A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Measurement of Mitochondrial Membrane Potential In Vivo using a Genetically Encoded Voltage Indicator
In This Article
Summary
This protocol describes the application of mitochondria-targeted genetically encoded voltage indicators (GEVIs). These GEVIs offer a significant advantage over traditional mitochondrial membrane potential dyes by enabling specific, in vivo, and real-time monitoring of mitochondrial membrane potential.
Abstract
Mitochondrial membrane potential (MMP, ΔΨm) is critical for mitochondrial functions, including ATP synthesis, ion transport, reactive oxygen species (ROS) generation, and the import of proteins encoded by the nucleus. Existing methods for measuring ΔΨm typically use lipophilic cation dyes, such as Rhodamine 800 and tetramethylrhodamine methyl ester (TMRM), but these are limited by low specificity and are not well-suited for in vivo applications. To address these limitations, we have developed a novel protocol utilizing genetically encoded voltage indicators (GEVIs). Genetically encoded voltage indicators (GEVIs), which generate fluorescent signals in response to membrane potential changes, have demonstrated significant potential for monitoring plasma membrane and neuronal potentials. However, their application to mitochondrial membranes remains unexplored. Here, we developed protein-based mitochondrial-targeted GEVIs capable of detecting ΔΨm fluctuations in cells and the motor cortex of living animals. The mitochondrial potential indicator (MPI)offers a non-invasive approach to study ΔΨm dynamics in real-time, providing a method to investigate mitochondrial function under both normal and pathological conditions.
Introduction
Mitochondria are essential organelles in eukaryotic cells, serving as the primary energy suppliers through adenosine triphosphate (ATP) generation while also performing a variety of other crucial functions, such as metabolite synthesis, calcium ions buffering, heat production, and regulation of cell survival1. Their roles are particularly critical in highly metabolic tissues like the brain and heart, where they help maintain cellular homeostasis. Mitochondrial membrane potential (MMP, Ψm) is central to these processes, including driving ATP synthesis via oxidative phosphorylation, facilitating the transport of metabolites and ions across the mitochondrial membranes, and contributing to the generation of reactive oxygen species (ROS)2,3. MMP also influences mitochondrial morphology and dynamics4, including mitophagy (the selective degradation of mitochondria)5and apoptosis (programmed cell death)6. Maintaining an appropriate Ψm is essential for cellular function; its dysregulation is linked to numerous pathologies, including neurodegenerative diseases, heart failure, and cancer. Current methods for measuring Ψm were primarily based on the use of lipophilic cationic dyes, including TMRM (tetramethylrhodamine methyl ester), TMRE (tetramethylrhodamine ethyl ester), Rhodamine 123, Safranin O, Rhodamine 800, DiOC6, JC-1, etc.7. However, these fluorescent molecules have several limitations. These dyes lack cell specificity, are susceptible to quenching, and some are toxic. Additionally, they can diffuse over time, and when mitochondrial ΔΨ is lost, they leak out, rendering them unable to indicate the membrane potential of depolarized mitochondria. Furthermore, rhodamine-based dyes like TMRM and TMRE are temperature-sensitive8, necessitating careful consideration of temperature effects on dye fluorescence, particularly when measuring mitochondrial membrane voltage during physiological activities involving cellular thermogenesis.
Genetically encoded voltage indicators (GEVIs), proteins capable of detecting membrane potential changes through fluorescent signals9,10, have emerged as powerful tools for monitoring membrane potentials in a variety of cellular contexts11. While GEVIs have been applied extensively to study plasma membranes, there has been little progress in adapting them to measure intracellular membrane potentials, particularly for mitochondria. This protocol seeks to address this gap by using mitochondrial-targeted GEVIs that could monitor mitochondrial membrane potential in vitro and in vivo. By adding mitochondrial signal sequence to the existing GEVIs, appropriate GEVI can be targeted to mitochondria12. These mitochondrial potential indicators (MPI) would provide new insights into mitochondrial physiology and offer significant potential for exploring mitochondrial function in various disease states in vivo, enhancing our understanding of how mitochondrial dynamics contribute to both normal and pathological cellular processes.
Protocol
All animal care and experiments were performed in accordance with the Institutional Animal Care and Use Committee guidelines of Zhengzhou University. Sterilize all surgical instruments before use. Follow aseptic techniques to prevent infection. After all the data had been acquired, the animals were euthanized using an overdose of inhalant anesthesia followed by decapitation.
1. In vitro applications
- Plasmid construction
- Obtain the accelerated sensor of action potentials 1 (ASAP1) and ASAP3 genes from Addgene or synthesizing from the sequence. (ASAP1, NCBI Accession ID: AHV90412.1, Addgene ID: 52519; ASAP3, Addgene ID: 132331).
- Use the primers listed in Table 1 for plasmid constructions.
- For expression in cells, use the EGFPN1 vector for construction.
- Initially, synthesize the cox8 gene sequence (NP_004065, amino acids 1-29) with four tandem repeats (4cox8) flanked by NheI and XhoI restriction sites. Digest both the synthesized 4cox8 fragment and the EGFPN1 vector with NheI and XhoI, followed by ligation to generate the 4cox8-EGFP vector.
- Subsequently, perform PCR amplification of the ASAP1 sequence with primer 1 and primer 2. Digest the ASAP1 fragment and the 4cox8-EGFP vector with SalI and NotI, followed by ligation to generate the final MPI-1 vector (MPI stands for mitochondrial potential indicator).
NOTE: The vector map of CMV-MPI-1 is depicted in Figure 1. The detailed sequence can be accessed from NCBI (Accession ID: PQ678920).
- For specific expression in neurons, perform PCR amplification of the hSyn promoter sequence from AAV-hSyn-EGFP plasmid (NCBI Accession ID: MH458079, Addgene ID: 50465) using primer 3 and primer 4.
- Digest both the hSyn sequence and the 4cox8-EGFP vector with AseI and NheI, followed by ligation to generate the hSyn-4cox8-EGFP vector. Subsequently, perform PCR amplification of ASAP3 sequence with primer 1 and primer 2.
- Digest the hSyn-4cox8-EGFP vector and the ASAP3 sequence with SalI and NotI, followed by ligation to generate the hSyn-MPI-2 vector.
NOTE: PCR conditions will vary based on the specific enzymes and primer melting temperatures (Tm). A typical PCR reaction involves an initial denaturation step at 95 °C for 30 s, followed by 25-35 cycles of denaturation (95 °C, 10 s), annealing (55 °C, 30 s), and extension (72°C, 30 s/kb). A final extension step at 72 °C for 5 min ensures complete product synthesis, followed by a hold at 4 °C. The annealing temperature should be approximately 5 °C below the Tm of the primers. Refer to the manufacturer's instructions for optimal PCR parameters. The primers for ASAP1 and ASAP3 are identical.
- Imaging in living cells
- Prepare DMEM culture medium by adding 10% fetal bovine serum and 100 mg/L penicillin/streptomycin to the medium.
- Prepare Ca2+ transfection reagents: 2.5 mol/L CaCl2, 2x HEBS (274 mmol/L NaCl, 10 mmol/L KCl, 1.4 mmol/L Na2HPO4, 15 mmol/L D-glucose, 42 mmol/L HEPES, pH 7.07).
- Prepare Tyrode's buffer for maintaining the cells during imaging (145 mmol/L NaCl, 3 mmol/L KCl, 10 mmol/L HEPES, 10 mmol/L Glucose, pH 7.4).
- Culture Hela cells in a 35 mm culture dish fed with DMEM culture medium at an incubator with a temperature of 37 °C and an atmosphere of 5% CO2/95% air. Add 2-3 coverslips per dish to allow cells to grow on them.
- Transfect the Hela cells via the calcium phosphate precipitation method.
- To transfect cells, start by replacing the old culture medium with 2 mL of serum-free DMEM. After 35 min, prepare the DNA/Ca2+ solution by mixing 10 µg of DNA, 4 µL of CaCl2, and ddH2O to a final volume of 40 µL in a microcentrifuge tube.
- Add 40 µL of 2x HEBS to the DNA/Ca2+ solution dropwise while stirring vigorously. Bubble air with a pipette into the mixture to facilitate calcium phosphate precipitation.
- After 25 min, add 80 µL of this precipitate solution to the dish with the cells and return it to the incubator for another 1-2 h. Then, remove the non-serum DMEM and rinse the cells with 15% glycerol (dissolved in PBS) for 2 min.
- After aspirating the glycerol, add 2 mL of DMEM containing 10% FBS and continue culturing the cells for 16-24 h.
- Place coverslips with cells in a 35 mm culture dish with 2 mL of Tyrode's buffer and stain the cells with rhodamine 800 dye by adding 20 µmol/L rhodamine 800 (dissolved in ddH2O) to the dish to a final concentration of 50 nmol/L.
- Prepare a fluorescence microscope and set it up with the appropriate filters and light source to excite the protein with a 488 nm light source and collect the emission between 490-540 nm. Excite the rhodamine 800 dye with a 633 nm light source and collect the emission between 650-720 nm.
- Prepare a carbonyl cyanide m-chlorophenyl hydrazone (CCCP) solution at 500 µM (stock) and add to the cells to a final concentration of 5 µM during imaging.
NOTE: The pH of the 2x HEBS solution is crucial for transfection efficiency. The pH should be carefully adjusted to 7.07 with solid sodium hydroxide. The pH cannot be lower than 7.01 or higher than 7.12.
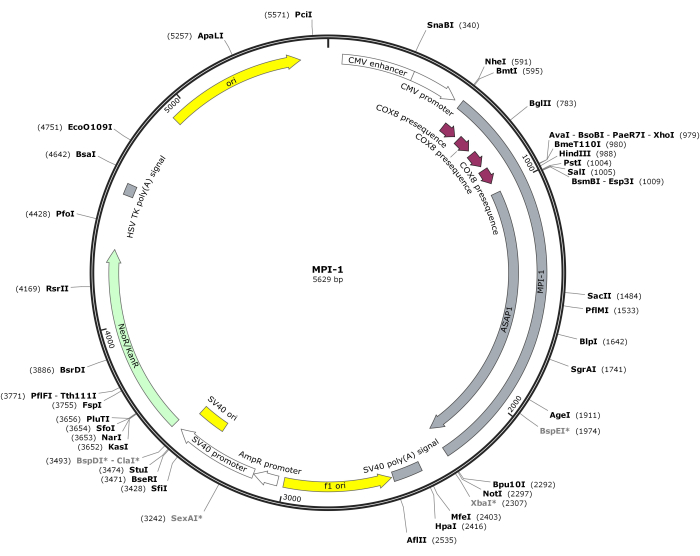
Figure 1: Vector map of CMV-MPI-1. Please click here to view a larger version of this figure.
2. In vivo applications
- Plasmid construct
- For in vivo expression, use AAV virus for transduction. Obtain the expressing backbone for AAV packaging (Addgene ID: 46954 or 26968), which contains inverted terminal repeats (ITR) sequences for AAV virus packaging.
- Perform PCR amplification of the hSyn-MPI-2 vector (from step 1.1.4) using primers 5 and 7 in Table 1.
- Perform a second PCR amplification using the same template and primers 6 and 8 in Table 1.
- Perform a final PCR using the product from the first and the second PCR (steps 2.1.2 and 2.1.3) as the template and primers 5 and 8 in Table 1, generating the final hSyn-MPI-2 sequence.
- Digest the hSyn-MPI-2 sequence and the pAAV backbone with MluI and EcoRI, followed by ligation to generate the final pAAV-hSyn-MPI-2 construct. The vector map of pAAV-hSyn-MPI-2 is illustrated in Figure 2. The detailed sequence can be accessed from NCBI (Accession ID: PQ678919).
NOTE: The hSyn-MPI-2 vector contains an internal EcoRI site originating from the original EGFPN1 backbone, which is removed using modified primers in an overlapping PCR strategy (steps 2.1.2-2.1.4).
- Virus preparation and transduction
- Culture HEK293t cells in a 100 mm culture dish fed with 10% FBS DMEM culture medium at an incubator with a temperature of 37 °C and an atmosphere of 5% CO2/95% air to reach a 70% confluence.
- Co-transfect the HEK293t cells with AAV gene plasmid, capsid (pAAV-DJ or pAAV9), and helper plasmids (pHelper) using the calcium phosphate precipitation method. The DNA amount for each plasmid was as follows: gene: 10 µg, pHelper: 11 µg, capsid: 9 µg. The transfection reagent amount: 12 µL of CaCl2, 120 µL of 2x HEBS.
- Incubate the transfected cells for a period of 72 h to enable virus production.
- After the incubation period, harvest the cells by centrifuge at 400 g. Rinse the cells with PBS 2 times and resuspend the cell in 400 µL of PBS.
- Harvest viruses from cell suspension with four cycles of freeze-thaw methods (37 °C and -80 °C ). Centrifuge the lysate at 13,500 × g at 4 °C and discard the pellet. The supernatant contains the AAV virus.
- Perform real-time PCR to determine the titration of the prepared virus. Detailed protocol for AAV titration determination can be found at https://www.addgene.org/protocols/aav-titration-qpcr-using-sybr-green-technology/.
- Aliquot and store the prepared AAV virus at -80 °C.
- Stereotactic virus injection
- Anesthetize an adult C57/BL male mouse weighing 24-26 g with 3% pentobarbital sodium via intraperitoneal injection at a dosage of 30 mg/kg. Once anesthetized, fix the mice on a stereotaxic apparatus to maintain a stable position throughout the procedure.
- Apply veterinary ophthalmic ointment to protect the eyes from light damage and to keep the eyes moist.
- Use depilatory cream to remove hair on the mouse's head and expose the scalp.
- Open the scalp using surgery scissors and expose the skull. Treat the skull with 3% hydrogen peroxide to sterilize and visualize key landmarks such as the bregma and lambda sites.
- Adjust the position of the mouse's head to ensure that the skull is horizontal.
- Use a programmable micropipette puller to pull glass pipettes to the desired shape and size for precise injections. Fill the glass micropipette with paraffin oil.
- Position the micropipette at the target site. In this protocol, the target is the M2 region on the right side, using the following coordinates: AP (anterior-posterior), +1.94 mm; ML (mediolateral), +0.75 mm; DV (dorsoventral), -1.5 mm.
- Use an electric drill to make a small hole in the skull at the target site.
- Aspire 500 nL of virus at a titer of 1.5 x 1011 Genome Copies per milliliter(GC/mL) through the tip of micropipette. Inject 500 nL of virus into the target site using an injection pump at a flow rate of 100 nL/min.
- After the injection, leave the pipette in place for approximately 5 min to allow for diffusion before carefully retracting it.
- Implant optical fiber (200 µm diameter, 0.37 numerical aperture, 2 mm length) into the injection site.
- Secure the optical fiber in place using dental cement.
- Suture the mouse's scalp with 5-0 nylon thread (aesthetic needle Δ1/2 4x12) and return it to its home cage.
- Place the home cage on a heating pad to maintain the mouse's body temperature until it fully recovers from anesthesia.
- Turn off the heating pad once the animal begins to move around. Allow for another 2 h of acclimation to the room temperature (RT) before transferring the mouse to the housing facility.
- House mouse in a 12-h light/12-h dark cycle at 22 °C with ad libitum access to food and water for 2-3 weeks to allow for viral expression.
- Fiber photometry
- Connect the optical fiber implanted in the mouse to an optic cable.
- Attach the other end of the optic cable to a detector that will be used to capture the imaging data.
- For imaging, use a light source to excite the proteins at a wavelength of 488 nm. Set up the detector to collect the emitted light at wavelengths ranging from 490 nm to 540 nm.
- Adjust the intensity of the excitation light to optimize the signal without causing damage to the brain tissue.
- Modify the gains of the fluorescent signal to ensure a clear and strong signal is captured by the detector.
- To avoid rapid bleaching of the fluorescence, which would reduce the quality of the imaging, fine-tune the light intensity and the gain of the fluorescent signal as necessary.
- Conduct mouse behavior test.
- Continuously monitor the imaging process during the behavior test.
- Data analysis
- Use Python and GNUplot to create a heatmap of experimental trials for a visual representation of the data. Source codes are provided in Supplementary File 1.
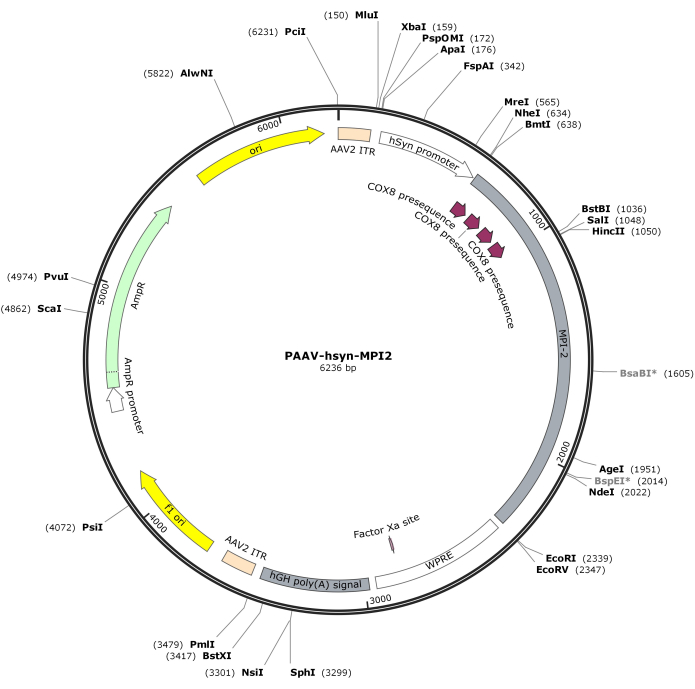
Figure 2: Vector map of AAV-hSyn-MPI-2. Please click here to view a larger version of this figure.
Results
After constructing the CMV-MPI-1 plasmid, its ability to target mitochondria was tested in Hela cells using the mitochondrial marker Rhodamine 800 for staining. Colocalization experiments showed a high degree of overlap between the fluorescence signal of MPI-1 and the signal from Rhodamine 800, indicating that MPI-1 was successfully localized to the mitochondria (Figure 3).
Discussion
The mitochondrial membrane voltage is maintained at -120--180 mV under resting conditions and fluctuates with changes in metabolic state. Currently, measurement of mitochondrial membrane potential can be performed using electrophysiological methods and fluorescence dye methods. Mitochondrial patch clamping requires the isolation of mitochondria and the destruction of cellular structures13. This approach may lead to measurements that deviate from physiological conditions. The fluorescence probe met...
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the support of the National Natural Science Foundation (NSF) of China: JSK (32071137 and 92054103) and Funding for the Scientific Research and Innovation Team of The First Affiliated Hospital of Zhengzhou University: JSK (ZYCXTD2023014).
Materials
| Name | Company | Catalog Number | Comments |
| BamHI | Thermo | FD0054 | |
| Calcium chloride | Sigma | C4901 | |
| CCCP | Sigma | C2759 | |
| Centrifuge | eppendorf | 5430R | |
| Centrifuge (cell culture) | eppendorf | 5810R | |
| CO2 cell incubator | ESCO | 170L IR Sensor | |
| Coverslips | Glaswarenfabrik Karl Hecht GmbH & Co.KG | 92100100030 | |
| Dental adhensive resin cement | Sun medical company, LTD. | Super-Bond C&B Kit | |
| D-glucose | Sigma | G7021 | |
| DNA Ligation Kit Ver. 2.1 | Takara | 6022 | |
| Dulbecco's modified Eagle medium | Gibco | 11965092 | |
| Electric drill | RWD Instruments | 78001 | |
| Fetal bovine serum | Gibco | A5670701 | |
| Fiber optic cannula | RWD Instruments | R-FOC-L200C-39NA | |
| Fiber photometry detector | Thinker | QAXK_FPS-TC-MC-LED | |
| Fluorescence microscope | Olympus | IX83 | |
| Glass pipette (for injection) | Drummond Scientific company | 3.5" Drummond # 3-000-203-G/X | |
| HEK293t | ATCC | Cat# CRL-3216 | |
| Hela cells | ATCC | Cat# CCL-2 | |
| HEPES | Sigma | H3375 | |
| Injection pump | Drummond Scientific company | 3-000-207 | |
| Isoflurane | RWD Instruments | R510-22 | |
| Laser scanning confocal microscope | Zeiss | LSM980 | |
| MluI | Thermo | FD0564 | |
| NheI | Thermo | FD0974 | |
| Optical fibers | RWD Instruments | R-FC-L-N3-200-L1 | |
| Paraffin oil | Sangon | B500301 | |
| PCR thermal Cycler | analytik jena | Biometra Tone 96G | |
| Pentobarbital sodium | Sinopharm Chemical Reagent Co.LTD | 57-33-0 | |
| Potassium chloride | Sigma | P5405 | |
| PrimeSTAR HS DNA Polymerase | Takara | R010A | |
| Programmable micropipette puller | Sutter Instruments | P2000 | |
| Quick self-curing acrylic resin | Yamahachi | V-PINK | |
| Real-time PCR thermal Cycler | analytik jena | qTOWER³ auto | |
| Rhodamine 800 | Sigma | 83701 | |
| SalI | Thermo | FD0644 | |
| Sodium chloride | Sigma | S9888 | |
| Sodium phosphate dibasic | Sigma | S9763 | |
| Stereotaxic apparatus | RWD Instruments | E06354 | |
| Veterinary ophthalmic ointment | Puralube | NA | |
| XhoI | Thermo | FD0694 |
References
- Vyas, S., Zaganjor, E., Haigis, M. C. Mitochondria and cancer. Cell. 166 (3), 555-566 (2016).
- Dzbek, J., Korzeniewski, B. Control over the contribution of the mitochondrial membrane potential (ΔΨ) and proton gradient (ΔpH) to the protonmotive force (Δp): IN SILICO STUDIES. J Biol Chem. 283 (48), 33232-33239 (2008).
- O'Rourke, B., Cortassa, S., Aon, M. A. Mitochondrial ion channels: Gatekeepers of life and death. Physiology. 20 (5), 303-315 (2005).
- Chan, D. C. Mitochondrial fusion and fission in mammals. Annu Rev Cell Dev Biol. 22 (1), 79-99 (2006).
- Jin, S. M., Lazarou, M., Wang, C., Kane, L. A., Narendra, D. P., Youle, R. J. Mitochondrial membrane potential regulates PINK1 import and proteolytic destabilization by PARL. J Cell Biol. 191 (5), 933-942 (2010).
- Ly, J. D., Grubb, D. R., Lawen, A. The mitochondrial membrane potential (deltapsi(m)) in apoptosis; an update. Apoptosis. 8 (2), 115-128 (2003).
- Perry, S. W., Norman, J. P., Barbieri, J., Brown, E. B., Gelbard, H. A. Mitochondrial membrane potential probes and the proton gradient: a practical usage guide. Biotechniques. 50 (2), 98-115 (2011).
- Meng, X. -. Y., et al. A sensitive mitochondrial thermometry 2.0 and the availability of thermogenic capacity of brown adipocyte. Front Physiol. 13, 977431 (2022).
- Kaestner, L., et al. Genetically encoded voltage indicators in circulation research. Int J Mol Sci. 16 (9), 21626-21642 (2015).
- Yang, H. H., St-Pierre, F. Genetically encoded voltage indicators: Opportunities and challenges. J Neurosci. 36 (39), 9977 (2016).
- Sepehri Rad, M., Cohen, L. B., Braubach, O., Baker, B. J. Monitoring voltage fluctuations of intracellular membranes. Sci Rep. 8 (1), 6911 (2018).
- Yang, R. -. Z., Wang, D. -. D., Li, S. -. M., Liu, P. -. P., Kang, J. -. S. Development and application of a mitochondrial genetically encoded voltage indicator in narcosis. Neurosci Bull. 40 (10), 1529-1544 (2024).
- Kumari, A., Nguyen, D. M., Garg, V. Patch-clamp technique to study mitochondrial membrane biophysics. J Gen Physiol. 155 (8), e202313347 (2023).
- Bayne, A. N., Dong, J., Amiri, S., Farhan, S. M. K., Trempe, J. -. F. MTSviewer: A database to visualize mitochondrial targeting sequences, cleavage sites, and mutations on protein structures. PLoS One. 18 (4), e0284541 (2023).
- Kim, B. B., et al. A red fluorescent protein with improved monomericity enables ratiometric voltage imaging with ASAP3. Sci Rep. 12 (1), 3678 (2022).
- Evans, S. W., et al. A positively tuned voltage indicator for extended electrical recordings in the brain. Nat Methods. 20 (7), 1104-1113 (2023).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved