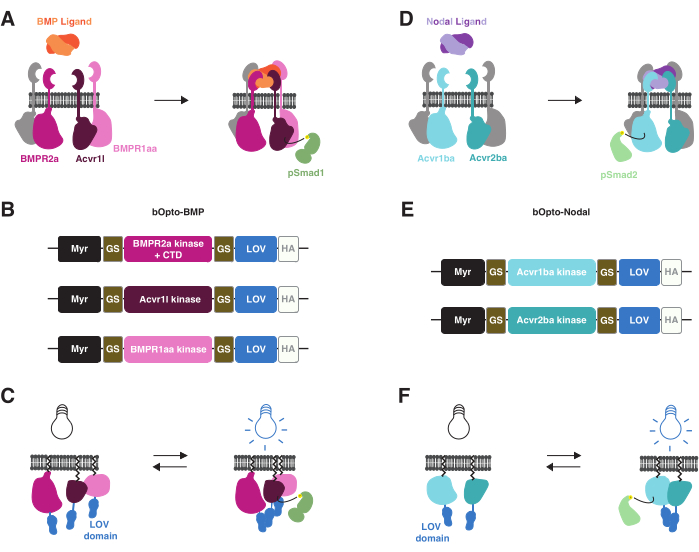
Figure 1: bOpto-BMP and -Nodal signaling activation strategy. (A) The endogenous BMP signaling pathway is activated by BMP ligand binding, leading to formation of a type I/II receptor complex, phosphorylation of Smad1/5/9, and expression of BMP target genes. The type I receptors BMPR1aa and Acvr1l are also known as Alk3 and Alk8, respectively. BMPR2a is a type II receptor. (B) bOpto-BMP constructs38. Putative kinase domains from BMPR1aa and Acvr1l are fused to LOV; the BMPR2a-LOV fusion contains the putative kinase domain and receptor C-terminal domain (CTD). All fusions are membrane-targeted with a myristoylation motif (Myr). Domains are separated by glycine-serine (GS) linkers. Constructs are tagged at the CTD with an HA epitope tag. This combination of three constructs was found to optimally activate BMP signaling. (C) bOpto-BMP-mediated BMP signaling activation. When exposed to blue light, LOV domains dimerize, which is thought to trigger complex formation and signaling activation. (D) The endogenous Nodal signaling pathway is activated by Nodal ligand binding, leading to formation of a type I/II receptor complex, phosphorylation of Smad2/3, and expression of Nodal target genes. The type I receptor Acvr1ba and the type II receptor Acvr2ba are also known as Acvr1b and Acvr2b, respectively. (E) bOpto-Nodal constructs39. Putative kinase domains from Acvr1ba and Acvr2ba are fused to LOV. All fusions are membrane-targeted with a myristoylation motif (Myr). Domains are separated by GS linkers. Constructs are tagged at the CTD with an HA epitope tag. (F) bOpto-Nodal-mediated Nodal signaling activation. When exposed to blue light, LOV domains dimerize, which is thought to trigger complex formation and signaling activation. Please click here to view a larger version of this figure.
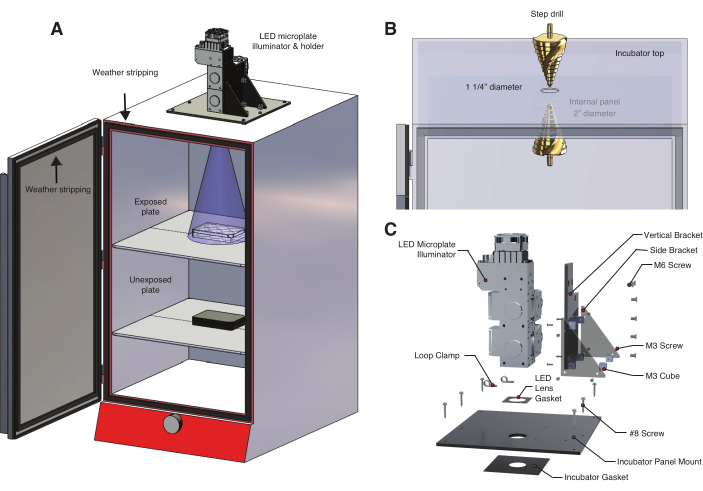
Figure 2: Temperature controlled light box for optogenetic experiments. (A) An LED microplate illuminator is mounted to the top of an incubator using a custom-built LED holder. Zebrafish embryos in a 6-well plate on the first shelf are exposed to light through a hole drilled into the top of the incubator. The lower shelf holds a second set of unexposed control embryos in an aluminum foil-wrapped 6-well plate. The incubator door is lined with weather stripping to prevent inadvertent exposure to room light or sunlight. (B) Detail of procedure to create a hole in the incubator using a step drill. The incubator model used here has an internal panel that required drilling a second, larger hole (Table of Materials). (C) Detail of the custom LED holder designed for a three-wavelength illumination system. Please click here to view a larger version of this figure.
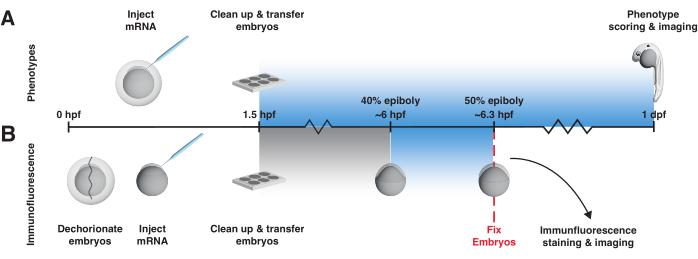
Figure 3: bOpto-BMP/Nodal experiment workflow. Phenotype assay and pSmad immunofluorescence staining to test activity of bOpto-BMP/Nodal. Embryos are injected with mRNA at the one-cell stage and transferred to a light box no later than 1.5 h post-fertilization (hpf). (A) Phenotype assay. Injected embryos and non-injected siblings are reared in the dark or exposed to uniform blue light starting at 1.5 hpf until 1-day post-fertilization (dpf). Optogenetic signaling activity can be evaluated by scoring embryos for phenotypes consistent with excess pathway activity. (B) pSmad immunofluorescence staining. Injected embryos and non-injected siblings are reared in the dark until 40% epiboly (~6 hpf). Half of the injected and half of the non-injected embryos are then exposed to uniform blue light for 20 min. After exposure, all embryos are fixed and subjected to immunofluorescence staining for pSmad. Elevated levels of pSmad1/5/9 or pSmad2/3 reflect optogenetic activation of BMP or Nodal signaling, respectively. Please click here to view a larger version of this figure.
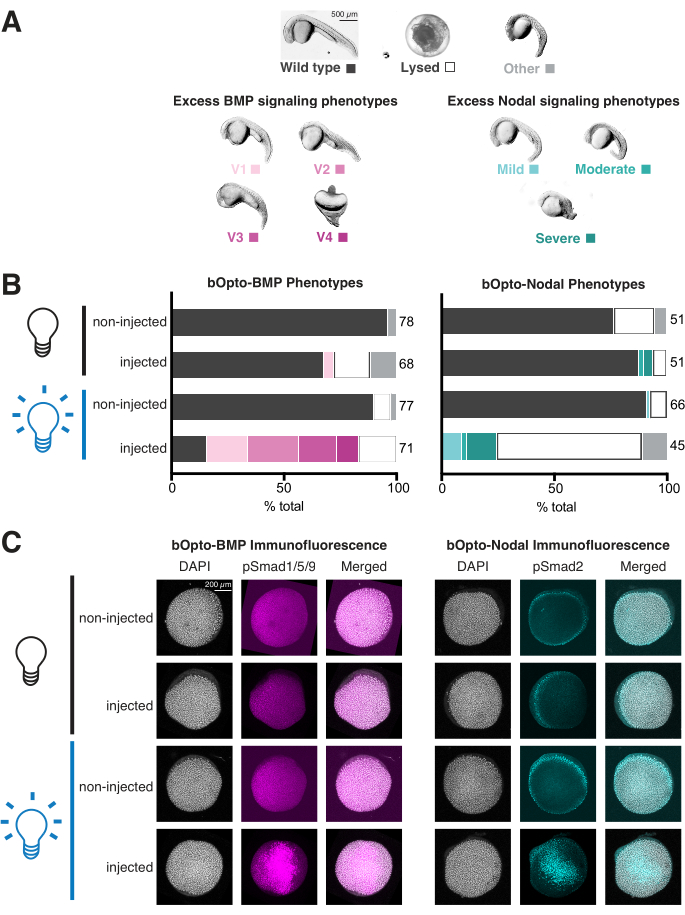
Figure 4: Assessing light-activated signaling responses in zebrafish embryos. Zebrafish embryos were injected at the one-cell stage with mRNA encoding bOpto-BMP/Nodal. (A) Embryos were either reared in the dark or exposed to uniform blue light starting at 1.5 h post fertilization (hpf). Phenotypes were scored at 1 day post fertilization (dpf). Representative phenotypes are shown. Excess BMP signaling leads to ventralization (left panel), while excess Nodal signaling causes developmental defects associated with extra mesendoderm (right panel). Scale bar = 500 µm. (B) Phenotype quantification. Injected embryos and non-injected siblings were reared in the dark starting at 1.5 hpf (black bulb). Half of the injected and half of the non-injected embryos were exposed to uniform blue light (blue bulb). (C) Injected embryos and non-injected siblings were reared in the dark starting at 1.5 hpf (black bulb). At 40% epiboly (~6 hpf), half of the injected and half of the non-injected embryos were exposed to uniform blue light (blue bulb). After 20 min, all embryos were fixed and subjected to immunofluorescence staining for either phosphorylated Smad1/5/9 or Smad2/3. Higher pSmad intensities indicate increased BMP/Nodal signaling, respectively. Scale bar = 200 µm. Please click here to view a larger version of this figure.
Supplementary File 1: Light box full assembly. 3D PDF file showing a 3D view of the full light box assembly. Please click here to download this File.
Supplementary File 2: Light box exploded view. 3D PDF file showing a 3D view of the exploded light box assembly. Please click here to download this File.
Supplementary File 3: Large light gasket. CAD drawing file (.DWG format) to fabricate the large light gasket for the LED holder using a laser cutter. Please click here to download this File.
Supplementary File 4: Small light gasket. CAD drawing file (.DWG format) to fabricate the small light gasket for the LED holder using a laser cutter. Please click here to download this File.
Supplementary File 5: Acrylic platform base. CAD drawing file (.DWG format) to fabricate the LED holder acrylic platform base using a laser cutter. Please click here to download this File.
Supplementary File 6: Acrylic platform vertical. CAD drawing file (.DWG format) to fabricate the LED holder acrylic platform vertical using a laser cutter. Please click here to download this File.
Supplementary File 7: Acrylic support left. CAD drawing file (.DWG format) to fabricate the LED holder acrylic left support using a laser cutter. Please click here to download this File.
Supplementary File 8: Acrylic support right. CAD drawing file (.DWG format) to fabricate the LED holder acrylic right support using a laser cutter. Please click here to download this File.
Discussion
Injection of mRNA is the current strategy to deliver bOpto-BMP/Nodal to zebrafish embryos. This method has several drawbacks. First, the appropriate amount of mRNA varies between labs. The amount used should be sufficient to activate signaling robustly with light exposure, but without inadvertent dark activation. It is a good idea to test several amounts to find optimal mRNA levels, and once established, create aliquots of a master mix to reproducibly introduce the same amount of mRNA. Second, uneven distribution of injected mRNA may lead to uneven signaling activation. Injecting into the center of the cell (not the yolk) is thought to promote even mRNA distribution. Finally, because injected mRNA degrades over time, this approach may not be suitable for experiments in older embryos. In the future, these problems could be addressed by transgenic zebrafish lines ubiquitously expressing bOpto-BMP/Nodal with a maternal or drug-inducible promoter. Although working with potentially light-sensitive adult zebrafish may be a challenge in this context, zebrafish61,62 and Drosophila22,34,35,63 transgenics harboring optogenetic tools have been successfully developed.
Avoiding inadvertent photoactivation is a general challenge with optogenetic tools. For simplicity, treat injected embryos older than 1.5 hpf as light sensitive. Inadvertent light exposure can often be avoided by simply wrapping plates or dishes with aluminum foil. However, for experiments requiring visual observation of live embryos older than 1.5 hpf, it is possible to use red light sources or to cover white light sources with inexpensive gel filter paper that blocks LOV-dimerizing wavelengths (Table of Materials).
The light box described here is designed for specific applications requiring precise control over light irradiance levels, dynamics, and wavelengths (Figure 2). Other benefits of this light box include uniform light exposure, negligible inadvertent sample heating, ample space for multiple 6-well plates, and long-lived, spectrally well-characterized light sources. However, different light exposure strategies may be preferable depending on the research application. Many labs have developed simpler and more cost-effective uniform light exposure systems with smaller footprints, including lining incubators with LED strips, suspending LED panels over samples, or incorporating LEDs in culture dish lids32,38,39,40,64,65,66. Importantly, the light box used in this protocol does not allow users to independently regulate individual wells (in contrast to Bugaj et al.52) or provide spatial control over light exposure. Spatially localized optogenetic activation has been demonstrated with bOpto-BMP38 and bOpto-Nodal39 using lasers in SPIM or confocal systems, respectively, and has also been realized with many other optogenetic strategies in a variety of model systems (discussed in Rogers and Müller12). Some approaches have even achieved sub-cellular spatial resolution29,30,31. Although the implementation of spatially localized light exposure systems is outside of the scope of this protocol, spatial activation experiments with bOpto-BMP/Nodal are theoretically possible with specialized equipment such as digital micromirror devices or masking approaches. Readers are encouraged to explore the extensive literature on DIY light boxes for optogenetic experiments before committing to a light exposure strategy (see e.g., Gerhardt et al.51, Bugaj et al.52, Kumar and Khammash53 and more at https://www.optobase.org/materials/).
Molecular optogenetic strategies often offer a higher degree of spatiotemporal control over biological processes compared to historical approaches such as mutants, ectopic gene expression, recombinant proteins, and drugs. Readers who are interested in the benefits of optogenetic approaches may explore other published tools available in zebrafish and other organisms. These include tools to manipulate additional signaling pathways32,65,67,68, regulate gene expression61,64,66,69,70,71, alter protein localization31,72, and activate apoptosis62. These tools and many others are conveniently cataloged at OptoBase, a curated web resource for molecular optogenetics approaches28. For those inspired to create novel optogenetic tools, the resource also features useful descriptions of light-responsive proteins that have been employed in a wide range of strategies, including light-responsive proteins that respond to green, red, and near-infrared wavelengths. We are excited for the scientific community to realize the full potential of molecular optogenetic approaches.
Disclosures
The authors have nothing to disclose.
Acknowledgements
Funding for this protocol was provided by the NICHD Intramural Program to KWR (ZIA HD009002-01). We thank Jeff Farrell and his lab for their illuminating feedback, Will Anderson for excellent technical support, Leanne Iannucci for stress testing the protocol and measuring irradiance, and the NIH Shared Zebrafish facility for their hard work keeping the zebrafish healthy.
Materials
| Name | Company | Catalog Number | Comments |
| Building a light box & Light exposure protocol | |||
| #8 x 1" Hex Self-drilling Screw | McMaster-Carr | 99663A222 | 1.4.5 |
| Digital Optical Power and Energy Meter | ThorLabs | PM100D | 1.7 4 |
| Incubator (142 liters) | Boekel Scientific | 139400 | 1.3.1 |
| Incubator Panel Mount (1/4" thick cast black acrylic) | Custom part / Piedmont Plastics | Incubator_panel | 1.4.4 |
| Large HSS Spiral Groove Step Drill Bit | CO-Z | SDB0001TA | 1.3.2 |
| LED lens gasket, Incubator gasket; 1/32" thick black silicone | McMaster-Carr | 5812T12 | 1.4.3 1.4.4 |
| LED microplate illuminator | Prizmatix | NA | 1.1 1.4.3 |
| M3 10mm Cube Standoff | Newark Eletronics | 005.60.533 | 1.4.1 |
| M3 x 10mm 316SS Flat Head Screw | McMaster-Carr | 91801A156 | 1.4.1 |
| M6 x 10mm 316SS Flat Head Screw | McMaster-Carr | 91801A305 | 1.4.3 |
| Memory card thermometer | Fisherbrand | 15-081-111 | 1.9 3.2.1 |
| Microscope Slide Power Meter Sensor Head (150 mW) | ThorLabs | S170C | 1.7 4 |
| Red gel filter paper #E106 | Rosco / B&H Foto & Electronics | 110084014805-E106 | 4.2.1 |
| Side Brackets (1/4" thick cast black acrylic) | Custom part / Piedmont Plastics | Side_bracket | 1.4.2 |
| Vertical Bracket (1/4" thick cast black acrylic) | Custom part / Piedmont Plastics | Vertical_bracket | 1.4.1 |
| Weather stripping: Light duty EPDM foam, 1/2" wd 1/4" tk | McMaster-Carr | 8694K12 | 1.8 |
| Generating mRNA | |||
| EZNA MicroElute Cycle Pure Kit | Omega | D6293-02 | 2.4 |
| GeneJET Miniprep Kit (250 rxns) | Thermo Scientific | K0503 | 2.2 |
| Microsample incubator (Hybex) | SciGene | 1057-30-0 | 2 |
| Microsample incubator 1.5 ml tube block (Hybex) | SciGene | 1057-34-0 | 2 |
| Nanodrop One Spectrophotometer | Thermo Scientific | ND-ONE-W | 2.4 |
| NotI-HF restriction enzyme | New England Biolabs (NEB) | R3189L | 2.1 |
| pCS2-Opto-Alk3 | Addgene | 207614 | 2 |
| pCS2-Opto-Alk8 | Addgene | 207615 | 2 |
| pCS2-Opto-BMPR2a | Addgene | 207616 | 2 |
| RNeasy Mini Kit (250 rxns) | Qiagen | 74106 | 2.3 |
| Injecting mRNA | |||
| Agarose (UltraPure) | Invitrogen / Thermo Fisher | 16500500 | 3.1.1 |
| 250 ml glass beakers | Fisherbrand | FB100250 | 3.3.2 |
| 6-well dishes (case of 50) | Falcon | 08 772 1B | 3.1.6 |
| B-8A ball joint | Narishige | B-8A | 3.3 |
| Back pressure unit (microinjection rig component) | Applied Scientific Instrumentation (ASI) | BPU | 3.3 |
| Foot switch (microinjection rig component) | Applied Scientific Instrumentation (ASI) | FWS | 3.3 |
| GJ-1 magnetic stand | Narishige | GJ-1 | 3.3 |
| Glass capillaries (4 in, OD 1 mm, filament) | World Precision Instruments | 1B100F-4 | 3.1.11 |
| Glass petri dish bottoms (for dechorionating) | Pyrex | 08-748A | 3.3.2 |
| Glass pipettes (5 3/4" with wide tip) | Kimble-Chase | 63A53WT | 3.1.9 |
| Injection dish molds | Adaptive Science Tools | tu1 | 3.1.3 |
| IP iron plate | Narishige | IP | 3.3 |
| M-152 micromanipulator | Narishige | M-152 | 3.3 |
| Micro pipette holder kit (microinjection rig component) | Applied Scientific Instrumentation (ASI) | MIMPH-MPIP-Kit | 3.3 |
| Micrometers | Meiji Techno America | MA285 | 3.3 |
| MPPI-2 pressure injector (microinjection rig component) | Applied Scientific Instrumentation (ASI) | MPPI-3 | 3.3 |
| Needle puller | World Precision Instruments | PUL-1000 | 3.1.11 |
| Petri dishes (100 mm x 15 mm, case of 500) | Falcon | 08-757-100D | 3.1.2 |
| Pipettor (10 ml, green) | Bel-Art | F37898-0000 | 3.3 |
| Pronase | Roche | 11459643001 | 3.3.2 |
| Squeeze bottles (500 ml) | Nalgene / Thermo Scientific | 2402-0500 | 3.3 |
References
- Jones, W. D., Mullins, M. C. Cell signaling pathways controlling an axis organizing center in the zebrafish. Current Topics in Developmental Biology. 150, 149-209 (2022).
- Hill, C. S. Establishment and interpretation of NODAL and BMP signaling gradients in early vertebrate development. Current Topics in Developmental Biology. 149, 311-340 (2022).
- Zinski, J., Tajer, B., Mullins, M. C. TGF-β Family Signaling in Early Vertebrate Development. Cold Spring Harbor Perspectives in Biology. 10 (6), a033274 (2018).
- Shore, E. M., Kaplan, F. S. Inherited human diseases of heterotopic bone formation. Nature Reviews. Rheumatology. 6 (9), 518-527 (2010).
- Hebron, K. E., Hernandez, E. R., Yohe, M. E. The RASopathies: from pathogenetics to therapeutics. Disease Models & Mechanisms. 15 (2), dmm049107 (2022).
- Grant, M. G., Patterson, V. L., Grimes, D. T., Burdine, R. D. Modeling Syndromic Congenital Heart Defects in Zebrafish. Current Topics in Developmental Biology. 124, 1-40 (2017).
- Nusse, R., Clevers, H. Wnt/beta-Catenin Signaling, Disease, and Emerging Therapeutic Modalities. Cell. 169 (6), 985-999 (2017).
- Farahani, P. E., Reed, E. H., Underhill, E. J., Aoki, K., Toettcher, J. E. Signaling, Deconstructed: Using Optogenetics to Dissect and Direct Information Flow in Biological Systems. Annual Review of Biomedical Engineering. 23, 61-87 (2021).
- Purvis, J. E., Lahav, G. Encoding and decoding cellular information through signaling dynamics. Cell. 152 (5), 945-956 (2013).
- Wibisana, J. N., Okada, M. Encoding and decoding NF-kappaB nuclear dynamics. Current Opinion in Cell Biology. 77, 102103 (2022).
- Friedel, L., Loewer, A. The guardian's choice: how p53 enables context-specific decision-making in individual cells. TheFEBS Journal. 289 (1), 40-52 (2022).
- Rogers, K. W., Müller, P. Optogenetic approaches to investigate spatiotemporal signaling during development. Current Topics in Developmental Biology. 137, 37-77 (2020).
- Johnson, H. E., Toettcher, J. E. Illuminating developmental biology with cellular optogenetics. Current Opinion in Biotechnology. 52, 42-48 (2018).
- Bosman, S. L., Sonnen, K. F. Signaling oscillations in embryonic development. Current Topics in Developmental Biology. 149, 341-372 (2022).
- Li, P., Elowitz, M. B. Communication codes in developmental signaling pathways. Development. 146 (12), dev170977 (2019).
- Tucker, J. A., Mintzer, K. A., Mullins, M. C. The BMP signaling gradient patterns dorsoventral tissues in a temporally progressive manner along the anteroposterior axis. Developmental Cell. 14 (1), 108-119 (2008).
- van Boxtel, A. L., et al. A temporal window for signal activation dictates the dimensions of a Nodal signaling domain. Developmental Cell. 35 (2), 175-185 (2015).
- Sorre, B., Warmflash, A., Brivanlou, A. H., Siggia, E. D. Encoding of temporal signals by the TGF-β pathway and implications for embryonic patterning. Developmental Cell. 30 (3), 334-342 (2014).
- Economou, A. D., Hill, C. S. Temporal dynamics in the formation and interpretation of Nodal and BMP morphogen gradients. Current Topics in Developmental Biology. 137, 363-389 (2020).
- Rogers, K. W., Schier, A. F. Morphogen gradients: from generation to interpretation. Annual Review of Cell and Developmental Biology. 27, 377-407 (2011).
- Barkai, N., Shilo, B. Z. Robust generation and decoding of morphogen gradients. Cold Spring Harbor Perspectives in Biology. 1 (5), a001990 (2009).
- Johnson, H. E., Djabrayan, N. J. V., Shvartsman, S. Y., Toettcher, J. E. Optogenetic Rescue of a Patterning Mutant. Current Biology. 30 (17), 3414-3424 (2020).
- Imayoshi, I., et al. Oscillatory control of factors determining multipotency and fate in mouse neural progenitors. Science. 342 (6163), 1203-1208 (2013).
- Lin, B., et al. Synthetic spatially graded Rac activation drives cell polarization and movement. Proceedings of the National Academy of Sciences of the United States of America. 109 (52), E3668-E3677 (2012).
- Cui, K. W., et al. Spatially controlled stem cell differentiation via morphogen gradients: A comparison of static and dynamic microfluidic platforms. Journal of Vacuum Science & Technology. A, Vaccum, Surfaces, and Films. 38 (3), 033205 (2020).
- Faden, F., Mielke, S., Lange, D., Dissmeyer, N. Generic tools for conditionally altering protein abundance and phenotypes on demand. Biological Chemistry. 395 (7-8), 737-762 (2014).
- Shoji, W., Sato-Maeda, M. Application of heat shock promoter in transgenic zebrafish. Development, Growth & Differentiation. 50 (6), 401-406 (2008).
- Kolar, K., Knobloch, C., Stork, H., Znidaric, M., Weber, W. OptoBase: A web platform for molecular optogenetics. ACS Synthetic Biology. 7 (7), 1825-1828 (2018).
- Benedetti, L., et al. Light-activated protein interaction with high spatial subcellular confinement. Proceedings of the National Academy of Sciences of the United States of America. 115 (10), E2238-E2245 (2018).
- Krueger, D., De Renzis, S. Optogenetic Methods to Control Tissue Mechanics in Drosophila. Methods in Molecular Biology. 2540, 269-283 (2022).
- Buckley, C. E. Optogenetic Control of Subcellular Protein Location and Signaling in Vertebrate Embryos. Methods in Molecular Biology. 1920, 143-162 (2019).
- Čapek, D., et al. Light-activated Frizzled7 reveals a permissive role of non-canonical wnt signaling in mesendoderm cell migration. Elife. 8, e42093 (2019).
- Krishnamurthy, V. V., et al. Reversible optogenetic control of kinase activity during differentiation and embryonic development. Development. 143 (21), 4085-4094 (2016).
- Huang, A., Amourda, C., Zhang, S., Tolwinski, N. S., Saunders, T. E. Decoding temporal interpretation of the morphogen Bicoid in the early Drosophila embryo. Elife. 6, e26258 (2017).
- Johnson, H. E., Toettcher, J. E. Signaling dynamics control cell fate in the early Drosophila embryo. Developmental Cell. 48 (3), 361.e3-370.e3 (2019).
- Aoki, K., et al. Stochastic ERK activation induced by noise and cell-to-cell propagation regulates cell density-dependent proliferation. Molecular Cell. 52 (4), 529-540 (2013).
- Chow, R. W., Vermot, J. The rise of photoresponsive protein technologies applications in vivo: a spotlight on zebrafish developmental and cell biology. F1000Research. , (2017).
- Rogers, K. W., ElGamacy, M., Jordan, B. M., Müller, P. Optogenetic investigation of BMP target gene expression diversity. Elife. 9, e58641 (2020).
- Sako, K., et al. Optogenetic control of Nodal signaling reveals a temporal pattern of Nodal signaling regulating cell fate specification during gastrulation. Cell Reports. 16 (3), 866-877 (2016).
- Grusch, M., et al. Spatio-temporally precise activation of engineered receptor tyrosine kinases by light. The EMBO Journal. 33 (15), 1713-1726 (2014).
- Crossman, S. H., Janovjak, H. Light-activated receptor tyrosine kinases: Designs and applications. Current Opinion in Pharmacology. 63, 102197 (2022).
- Kainrath, S., Janovjak, H. Design and Application of Light-Regulated Receptor Tyrosine Kinases. Methods in Molecular Biology. 2173, 233-246 (2020).
- Takahashi, F., et al. AUREOCHROME, a photoreceptor required for photomorphogenesis in stramenopiles. Proceedings of the National Academy of Sciences of the United States of America. 104 (49), 19625-19630 (2007).
- Toyooka, T., Hisatomi, O., Takahashi, F., Kataoka, H., Terazima, M. Photoreactions of aureochrome-1. Biophysical Journal. 100 (11), 2801-2809 (2011).
- Vopalensky, P., Pralow, S., Vastenhouw, N. L. Reduced expression of the Nodal co-receptor Oep causes loss of mesendodermal competence in zebrafish. Development. 145 (5), dev.158832 (2018).
- Kishimoto, Y., Lee, K. H., Zon, L., Hammerschmidt, M., Schulte-Merker, S. The molecular nature of zebrafish swirl: BMP2 function is essential during early dorsoventral patterning. Development. 124 (22), 4457-4466 (1997).
- Rogers, K. W., et al. Nodal patterning without Lefty inhibitory feedback is functional but fragile. Elife. 6, e28785 (2017).
- Dubrulle, J., et al. Response to Nodal morphogen gradient is determined by the kinetics of target gene induction. Elife. 4, e05042 (2015).
- Harvey, S. A., Smith, J. C. Visualisation and quantification of morphogen gradient formation in the zebrafish. PLoS Biology. 7 (5), e1000101 (2009).
- Zinski, J., Tuazon, F., Huang, Y., Mullins, M., Umulis, D. Imaging and Quantification of P-Smad1/5 in Zebrafish Blastula and Gastrula Embryos. Methods in Molecular Biology. 1891, 135-154 (2019).
- Gerhardt, K. P., Castillo-Hair, S. M., Tabor, J. J. DIY optogenetics: Building, programming, and using the Light Plate Apparatus. Methods in Enzymology. 624, 197-226 (2019).
- Bugaj, L. J., Lim, W. A. High-throughput multicolor optogenetics in microwell plates. Nature Protocols. 14 (7), 2205-2228 (2019).
- Kumar, S., Khammash, M. Platforms for Optogenetic Stimulation and Feedback Control. Frontiers in Bioengineering and Biotechnology. 10, 918917 (2022).
- Urushibata, H., et al. Control of Developmental Speed in Zebrafish Embryos Using Different Incubation Temperatures. Zebrafish. 18 (5), 316-325 (2021).
- Rogers, K. W., Bläßle, A., Schier, A. F., Müller, P. Measuring protein stability in living zebrafish embryos using fluorescence decay after photoconversion (FDAP). Journal of Visualized Experiments. (95), e52266 (2015).
- Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B., Schilling, T. F. Stages of embryonic development of the zebrafish. Developmental Dynamics. 203 (3), 253-310 (1995).
- Feldman, B., et al. Zebrafish organizer development and germ-layer formation require nodal-related signals. Nature. 395 (6698), 181-185 (1998).
- Shimizu, T., et al. Cooperative roles of Bozozok/Dharma and Nodal-related proteins in the formation of the dorsal organizer in zebrafish. Mechanisms of Development. 91 (1-2), 293-303 (2000).
- Rebagliati, M. R., Toyama, R., Fricke, C., Haffter, P., Dawid, I. B. Zebrafish nodal-related genes are implicated in axial patterning and establishing left-right asymmetry. Developmental Biology. 199 (2), 261-272 (1998).
- Gritsman, K., Talbot, W. S., Schier, A. F. Nodal signaling patterns the organizer. Development. 127 (5), 921-932 (2000).
- LaBelle, J., et al. TAEL 2.0: An Improved Optogenetic Expression System for Zebrafish. Zebrafish. 18 (1), 20-28 (2021).
- Mruk, K., Ciepla, P., Piza, P. A., Alnaqib, M. A., Chen, J. K. Targeted cell ablation in zebrafish using optogenetic transcriptional control. Development. 147 (12), dev183640 (2020).
- Johnson, H. E., et al. The spatiotemporal limits of developmental Erk signaling. Developmental Cell. 40 (2), 185-192 (2017).
- Motta-Mena, L. B., et al. An optogenetic gene expression system with rapid activation and deactivation kinetics. Nature Chemical Biology. 10 (3), 196-202 (2014).
- Patel, A. L., et al. Optimizing photoswitchable MEK. Proceedings of the National Academy of Sciences of the United States of America. 116 (51), 25756-25763 (2019).
- LaBelle, J., Woo, S. Light-Induced GFP Expression in Zebrafish Embryos using the Optogenetic TAEL/C120 System. Journal of Visualized Experiments. (174), e62818 (2021).
- Kainrath, S., Stadler, M., Reichhart, E., Distel, M., Janovjak, H. Green-light-induced inactivation of receptor signaling using cobalamin-binding domains. Angewandte Chemie. 56 (16), 4608-4611 (2017).
- Benman, W., et al. Temperature-responsive optogenetic probes of cell signaling. Nat Chem Biol. 18 (2), 152-160 (2022).
- Reade, A., et al. TAEL: a zebrafish-optimized optogenetic gene expression system with fine spatial and temporal control. Development. 144 (2), 345-355 (2017).
- Putri, R. R., Chen, L. Spatiotemporal control of zebrafish (Danio rerio) gene expression using a light-activated CRISPR activation system. Gene. 677, 273-279 (2018).
- Liu, H., Gomez, G., Lin, S., Lin, S., Lin, C. Optogenetic control of transcription in zebrafish. PLoS One. 7 (11), e50738 (2012).
- Buckley, C. E., et al. Reversible optogenetic control of subcellular protein localization in a live vertebrate embryo. Developmental Cell. 36 (1), 117-126 (2016).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved