A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Preparation of Multifunctional Silk-Based Microcapsules Loaded with DNA Plasmids Encoding RNA Aptamers and Riboswitches
In This Article
Summary
The protocol describes the formation of robust and biocompatible DNA-laden microcapsules as multiplexed in vitro biosensors capable of tracking several ligands.
Abstract
We introduce a protocol for the preparation of DNA-laden silk fibroin microcapsules via the Layer-by-Layer (LbL) assembly method on sacrificial spherical cores. Following adsorption of a prime layer and DNA plasmids, the formation of robust microcapsules was facilitated by inducing β-sheets in silk secondary structure during acute dehydration of a single silk layer. Hence, the layering occurred via multiple hydrogen bonding and hydrophobic interactions. Upon adsorption of multilayered shells, the core-shell structures can be further functionalized with gold nanoparticles (AuNPs) and/or antibodies (IgG) to be used for remote sensing and/or targeted delivery. Adjusting several key parameters during sequential deposition of key macromolecules on silica cores such as the presence of a polymer primer, the concentration of DNA and silk protein, as well as a number of adsorbed layers resulted in biocompatible, DNA-laden microcapsules with variable permeability and DNA loadings. Upon dissolution of silica cores, the protocol demonstrated the formation of hollow and robust microcapsules with DNA plasmids immobilized to the inner surface of the capsule membrane. Creating a selectively permeable biocompatible membrane between the DNA plasmids and the external environment preserved the DNA during long-term storage and played an important role in the improved output response from spatially confined plasmids. The activity of DNA templates and their accessibility were tested during in vitro transcription and translation reactions (cell-free systems). DNA plasmids encoding RNA light-up aptamers and riboswitches were successfully activated with corresponding analytes, as was visualized during localization of fluorescently labeled RNA transcripts or GFPa1 protein in the shell membranes.
Introduction
The field of synthetic biology offers unique opportunities to develop sensing capabilities by exploiting natural mechanisms evolved by microorganisms to monitor their environment and potential threats. Importantly, these sensing mechanisms are typically linked to a response that protects these microorganisms from harmful exposure, regulating gene expression to mitigate negative effects or prevent intake of toxic materials. There have been significant efforts to engineer these microorganisms to create whole-cell sensors taking advantage of these natural responses but re-directing them to recognize novel targets and/or to produce a measurable signal that can be measured for quantification purposes (typically fluorescence)1,2. Currently, concerns with the use of genetically modified microorganisms (GMOs), especially when releasing in the environment or the human body, due to leakage of whole cells or some of their genetic material, even if encapsulated in a polymer matrix, suggest that alternative ways to exploit these sensing approaches are needed3.
A powerful approach to exploit the benefits of microorganisms-based sensing without the concern for the deployment of GMOs is the use of in vitro transcription/translation (IVTT) systems. From a practical perspective, IVTT systems consist of a mixture containing most of the cell components in an active state that has been "extracted" from cells by different means, including sonication, bead-beating, or others4. The final product of this process is a biochemical reaction mixture already optimized to perform transcription and translation that can be used to test different sensors in an "open vessel" format, without the constraints associated with the use of whole cells (membrane diffusion, transformation efficiency, cell toxicity, etc.). Importantly, different sensor components can be quantitatively added, and their effect studied by different optical and spectrometric techniques, as we have demonstrated5. It has been noticed that the performance of IVTT systems can be inconsistent; however, recent studies have shown approaches to standardize their preparation and characterization, which is of great help when studying their performance in sensor design6. Recently, many examples of IVTT systems using to create paper-based assays through the lyophilization of their components in paper matrices have been demonstrated, including detection of heavy metal ions, drugs, quorum sensing elements, and others7,8,9. An exciting application space for IVTT-based sensors is their use in sensing applications in different types of environments, including soil, water, and the human body. In order to deploy these IVTT systems to these challenging environments, an encapsulation approach need to be implemented to contain the IVTT components and protect them from degradation.
The most common encapsulation approaches for IVTT systems include the use of lipid capsules, micelles, polymersomes, and other tightly enclosed microcontainers10,11,12. One disadvantage of this approach is the need to incorporate either passive or active mechanisms to transport materials in and out of the containers to allow communication with the external environment and provide sensing capabilities. To overcome some of these issues, the study here reports a method that provides a simple yet effective approach to encapsulate the encoding materials for different sensor designs to be expressed in IVTT systems. This approach is based on the use of Layer-by-Layer (LbL) deposition of a biopolymer in the presence of the plasmids of interest to create hollow microcapsules with high porosity, which allows the protected genetic material to interact with the different components of the IVTT of choice. The study demonstrated that encapsulated plasmids could direct transcription and translation when activated within this polymeric matrix, as shown with the response of a plasmid-encoded aptamer and a riboswitch to their corresponding targets. Additionally, this LbL coating protects the plasmids for months without any special storage conditions.
Protocol
1. Construction of plasmid vector.
- Construct a plasmid vector (pSALv-RS-GFPa1, 3.4 kb) by amplification of the coding sequence of a theophylline riboswitch (ThyRS) coupled with GFPa1 from pJ201:23976-RS-GFPa1 vector (designed and created by DNA2.0) and insertion into E. coli expression vector, pSAL13. Use forward (5'-CGTGGTACCGGTGATACCAGCATCGTCTTGATG-3') and reverse (5'-CGTGCTCAGCTTAAGCCAGCTCGTAG-3') primers to amplify the coding sequence of ThyRS coupled with GFPa1 and perform a PCR reaction in 50 µL volume using DNA polymerase according to the manufacturer's protocol14.
- Prepare a 1% agarose gel from 0.5 g of agarose, 50 mL of TAE buffer (40 mM Tris Acetate, 1 mM EDTA, pH 8.0), and 3 µL of DNA stain.
- Mix 5 µL aliquot of the PCR-amplified product with 5 µL of RNase/DNase-free water and 2 µL of 6x gel loading dye and analyze by agarose gel electrophoresis. Load a DNA ladder (0.1-10.0 kb) as a reference. Run the gel at 120 V until the dye line has almost reached the bottom of the gel.
- Visualize the DNA fragments using a UV transilluminator imaging system to check for the correct size of DNA15.
- Purify the PCR product using a PCR purification kit according to the manufacturer's protocol16.
- Digest the PCR product and pSAL expression vector with KpnI and BlpI restriction enzymes in a 15 µL reaction, containing 10 µL of PCR product or plasmid vector (concentration 20-50 ng/µL), 1.5 µL of 10x enzyme buffer, 1 µL of each enzyme, and 1.5 µL of RNase/DNase-free water, at 37 °C for 2 h.
- Add 3 µL of 6x gel loading dye to the reaction mixture and separate the digested fragments on a 1% agarose gel as described in steps 1.3-1.5.
- Purify the DNA fragments using a gel extraction kit according to the manufacturer's protocol16.
- Ligate the digested PCR product into a digested linearized plasmid vector, pSAL, using T4 DNA ligase and supplemented ligase buffer in a 10 µL reaction containing 3-20 fmol of the digested vector, 9-60 fmol of the digested PCR product, 2 µL of ligase buffer, 1 µL (1 unit) of T4 DNA Ligase, and DNase/RNase-free water. Incubate the ligation reaction at 25 °C for 3 h.
NOTE: Ensure that the total DNA content in the reaction mixture is 0.01-0.1 µg. - Transform E. coli DH5α competent cells with 10 ng of the ligation reaction mixture according to the manufacturer's protocol17.
- Grow the transformed cells at 37 °C overnight on LB-agar plates supplemented with ampicillin (100 µg/mL).
- Pick 3-4 bacterial colonies from the plate and aseptically transfer each of them into 5 mL of LB media supplemented with ampicillin (100 µg/mL). Grow the cultures overnight at 37 °C in a shaking incubator at 225 rpm.
- Pellet the overnight cultures by centrifugation at 11 x g for 3 min at room temperature.
- Use a purification kit to purify the plasmids according to the manufacturer's protocol16.
- Verify the sequences of the purified plasmids by DNA sequencing. The plasmid map and the sequence of the resulting construct (pSALv-RS-GFPa1) are shown in Figure 1.
2. Large-scale DNA purification.
- Transform the plasmid vector pSALv-RS-GFPa1 (3.4 kb) (encoding theophylline riboswitch coupled with GFPa1 reporter gene) or pET28c-F30-2xBroccoli (5.4 kb) (encoding Broccoli aptamer) into E. coli DH5α competent cells according to the manufacturer's protocol17.
- Grow the transformed cells at 37 °C overnight on LB-agar plates supplemented with ampicillin (100 µg/mL) for cells transformed with pSALv-RS-GFPa1 or kanamycin (50 µg/mL) for cells transformed with pET28c-F30-2x Broccoli.
- Pick 3-4 bacterial colonies from the plate and aseptically transfer each colony into 5 mL of LB media supplemented with appropriate antibiotic (100 µg/mL ampicillin or 50 µg/mL kanamycin). Grow the cultures overnight at 37 °C in a shaking incubator at 225 rpm.
- Use 3 mL of the overnight culture to inoculate into 150 mL of LB supplemented with appropriate antibiotic (100 µg/mL ampicillin or 50 µg/mL kanamycin) and grow the cultures overnight at 37 °C in a shaking incubator at 225 rpm.
- Pellet the cells by centrifugation at ≥3400 x g for 10 min at 4 °C.
- Use a purification kit to purify the plasmids according to the manufacturer's protocol16.
- Elute the DNA with 0.5 mL of pure DNase/RNase-free water. Measure the DNA concentration and prepare 1 mL of DNA stock solutions (100 ng/µL). Store the tubes with DNA at 4 °C until further use.
3. Extraction of silk fibroin and preparation of initial materials.
- Prepare an aqueous solution of reconstituted silk fibroin (SF) protein from Bombyx mori silkworm cocoons according to the procedure described in detail elsewhere to account for 10% of Silk-LiBr solution18.
- Determine the final concentration of the aqueous SF solution. Pipette 0.5 mL of silk solution to a 60 mm Petri dish, let it dry at 60 °C, and measure the weight of dry silk film. Divide the dry weight by 0.5 mL to calculate the weight per volume percentage.
- Dilute the concentrated silk solution with DNase/RNase-free distilled water by slowly adding water via serological pipette to obtain 1 mg/mL final concentration. Store the solution at 4 °C for future use.
- Prepare fluorescently labeled silk fibroin using an antibody labeling kit. Use 1 mL of 2 mg/mL silk fibroin solution to couple the N-terminal α-amino groups of the protein with an NHS ester-activated derivative dye according to the manufacturer's protocol19.
- Prepare 50 mL of polyethyleneimine (PEI) aqueous solution with a concentration of 6 mg/mL, adjust pH to 4 with HCl (1 M). Filter the solution through a sterile 0.2 µm membrane. Storage is possible at ambient conditions for months.
- Prepare SiO2 cores. Pipette 300 µL of SiO2 particles into a 2 mL microcentrifuge tube. Wash the microparticles two times with 1 mL of DNase/RNase-free water by centrifugation at 0.2 x g for 1 min.
4. Perform Layer-by-Layer deposition of a prime layer, DNA plasmids, and silk layers.
- To deposit the PEI prime layer onto the SiO2 microparticles, add 1 mL of PEI solution to the spun down pellet from step 3.6 and agitate the mixture at ambient conditions on a thermomixer at 800 rpm for 15 min. Wash the particles four times with 1 mL of DNase/RNase-free deionized water by centrifugation at 0.2 x g for 1 min.
- To perform deposition of the DNA layer, add 1 mL of the aqueous solution of DNA plasmids from step 2.7 to the PEI-primed microparticles and gently agitate the mixture at 4 °C on a thermomixer at 800 rpm for 15 min. To prepare microcapsules with different DNA loads, adjust the concentration of DNA plasmids from 50-200 ng/µL using DNase/RNase-free distilled water and use 1 mL of these solutions to deposit the DNA. Collect the microparticles by centrifugation at 0.2 x g for 1 min.
- Mark the tubes for DNA plasmids encoding theophylline riboswitch coupled with GFPa1 as ThyRS-GFPa1, and DNA plasmids encoding Broccoli aptamer as BrocApt.
NOTE: Keep the microcentrifuge tubes with DNA on ice. - Carefully remove the supernatant and wash the microparticles four times with 1 mL of DNase/RNase-free distilled water, each time discarding the supernatant after centrifugation at 0.2 x g for 1 min. Perform all experiments at room temperature (RT) unless it is specified otherwise.
- To perform deposition of the silk fibroin layer add 1 mL of the reconstituted aqueous SF solution from step 3.3 to the DNA-adsorbed microparticles, gently vortex and agitate the mixture at 10 °C on the thermomixer at 750 rpm for 15 min. Collect the microparticles by centrifugation at 0.2 x g for 1 min at 4 °C, remove the supernatant, and then wash them once with 1 mL of DNase/RNase-free distilled water. Repeat the centrifugation and discard the supernatant.
NOTE: During the experiment, keep the solution of silk on ice to avoid temperature-induced gelation. - Gradually treat the particles with methanol to induce β-sheet formation in silk protein structure. First, add 0.5 mL of DNase/RNase distilled water, vortex the microcentrifuge tube, then add 0.5 mL of 100% methanol. Gently shake the particles on the thermomixer at 10 °C for 5 min. Collect the particles by centrifugation at 0.2 x g for 1 min. Remove the supernatant.
- Treat the particles with methanol to promote the formation of β-sheets and ensure strong physical adsorption of the silk layer. Add 1 mL of 100% methanol. Gently shake the particles on the thermomixer at 750 rpm for 10 min at 10 °C.
- Collect the particles by centrifugation at 0.2 x g for 1 min at 4 °C and wash them twice with 1 mL of DNase/RNase-free distilled water each time, discarding the supernatant and gently vortexing before the next centrifugation.
- Repeat steps 4.5-4.8 20 times to obtain silk multilayered core-shell structures. For the last deposition step, use fluorescently labeled silk from step 3.4 (Silk-DyLight550, 1 mL).
- Perform the last washing step and keep the microparticles in 1 mL of DNase/RNase-free distilled water at ambient conditions.
NOTE: To avoid aggregation of particles during deposition of silk layers, perform a visual inspection of the particles suspension and pipette it up and down using a 1 mL pipette tip to promote homogeneous particle distribution. - Calculate the number of DNA plasmid copies encapsulated in each microcapsule, NDNA using Equation 1:
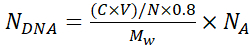 (1)
(1)
Where N = 6.769 × 1011 - the number of SiO2 cores used for encapsulation. Calculate it from a standard curve for known concentrations of silica particles using serial dilutions and absorption A320 at λ = 320 nm;
C- initial concentration of DNA used for adsorption
V- the volume of DNA used for adsorption
0.8- DNA adsorption efficiency on the cores
Mw- Molecular weight of DNA plasmid
NA- Avogadro's number (6.022 × 1023)
5. Dissolution of cores to obtain silk microcapsules.
- Prepare 8% hydrofluoric acid (HF) solution, pH 5.5, by diluting stock solution (48%) with distilled water. Acquire a 50 mL centrifuge tube. Carefully pipette 5 mL of HF and add 25 mL of distilled water to obtain 8% HF solution.
CAUTION: HF is a highly corrosive acid and may cause severe burns to the tissues. Extreme caution must be taken during the handling and use of HF for the experiments. Adhere to Standard Operating Procedure (SOP) for the proper usage and handling of HF developed by the organization to avoid undesirable spillage accidents. Do not use glass containers to dilute HF acid. Use the chemical hood to perform this step of the protocol. - Dissolve SiO2 cores by adding 1.5 mL of 8% HF solution to pelleted core-shell microparticles from step 4.10. Gently vortex and let cores dissolve overnight at ambient conditions with gentle shaking at 450 rpm.
NOTE: To avoid spillage of HF, use grafting tape to seal the microcentrifuge tube. Use a chemical hood to perform this step of the protocol. - Prepare a 2 L glass beaker filled with 2 L of deionized water. Transfer microcapsules solution to dialysis devices (50 kDa MWCO) and dialyze them against deionized water with a repeated change of the water every 3 h for the next 3 days.
NOTE: Collect the supernatant during the first three exchanges of water and discard the solution according to the established protocol for hazardous waste materials. - Use a 1 mL pipette to transfer the suspension from dialysis devices into new 2 mL microcentrifuge tubes to collect the microcapsules.
NOTE: Store the aqueous solutions of microcapsules at ambient conditions for several years.
6. Imaging of silk fibroin microcapsules using confocal laser scanning microscope (CLSM).
- Perform DNA staining using a DNA dye.
- Transfer 300 µL of hollow silk fibroin microcapsules into a fresh 1 mL microcentrifuge tube. Add 500 µL of RNase/DNase-free distilled water.
- Add 5 µL of the DNA staining dye, briefly vortex, and incubate at RT for 2 h protected from light.
- Perform four washing steps by centrifugation at 0.1 x g for 20 min at 4 °C each time, carefully removing 400 µL of the supernatant and replenishing it with 400 µL of RNase/DNase-free distilled water.
- Perform the imaging of silk capsules on inverted confocal systems equipped with three major lasers (405 nm, 488 nm, 561 nm) using 100x oil-immersion objective (NA 1.49). Transfer 100 µL of the capsule sample into a single well of 8-well chambered glass slides, allow for capsules to sediment for 20-30 min prior to imaging.
NOTE: Dyes are very sensitive to photobleaching. Protect the samples by covering the slides with aluminum foil.
7. Estimation of the permeability of hollow microcapsules using molecular weight cut-off (MWCO) method.
- Prepare 2 mL each of FITC-labeled dextran fluorophore solutions (20 µM, diH2O) of different Mw (4 kDa, 20 kDa, 40 kDa, 70 kDa, 150 kDa, 250 kDa, 500 kDa, and 2 MDa).
- Pipette 100 µL of the suspension of capsules into a single well of a chambered glass slide. Analyze each microcapsule design (concentration of PEI, loading number of DNA plasmids, the concentration of silk fibroin, and the number of layers) separately.
- To each well, add 300 µL of the specific fluorophore solution starting from the lowest Mw up to the highest, so each well will correspond to the specific fluorophore solution. Mix by pipetting up and down and let the mixture incubate for 1 h at RT until the diffusion of fluorophore solutions reaches equilibrium.
- Transfer the slide to a confocal laser scanning microscope (CLSM) and image each well using 100x oil-immersion objective at excitation λ = 488 nm.
- Identify the area of interest by adjusting the focal plane to make sure that the capsules appear in the form of circles of the largest diameter. This typically happens when viewing the samples closer to the bottom of the well when capsules sediment due to gravity.
- Collect several images of microcapsule samples by moving the slide in XY direction. Capture images to account for up to 100-150 capsules for each sample.
- Use ImageJ software to analyze the permeability of the capsule's membrane in each Mw fluorophore solution by comparing fluorescence intensities inside and outside of the capsules. For that, draw a region of interest (ROI) in the form of a circle to outline the circumference of the capsule and click Analyze/ Measure to measure the fluorescence intensity inside. Tabulate the data into a spreadsheet. Perform this operation for each microcapsule for a total of 200-300 capsules.
- Assess the outside fluorescence intensity in the same way by outlining the ROI and measuring the intensity away from the capsules. Perform 3-5 measurements for statistical analysis.
- To perform statistical analysis, compare the fluorescence intensities inside and outside of the capsules using paired t-test (p < 0.05).
- Use the conversion Table 2 to estimate the permeability of microcapsules based on the hydrodynamic radii for FITC-Dextran with variable Mw.
8. In vitro activation of synthetic theophylline riboswitch in silk microcapsules
- Prepare 1 mL of theophylline stock solution (100 mM, DMSO). Prepare E. coli S30 extract system for circular DNA by thawing the components on ice for 40 min.
- Obtain a 0.5 mL DNase/RNase-free microcentrifuge tube. Perform in vitro transcription/translation reaction, combine the cell-free components with a sample of microcapsules in the following order (50 µL total volume): S30 premix without amino acids (20 µL); S30 extract, circular (15 µL); complete amino acids mixture (5 µL); hollow microcapsules containing ThyRS-GFPa1 plasmids from step 4.10 (9 µL); and theophylline, 100 mM DMSO (1 µL).
NOTE: After adding all components, briefly vortex the tube and collect the sample during brief centrifugation at 0.2 × g for a couple of seconds. - Incubate the tube at 30 °C for 4 h and check the fluorescence on a plate reader using excitation at λ = 488 nm and emission for GFP/FITC filter (510 nm ± 20 nm).
- Image the capsules on any LCSM system using 488 nm and 561 nm lasers. Obtain the best quality images using 100x oil immersion objective and 8-well chambered slides.
9. In vitro activation of broccoli aptamer in silk microcapsules
- Prepare 1 mL of stock solution of DFHBI-1T dye (30 µM, diH2O). Prepare the PURE (protein synthesis using recombinant elements) cell-free system reaction kit by thawing the components on ice for 40 min.
- Obtain a 0.5 mL DNase/RNase-free microcentrifuge tube. Perform in vitro transcription reaction by combining cell-free reaction components with a sample of microcapsules in the following order (50 µL total volume): solution A (20 µL); solution B (15 µL); hollow microcapsules containing BrocApt plasmids from step 4.10 (14 µL); and DFHBI-1T dye (1 µL).
NOTE: After adding all components, briefly vortex the tube and collect the sample during brief centrifugation at 0.2 × g for a couple of seconds. - Incubate the tube at 37 °C for 6 h and check the fluorescence on a plate reader using excitation at λex = 470 nm and emission at λem = 510 nm ± 20 nm.
- Image the capsules on any LCSM system using 488 nm and 561 nm lasers. Obtain the best quality images using 100x oil-immersion objective, and 8-well coverglass chambered slides.
Results
Here, the study addresses the functionality of DNA templates encoding different sensor designs (two types of RNA-regulated transcription/translation elements) after encapsulation in silk protein capsules. Microcapsules were prepared via templated Layer-by-Layer (LbL) assembly of the key components: A prime layer, DNA plasmids encoding sensor designs, and silk fibroin biopolymer (Figure 2). Deposition of macromolecules in a layered fashion allows controlling the permeability of the capsule me...
Discussion
Selectively permeable hydrogel microcapsules loaded with various types of DNA-encoded sensor designs can be prepared following this protocol. One of the distinctive features of the LbL approach is the ability to tailor the complexity of microcapsules during the bottom-up assembly, which usually starts with the adsorption of molecular species on sacrificial templates. By carefully adjusting concentrations of the initial components, pH conditions, and the number of layers, microcapsules with different DNA loading parameter...
Disclosures
The views and opinions presented herein are those of the authors and do not necessarily represent the views of DoD or its Components
Acknowledgements
This work was supported by LRIR 16RH3003J grant from the Air Force Office of Scientific Research, as well as the Synthetic Biology for Military Environments Applied Research for the Advancement of S&T Priorities (ARAP) program of the U.S. Office of the Under Secretary of Defense for Research and Engineering.
The plasmid vector sequence for ThyRS (pSALv-RS-GFPa1, 3.4 kb) was generously provided by Dr. J. Gallivan. Silkworm cocoons from Bombyx mori were generously donated by Dr. D.L. Kaplan from Tufts University, MA.
Materials
| Name | Company | Catalog Number | Comments |
| (Z)-4-(3,5-difluoro-4-hydroxybenzylidene)-2-methyl-1-(2,2,2-trifluoroethyl)-1H-imidazol-5(4 H)-one (DFHBI-1T) | Lucerna | DFHBI-1T | |
| 5x T4 DNA Ligase Buffer | ThermoFisher Scientific | 46300-018 | |
| 6x Blue Gel Loading Dye | New England BioLabs | B7021S | |
| 96-well plates, black circular | Corning | 3601 | |
| Agarose | Sigma-Aldrich | A9539 | BioReagent, for molecular biology, low EEO |
| Ampicillin sodium salt | Sigma-Aldrich | A0166 | powder or crystals, BioReagent, suitable for cell culture |
| BlpI restriction enzymes | New England BioLabs | R0585S | |
| Corning Disposable Vacuum Filter/Storage Systems | FisherScientific | 09-761-1 | |
| Dimethyl sulfoxide, DMSO | Sigma-Aldrich | 472301 | ACS reagent, ≥99.9% |
| DNA Plasmid, pET28c-F30-2x Broccoli (5.4 kb), BrocApt. | Addgene | Plasmid #66788 | |
| DyLightTM550 Antibody Labeling kit (Invitrogen) | ThermoFisher Scientific | 84530 | |
| E. coli S30 extract system for circular DNA | Promega | L1020 | |
| Falcon Conical centrifuge tubes, 15 mL | FisherScientific | 14-959-53A | |
| Falcon Conical centrifuge tubes, 50 mL | 14-432-22 | ||
| Fisherbrand Microcentrifuge tubes, 1.5 mL | FisherScientific | 05-408-129 | |
| Hydrofluoric acid, HF | Sigma-Aldrich | 695068 | ACS reagent, 48% |
| Kanamycin sulfate | Sigma-Aldrich | 60615 | mixture of Kanamycin A (main component) and Kanamycin B and C |
| KpnI restriction enzymes | New England BioLabs | R0142S | |
| LB agar plate supplemented with 100 µg/mL ampicillin | Sigma-Aldrich | L5667 | pre-poured agar plates with 100 µg/mL ampicillin |
| LB agar plate supplemented with 50 µg/mL kanamycin | Sigma-Aldrich | L0543 | pre-poured agar plates with 50 µg/mL kanamycin |
| LB broth (Lennox grade) | Sigma-Aldrich | L3022 | |
| Lithium bromide, LiBr | Sigma-Aldrich | 213225 | ReagentPlus, ≥99% |
| Max Efficiency DH5-α competent E. coli strain | ThermoFisher Scientific | 18258012 | |
| Methanol | MilliporeSigma | 322415 | anhydrous, 99.8% |
| MilliQ-water | EMD MilliPore | Milli-Q Reference Water Purification System | |
| MinElute PCR Purification Kit | Qiagen | 28004 | |
| N-(3-Dimethylaminopropyl)-N′-ethylcarbodiimide hydrochloride, EDC | Sigma-Aldrich | E1769 | |
| PBS (phosphate buffered saline) | ThermoFisher Scientific | 10010023 | 1x PBS, pH 7.4 |
| Phusion High-Fidelity DNA Polymerase | New England Biolabs | M0530S | |
| Polyethylenimine, branched | Sigma-Aldrich | 408727 | average Mw ~25,000 |
| PURExpress In Vitro Protein Synthesis Kit | New England BioLabs | E6800S | |
| QIAEX II Gel Extraction Kit | Qiagen | 20021 | |
| QIAprep Spin Miniprep Kit | Qiagen | 27104 | |
| Quick-Load 2-Log DNA Ladder (0.1-10.0 kb) | New England BioLabs | N0469S | |
| SiO? silica microspheres, 4.0 µm | Polysciences, Inc. | 24331-15 | 10% aqueous solution |
| Slide-A-Lyzer G2 Dialysis Cassettes, 3.5K MWCO, 15 mL | ThermoFisher Scientific | 87724 | |
| Sodium carbonate, Na?CO? | Sigma-Aldrich | 222321 | ACS reagent, anhydrous, ≥99.5%, powder |
| Spectrum Spectra/Por Float-A-Lyzer G2 Dialysis Devices | FisherScientific | 08-607-008 | Spectrum G235058 |
| SYBR Safe DNA gel stain | ThermoFisher Scientific | S33102 | |
| T4 DNA Ligase (5 U/µL) | ThermoFisher Scientific | EL0011 | |
| Theophylline | Sigma-Aldrich | T1633 | anhydrous, ≥99%, powder |
| Tris Acetate-EDTA buffer (TAE buffer) | Sigma-Aldrich | T6025 | Contains 40 mM Tris-acetate and 1 mM EDTA, pH 8.3. |
| UltraPure DNase/RNase-Free Distilled Water | FisherScientific | 10-977-023 | |
| ZymoPURE II Plasmid MaxiPrep kit | ZymoResearch | D4202 |
References
- Slomovic, S., Pardee, K., Collins, J. J. Synthetic biology devices for in vitro and in vivo diagnostics. Proceedings of the National Academy of Sciences of the United States of America. 112 (47), 14429-14435 (2015).
- Harbaugh, S. V., Goodson, M. S., Dillon, K., Zabarnick, S., Kelley-Loughnane, N. Riboswitch-based reversible dual-color sensor. ACS Synthetic Biology. 6 (5), 766-781 (2017).
- König, H., Frank, D., Heil, R., Coenen, C. Synthetic genomics and synthetic biology applications between hopes and concerns. Current Genomics. 14 (1), 11-24 (2013).
- Silverman, A. D., Karim, A. S., Jewett, M. C. Cell-free gene expression: An expanded repertoire of applications. Nature Reviews Genetics. 21, 151-170 (2020).
- Chushak, Y., et al. Characterization of synthetic riboswitch in cell-free protein expression systems. RNA Biology. , 1-12 (2021).
- Cole, S. D., et al. Quantification of interlaboratory cell-free protein synthesis variability. ACS Synthetic Biology. 8 (9), 2080-2091 (2019).
- Thavarajah, W., et al. Point-of-use detection of environmental fluoride via a cell-free riboswitch-based biosensor. ACS Synthetic Biology. 9 (1), 10-18 (2020).
- Grӓwe, A., et al. A paper-based, cell-free biosensor system for the detection of heavy metals and date rape drugs. PLoS One. 14 (3), 0210940 (2019).
- Lin, X., et al. Portable environment-signal detection biosensors with cell-free synthetic biosystems. RSC Advances. 10 (64), 39261-39265 (2020).
- Caschera, F., Lee, J. W., Ho, K. K. Y., Liu, A. P., Jewett, M. C. Cell-free compartmentalized protein synthesis inside double emulsion templated liposomes with in vitro synthesized and assembled ribosomes. Chemical Communications. 52 (31), 5467-5469 (2016).
- Niederholtmeyer, H., Chaggan, C., Devaraj, N. K. Communication and quorum sensing in non-living mimics of eukaryotic cells. Nature Communications. 9, 5027 (2018).
- Timin, A. S., Gould, D. J., Sukkhorukov, G. B. Multi-layer microcapsules: Fresh insights and new applications. Expert Opinion on Drug Delivery. 14 (5), 583-587 (2017).
- Bomati, E. K., Haley, J. E., Noel, J. P., Deheyn, D. D. Spectral and structural comparison between bright and dim green fluorescent proteins in Amphioxus. Scientific Reports. 4, 5469 (2014).
- Frey, B., Reischl, U. Amplification of Genomic DNA by PCR. Molecular Diagnosis of Infectious Diseases. Methods in Molecular Medicine. 13, 143-156 (1998).
- Lee, P. Y., Costumbrado, J., Hsu, C. -. Y., Kim, Y. H. Agarose gel electrophoresis for the separation of DNA fragments. Journal of Visualized Experiments. (62), e3923 (2012).
- Zhou, Y., et al. Rapid regeneration and reuse of silica columns from PCR purification and gel extraction kits. Scientific Reports. 8, 12870 (2018).
- Kostylev, M., Otwell, A. E., Richardson, R. E., Suzuki, Y. Cloning should be simple: Escherichia coli DH5α-mediated assembly of multiple DNA fragments with short end homologies. PLoS One. 10 (9), 0137466 (2015).
- Rockwood, D. N., et al. Materials fabrication from Bombyx mori silk fibroin. Nature Protocols. 6 (10), 1612-1631 (2011).
- Drachuk, I., et al. Silk macromolecules with amino acid-Poly(Ethylene Glycol) grafts for controlling layer-by-layer encapsulation and aggregation of recombinant bacterial cells. ACS Nano. 9 (2), 1219-1235 (2015).
- Antipov, A. A., Sukhorukov, G. B. Polyelectrolyte multilayer capsules as vehicles with tunable permeability. Advances in Colloid and Interface Science. 111 (1-2), 49-61 (2004).
- Drachuk, I., Harbaugh, S., Chávez, J. L., Kelley-Loughnane, N. Improving the activity of DNA-encoded sensing elements through confinement in silk microcapsules. ACS Applied Materials & Interfaces. 12 (43), 48329-48339 (2020).
- Melnikov, S., Ben-Shem, A., Garreau de Loubresse, N., Jenner, L., Yusupova, G., Yusupov, M. Structural basis for the inhibition of the eukaryotic ribosome. Nature Structural & Molecular Biology. 19 (6), 560-567 (2012).
- Zhao, S., et al. The future of layer-by-layer assembly: A tribute to ACS Nano associate editor Helmuth Möhwald. ACS Nano. 13 (6), 6151-6169 (2019).
- Main, K. H. S., Provan, J. I., Haynes, P. J., Wells, G., Hartley, J. A., Pyne, A. L. B. Atomic force microscopy-A tool for structural and translational DNA research. APL Bioengineering. 5, 031504 (2021).
- Riera, R., Feiner-Gracia, N., Fornaguera, C., Cascante, A., Borrós, S., Albertazzi, L. Tracking the DNA complexation state of pBAE polyplexes in cells with super resolution microscopy. Nanoscale. 11 (38), 17869-17877 (2019).
- Bilokapic, S., Strauss, M., Halic, M. Cryo-EM of nucleosome core particle interactions in trans. Scientific Reports. 8, 7046 (2018).
- Pritchard, E. M., Dennis, P. B., Omenetto, F., Naik, R. R., Kaplan, D. L. Physical and chemical aspects of stabilization of compounds in silk. Biopolymers. 97 (6), 479-498 (2012).
- Fritz, B. R., Jamil, O. K., Jewett, M. C. Implications of macromolecular crowding and reducing conditions for in vitro ribosome construction. Nucleic Acids Research. 43 (9), 4774-4784 (2015).
- Ge, X., Luo, D., Xu, J. Cell-free protein expression under macromolecular crowding conditions. PLoS One. 6 (12), 28707 (2011).
- Cawte, A. D., Unrau, P. J., Rueda, D. S. Live cell imaging of single RNA molecules with fluorogenic mango II arrays. Nature Communications. 11, 1283 (2020).
- Chen, X., et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs. Nature Biotechnology. 37 (11), 1287-1293 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved