A subscription to JoVE is required to view this content. Sign in or start your free trial.
Methods Article
Continuous Fluorescence-Based Endonuclease-Coupled DNA Methylation Assay to Screen for DNA Methyltransferase Inhibitors
In This Article
Summary
DNA methyltransferases are potential cancer drug targets. Here, a protocol is presented to assess small molecules for DNA methyltransferase inhibition. This assay utilizes an endonuclease to couple DNA methylation to fluorescence generation and allows for enzyme activity to be monitored in real time.
Abstract
DNA methylation, a form of epigenetic gene regulation, is important for normal cellular function. In cells, proteins called DNA methyltransferases (DNMTs) establish and maintain the DNA methylation pattern. Changes to the normal DNA methylation pattern are linked to cancer development and progression, making DNMTs potential cancer drug targets. Thus, identifying and characterizing novel small molecule inhibitors of these enzymes is of great importance. This paper presents a protocol that can be used to screen for DNA methyltransferase inhibitors. The continuous coupled kinetics assay allows for initial velocities of DNA methylation to be determined in the presence and absence of potential small molecule inhibitors. The assay uses the methyl-sensitive endonuclease Gla I to couple methylation of a hemimethylated DNA substrate to fluorescence generation.
This continuous assay allows for enzyme activity to be monitored in real time. Conducting the assay in small volumes in microtiter plates reduces the cost of reagents. Using this assay, a small example screen was conducted for inhibitors of DNMT1, the most abundant DNMT isozyme in humans. The highly substituted anthraquinone natural product, laccaic acid A, is a potent, DNA-competitive inhibitor of DNMT1. Here, we examine three potential small molecule inhibitors — anthraquinones or anthraquinone-like molecules with one to three substituents — at two concentrations to describe the assay protocol. Initial velocities are used to calculate the percent activity observed in the presence of each molecule. One of three compounds examined exhibits concentration-dependent inhibition of DNMT1 activity, indicating that it is a potential inhibitor of DNMT1.
Introduction
DNA methylation is an important epigenetic mark that regulates gene expression and chromatin structure. Methylation occurs predominately in CpG dinucleotides — cytosine followed by guanosine; the methyl group is added to the 5-position of cytosine. Correct DNA methylation patterns, and thus proper gene expression, are needed for appropriate cellular development and function. Many disease states have been associated with changes to the normal methylation pattern1,2,3. For example, there is a link between cancer initiation and progression and alterations to the DNA methylation pattern. Typically, cancer cells exhibit lower overall levels of methylcytosine, which contributes to genome instability. At the same time, the methylcytosine that is present in the genome is concentrated in the promoter regions of tumor suppressor genes, which leads to gene silencing of these important proteins. Notably, epigenetic changes are dynamic and reversible, unlike the DNA mutations associated with tumorigenesis. This has made the proteins involved in epigenetic gene regulation interesting drug targets2,4.
DNA methyltransferases (DNMTs) are the proteins responsible for generating and maintaining DNA methylation patterns. Three catalytically active isozymes, DNMT1, DNMT3a, and DNMT3b, exist in humans. During development and differentiation, the de novo methyltransferases, DNMT3a and DNMT3b, establish methylation patterns. Both enzymes can bind the catalytically inactive DNMT3L protein to form complexes that exhibit increased activity1,5. Following cell division, daughter cells contain hemimethylated DNA — DNA containing methylcytosine in only one strand of the duplex — because the newly synthesized DNA is devoid of methylation marks. The major function of DNMT1 is to methylate this hemimethylated DNA, thus re-establishing the full methylation pattern1,5.
Links between DNMT activity and cancer are well established. Overexpression of DNMT1, either by transcriptional or post-translational mechanisms, is a consequence of several common oncogenic pathways6,7,8,9. Genetic approaches to lower DNMT1 activity using hypomorphic alleles result in decreased tumor formation in Apc(Min)mice10. Antisense oligonucleotides that knockdown DNMT1 inhibit neoplasia in cell culture and mouse tumor models11,12. Thus, inhibiting DNMT1 activity seems like a promising cancer therapy approach. However, the roles the DNMT3 isozymes play are not so straightforward. DNMT3a mutations are found in acute myeloid leukemia13 and myelodysplastic syndrome14. At least one of the identified mutations has been shown to decrease the DNA methylation activity of the enzyme15. However, DNMT3b is overexpressed in breast cancer16 and colorectal cancer17. With the various DNMT isozymes playing different roles in carcinogenesis, identifying isozyme-specific inhibitors will be critical. Not only will these compounds be useful for the development of therapeutics, but isozyme-specific inhibitors would also be an invaluable tool to dissect the role of each DNMT isozyme in cancer etiology.
Several DNMT inhibitors have been reported in the literature. Known DNMT inhibitors can be divided into two classes: nucleoside and non-nucleoside. Nucleoside inhibitors are typically cytidine analogs. These compounds are incorporated into DNA and covalently trap DNMTs. 5-azacytidine and 5-aza-2'-deoxycytidine have been approved for the treatment of myelodysplastic syndrome and acute myeloid leukemia4,18. The high toxicity, low bioavailability, and chemical instability of these compounds present problems. Ongoing work is examining the efficacy of the next generation of nucleoside inhibitors; SGI-110, derived from 5-aza-2'-deoxycytidine, is one example19,20. Nucleoside inhibitors are not isozyme-specific and will inactivate any DNMT isozyme encountered. Therefore, treatment with a nucleoside-demethylating agent results in the depletion of all DNMT isozymes4,18. Non-nucleoside inhibitors do not need to be incorporated into DNA to exert their inhibitory effects. Instead, these molecules bind directly to DNMTs, introducing the possibility for isozyme-specific inhibition. Several non-nucleoside inhibitors have been discovered to date, including SGI-102721, hydralazine22, procainamide23, RG108 and derivatives24, and natural products, (−)-epigallocatechin 3-gallate (EGCG)25 and laccaic acid A26,27. Most of the non-nucleoside inhibitors discovered to date are not isozyme-selective or display weak preferences for one DNMT isozyme. In addition, the potency of these molecules needs to be improved, especially in cells4,18. Thus, there is a need to discover or develop more potent, isozyme-selective DNMT inhibitors.
A hurdle to discovering new small molecule inhibitors of DNMTs is the laborious assays traditionally used to examine DNMT activity28. Assays are usually discontinuous with multiple steps. The enzymatic activity of DNMTs is still routinely assayed using radioactive S-adenosyl methionine (SAM)29,30,31,32,33,34. Non-radioactive assays for DNA methylation have been developed as well. For example, assays utilizing methyl-sensitive restriction endonucleases and electrophoresis to separate the digestion products have been described35,36. These types of discontinuous, multistep assays are not readily amenable to drug discovery. Since the mid-2000s, several DNA methylation assays with a higher throughput have been developed28. A scintillation proximity assay was used to screen for DNMT1 inhibitors37. Another assay utilizing a methyl-sensitive restriction endonuclease was used to screen for DNMT3a inhibitors25,38. While both assays allowed for higher throughput than traditional DNA methylation assays, the assays require multiple steps and do not allow the observation of methylation activity in real time. More recently, a continuous kinetics assay has been described that couples the formation of S-adenosylhomocysteine (SAH), one product of the methylation reaction, to the spectroscopic change at 340 nm associated with NADPH oxidation39. This assay utilizes three coupling enzymes to generate a spectroscopic signal.
We developed a fluorescence-based endonuclease-coupled DNA methylation assay that utilizes a single commercially available coupling enzyme and can generate data in real time (Figure 1). A hairpin oligonucleotide containing three methylcytosines is used as a substrate. The substrate DNA contains a fluorophore on the 5' end and a quencher on the 3' end. Methylation of the hemimethylated CpG site generates the cleavage site for the endonuclease Gla I — fully methylated GCGC. Gla I cleavage of the product oligonucleotide releases the fluorophore from the quencher and generates fluorescence in real time. The assay can be used to examine the activity of any isoform of DNMT; however, higher activity is observed with DNMT1 as this isozyme preferentially methylates hemimethylated DNA1,5. Even more robust activity is observed if the autoinhibitory Replication Foci Targeting Sequence (RFTS) domain is removed from DNMT1. This domain, found in the N-terminal regulatory region, binds to the catalytic site and prevents DNA binding. Removal of the first ~600 amino acids results in a truncated enzyme that is significantly more active than the full-length enzyme (~640-fold increase in kcat/Km)40. This activated form of the enzyme, referred to as RFTS-lacking DNMT1 (amino acids 621–1616), allows for the easier identification of inhibitors due to its increased catalytic power. This paper presents a protocol to utilize RFTS-lacking DNMT1 in assays to screen for potential small molecule inhibitors. Using the endonuclease-coupled continuous assay, the initial velocity is determined in the presence and absence of a few small molecules. Each potential inhibitor is examined at two concentrations to look for concentration-dependent DNMT1 inhibition. The percent activity observed in the presence of the small molecules was calculated in each case.

Figure 1: DNA methylation assay. A hemimethylated hairpin DNA with a fluorophore on the 5' end and a quencher on the 3' end is used as a substrate. DNMT1 catalyzes the transfer of the methyl group from S-adenosylmethionine to the nonmethylated CpG site, generating S-adenosylhomocysteine and fully methylated DNA. The DNA product contains the cleavage site for the endonuclease Gla I, which cleaves fully methylated GCGC sites. Cleavage of the product DNA releases the 5' fluorophore from the 3' quencher, generating fluorescence. Abbreviations: Fl = fluorophore; Q =quencher; DNMT1 = DNA methyltransferase 1; SAM = S-adenosylmethionine; SAH = S-adenosylhomocysteine. Please click here to view a larger version of this figure.
Protocol
1. Prepare assay solutions for the screen
NOTE: The concentrations of substrates used in this assay can be adapted. For RFTS-lacking DNMT1, the experimentally determined Km values for the hairpin DNA substrate and SAM are 1–2 nM and 2 µM, respectively26,40.
- Prepare 600 µL of each assay condition being tested in microcentrifuge tubes on ice.
NOTE: The total volume of assay solution needed depends on the number of replicates being conducted. Each assay condition was run in triplicate for this protocol.- Add ddH2O and 5x methylation buffer (250 mM Tris pH 7.5, 5 mM MgCl2, 500 mM potassium glutamate, 5 mM dithiothreitol (DTT), 25% glycerol) to each tube to achieve a final concentration of 1x buffer. For assays containing 20 µM compound, add 472 µL of ddH2O and 120 µL 5x buffer. For assays containing 50 µM compound, add 470 µL of ddH2O and 120 µL of 5x buffer.
- Add 3.15 µL of 20 mg/mL bovine serum albumin (BSA) to each sample.
- Add 2 µL of 3.15 µM hairpin DNA substrate and 1.33 µL of 4.75 mM SAM to each sample.
NOTE: The concentration of substrates in the assay solution mixture is 1.05x the desired final concentrations. - Add inhibitor to a final concentration of 21 µM (1.26 µL of 10 mM) or 52.5 µM (3.15 µL of 10 mM) to the appropriate samples. Add an equivalent amount of dimethylsulfoxide (DMSO) to a control sample.
NOTE: The concentration of inhibitor used in the screen can be varied. The maximum final DMSO concentration should be ≤5% (v/v). DMSO concentrations up to 5% (v/v) have no effect on the activity observed in the assay26.
- Mix the solutions by vortexing (3,000 rpm) for 3 s. Spin the samples for a few seconds in a tabletop mini centrifuge (1,200 × g) to ensure that the solution is collected at the bottom of the tube.
- Aliquot 95 µL of each assay solution into 6 consecutive wells in a black half-area 96-well plate (Figure 2).
NOTE: These screening assays were performed in two batches. First, 20 µM inhibitor samples (4 total conditions) were examined, followed by the 50 µM inhibitor samples (4 total conditions).
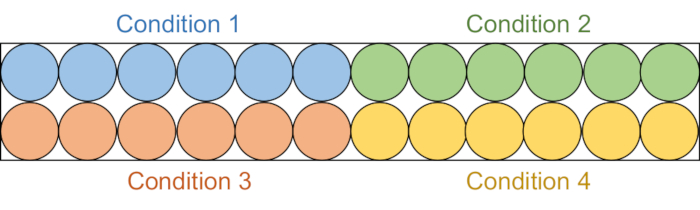
Figure 2: Assay plate setup. Each assay solution is aliquoted into six wells in the black 96-well plate: DMSO control (blue), compound 1 (green), compound 2 (red), and compound 3 (yellow). Both RFTS-lacking DNMT1 and Gla I will be added to three wells. As a control, Gla I alone will be added to the other three wells. Abbreviations: RFTS = Replication Foci Targeting Sequence; DNMT1 = DNA methyltransferase 1; DMSO = dimethylsulfoxide. Please click here to view a larger version of this figure.
2. Prepare enzyme solutions for screen
NOTE: RFTS-lacking DNMT1 can be expressed in E. coli and purified to homogeneity. Expression and purification procedures for RFTS-lacking DNMT1 have been described previously41. The volume of enzyme needed depends on the number of assays being conducted. Here, four different assays are being performed in each set; each assay is completed in triplicate.
- Prepare 75 µL of each enzyme solution in microcentrifuge tubes on ice.
NOTE: Two enzyme solutions will be needed. One solution will contain DNMT1 and Gla I; the other will contain only Gla I.- Add ddH2O and 5x methylation buffer (250 mM Tris pH 7.5, 5 mM MgCl2, 500 mM potassium glutamate, 5 mM DTT, 25% glycerol) to each tube to achieve a final concentration of 1x buffer. For the Gla I enzyme solution, add 15 µL of 5x buffer and 58.8 µL ddH2O. For the DNMT1+Gla I enzyme solution, add 15 µL of 5x buffer and 58.2 µL ddH2O.
- Add 1.2 µL of 10 U/µL Gla I to each solution for a final concentration of 0.16 U/µL.
- Add 0.6 µL of 5 µM RFTS-lacking DNMT1 for a final concentration of 40 nM to the DNMT1+Gla I solution.
- Tap gently to mix the solutions. Spin the samples for a few seconds in a tabletop mini centrifuge (1,200 × g) to ensure that the solution is collected at the bottom of the tube.
- Aliquot 12 µL of each enzyme solution into 6 wells in a conical-bottomed 96-well plate (Figure 3).

Figure 3: Enzyme plate setup. The Gla I (grey) and DNMT1+Gla I (blue) solutions are each aliquoted into six wells in the 96-well plate. Using a multichannel pipet, the enzyme can be added to a row of assay solutions simultaneously. For each assay condition (six wells), three wells will receive DNMT1+Gla I, and three wells will receive Gla I alone. Abbreviation: DNMT1 = DNA methyltransferase 1. Please click here to view a larger version of this figure.
3. Run the assay and analyze the data.
- Preheat the plate reader to 37 °C. Insert the black plate containing the assay solutions. Shake the plate (double orbital at 425 cpm for 5 s) and read the fluorescence with an excitation wavelength of 485 nm and an emission wavelength of 528 nm. Incubate the plate at 37 °C for 5 min.
NOTE: Alternatively, filters with proper wavelength cutoffs can be used for the fluorescence readings. - Remove the assay plate. Add 5 µL of enzyme solution to each well and mix by pipetting up and down.
NOTE: Use multichannel pipets so that a row of samples can be initiated simultaneously. - Insert the plate into the plate reader. Record fluorescence every 53 s for 30 min. Shake the plate (double orbital at 425 cpm for 4 s) before each reading.
- Average triplicate Gla I control assays for each condition. Subtract the average Gla I-containing control reaction trace from the triplicate traces obtained in the presence of RFTS-lacking DNMT1. Determine the average and standard error of the mean for the corrected replicates.
- Plot the average corrected reaction trace and fit the initial linear portion to a line to determine the initial velocity.
- Determine the percent activity by dividing the velocity observed in the presence of a compound by the velocity observed in the DMSO-containing control reaction and multiplying by 100.
4. Additional assay control — sequential addition of enzymes
- Prepare 550 µL of assay solution in a microcentrifuge tube on ice. Add 110 µL of 5x methylation buffer, 3.88 µL of 3.15 µM hairpin DNA substrate, 6.43 µL of 47.5 mM SAM, 3.06 µL of 20 mg/mL BSA, and 426.6 µL of ddH2O.
- Mix the solution by vortexing (3,000 rpm) for 3 s. Spin the samples for a few seconds in a tabletop mini centrifuge (1,200 × g) to ensure that the solution is collected at the bottom of the tube.
- Aliquot 90 µL of the assay solution into 6 consecutive wells in a black half-area 96-well plate.
- Prepare the DNMT1 enzyme solutions on ice. Add 4 µL of 5x methylation buffer, 0.76 µL of 5 µM RFTS-lacking DNMT1, and 15.24 µL of ddH2O to one tube. Add 4 µL of 5x methylation buffer and 16 µL of ddH2O to another tube.
- Tap gently to mix the solutions. Spin the samples for a few seconds in a tabletop mini centrifuge (1,200 × g) to ensure that the solution is collected at the bottom of the tube.
- Add 5 µL of the DNMT1-containing solution to three wells in the black half-area 96-well plate. Add 5 µL of the buffer control solution to the other three wells.
- Place the microtiter plate into the plate reader preheated to 37 °C. Shake the plate (double orbital at 425 cpm for 3 s) and read the fluorescence with an excitation wavelength of 485 nm and an emission wavelength of 528 nm. Repeat the shake and read every 60 s for 30 min.
- While the assay plate is in the plate reader, prepare the Gla I solution on ice. Add 8 µL of 5x methylation buffer, 0.64 µL of 10 U/µL Gla I, and 31.4 µL of ddH2O to a microcentrifuge tube. Tap gently to mix the solution. Spin the sample for a few seconds in a tabletop mini centrifuge (1,200 × g) to ensure that the solution is collected at the bottom of the tube.
- Aliquot 6 µL of the Gla I solution into 6 wells in a conical-bottomed 96-well plate.
- Following the initial 30 min read, remove the black plate from the plate reader. Add 5 µL of Gla I solution to all 6 wells and mix by pipetting up and down.
NOTE: Use a multichannel pipet so that all wells can be initiated simultaneously. - Place the black plate back into the plate reader. Record fluorescence every 35 s for 35 min. Shake the plate (double orbital at 425 cpm for 3 s) before each reading.
Results
Active DNMT1 is a prerequisite for this analysis. RFTS-lacking DNMT1 was expressed in E.coli and purified to homogeneity following previously published procedures41. To ensure the purified enzyme was active, a discontinuous endonuclease-coupled assay was used to examine DNA methylation activity36. This assay utilizes a 32 base pair duplex DNA with a single hemimethylated CpG positioned in a Sau3A1 cleavage site. Sau3A1 can cleave the hemimethylated substrate DNA; h...
Discussion
To identify and characterize inhibitors of DNA methyltransferases, the activity of the enzyme must be measured. Several methods for examining DNA methyltransferase activity exist. Activity is commonly monitored using radioactivity; transfer of the labeled methyl group of SAM can be quantified29,30,31,32,33,34. Gel-based assay...
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
The authors thank Bucknell University and the Department of Chemistry for their support of this work.
Materials
| Name | Company | Catalog Number | Comments |
| 96-well Half Area Black Flat Bottom Polystyrene Not Treated Microplate | Corning | 3694 | |
| 96-Well Polystyrene Conical Bottom Plates | ThermoFisher | 249570 | |
| Bovine Serum Albumin | NEB | B9000S | |
| compound 1 | ChemBridge | 5812086 | screening compound; resuspended in DMSO to 10 mM |
| compound 2 | ChemBridge | 6722175 | screening compound; resuspended in DMSO to 10 mM |
| compound 3 | ChemBridge | 5249376 | screening compound; resuspended in DMSO to 10 mM |
| Dithiothreitol | Sigma | D0632 | |
| Gla I | SibEnzyme | E494 | methyl-sensitive endonuclease |
| Glycerol | RPI | G22025 | |
| Magnesium Chloride | Sigma | M0250 | |
| Oligonucleotide (5'-FAM-CCTATGCGmCATCAGTTTTCTGATGmCGmCATAGG-3'-Iowa Black Quencher) | IDT | custom synthesized | internally quenched hairpin DNA (substrate) |
| Potassium Glutamate | Sigma | G1501 | |
| S-adenosylmethionine | Sigma | A4377 | methyl-donating co-factor (substrate) |
| Tris Base | RPI | T60040 |
References
- Jurkowska, R. Z., Jurkowski, T. P., Jeltsch, A. Structure and function of mammalian DNA methyltransferases. Chembiochem. 12 (2), 206-222 (2011).
- Hamidi, T., Singh, A. K., Chen, T. Genetic alterations of DNA methylation machinery in human diseases. Epigenomics. 7 (2), 247-265 (2015).
- Norvil, A. B., Saha, D., Dar, M. S., Gowher, H. Effect of disease-associated germline mutations on structure function relationship of DNA methyltransferases. Genes. 10 (5), 369 (2019).
- Foulks, J. M., et al. Epigenetic drug discovery: targeting DNA methyltransferases. Journal of Biomolecular Screening. 17 (1), 2-17 (2012).
- Goll, M. G., Bestor, T. H. Eukaryotic cytosine methyltransferases. Annual Review of Biochemistry. 74, 481-514 (2005).
- Bigey, P., Ramchandani, S., Theberge, J., Araujo, F. D., Szyf, M. Transcriptional regulation of the human DNA Methyltransferase (dnmt1) gene. Gene. 242 (1-2), 407-418 (2000).
- Detich, N., Ramchandani, S., Szyf, M. A conserved 3'-untranslated element mediates growth regulation of DNA methyltransferase 1 and inhibits its transforming activity. Journal of Biological Chemistry. 276 (27), 24881-24890 (2001).
- MacLeod, A. R., Rouleau, J., Szyf, M. Regulation of DNA methylation by the Ras signaling pathway. Journal of Biological Chemistry. 270 (19), 11327-11337 (1995).
- Slack, A., Cervoni, N., Pinard, M., Szyf, M. DNA methyltransferase is a downstream effector of cellular transformation triggered by simian virus 40 large T antigen. Journal of Biological Chemistry. 274 (15), 10105-10112 (1999).
- Eads, C. A., Nickel, A. E., Laird, P. W. Complete genetic suppression of polyp formation and reduction of CpG-island hypermethylation in Apc(Min/+) Dnmt1-hypomorphic mice. Cancer Research. 62, 1296-1299 (2002).
- MacLeod, A. R., Szyf, M. Expression of antisense to DNA methyltransferase mRNA induces DNA demethylation and inhibits tumorigenesis. Journal of Biological Chemistry. 270 (14), 8037-8043 (1995).
- Ramchandani, S., MacLeod, A. R., Pinard, M., von Hofe, E., Szyf, M. Inhibition of tumorigenesis by a cytosine-DNA, methyltransferase, antisense oligodeoxynucleotide. Proceedings of the National Academy Sciences of the United States of America. 94 (2), 684-689 (1997).
- Ley, T. J., et al. DNMT3A mutations in acute myeloid leukemia. New England Journal of Medicine. 363, 2424-2433 (2010).
- Walter, M. J., et al. Recurrent DNMT3A mutations in patients with myelodysplastic syndromes. Leukemia. 25 (7), 1153-1158 (2011).
- Russler-Germain, D. A., et al. The R882H DNMT3A mutation associated with AML dominantly inhibits wild-type DNMT3A by blocking its ability to form active tetramers. Cancer Cell. 25 (4), 442-454 (2014).
- Roll, J. D., Rivenbark, A. G., Jones, W. D., Coleman, W. B. DNMT3b overexpression contributes to a hypermethylator phenotype in human breast cancer cell lines. Molecular Cancer. 7, 15 (2008).
- Nosho, K., et al. DNMT3B expression might contribute to CpG island methylator phenotype in colorectal cancer. Clinical Cancer Research. 15 (11), 3663-3671 (2009).
- Erdmann, A., Halby, L., Fahy, J., Arimondo, P. B. Targeting DNA methylation with small molecules: what's next. Journal of Medicinal Chemistry. 58 (6), 2569-2583 (2015).
- Chuang, J. C., et al. S110, a 5-Aza-2'-deoxycytidine-containing dinucleotide, is an effective DNA methylation inhibitor in vivo and can reduce tumor growth. Molecular Cancer Therapeutics. 9 (5), 1443-1450 (2010).
- Issa, J. J., et al. Safety and tolerability of guadecitabine (SGI-110) in patients with myelodysplastic syndrome and acute myeloid leukaemia: a multicentre, randomised, dose-escalation phase 1 study. Lancet Oncology. 16 (9), 1099-1110 (2015).
- Datta, J., et al. A new class of quinoline-based DNA hypomethylating agents reactivates tumor suppressor genes by blocking DNA methyltransferase 1 activity and inducing its degradation. Cancer Research. 69 (10), 4277-4285 (2009).
- Zambrano, P., et al. A phase I study of hydralazine to demethylate and reactivate the expression of tumor suppressor genes. BMC Cancer. 5, 44 (2005).
- Lee, B. H., Yegnasubramanian, S., Lin, X., Nelson, W. G. Procainamide is a specific inhibitor of DNA methyltransferase 1. Journal of Biological Chemistry. 280 (49), 40749-40756 (2005).
- Asgatay, S., et al. Synthesis and evaluation of analogues of N-phthaloyl-l-tryptophan (RG108) as inhibitors of DNA methyltransferase 1. Journal of Medicinal Chemstry. 57 (2), 421-434 (2014).
- Ceccaldi, A., et al. C5-DNA methyltransferase inhibitors: from screening to effects on zebrafish embryo development. Chembiochem. 12 (9), 1337-1345 (2011).
- Fagan, R. L., Wu, M., Chédin, F., Brenner, C. An ultrasensitive high throughput screen for DNA methyltransferase 1-targeted molecular probes. PLoS One. 8 (11), 78752 (2013).
- Fagan, R. L., Cryderman, D. E., Kopelovich, L., Wallrath, L. L., Brenner, C. Laccaic acid A is a direct, DNA-competitive inhibitor of DNA methyltransferase 1. Journal of Biological Chemistry. 288 (33), 23858-23867 (2013).
- Eglen, R. M., Reisine, T. Screening for compounds that modulate epigenetic regulation of the transcriptome: an overview. Journal of Biomolecular Screening. 16 (10), 1137-1152 (2011).
- Holz-Schietinger, C., Matje, D. M., Reich, N. O. Mutations in DNA methyltransferase (DNMT3A) observed in acute myeloid leukemia patients disrupt processive methylation. Journal of Biological Chemistry. 287 (37), 30941-30951 (2012).
- Norvil, A. B., et al. Dnmt3b methylates DNA by a noncooperative mechanism, and its activity is unaffected by manipulations at the predicted dimer interface. Biochemistry. 57 (29), 4312-4324 (2018).
- Bashtrykov, P., Ragozin, S., Jeltsch, A. Mechanistic details of the DNA recognition by the Dnmt1 DNA methyltransferase. FEBS Letters. 586 (13), 1821-1823 (2012).
- Bashtrykov, P., et al. Targeted mutagenesis results in an activation of DNA methyltransferase 1 and confirms an autoinhibitory role of its RFTS domain. Chembiochem. 15 (5), 743-748 (2014).
- Berkyurek, A. C., et al. The DNA methyltransferase Dnmt1 directly interacts with the SET and RING finger-associated (SRA) domain of the multifunctional protein Uhrf1 to facilitate accession of the catalytic center to hemi-methylated DNA. Journal of Biological Chemistry. 289 (1), 379-386 (2014).
- Kanada, K., Takeshita, K., Suetake, I., Tajima, S., Nakagawa, A. Conserved threonine 1505 in the catalytic domain stabilizes mouse DNA methyltransferase 1. Journal of Biochemistry. 162 (4), 271-278 (2017).
- Bashtrykov, P., et al. Specificity of Dnmt1 for methylation of hemimethylated CpG sites resides in its catalytic domain. Chemistry & Biology. 19 (5), 572-578 (2012).
- Dolen, E. K., McGinnis, J. H., Tavory, R. N., Weiss, J. A., Switzer, R. L. Disease-associated mutations G589A and V590F relieve replication focus targeting sequence-mediated autoinhibition of DNA methyltransferase 1. Biochemistry. 58 (51), 5151-5159 (2019).
- Kilgore, J. A., et al. Identification of DNMT1 selective antagonists using a novel scintillation proximity assay. Journal of Biological Chemistry. 288 (27), 19673-19684 (2013).
- Ceccaldi, A., et al. Identification of novel inhibitors of DNA methylation by screening of a chemical library. ACS Chemical Biology. 8 (3), 543-548 (2013).
- Duchin, S., Vershinin, Z., Levy, D., Aharoni, A. A continuous kinetic assay for protein and DNA methyltransferase enzymatic activities. Epigenetics & Chromatin. 8, 56 (2015).
- Syeda, F., et al. The replication focus targeting sequence (RFTS) domain is a DNA-competitive inhibitor of Dnmt1. Journal of Biological Chemistry. 286 (17), 15344-15351 (2011).
- Switzer, R. L., Medrano, J., Reedel, D. A., Weiss, J. Substituted anthraquinones represent a potential scaffold for DNA methyltransferase 1-specific inhibitors. PLoS One. 14 (7), 0219830 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved