Method Article
Protein Engineering by Yeast Surface Display
In This Article
Summary
This protocol describes the essential steps for conducting yeast surface display selection campaigns to enrich protein variants binding to an antigen of interest.
Abstract
Protein engineering enables the improvement of existing functions of a given protein or the generation of novel functions. One of the most widely used and versatile tools in the protein engineering field is yeast surface display, where a pool of randomized proteins is expressed on the surface of yeast. The linkage of phenotype (e.g., binding of the yeast-displayed protein to the antigen of interest) and genotype (the plasmid encoding for the protein variant) enables selection of this library for desired properties and subsequent sequencing of enriched variants. By combining magnetic bead selection with flow cytometric sorting, protein variants with enhanced binding to a target antigen can be selected and enriched. Notably, in addition to affinity maturation, binding to a target can also be achieved without any initial binding affinity. Here, we provide a step-by-step protocol that covers all essential parts of a yeast surface display selection campaign and gives examples of typical yeast surface display results. We demonstrate that yeast surface display is a broadly applicable and robust method that can be established in any molecular biology laboratory with access to flow cytometry.
Introduction
Yeast surface display is one of the key technologies in the field of protein engineering. It enables the selection of protein variants with desired properties such as improved affinity or stability. First introduced in 19971, it is one of the most commonly used display technologies besides phage display2,3, ribosome display4, and mammalian cell display5,6,7. The protein of interest (POI) is displayed on the surface of yeast cells by fusing it to anchor proteins. A range of different anchor proteins is available, and most commonly, the POI is fused to the C-terminus of the yeast agglutinin mating protein Aga2p1,8. Additionally, the POI is typically flanked by two tags, such as a hemagglutinin tag (HA-tag) and c-myc tag, which enables detection of the display level by using fluorescently labeled antibodies and flow cytometry (Figure 1A). Typical yeast selection campaigns involve a combination of magnetic bead selections and flow cytometric sorting. The bead selections enable the handling of large cell numbers and enrichment of protein variants binding to the target antigen also with low affinities since multivalent interactions with the antigen-loaded beads lead to avidity effects and, therefore, prevent the loss of low-affinity variants (Figure 1B). Flow cytometric analysis and selection offer the advantage of visualizing the binding of the displayed POI variants to the labeled antigen. Consequently, the binding populations can be sorted and cultivated, leading to the enrichment of protein variants with desired characteristics throughout several sorting rounds. Moreover, additional rounds of random mutagenesis can be performed to further increase the diversity and, hence, the likelihood of finding additional mutations that contribute to the affinity and/or stability of the protein.
Yeast surface display presents certain advantages, such as (a) eukaryotic expression machinery, enabling oxidative protein folding as well as eukaryotic post-translational modifications (such as N-glycosylation), (b) expression normalization due to the detection of the two peptide tags flanking the protein, (c) visual inspection of the selection progress by flow cytometry (e.g., percentage of binding cells and binding intensity) and (d) the possibility of analyzing individual protein mutants on yeast (e.g., analyzing thermostability as well as affinity), presenting a time-saving alternative to laborious protein expression and purification9. In fact, both affinities (KD values) as well as stabilities (T50 values) of yeast surface displayed proteins have shown good correlations with data obtained using biophysical methods and soluble proteins9,10,11,12. Yeast surface display has been employed for the engineering of a variety of proteins, e.g., antibody fragments13,14,15,16, the 10th type III fibronectin domain17,18, rcSso7d19,20, or knottins21. Similarly, extensive research has been undertaken to optimize yeast library designs by altering the randomized positions as well as amino acid codon usage17,22,23. Yeast surface display has been proven successful for the engineering of stability14,15,24,25, affinity18,26,27, enzymatic activity28,29,30,31, and protein expression32. Additionally, more sophisticated applications like conditional binding in the presence or absence of a small molecule were accomplished using yeast surface display20.
In this protocol, we describe all essential steps for a selection campaign with yeast surface display with the example of the G4 library (based on the 10th type III fibronectin domain, Fn3) selected against the antigen human retinol-binding protein 4 (hRBP4) in the presence of the small molecule A112020. This selection was conducted to yield a protein-protein interaction which is dependent on a small molecule that can be used as a molecular switch. Of note, while alternative approaches are possible with yeast surface display, typical yeast selections usually aim for binding to a target antigen without any previous binding affinity. We cover all steps of a yeast selection campaign, involving the cultivation of a yeast library, bead selections, flow cytometric sorting, and affinity maturation by error-prone PCR (epPCR). Therefore, this protocol complements previous yeast surface display protocols33,34 and can be used as a basis for yeast surface display selections (Figure 1) with any given yeast library and target antigen of choice.
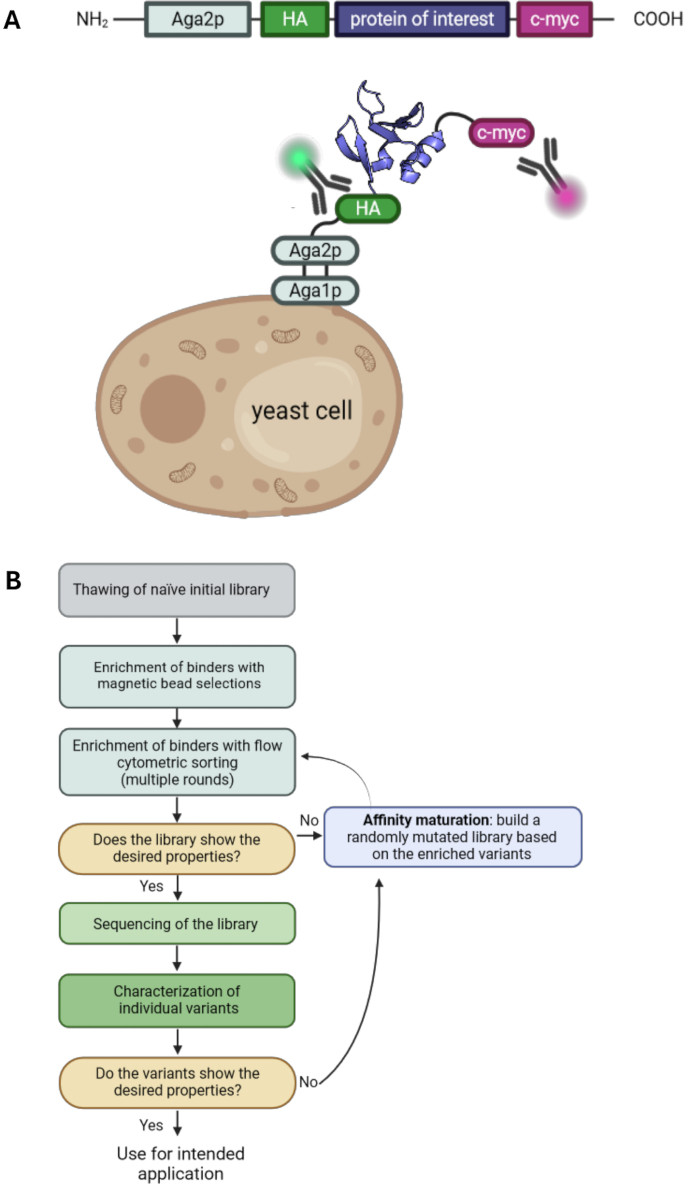
Figure 1: Principle of yeast surface display and a typical workflow for yeast surface display selections. (A) The POI is cloned into a yeast surface display vector and typically flanked by an N-terminal HA- and a C-terminal c-myc-tag. The construct is fused to the yeast mating protein Aga2p for display on the surface. The protein depicted is the engineered binder "RS3" from PDB ID: 6QBA20. (B) Flowchart illustrating a typical workflow for yeast surface display selection campaigns, which combine enrichment of protein variants with desired properties by bead selections and flow cytometric sorting, as well as epPCR for affinity maturation. Please click here to view a larger version of this figure.
Protocol
1. Thawing and cultivation of yeast libraries
- Thaw sufficient aliquots of the frozen yeast library at RT and dilute immediately in SD-CAA (see Table 1) to an OD600 of approximately 1 (this corresponds to 1 x 107 cells/mL).
NOTE: Make sure to cover the diversity of the library by at least 10x. Thus, if the library size accounts for 2.5 x 108 clones (this is the diversity of the G4 library used in this example), inoculate a minimum of 2.5 x 109 cells in 250 mL of SD-CAA to reach an OD600 of 1. This protocol can also be started from liquid cultures (yeast cells freshly transformed with a library). - Assess the number of viable cells.
- Take a 100 µL aliquot from this yeast suspension (at OD600 of 1) and perform a 1:10 dilution series with 900 µL of SD-CAA medium in microcentrifuge tubes.
- Prepare dilutions up to 10-5 and plate 100 µL of dilutions 10-3, 10-4, and 10-5 on SD-CAA plates (Table 1). Incubate the plates at 30 °C for 2-3 days. Ensure that the number of viable cells after thawing exceeds the library diversity by at least 10-fold.
- Meanwhile, continue with the remaining cell suspension and cultivate the yeast culture containing the G4 library in glass flasks overnight at 30 °C while shaking (180 rpm). After the overnight incubation, continue with section 2 (Induction of protein expression on the yeast surface).
- After 2-3 days of incubation, count the colonies on the SD-CAA plates by visual inspection. When calculating the final number of viable cells after thawing, consider the dilution factor and the ratio of the plated (i.e., 100 µL) vs. the original culture volume (i.e., 250 mL).
Viable cells/mL = number of colonies on plate × dilution factor
Viable cells (total) = (viable cells/mL) × culture volume
| Medium/buffer | Component | Concentration [g/L] | Comments/Description | |||
| SD-CAA | D-glucose | 20 | Dissolve all media components in 1000 mL ddH2O and sterile filtrate with disposable 0.22 µm sterile filters. | |||
| Yeast nitrogen base | 6.7 | |||||
| Casmino acids | 5 | |||||
| Citric acid monohydrate | 7.4 | |||||
| Tri-sodium citrate dihydrate | 10.83 | |||||
| SG-CAA | D-galactose | 20 | Dissolve all media components in 1000 mL ddH2O and sterile filtrate with disposable 0.22 µm sterile filters. | |||
| D-glucose | 2 | |||||
| Yeast nitrogen base | 6.7 | |||||
| Casamino acids | 5 | |||||
| di-sodium hydrogen phosphate heptahydrate | 10.2 | |||||
| Sodium dihydrogen phosphate monohydrate | 8.56 | |||||
| SD-CAA plates | Sorbitol | 182 | Dissolve sorbitol, di-sodium hydrogen phosphate heptahydrate, sodium dihydrogen phosphate monohydrate and agar-agar in 900 mL ddH2O and autoclave. Dissolve and sterile filtrate the remaining components in 100 mL ddH2O and add when the autoclaved media is lukewarm. | |||
| di-sodium hydrogen phosphate heptahydrate | 10.2 | |||||
| Sodium dihydrogen phosphate monohydrate | 7.44 | |||||
| Agar-agar | 15 | |||||
| D-glucose | 20 | |||||
| Yeast nitrogen base | 6.7 | |||||
| Casamino acids | 5 | |||||
| YPD | Peptone | 20 | Prepare a 10x D-glucose stock (200 g/L) and sterile filtrate with disposable 0.22 µm sterile filters. Dissolve peptone and yeast extract in 900 mL ddH2O and autoclave. When lukewarm, add 100 mL 10x D-glucose. | |||
| Yeast extract | 10 | |||||
| D-glucose | 20 | |||||
| YPD plates | Peptone | 20 | Prepare a 10x D-glucose stock (200 g/L) and sterile filtrate with disposable 0.22 µm sterile filters. Dissolve peptone, yeast extract and agar-agar in 900 mL ddH2O and autoclave. When lukewarm, add 100 mL 10x D-glucose. | |||
| Yeast extract | 10 | |||||
| D-glucose | 20 | |||||
| Agar-agar | 15 | |||||
| PBSA | BSA | 1 | Dissolve BSA in PBS and sterile filtrate with disposable 0.22 µm sterile filters. | |||
Table 1: Media and buffer composition.
2. Induction of protein expression on the yeast surface
- After overnight incubation, the yeast cultures typically reach an OD600 between 2-20. Measure the OD600 with a photometer to determine the density of the yeast culture.
NOTE: OD measurements are only linear in the range of 0.1-1. Therefore, perform dilutions (typically 1:20 and 1:50) in SD-CAA to determine the OD600.- Dilute the cells in SD-CAA to an OD600 of 1 and ensure that the diversity is covered 10x (i.e., 2.5 x 109 cells are used for the dilution of the G4 library). Continue incubation at 30 °C while shaking to allow the cells to reach an exponential growth phase.
- Prepare glycerol stocks (covering the diversity 50x) of yeast libraries (in SD-CAA containing 15% glycerol) from the overnight culture. For this purpose, centrifuge the required number of yeast cells (2000 x g, 3 min, 4 °C), discard the supernatant, and resuspend the cell pellet in 1 mL of SD-CAA medium supplemented with 15% glycerol. Immediately freeze glycerol stocks at -80 °C.
NOTE: The viability of yeast cells is increased if yeast cultures are grown to a stationary phase before freezing. In later selection rounds, when the diversity of the libraries is lower, it typically is sufficient to use 700 µL of the yeast culture and mix it with 300 µL of SD-CAA supplemented with 50% glycerol. Here, yeast cultures typically show OD600 values of 15 in the stationary phase, but the actual value may depend on the photometer in use. Thus, the stationary phase can be determined by repeated OD600 measurements. If the OD600 value does not increase over an incubation period of 2 h, a stationary phase is reached.
- Centrifuge an appropriate volume of cells to cover the diversity 10x at 2000 x g for 5 min, 4-6 h after the dilution of the yeast culture (typical OD600 values are between 4-6), and discard the supernatant.
- Resuspend the cell pellet in SG-CAA (see Table 1) to yield an OD600 of 1 and transfer it to a fresh glass flask.
NOTE: Resuspension in a galactose-containing medium induces expression of the protein on the surface by the galactose-inducible promotor. - Incubate yeast cultures overnight at 20 °C while shaking at 180 rpm to induce surface expression of the POI. Following overnight induction, yeast cultures will typically reach OD600 values between 2-5; if the temperature of induction is raised to 37 °C, expect OD600 values of 4-10.
NOTE: Reducing the temperature to 20°C slows down cell growth and typically yields better display levels. However, induction at 37 °C might be useful in order to increase the selection pressure towards stabilized mutants35.
3. First bead selection round of yeast libraries (positive selection)
NOTE: A standard bead selection procedure involves 6 steps (Table 2).
| Day | Step | |
| 0 | Overnight culture | |
| 1 | Induction of protein expression on the surface of the yeast cells | |
| 2 | First bead selection with 1 positive selection | |
| 3 | Removal of the beads, passaging, induction of protein expression on the surface of the yeast cells and freezing of the library | |
| 4 | Second bead selection with 3 negative and 1 positive selection | |
| 5 | Removal of the beads and freezing of the library | |
Table 2: Typical timeline for the conduction of bead selections of a yeast library.
- Preparation of the beads
- Prepare the beads (see Table of Materials) for the first bead selection: For each tube, resuspend 10 µL of biotin binder magnetic beads (4 × 105 beads/µL) in 990 µL of PBSA (Table 1) for washing, place the tube on a magnetic rack for 2 min with the lid open. Carefully remove the supernatant. Repeat the washing step with 1 mL PBSA.
NOTE: If multiple tubes of beads with the same biotinylated antigen are needed, they can be prepared in one tube at this step to yield a uniform antigen-bead solution. In this case, the amount of antigen needs to be increased accordingly in the next step. - Resuspend the beads in a total volume of 1 mL of PBSA with 6.7-33 pmoles of biotinylated antigen (here: 6.86 µL of hRBP4 with a concentration of 0.125 mg/mL) in a 1.5 mL microcentrifuge tube.
NOTE: While 6.7 pmoles of antigen are sufficient, 33 pmoles should be used if the antigen is not limiting. If the selection is performed for an antigen bound to a small molecule drug or ligand, as in this case, this drug/ligand must be present during the selection and should be added in this step (here: 5 µM A1120). - Incubate at 4 °C for 2 h with gentle shaking or rotation.
NOTE: Cells can be prepared during this step. - After incubation and before addition to the cells, place the tube on a magnetic rack for 2 min with the lid open. Remove the supernatant and wash the antigen-loaded beads with 1 mL of PBSA as described above. Repeat the washing step.
NOTE: If an antigen bound to a drug or ligand is selected, the washing buffer should contain this drug or ligand in excess (here: 5 µM A1120). - Resuspend the antigen-loaded beads in 50 µL of PBSA (in the presence of the drug or ligand, if applicable).
NOTE: If the same antigen-loaded beads were prepared for multiple tubes, adjust the volume accordingly (for the G4 library, the final volume here is 100 µL yielding beads for two tubes).
- Prepare the beads (see Table of Materials) for the first bead selection: For each tube, resuspend 10 µL of biotin binder magnetic beads (4 × 105 beads/µL) in 990 µL of PBSA (Table 1) for washing, place the tube on a magnetic rack for 2 min with the lid open. Carefully remove the supernatant. Repeat the washing step with 1 mL PBSA.
- Preparation of the yeast cells
- Measure OD600 of the induced yeast culture as described above.
- Pellet sufficient cells to cover 10x the diversity of the library (2000 x g, 10 min, 4 °C). For the G4 library, use 2.5 x 109 cells for centrifugation. Remove the supernatant and wash the cells with 10 mL of PBSA.
- Centrifuge again (2000 x g, 10 min, 4 °C) and resuspend the cells to a concentration of 1.25 x 109 cells/mL in the appropriate volume of PBSA (here: 2 mL) and transfer to a microcentrifuge tube.
NOTE: The volume of PBSA depends on the number of cells. - Centrifuge (2000 x g, 3 min, 4 °C), and resuspend in 950 µL of PBSA.
- Incubation of yeast cells with antigen-loaded beads
- Add 50 µL of antigen-loaded beads to 950 µL of cell suspension and incubate for 2 h at 4 °C while shaking or rotation.
- After the incubation, place the cell-antigen-bead solution on a magnetic rack with the lid open. Pipette liquid from the lid to the tube to subject the whole yeast culture to magnetic selection. Wait for 2 min before carefully discarding unbound cells. Immediately wash with 1 mL of PBSA.
NOTE: If selections are conducted in the presence of a drug/ligand - in this example A1120 - this needs to be present in PBSA. - After the washing step with PBSA, resuspend the cells rapidly in 1 mL of SD-CAA to prevent them from drying out.
- Add 200 mL of SD-CAA and perform serial dilutions as described above. Plate 100 µL on SD-CAA plates and incubate for 2-3 days at 30 °C to determine the new maximum diversity. Here, the diversity of the G4 library after the first bead selection is 1.25 x106 cells.
NOTE: Since the original library was oversampled 10-fold in this selection round, it is reasonable to assume that most clones will be present multiple times in the enriched pool after selection. As a consequence, the real diversity will be lower, and this number should be considered as a maximum value. Usually, the diversity is reduced by at least 100x within this first selection round. Incubate the remaining culture (200 mL) at 30 °C with shaking (180 rpm) for at least 16 h. The protocol can be paused after overnight incubation by placing the cells at 4 °C.
4. Removal of the beads and cultivation prior to the next bead selection round
- Measure OD600 as described above. Pellet sufficient cells to cover the diversity by at least 10x (2000 x g, 3 min, RT). Discard the supernatant and resuspend cells in 1 mL of SD-CAA.
- Transfer to a microcentrifuge tube and place on a magnetic rack for 2 min with the lid open. Transfer the unbound cells to a new microcentrifuge tube and repeat the separation process.
- Dilute the unbound cells in 100 mL of SD-CAA and incubate at 30 °C with shaking at 180 rpm until an OD600 of 3-4 is reached.
NOTE: If the OD600 is lower than 4 before the removal of the beads, the cells can be induced directly afterward without incubation. - When an OD600 of 3-4 is reached, centrifuge a sufficient number of cells to cover the diversity at least 10x (2000 x g, 3 min, RT) and resuspend in SG-CAA to an OD600 of 1 for induction of protein expression and incubate at 20 °C while shaking at 180 rpm overnight.
- When the remaining SD-CAA yeast culture has reached stationary growth phase (OD600 above 15), pellet sufficient cells to cover the diversity 50x (2000 x g, 3 min, RT). Freeze the cells by centrifugation (2000 x g, 3 min), followed by resuspension of the cell pellet in SD-CAA containing 15% glycerol and storage at -80 °C.
5. Second bead selection round with 3 negative and 1 positive selection
- Prepare the cells and the antigen beads as described above. Prepare a solution of bare beads (without adding the antigen) for the negative selections.
NOTE: Since 3 negative selections are performed, the beads can be prepared together for all three rounds. - After washing, resuspend the antigen beads in 50 µL of PBSA and resuspend bare beads in 150 µL of PBSA (if they were prepared for all three negative selections).
- For the first negative selection, add 50 µL of washed bare beads to 950 µL of washed cells in PBSA and incubate for 1.5 h at 4 °C.
- After the incubation, place the tubes containing the bare bead-cell suspensions on a magnetic rack with the lid open. Pipette any liquid in the lid into the tube and wait for 2 min. Transfer the unbound cells to a fresh microcentrifuge tube and add 50 µL of washed bare beads.
- Repeat the procedure for two more rounds of negative selection. Meanwhile, prepare antigen loaded beads for the positive selection.
- After 3 negative selection rounds, add 50 µL of antigen-loaded bead solution to the cells and incubate for 2 h at 4 °C.
- Place the cells containing the antigen-loaded beads on a magnetic rack with the lid open, and pipette any liquid that is in the lid into the tube. Wait for 2 min before discarding unbound cells.
- Perform all remaining steps as described for the first antigen-bead selection. The only difference to the first positive selection (step 3.3.2) is the number of washing steps: Instead of washing once with PBSA, wash the bead-cell suspension twice with PBSA to increase the stringency of selection.
NOTE: Usually, the diversity is again reduced by 10x-100x. The maximum diversity of the G4 library after this second bead selection was 3.45 x 105 cells. As noted above, the actual diversity is expected to be lower due to oversampling of the library.
6. Selection of libraries via flow cytometric sorting
- After overnight induction of surface expression in SG-CAA, measure OD600 as described above. Pellet sufficient cells to cover 10x the diversity (2000 x g, 5 min, RT). Discard the supernatant.
- Resuspend the pellet in PBSA and transfer it to microcentrifuge tubes. Use 3 x 107 cells for staining for each tube. Prepare as many tubes as required, depending on the diversity. Prepare one control tube for staining without the antigen.
NOTE: For example, if 30 x 106 cells should be sorted, pellet 300 x 106 cells to achieve a visible cell pellet and prevent loss of cells in the subsequent steps. Resuspend in 5 mL of PBSA and transfer 3 to 4 aliquots of 500 µL each into microcentrifuge tubes. Prepare approximately 3-fold more cells than the number that should be sorted because a fraction of cells will be lost during the staining process. If the selection is performed in the presence of a drug/ligand, prepare an additional control with the antigen only but without the drug/ligand. - Centrifuge again (2000 x g, 5 min, RT), resuspend the pellet in 200 µL of PBSA containing the antigen (hRBP4 with 5 µM A1120) and incubate for 1 h at 4 °C.
NOTE: All steps from this point on should be performed on ice and with cold reagents. To prevent antibody or antigen depletion, all reagents should be added in stoichiometric excess compared to the total number of yeast-displayed proteins in the tube. Otherwise, there might be the risk of limiting the binding of yeast-displayed variants due to the limited availability of antigens and/or antibodies. Usually, 5 x 104 molecules per yeast cell are used for calculation, and adding at least a 10-fold stochiometric excess of antigen and antibodies is advised. If the antigen is very limited, the excess can be reduced to 5-fold. A more detailed explanation, including examples for calculations to avoid ligand depletion, is provided in a previous protocol9. - After incubation, centrifuge cells (2000 x g, 5 min, 4 °C) and wash with 1 mL of PBSA (if a drug/ligand is present during selections - as A1120 in this case - this needs to be present in the washing buffer).
- Resuspend the cells in 100 µL of cold PBSA containing the antibodies for display staining and detection of the bound antigen and incubate for 30 min at 4 °C.
- Use anti-c-myc antibody (final dilution 1:100) to detect full-length displayed POI. Use anti-HA antibody (final dilution 1:50) to detect the total amount of displayed proteins. Use Penta-His antibody (final dilution 1:20) or streptavidin (final dilution 1:200) to select yeast cells binding to the soluble antigen containing either a 6x HIS tag or biotin, respectively.
- Alternate between the detection reagents between sorting rounds to prevent enrichment of protein variants that bind to the secondary reagent.
NOTE: Different fluorophore combinations can be used, but Alexa Fluor 488 and Alexa Fluor 647 are good options as they do not require compensation in flow cytometry. To prevent antibody depletion, it should be added in stoichiometric excess compared to the total number of yeast-displayed proteins in the tube. Usually, calculations are based on 5 x 104 molecules per yeast cell. A more detailed explanation is provided in previous protocols9.
- After incubation, centrifuge cells (2000 x g, 5 min, 4 °C) and add 1 mL of PBSA (containing the drug/ligand, for the hRBP4-selected G4 library 5 µM of A1120) to the pellet and centrifuge again. Remove most of the supernatant and keep only 20-30 µL to prevent the pellet from drying out.
- Resuspend the pellet in cold PBSA only right before the sorting and sort for the yeast cells displaying Fn3 molecules with the desired properties (i.e., binding to hRBP4 in the presence of A1120).
NOTE: The gating strategy is depicted in Figure 2A. - Sort the cells directly into SD-CAA medium, add more SD-CAA medium after the sort, and incubate at 30 °C with shaking at 180 rpm.
NOTE: In case the volume increases significantly after sorting, a centrifugation step should be performed (2000 x g, 5 min, RT) to remove the excess sheath fluid, and the cells are resuspended in a fresh SD-CAA medium. Write down the number of cells sorted into the new tube, as this becomes the new maximum diversity for the next round of sorting. As noted above, the actual diversity of the library will be lower due to oversampling of the library. - Repeat flow cytometric sorts for several rounds to see an enrichment of binding protein variants in the yeast library (Figure 2B).
NOTE: It can be useful to compare the libraries side-by-side in an analytical flow cytometric experiment. In later selection rounds, lowering the concentration of the antigen can be beneficial in discriminating between lower and higher affinity binders and sorting specifically for the protein variants that show a higher binding signal to the target at low concentrations (Figure 2C). In later selection rounds, it is also possible to sort for increased thermostability. For this purpose, random mutagenesis of the library by epPCR was performed, as described in Section 7. Prepare cells similar to other flow cytometric sorting rounds by induction, determination of OD600, and washing of cells. Place yeast cells in PBSA on ice for 10 min, followed by a 10 min incubation in a thermocycler for the heat shock at an elevated temperature. Afterward, place them again on ice for 10 min and proceed with the staining protocol. Stabilized variants will resist thermal denaturation and, therefore, show maintained binding after the heat incubation. Different temperatures can be tested for heat shock, depending on the initial thermostability of the wildtype protein. Typically, temperatures ranging from 40-80 °C are suitable. However, above 48 °C, the viability of yeast cells will decrease drastically. Therefore, it is necessary to extract the plasmids after sorting the heat-shocked yeast library, as discussed in step 7.1. In this case, EBY100 cells need to be freshly transformed with enriched variants before proceeding to the next sorting round36. A more detailed protocol for increasing the thermostability can be found elsewhere37. If the denaturation of the displayed protein is reversible, this method will not work. Alternatively, induce protein expression at 37 °C instead of 20 °C the day prior to sorting and sort for protein variants still showing efficient full-length display.
7. Affinity maturation with epPCR to introduce random mutations
NOTE: Affinity maturation by using epPCR can either be performed before the first flow cytometric sorting round or in between flow cytometric sorting rounds. For the selection of the G4 library with hRBP4 in the presence of A1120, affinity maturation was conducted before the first round of flow cytometric sorting. This also depends on the library size after bead selection and the binding signal that can be detected with flow cytometry. In particular, in cases where affinities after bead selections are not sufficient to obtain a signal in flow cytometric experiments (because the antigen dissociates quickly during the washing steps), epPCR can generate improved variants that can subsequently be detected and selected via flow cytometry.
- Perform a yeast miniprep DNA extraction from the enriched yeast library using the commercial plasmid miniprep II kit (see Table of Materials) with the following modifications:
- Measure the cell density of overnight culture, dilute to an OD600 of 0.2 in the morning, and incubate at 30 °C with shaking.
- When OD600 reaches 0.6, harvest 3 mL of culture by centrifuging the cells at 600 x g for 2 min.
- Add 200 µL of Solution 1 to the pellet. Add 6 µL of Zymolyase. Resuspend the pellet by flicking or mild vortexing and incubate at 37 °C for 3 h.
- Add 200 µL of Solution 2 and mix gently. Add 400 µL of Solution 3 and mix gently.
- Centrifuge at maximum speed for 3 min.
- Transfer the supernatant to the spin-I column, centrifuge at maximum speed for 30 s, and discard the flowthrough.
- Wash with 550 µL of Wash Buffer containing ethanol and centrifuge at maximum speed for 2 min.
- Transfer the column to a clean microcentrifuge tube and centrifuge at maximum speed for 2 min to remove residual wash buffer.
- Transfer the column to a new microcentrifuge tube, add 10 µL of nuclease-free water, and spin for 1 min at maximum speed to elute the DNA.
- Perform an initial PCR to amplify the template for epPCR.
- Amplify the DNA by PCR using primers annealing on the outside of the insert on the yeast surface display vector.
NOTE: As a lot of genomic yeast DNA is eluted along with the plasmid of interest during the yeast miniprep, this step is recommended to obtain a clean DNA fragment to be used as a template for the epPCR. - For pCTCON2, use primers fwd 5'- GGCTCTGGTGGAGGCGGTAGCGGAGGCGGAG
GGTCGGCTAGC and rev 5'- CTATTACAAGTCC
TCTTCAGAAATAAGCTTTTGTTCGGATCC with the reaction components described in Table 3 and cycling conditions depicted in Table 4. - Purify the PCR with a PCR and DNA cleanup kit. Elute in 20 µL of nuclease-free H2O.
- Amplify the DNA by PCR using primers annealing on the outside of the insert on the yeast surface display vector.
- Perform epPCR to introduce random mutations.
- Perform an epPCR with nucleotide analogs with the indicated ingredients (Table 5) and conditions (Table 6).
- Run the entire epPCR product on a preparative 1% agarose gel at 120 V for 45 min. Include a marker and leave one lane between the epPCR product and the ladder. Excise the desired band corresponding to the expected size of the insert, purify it using a gel extraction kit, and measure the DNA concentration.
- Amplify the randomized DNA by PCR.
- Amplify 50 ng of the DNA with a Q5 polymerase. Set up 2 x 100 µL of PCR reaction for each electroporation (Table 7) with the indicated conditions (Table 8). Use the same primers that were used for the first PCR.
NOTE: Large amounts of DNA are required for electroporation into yeast. - Analyze 5 µL of the PCR product on a 1% agarose gel, run with 120 V for 45 min.
- Amplify 50 ng of the DNA with a Q5 polymerase. Set up 2 x 100 µL of PCR reaction for each electroporation (Table 7) with the indicated conditions (Table 8). Use the same primers that were used for the first PCR.
- Purify the amplified DNA with ethanol precipitation.
- Pool 2 x 100 µL together. Add 20 µL (10% of the volume) of sterile filtered 3 M sodium acetate, pH 5.2.
NOTE: This buffer can be prepared in advance and stored for a few months at RT. - Add 440 µL of 100% ethanol (or at least 2 x volume) and incubate at RT for 2 min.
- Centrifuge at 20,000 x g for 5 min at RT. Remove the supernatant, add 500 µL of 70% ethanol and mix briefly.
- Centrifuge at 20,000 x g for 5 min at RT. Remove the supernatant, add 500 µL of 100% ethanol and mix briefly.
- Centrifuge at 20,000 x g for 5 min at RT. Remove the supernatant and let the DNA pellet dry under a laminar flow hood until all the ethanol evaporated.
NOTE: Use a heat block at 48 °C to speed up this process. The DNA pellet might not be visible at all time. - Dissolve the dried DNA pellet in 10 µL of nuclease-free H2O.
NOTE: DNA should be dissolved in as little volume as possible. Do not measure the DNA concentration.
- Pool 2 x 100 µL together. Add 20 µL (10% of the volume) of sterile filtered 3 M sodium acetate, pH 5.2.
| Volume [µL] | Final concentration | |
| 5x Q5 enhancer | 10 | 1x |
| 5x Q5 buffer | 10 | 1x |
| Primer fwd 10 µM | 2.5 | 0.5 µM |
| Primer rev 10 µM | 2.5 | 0.5 µM |
| dNTPs 10 mM | 1 | 200 µM |
| Q5 polymerase | 0.5 | 20 U/mL |
| DNA from yeast miniprep | 10 | |
| Nuclease-free H2O | 13.5 |
Table 3: Conditions for the 1st step PCR for amplification of the POI genes from the isolated yeast miniprep.
| Step | Temperature | Time |
| Initial denaturation | 98 °C | 30 s |
| 25 cycles | 98 °C | 10 s |
| 72 °C | 30 s | |
| 72 °C | 30 s | |
| Final extension | 72 °C | 2 min |
| Hold | 4 °C |
Table 4: Cycling conditions for the 1st step PCR for amplification of the POI genes from the isolated yeast miniprep.
| Volume [µL] | Final concentration | |
| Nuclease-free H2O | up to 50 | |
| 10x Thermopol buffer | 5 | 1x |
| Primer_fwd (10 µM) | 2.5 | 0.5 µM |
| Primer_rev (10 µM) | 2.5 | 0.5 µM |
| dNTPs (10 mM) | 1 | 200 µM |
| 8-oxo-dGTP (100 µM) | 1 | 2 µM |
| dPTP (100 µM) | 1 | 2 µM |
| PCR product from 1st PCR | XX | 50 ng |
| Taq DNA polymerase | 0.5 | 0.05 U/µL |
Table 5: Conditions for the epPCR which is performed after amplification of the POI DNA with the 1st step PCR.
| Step | Temperature | Time |
| Initial denaturation | 94 °C | 30 s |
| 15 cycles | 94 °C | 45 s |
| 60 °C | 30 s | |
| 72 °C | 1 min | |
| Final extension | 72 °C | 10 min |
| Hold | 4 °C |
Table 6: Cycling conditions for the epPCR.
| Volume [µL] | Final concentration | |
| 5x Q5 enhancer | 20 | 1x |
| 5x Q5 buffer | 20 | 1x |
| Primer fwd 10 µM | 5 | 0.5 µM |
| Primer rev 10 µM | 5 | 0.5 µM |
| dNTPs 10 mM | 1 | 200 µM |
| Q5 polymerase | 1 | 20 U/mL |
| 50 ng DNA | XX | |
| ddH20 | up to 100 |
Table 7: Conditions for the 2nd step PCR for amplification of the epPCR product before electroporation of EBY100 cells.
| Step | Temperature | Time |
| Initial denaturation | 98 °C | 30 s |
| 25 cycles | 98 °C | 10 s |
| 72 °C | 30 s | |
| 72 °C | 30 s | |
| Final extension | 72 °C | 2 min |
| Hold | 4 °C |
Table 8: Cycling conditions for the 2nd step PCR for amplification of the epPCR product.
8. Linearization of the yeast-display vector for electroporation
- Linearize the pCTCON2 yeast surface display vector (Addgene plasmid #41843).
- Perform a large-scale digest and use this digested linearized vector for all further electroporations.
- Digest 2 x 200 µg of pCTCON2 plasmid DNA with Sal I for 24 h at 37 °C for an initial linearization (Table 9).
- Digest with NheI and BamHI for 24 h at 37 °C to excise the insert (Table 10).
- Finally, add 5 µL of each enzyme to the tube (Sal I, NheI, BamHI) and incubate again for 24 h at 37 °C.
- Run the digested vector on a preparative agarose gel. Excise the bands and purify with a gel extraction kit. Elute the vector in 20 µL, measure the DNA concentration and adjust it to roughly 2 µg/µL.
NOTE: Digesting the plasmid for a prolonged time ensures sufficient cleavage of the insert and will result in less background from the vector after electroporation.
| DNA | 200 µg |
| 10x CutSmartBuffer | 50 µL |
| Sal I-HF (NEB) | 30 µL (60 U) |
| H2O | Up to 500 µL |
Table 9: Conditions for the first step of the large-scale digest of the yeast surface display vector pCTCON2.
| pCTCON2 (Sal I digested) | 500 µL |
| 10x CutSmartBuffer | 37.5 µL |
| NheI-HF (NEB) | 15 µL (30 U) |
| BamHI-HF (NEB) | 15 µL (30 U) |
| H2O | up to 875 µL |
Table 10: Conditions for the second step of the large-scale digest of the yeast surface display vector pCTCON2.
9. Electroporation of EBY100 with randomized DNA and linearized vector
- Prepare the EBY100 yeast cells (ATCC, see Table of Materials) by striking them from a glycerol stock 3 days prior to electroporation. Since non-transformed EBY100 cells do not contain the yeast surface display vector, use full media YPD plates (Table 1). Incubate at 30 °C.
- One day prior to electroporation, inoculate one single EBY100 colony into 30 mL freshly prepared YPD medium (Table 1). Incubate overnight at 30 °C while shaking at 180 rpm.
NOTE: The YPD medium should not be older than 1 month. - On the next morning, dilute the EBY100 overnight culture to an OD600 of 0.2 in 100 mL of YPD medium and incubate at 30 °C while shaking at 180 rpm.
NOTE: A culture volume of 50 mL is sufficient for 2 electroporations. Typically, 100 mL are prepared to have sufficient cells for the controls (cells only and cells plus vector). - When an OD600 of 1.3-1.5 is reached, pellet the cells in 50 mL tubes at 2000 x g for 3 min and remove the supernatant.
- Resuspend the pellet in 25 mL of 100 mM sterile filtered lithium acetate (see Table of Materials). Add freshly prepared sterile filtered DTT (1 M) (see Table of Materials) to a final concentration of 10 mM.
NOTE: The lithium acetate volume should be half the original volume of YPD. Scale down or up if necessary. Lithium acetate (100 mM) can be prepared in advance and stored for a few months at RT, but the 1 M DTT stock should be prepared right before use and kept on ice. - Incubate the cells at 30 °C while shaking for 10 min.
- Centrifuge at 2000 x g for 3 min, discard the supernatant, and place cells on ice.
NOTE: All following steps should be performed on ice and with cooled reagents, cuvettes, and microcentrifuge tubes. - Resuspend the pellet in 25 mL of cool sterile H2O. Centrifuge at 2000 x g for 3 min.
NOTE: The H2O volume used should be half of the original YPD volume. - Resuspend in 350 µL of sterile H2O.
NOTE: The total volume after resuspension should be 500 µL. Add more H2O if required. - Prepare the DNA library in parallel: Place the microcentrifuge tubes containing the ethanol-purified inserts on ice and add 4 µg of linearized pCTCON2 vector.
- Add 250 µL of the cells to the DNA and mix gently by snipping. Transfer to a pre-chilled 2 mm electroporation cuvette and keep on ice until the electroporation.
NOTE: Add controls (cells only without DNA and cells with linearized vector only without insert) to determine the success of electroporation. - Electroporate using 2 mm electroporation cuvettes (see Table of Materials) with the following conditions: square wave protocol, single pulse, 500 V, 15 ms pulse duration.
NOTE: Typical droops are 3%- 9%. Other electroporators can be used, but the conditions indicated are optimized for the instrument used in this study. - Rescue the cells with 1 mL of pre-warmed YPD, transfer to a pre-warmed microcentrifuge tube, and incubate without shaking for 1 h at 30 °C.
NOTE: Place the heating block next to the electroporator to ensure that the recovery tubes stay at the proper temperature. - Plate 100 µL of 10-3, 10-4, 10-5, and 10-6 dilutions of cells on SD-CAA plates (see Table 1) and allow for growth of transformants at 30 °C.
NOTE: The theoretical library diversity is obtained by calculating back the number of transformants to the original culture volume, here 1 mL. Typically, this transformation protocol yields approximately 107-108 transformed cells. - Pellet the remaining cells at 2000 x g for 3 min, resuspend in 200 mL SD-CAA, and incubate at 30 °C with shaking at 180 rpm overnight. The transformed cells containing the epPCR-randomized library can either be passaged in SD-CAA and induced for surface expression in SG-CAA on the next day or stored at 4 °C until further use. Freeze glycerol stocks of the library after epPCR after determining the diversity. Since on the next day a fraction of cells will still be non-transformed cells that were carried over in the transformation procedure without obtaining a plasmid, it is recommended to passage at least 20-fold of the library diversity in the first passaging step to compensate for this effect.
10. Sequencing of yeast libraries after several selection rounds
- Extract the yeast DNA by performing a yeast miniprep as described above.
- Electroporate commercial E. coli 10-beta electro-competent cells (see Table of Materials) with the isolated DNA.
- Place the electroporation cuvettes (1 mm, see Table of Materials) and microcentrifuge tubes on ice. Pre-warm microcentrifuge tubes with 950 µL of outgrowth medium at 37 °C to recover the cells after electroporation.
- Thaw the cells on ice and mix by flicking the tube gently. Transfer 50 µL of the cells to a chilled microcentrifuge tube. Add 1 µL of the yeast miniprep DNA.
- Carefully transfer the cell-DNA mix into a chilled electroporation cuvette.
NOTE: Avoid bubbles and ensure that the cells deposit on the bottom of the cuvette. - Electroporate with an electroporation device using the following conditions: 2.0 kV; 200 Ω; 25 µF.
NOTE: Typical time constants are 4.8-5.1 ms. Other electroporators can be used, but the conditions indicated are optimized for the instrument used in this study. - Immediately add 950 µL of 37 °C pre-warmed outgrowth medium (see Table of Materials) to the cuvettes, mix gently, and transfer back to the warm microcentrifuge tubes.
NOTE: Place the heating block next to the electroporator to ensure that the recovery tubes stay at the proper temperature. - Incubate for 40 min at 37 °C and 300 rpm. Plate 10 µL, 100 µL, and the rest of the cells (after centrifugation) onto a pre-warmed selective LB plate (see Table 1). Incubate the plates overnight at 37 °C.
NOTE: For the pCTCON2 vector, ampicillin is used for antibiotic selection. - Prepare a copy-plate with LB agar and the required antibiotic (here ampicillin is used): pipette 200 µL of LB agar with 100 µg/mL ampicillin in each well of a 96 well plate and let it solidify.
- Prepare a 96 well sequencing plate with 50 µL of LB with 100 µg/mL ampicillin per well.
- Pick a single colony from the plates, and first inoculate the copy plate and the same well of the sequencing plate.
NOTE: The copy plate ensures that the desired plasmids can be easily obtained by inoculating a liquid culture and performing a miniprep. - Incubate the copy plate overnight at 37 °C and then store at 4 °C.
- Incubate the sequencing plate for 3-4 h at 37 °C while shaking (180 rpm) covered with a plastic or aluminum foil.
- Seal the sequencing plate and send it for sequencing along with 500 µL of 10 µM pCTCON2 sequencing primer (5'-CGTTTGTCAGTAATTGCGGTTCTC).
- For analysis of the sequences, translate the obtained DNA sequences to amino acid sequences using a suitable program of choice, for example, EMBOSS Transeq38. Next, use Multiple Sequence Alignment (MUSCLE)39 or a similar alignment tool to align the variant sequences to each other and to the parental wildtype protein.
NOTE: By performing sequencing of libraries of different selection rounds (for example, after an earlier round of selection, as well as after additional rounds of selection with lower antigen concentrations to enrich high-affinity variants), it is possible to identify different variants. This is highly dependent on the stringency of the gates applied during sorting rounds and, hence, the diversities of the libraries.
Results
The G4 library was selected against the antigen hRBP4 bound to the small molecule drug A1120. The staining of the libraries for flow cytometric sorting was performed as described in Method 6, and the applied gating strategy is shown in Figure 2A. An initial gate included all cells based on cell morphology, and the second gate (histogram of FSC-Width) showed a stringent gating strategy that was applied to select single cells and remove cell aggregates. The third and final gate showed the display of protein variants (x-axis) versus antigen binding (y-axis). Yeast cells showing both display and binding signals were sorted. Importantly, the sorting gate was set in a stringent way to enrich binding domains with high binding signal and thus high affinity. This stringent selection yielded an enrichment of displaying yeast cells that specifically bind to the target antigen throughout the selection campaign (Figure 2B). In later flow cytometric sorting rounds, the antigen concentration was decreased 10-fold (from 100 nM to 10 nM). Therefore, the overall binding signal was reduced, and only binders with a high affinity were still detectable and sorted (Figure 2C).
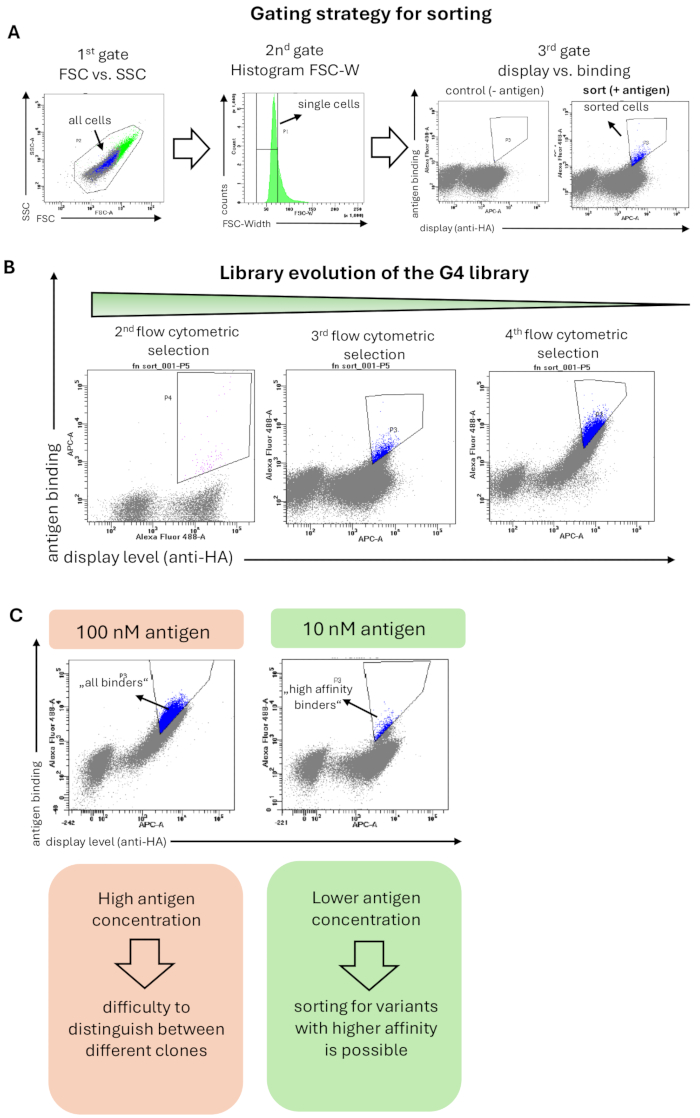
Figure 2: Representative results from a yeast surface display selection of the Fn3-based G4 library for binding to the antigen (hRBP4 in the presence of A1120). (A) The general gating strategy for sorting of yeast libraries. The first gate (FSC vs. SSC) is to select all yeast cells and exclude scatter events; the second gate (histogram of FSC-W) aims to remove cell aggregates and only select single yeast cells. The third gate plots the surface display level (detection of the HA- or c-myc-tag) vs. binding to the antigen (here hRBP4 in the presence of 5 µM A1120, detected by anti-His antibody). The library was additionally stained with secondary antibodies only (without antigen), where no antigen binding is expected. Sorted cells are highlighted in blue. (B) Evolution of the G4 library throughout 3 rounds of flow cytometric sorting. Enrichment of the binding population can be observed with each selection round. (C) The use of lower antigen concentrations enables the selection of protein variants with a higher affinity towards the target antigen. Upon reduction of the antigen concentration (here hRBP4) by 10-fold, different diagonals appear, indicating the presence of clones with higher (sorted cells, blue) or lower affinity. Please click here to view a larger version of this figure.
Discussion
Yeast surface display has evolved as one of the key methods used in protein engineering. Although it is commonly employed for the engineering of affinity1,18,40,41, expression/stability24,27,42,43 and activity28,44, further uses like epitope mapping45,46 or characterization of the individual mutants on the surface of yeast cells9 are possible as well. In this protocol, we provide the basic steps for starting a yeast surface display selection campaign, including the selection with magnetic beads and by flow cytometric sorting as well as diversification of the yeast library by epPCR for affinity maturation.
One essential requirement for conventional yeast surface display selections is the availability of soluble protein of sufficient quality. Starting with a well-folded target protein with high purity and a defined oligomerization state (i.e., monomeric protein should only be present as monomer) provides the highest success rate to select for a protein variant binding to the target antigen with high affinity. An alternative for difficult-to-express target proteins is cell-based selections, which present a reasonable strategy to circumvent this limitation47. However, yeast surface display offers many advantages, such as the possibility of characterizing resulting protein variants directly on the surface of yeast without the necessity to perform laborious and time-intensive cloning, expression in a soluble format, and protein purification. Both the affinity and the stability of the variants can be analyzed directly on the yeast surface9.
In this protocol, we show how the G4 library of protein variants, more specifically of the 10th type III domain of human fibronectin, was selected for binding to the antigen hRBP4 in the presence of the small molecule A1120. The combination of bead selections and flow cytometric sorting yielded an enrichment of variants, which showed an increased binding to the target antigen throughout selection rounds (Figure 2B). We showed that using lower concentrations of antigen enables the selection of high-affinity protein variants (Figure 2C). Typically, affinities that can be achieved with yeast display selections are in the nanomolar or even picomolar range18. The final affinities depend on the target antigen, the number of selection rounds and affinity maturation, the binding scaffold used, and the applied gating strategy. The characterization of individual protein variants is not covered in this protocol but is explained in detail in our previous work9. Although yeast display was originally employed for the engineering of antibody fragments such as scFvs1,40, the method has been widely used for non-antibody-based proteins as well10.
To sum up, yeast surface display is a powerful protein engineering tool that enables the generation of protein variants with novel or improved properties, such as binding to almost any target protein and/or increased stability.
Disclosures
M.W.T. receives funding from Miltenyi Biotec. All authors are inventors on patent applications for technologies and engineered proteins that were developed by using yeast surface display.
Acknowledgements
This work was supported by the Austrian Science Fund (FWF Project W1224 - Doctoral Program on Biomolecular Technology of Proteins - BioToP and FWF Project ESP 465-B), the Federal Ministry for Digital and Economic Affairs of Austria, the National Foundation for Research, Technology and Development of Austria to the Christian Doppler Research Association (Christian Doppler Laboratory for Next Generation CAR T Cells), and by private donations to the St. Anna Children's Cancer Research Institute (Vienna, Austria). E.S. is a recipient of a DOC Fellowship of the Austrian Academy of Sciences at the St. Anna Children's Cancer Research Institute. Figures were created with BioRender.com.
Materials
| Name | Company | Catalog Number | Comments |
| 10-beta electrocompetent E. coli | NEB | C3020K | |
| Agar-Agar, Kobe I | Carl Roth | 5210.5 | |
| Ampicillin sodium salt | Carl Roth | K029.2 | |
| Anti-c-myc antibody, clone 9E10, AF488 | Invitrogen | MA1-980-A488 (Thermo Fisher) | |
| Anti-c-myc antibody, clone 9E10, AF647 | Invitrogen | MA1-980-A647 (Thermo Fisher) | |
| Anti-HA antibody, clone 16B12, AF488 | BioLegend | 901509 (Biozym) | |
| Anti-HA antibody, clone 16B12, AF647 | BioLegend | 682404 (Biozym) | |
| BamHI-HF | NEB | R3136S | |
| Bovine serum albumin, cold ethanol fraction | Sigma-Aldrich | A4503 | |
| Citric acid monohydrate | Sigma-Aldrich | C1909 | |
| D-Galactose | Carl Roth | 4987.2 | |
| D-Glucose | Sigma-Aldrich | G8270 | |
| Difco yeast nitrogen base | Becton Dickinson (BD) | 291940 | |
| Di-Sodium hydrogen phosphate heptahydrate | Carl Roth | X987.3 | |
| DL-Dithiothreitol | Sigma-Aldrich | D0632 | |
| D-Sorbitol | Carl Roth | 6213.1 | |
| Dulbecco’s phosphate buffered saline (10x) | Thermo Scientific | 14190169 | |
| Dynabeads Biotin Binder | Invitrogen | 11047 (Fisher Scientific) | |
| DynaMag-2 Magnet | Thermo Fisher | 12321D | |
| EBY100 | ATCC | MYA-4941 | |
| Electroporation cuvette 1 mm (for E.coli) | VWR | 732-1135 | |
| Electroporation cuvette 2 mm (for yeast) | VWR | 732-1136 | |
| Ethanol absolute | MERCK | 1070172511 | |
| GeneMorph II Random Mutagenesis Kit | Agilent Technologies | 200550 | |
| Gibco Bacto Casamino Acids | Becton Dickinson (BD) | 223120 | |
| Glycerol | AppliChem | 131339.1211 | |
| LE agarose | Biozym | 840004 | |
| Lithium acetate dihydrate | Sigma-Aldrich | L4158 | |
| Monarch DNA Gel Extraction Kit | NEB | T1020S | |
| Monarch PCR & DNA Cleanup Kit | NEB | T1030S | |
| Multifuge 1S-R | Heraeus | ||
| NheI-HF | NEB | R3131S | |
| Outgrowth medium | NEB | B9035S | |
| pCTCON2 | Addgene | #41843 | |
| Penicillin G sodium salt | Sigma-Aldrich | P3032 | |
| Penta-His antibody, AF488 | Qiagen | 35310 | |
| Penta-His antibody, AF647 | Qiagen | 35370 | |
| Peptone ex casein tryptically digested | Carl Roth | 8986.3 | |
| Q5 High-Fidelity DNA Polymerase | NEB | M0491S | |
| SaII-HF | NEB | R3138S | |
| Sodium acetate | Sigma-Aldrich | S8750-1KG | |
| Sodium chloride | Carl Roth | 3957.2 | |
| Sodium dihydrogen phosphate monohydrate | Carl Roth | K300.2 | |
| Steritop threaded bottle top filter | MERCK | S2GPT01RE | |
| Streptavidin, AF488 | Invitrogen | S32354 (Thermo Fisher) | |
| Streptavidin, AF647 | Invitrogen | S32357 (Thermo Fisher) | |
| Streptomycin sulfate | Sigma-Aldrich | S6501 | |
| Tri-Sodium citrate dihydrate | Carl Roth | 4088.1 | |
| UV-Vis spectrophotometer | Agilent | 8453 | |
| Yeast extract, micro-granulated | Carl Roth | 2904.4 | |
| Zymoprep Yeast Plasmid Miniprep II | Zymo Research | D2004 |
References
- Boder, E. T., Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat Biotechnol. 15 (6), 553-557 (1997).
- Scott, J. K., Smith, G. P. Searching for peptide ligands with an epitope library. Science. 249 (4967), 386-390 (1990).
- Smith, G. P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science. 228 (4705), 1315-1317 (1985).
- Schaffitzel, C., Hanes, J., Jermutus, L., Pluckthun, A. Ribosome display: an in vitro method for selection and evolution of antibodies from libraries. J Immunol Methods. 231 (1-2), 119-135 (1999).
- Ho, M., Nagata, S., Pastan, I. Isolation of anti-CD22 Fv with high affinity by Fv display on human cells. Proc Natl Acad Sci U S A. 103 (25), 9637-9642 (2006).
- Wagner, A., et al. Identification of activating mutations in the transmembrane and extracellular domains of EGFR. Biochemistry. 61 (19), 2049-2062 (2022).
- Wagner, A., et al. PhosphoFlowSeq - A high-throughput kinase activity assay for screening drug resistance mutations in EGFR. J Mol Biol. 433 (22), 167210(2021).
- Cherf, G. M., Cochran, J. R. Applications of yeast surface display for protein engineering. Methods Mol Biol. 1319, 155-175 (2015).
- Zajc, C. U., Teufl, M., Traxlmayr, M. W. Affinity and stability analysis of yeast displayed proteins. Methods Mol Biol. 2491, 155-173 (2022).
- Gai, S. A., Wittrup, K. D. Yeast surface display for protein engineering and characterization. Curr Opin Struct Biol. 17 (4), 467-473 (2007).
- Orr, B. A., Carr, L. M., Wittrup, K. D., Roy, E. J., Kranz, D. M. Rapid method for measuring ScFv thermal stability by yeast surface display. Biotechnol Prog. 19 (2), 631-638 (2003).
- Teufl, M., Zajc, C. U., Traxlmayr, M. W. Engineering strategies to overcome the stability-function trade-off in proteins. ACS Synth Biol. 11 (3), 1030-1039 (2022).
- Wozniak-Knopp, G., et al. Introducing antigen-binding sites in structural loops of immunoglobulin constant domains: Fc fragments with engineered HER2/neu-binding sites and antibody properties. Protein Eng Des Sel. 23 (4), 289-297 (2010).
- Traxlmayr, M. W., et al. Directed evolution of stabilized IgG1-Fc scaffolds by application of strong heat shock to libraries displayed on yeast. Biochim Biophys Acta. 1824 (4), 542-549 (2012).
- Traxlmayr, M. W., et al. Directed evolution of Her2/neu-binding IgG1-Fc for improved stability and resistance to aggregation by using yeast surface display. Protein Eng Des Sel. 26 (4), 255-265 (2013).
- Hasenhindl, C., et al. Stability assessment on a library scale: a rapid method for the evaluation of the commutability and insertion of residues in C-terminal loops of the CH3 domains of IgG1-Fc. Protein Eng Des Sel. 26 (10), 675-682 (2013).
- Koide, A., Wojcik, J., Gilbreth, R. N., Hoey, R. J., Koide, S. Teaching an old scaffold new tricks: monobodies constructed using alternative surfaces of the FN3 scaffold. J Mol Biol. 415 (2), 393-405 (2012).
- Hackel, B. J., Kapila, A., Wittrup, K. D. Picomolar affinity fibronectin domains engineered utilizing loop length diversity, recursive mutagenesis, and loop shuffling. J Mol Biol. 381 (5), 1238-1252 (2008).
- Traxlmayr, M. W., et al. Strong enrichment of aromatic residues in binding sites from a charge-neutralized hyperthermostable Sso7d scaffold library. J Biol Chem. 291 (43), 22496-22508 (2016).
- Zajc, C. U., et al. A conformation-specific ON-switch for controlling CAR T cells with an orally available drug. Proc Natl Acad Sci U S A. 117 (26), 14926-14935 (2020).
- Moore, S. J., Cochran, J. R. Engineering knottins as novel binding agents. Methods Enzymol. 503, 223-251 (2012).
- Hackel, B. J., Ackerman, M. E., Howland, S. W., Wittrup, K. D. Stability and CDR composition biases enrich binder functionality landscapes. J Mol Biol. 401 (1), 84-96 (2010).
- Hackel, B. J., Wittrup, K. D. The full amino acid repertoire is superior to serine/tyrosine for selection of high affinity immunoglobulin G binders from the fibronectin scaffold. Protein Eng Des Sel. 23 (4), 211-219 (2010).
- Laurent, E., et al. Directed evolution of stabilized monomeric CD19 for monovalent CAR interaction studies and monitoring of CAR-T cell patients. ACS Synth Biol. 10 (5), 1184-1198 (2021).
- Julian, M. C., et al. Co-evolution of affinity and stability of grafted amyloid-motif domain antibodies. Protein Eng Des Sel. 28 (10), 339-350 (2015).
- Cochran, J. R., Kim, Y. S., Lippow, S. M., Rao, B., Wittrup, K. D. Improved mutants from directed evolution are biased to orthologous substitutions. Protein Eng Des Sel. 19 (6), 245-253 (2006).
- Kieke, M. C., et al. High affinity T cell receptors from yeast display libraries block T cell activation by superantigens. J Mol Biol. 307 (5), 1305-1315 (2001).
- Lipovsek, D., et al. Selection of horseradish peroxidase variants with enhanced enantioselectivity by yeast surface display. Chem Biol. 14 (10), 1176-1185 (2007).
- Fushimi, T., et al. Mutant firefly luciferases with improved specific activity and dATP discrimination constructed by yeast cell surface engineering. Appl Microbiol Biotechnol. 97 (9), 4003-4011 (2013).
- Lam, S. S., et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat Methods. 12 (1), 51-54 (2015).
- Park, S. W., Lee, D. S., Kim, Y. S. Engineering a U-box of E3 ligase E4B through yeast surface display-based functional screening generates a variant with enhanced ubiquitin ligase activity. Biochem Biophys Res Commun. 612, 147-153 (2022).
- Schutz, M., et al. Directed evolution of G protein-coupled receptors in yeast for higher functional production in eukaryotic expression hosts. Sci Rep. 6, 21508(2016).
- Angelini, A., et al. Protein engineering and selection using yeast surface display. Methods Mol Biol. 1319, 3-36 (2015).
- Chen, T. F., de Picciotto, S., Hackel, B. J., Wittrup, K. D. Engineering fibronectin-based binding proteins by yeast surface display. Methods Enzymol. 523, 303-326 (2013).
- Shusta, E. V., Holler, P. D., Kieke, M. C., Kranz, D. M., Wittrup, K. D. Directed evolution of a stable scaffold for T-cell receptor engineering. Nat Biotechnol. 18 (7), 754-759 (2000).
- Traxlmayr, M. W., Obinger, C. Directed evolution of proteins for increased stability and expression using yeast display. Arch Biochem Biophys. 526 (2), 174-180 (2012).
- Traxlmayr, M. W., Shusta, E. V. Directed evolution of protein thermal stability using yeast surface display. Methods Mol Biol. 1575, 45-65 (2017).
- Madeira, F., et al. The EMBL-EBI Job Dispatcher sequence analysis tools framework in 2024. Nucleic Acids Res. 52 (W1), W521-W525 (2024).
- Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32 (5), 1792-1797 (2004).
- Chao, G., et al. Isolating and engineering human antibodies using yeast surface display. Nat Protoc. 1 (2), 755-768 (2006).
- VanAntwerp, J. J., Wittrup, K. D. Fine affinity discrimination by yeast surface display and flow cytometry. Biotechnol Prog. 16 (1), 31-37 (2000).
- McConnell, A. D., et al. A general approach to antibody thermostabilization. MAbs. 6 (5), 1274-1282 (2014).
- Pavoor, T. V., Wheasler, J. A., Kamat, V., Shusta, E. V. An enhanced approach for engineering thermally stable proteins using yeast display. Protein Eng Des Sel. 25 (10), 625-630 (2012).
- Chen, I., Dorr, B. M., Liu, D. R. A general strategy for the evolution of bond-forming enzymes using yeast display. Proc Natl Acad Sci U S A. 108 (28), 11399-11404 (2011).
- Traxlmayr, M. W., et al. Construction of a stability landscape of the CH3 domain of human IgG1 by combining directed evolution with high throughput sequencing. J Mol Biol. 423 (3), 397-412 (2012).
- Chao, G., Cochran, J. R., Wittrup, K. D. Fine epitope mapping of anti-epidermal growth factor receptor antibodies through random mutagenesis and yeast surface display. J Mol Biol. 342 (2), 539-550 (2004).
- Stern, L. A., et al. Cellular-based selections aid yeast-display discovery of genuine cell-binding ligands: Targeting oncology vascular biomarker CD276. ACS Comb Sci. 21 (3), 207-222 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved