Aby wyświetlić tę treść, wymagana jest subskrypcja JoVE. Zaloguj się lub rozpocznij bezpłatny okres próbny.
Method Article
Study of Dendritic Cell Development by Short Hairpin RNA-Mediated Gene Knockdown in a Hematopoietic Stem and Progenitor Cell Line In vitro
W tym Artykule
Podsumowanie
Here we provide a protocol for screening potential transcription factors involved in the development of dendritic cell (DC) using lentiviral transduction of shRNA to obtain stable knockdown cell lines for in vitro DC differentiation.
Streszczenie
Dendritic cells (DCs) are important antigen-presenting cells that connect innate and adaptive immune responses. DCs are heterogeneous and can be divided into conventional DCs (cDCs) and plasmacytoid DCs (pDCs). cDCs specializes in presenting antigens to and activate naïve T cells. On the other hand, pDCs can produce large quantities of type I interferons (IFN-I) during viral infection. The specification of DCs occurs at an early stage of DC progenitors in the bone marrow (BM) and is defined by a network of transcription factors (TFs). For example, cDCs highly express ID2, while pDCs highly express E2-2. Since more and more subsets of DCs are being identified, there is a growing interest in understanding specific TFs controlling DC development. Here, we establish a method to screen TFs critical for DCs differentiation in vitro by delivering lentivirus carrying short hairpin RNA (shRNA) into an immortalized hematopoietic stem and progenitor cell (iHSPCs) line. After the selection and in vitro differentiation, cDC and pDC potential of the stable knockdown cell lines are analyzed by flow cytometry. This approach provides a platform to identify genes potentially governing DC fates from progenitors in vitro.
Wprowadzenie
DCs are key regulators of innate and adaptive immunity1. DCs are mainly classified into two functionally distinct populations, namely pDCs and cDCs. Moreover, cDCs comprise two subsets, namely, type I and type II cDCs or cDC1s and cDC2s, respectively2. pDCs, expressing BST2, Siglec-H, and intermediate levels of CD11c in mice3,4, are the cells that can secrete large amounts of IFN-I during inflammation and viral infection5. Due to their robust IFN-I-producing ability, they are also suspected to play a key role in the progression of autoimmune diseases, including systemic lupus erythematosus (SLE)6. cDC1s, defined by the surface expression of XCR1, CD8a, CLEC9A, and CD103 in mice7, are specialized in the activation and polarization of cytotoxic CD8+ T cells (CTLs) through the antigen cross-presentation, thereby initiating type I immunity in response to intracellular pathogens and cancer8,9. On the other hand, cDC2s, expressing CD11b and CD172α (also known as Sirpα) in both humans and mice, can activate CD4+ T cells and promote type II immune response against allergen and parasites10, as well as modulate type III immunity following extracellular bacteria and microbiota recognition11,12.
Diversification of DCs is determined by a group of TFs from hematopoietic stem and progenitor cells (HSPCs) in the BM. E2-2 (encoded by Tcf4) is a master regulator for differentiation and function of pDCs13,14. In contrast, the inhibitor of DNA binding 2 (ID2) drives cDC specification and inhibits pDC development through blocking E protein activity15. Moreover, the development of cDC1s requires IRF8 and BATF3, while differentiation of cDC2s highly depends on IRF416. Recent works have explored the heterogeneity of pDCs17 and cDCs and their transcriptional regulation18. Because of the complexity of DC network, there is an unmet need to establish a platform to identify other TFs controlling the development and functionality of DCs.
Here, we used an iHSPC that was generated by expressing estrogen-regulated nuclear translocation of Hoxb8 in BM cells (also referred to as Hoxb8-FL cells)19. iHSPCs can proliferate and remain in an undifferentiated stage in the presence of β-estradiol and Flt3 ligand (FL), whereas they start to differentiate into different DC types in the presence of FL upon withdrawal of β-estradiol19. Based on this feature, we can knock down genes of interest at the progenitor stage, followed by examining the effect on in vitro differentiation of pDCs and cDCs. Therefore, this method is a powerful tool to discover the genes that regulate the development and function of DCs.
Protokół
Handling of lentivirus is performed as per the regulation of the Department of Environmental Health and Safety of National Taiwan University College of Medicine.
1. Preparation of immortalized hematopoietic stem and progenitor cell lines (iHSPCs)
- Maintain iHSPC cell line in complete RPMI 1640 medium containing 100 ng/mL FL and 1 µM β-estradiol.
- Passage the cells at a ratio of 1:10 every 3 days.
NOTE: Make complete RPMI 1640 medium by supplementing with 10% fetal bovine serum (FBS), 5 x 10-5 M β-mercaptoethanol, and 10 µg/mL gentamicin. Recombinant murine FL is also commercially available.
2. Lentiviral transduction
- Plate iHSPCs at a density of 1 x 105 cells/well in 12-well plates in 1 mL of complete medium containing 100 ng/mL FL, 1 µM β-estradiol, and 8 µg/mL polybrene.
NOTE: The concentration of polybrene depends on the cell types and is usually in the range of 4-8 µg/mL. - Add shRNA-carrying lentivirus in each well at a multiplicity of infection (MOI) of 100.
NOTE: The lentiviral vector is pLKO.1-Puro with a puromycin selection marker (Figure 1).The target sequences of shRNAs against LacZ, Tcf4, and Id2, respectively, are listed in the Table of Materials. The MOI is defined by the number of virions that are added per cell during infection.
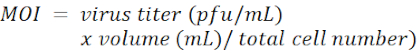
- Spin the plates at 1,100 x g for 90 min at 37 °C.
NOTE: Spin infection is carried out using a swing bucket rotor. - Incubate the plate containing infected cells for overnight at 37 °C in an incubator.
NOTE: If the cells are sensitive to polybrene, then refresh the cells with the complete medium without polybrene after the spin infection. - Refresh cells with complete medium containing 100 ng/mL FL and 1 µM β-estradiol 24 h after the infection.
- Add 6 µg/mL of puromycin to the medium to select the infected cells after an additional 24 h.
NOTE: Transduced lentiviral vector usually takes 48 h for expressing genes, including puromycin resistant gene. Ensure that the concentration of puromycin is optimized for each cell line. - Refresh the selection medium containing 100 ng/mL FL, 1 µM β-estradiol, and 6 µg/mL puromycin every 3 days and maintain the cells for at least one week to expand the stably transduced iHSPC cells.
NOTE: Puromycin selection usually takes effect 48 h later, and the period of selection depends on cell types.
3. Measurement of knockdown efficiency by reverse transcription and real-time PCR (RT-PCR)
- Extract total RNA from 1 x 107 shLacZ, shTcf4, and shId2 stable knockdown iHSPC cells using commercial RNA extraction reagent and precipitate RNA from the aqueous layer with isopropanol, followed by washing the RNA precipitate with 75% ethanol.
- Dissolve the RNA (~ 5 µg) with 5 µL of DEPC-treated H2O and adjust the concentration to 1 µg/µL.
- Take 1-3 µg of RNA, mix with DEPC-treated H2O to a final volume of 17.4 µL, add 1 µL of 1 unit/µL RNase-free DNase I and incubate for 20 min at 37 °C.
NOTE: This step is to digest the genomic DNA in the RNA samples. - Add 1 µL of 20 mM EDTA to the RNA samples, incubate at 65 °C for 10 min to inactivate DNase I, and put the RNA samples immediately at 4 °C.
- Add 11.6 µL of the reaction mix containing 1 µL of oligo (dT) primers (45 µM), 6 µL of 5x 1st strand buffer, 3 µL of dNTP (2 mM), 0.6 µL of RNase inhibitor (50 Unit/µL) and 1 µL of reverse transcriptase (200 Unit/µL) to the RNA samples and incubate at 40 °C for 1 h.
- Stop the reaction by heating at 70 °C for 10 min and dilute the reaction mix with 30 µL of H2O.
- Take 2 µL of the diluted RT reaction mix as a DNA template and PCR amplify it using primers against Tcf4 or Id2 (see Table 1 for thermocycler conditions).
NOTE: The primer sequences are included in the Table of Materials.
4. In vitro differentiation of the stable knockdown iHSPC cell lines
- Maintain the stable single knockdown of LacZ (shLacZ), Tcf4 (shTcf4), or Id2 (shId2) in iHSPCs in complete medium containing 100 ng/mL of FL and 1 µM β-estradiol.
- Collect shLacZ, shTcf4, and shId2 un-differentiated iHSPCs into a 15 mL tube and centrifuge for 5 min at 500 x g to pellet the cells.
- Discard the supernatant and add 10 mL of PBS to wash the cells. Repeat this step twice.
- Resuspend and seed the iHSPC cells at a density of 2 x 105 cells/mL into a 12-well plate in 1 mL of complete medium containing 100 ng/mL FL only.
- Add 1 mL of fresh complete medium containing 100 ng/mL FL three days later.
- Analyze the differentiated cells (shLacZ, shTcf4, and shId2 iHSPCs) by flow cytometry two days later.
5. Flow cytometric analysis of the differentiated DCs
- Collect the cells into 1.5 mL tubes by pipetting the cells up and down 2-3 times in the plate, and centrifuge for 5 min at 500 x g at room temperature (RT).
NOTE: In vitro differentiated DCs will slightly attach to the plates. Gentle pipetting will help recover DCs from the plates. - Discard the supernatant and resuspend the cells in 50 µL of FACS buffer. Next, add 50 µL of anti-CD16/32 hybridoma supernatant and incubate for 5-10 min on ice.
NOTE: The Fc blocking antibody prevents nonspecific binding of antibodies to Fc receptors expressing on some myeloid cells and B cells and is commercially available through various vendors. - Add fluorescent dye-conjugated antibodies (0.04 µg for each antibody) directly to the cells and incubate for 15 min on ice in the dark. The antibodies used are APC/cy7 anti-mouse CD11c, FITC anti-mouse CD11b, and PE anti-mouse B220.
NOTE: pDCs are defined as CD11c+CD11b-B220+, and cDCs are defined as CD11c+CD11b+B220-. - Wash cells with 1 mL of FACS buffer and centrifuge for 5 min at 500 x g at RT.
- Resuspend the cells in 100 µL of FACS buffer and analyze by flow cytometry.
Wyniki
The map of lentiviral vector pLKO.1-Puro is shown (Figure 1). After the delivery of lentivirus expressing shRNA against LacZ (a non-targeting control), Tcf4, and Id2 in iHSPCs, the knockdown efficiency confirmed by RT-qPCR revealed that the expression of Tcf4 was reduced in shTcf4 iHSPCs, compared to shLacZ iHSPCs (Figure 2A). On the other hand, the decreased expression of Id2 was also observed in sh...
Dyskusje
Lentivirus-based shRNA vectors are often used for gene silencing by viral transduction into cells and permit stable integration into the host genome. However, various transduction efficiency in different cell types needs to be considered, and a number of approaches have been taken to overcome this problem.
Polybrene is a polycationic polymer that can neutralize the charges on the cell membrane, thereby enhancing the binding of the virion to the cells during transduction20
Ujawnienia
The authors have nothing to disclose.
Podziękowania
We are grateful for technical support from Dr. Tz-Ling Chen. We thank the National RNAi Core Facility (Academia Sinica, Taiwan) for providing shRNA lentivirus (http://rnai.genmed.sinica.edu.tw). This work was supported by the Ministry of Science and Technology, Taiwan (MOST 108-2320-B-002-037-MY3 and MOST 109-2320-B-002-054-MY3).
Materiały
| Name | Company | Catalog Number | Comments |
| Antibodies | |||
| APC/Cy7 anti-mouse CD11c Antibody | Biolegend | 117324 | (Clone: N418) |
| FITC anti-mouse/human CD11b Antibody | Biolegend | 101206 | (Clone: M1/70) |
| PE anti-mouse/human B220 Antibody | Biolegend | 103208 | (Clone: RA3-6B2) |
| Cell culture | |||
| 1.5 mL Micro tube | ExtraGene | TUBE-170-C | |
| 12-well tissue culture-treated plate | Falcon | 353043 | |
| Fetal bovine serum (FBS) | Corning | 35-010-CV | |
| RPMI 1640 medium | gibco | 11875-085 | |
| Reagent | |||
| β-estradiol | Sigma-Aldrich | E2758-250MG | |
| β-mercaptoethanol (β-ME) | Sigma-Aldrich | M6250 | |
| FACS buffer | home-made | Formula: 1xPBS+0.5 %FBS+0.1%NaN3 | |
| Flt3 ligand (FL) | home-made | ||
| Polybrene | Sigma-Aldrich | TR-1003-G | |
| Puromycin | Invivogen | ant-pr-1 | |
| TRIsure | BIOLINE | BIO-38032 | |
| shRNA (Taregt sequence/clone ID) | Company | ||
| shId2 (GCTTATGTCGAATGATAGCAA/TRCN0000054390) | The RNAi Consortium (TRC) | ||
| shLacZ (CGCGATCGTAATCACCCGAGT/TRCN0000072224) | The RNAi Consortium (TRC) | ||
| shTcf4 (GCTGAGTGATTTACTGGATTT/TRCN0000012094) | The RNAi Consortium (TRC) | ||
Odniesienia
- Steinman, R. M., Witmer, M. D. Lymphoid dendritic cells are potent stimulators of the primary mixed leukocyte reaction in mice. Proceedings of the National Academy of Sciences. 75 (10), 5132-5136 (1978).
- Guilliams, M., et al. Dendritic cells, monocytes and macrophages: a unified nomenclature based on ontogeny. Nature Reviews Immunology. 14 (8), 571-578 (2014).
- Blasius, A. L., Cella, M., Maldonado, J., Takai, T., Colonna, M. Siglec-H is an IPC-specific receptor that modulates type I IFN secretion through DAP12. Blood. 107 (6), 2474-2476 (2006).
- Swiecki, M., Colonna, M. Unraveling the functions of plasmacytoid dendritic cells during viral infections, autoimmunity, and tolerance. Immunology Reviews. 234 (1), 142-162 (2010).
- Liu, Y. -. J. IPC: Professional Type 1 interferon-producing cells and plasmacytoid dendritic cell precursors. Annual Review of Immunology. 23 (1), 275-306 (2005).
- Panda, S. K., Kolbeck, R., Sanjuan, M. A. Plasmacytoid dendritic cells in autoimmunity. Current Opinion in Immunology. 44, 20-25 (2017).
- Durai, V., Murphy, K. M. Functions of murine dendritic cells. Immunity. 45 (4), 719-736 (2016).
- Mashayekhi, M., et al. CD8α(+) dendritic cells are the critical source of interleukin-12 that controls acute infection by Toxoplasma gondii tachyzoites. Immunity. 35 (2), 249-259 (2011).
- Anderson, D. A., Dutertre, C. -. A., Ginhoux, F., Murphy, K. M. Genetic models of human and mouse dendritic cell development and function. Nature Reviews Immunology. 21 (2), 101-115 (2021).
- Plantinga, M., et al. Conventional and monocyte-derived CD11b(+) dendritic cells initiate and maintain T helper 2 cell-mediated immunity to house dust mite allergen. Immunity. 38 (2), 322-335 (2013).
- Persson, E. K., et al. IRF4 transcription-factor-dependent CD103(+)CD11b(+) dendritic cells drive mucosal T helper 17 cell differentiation. Immunity. 38 (5), 958-969 (2013).
- Kumamoto, Y., et al. CD301b+ dermal dendritic cells drive T helper 2 cell-mediated immunity. Immunity. 39 (4), 733-743 (2013).
- Cisse, B., et al. Transcription factor E2-2 is an essential and specific regulator of plasmacytoid dendritic cell development. Cell. 135 (1), 37-48 (2008).
- Grajkowska, L. T., et al. Isoform-specific expression and feedback regulation of E protein TCF4 control dendritic cell lineage specification. Immunity. 46 (1), 65-77 (2017).
- Jackson, J. T., et al. Id2 expression delineates differential checkpoints in the genetic program of CD8α+ and CD103+ dendritic cell lineages. The EMBO Journal. 30 (13), 2690-2704 (2011).
- Suzuki, S., et al. Critical roles of interferon regulatory factor 4 in CD11bhighCD8alpha- dendritic cell development. Proceedings of the National Academy of Science U. S. A. 101 (24), 8981-8986 (2004).
- Rodrigues, P. F., et al. Distinct progenitor lineages contribute to the heterogeneity of plasmacytoid dendritic cells. Nature Immunology. 19 (7), 711-722 (2018).
- Brown, C. C., et al. Transcriptional basis of mouse and human dendritic cell heterogeneity. Cell. 179 (4), 846-863 (2019).
- Redecke, V., et al. Hematopoietic progenitor cell lines with myeloid and lymphoid potential. Nature Methods. 10 (8), 795-803 (2013).
- Abe, A., Miyanohara, A., Friedmann, T. Polybrene increases the efficiency of gene transfer by lipofection. Gene Therapy. 5 (5), 708-711 (1998).
- Sorgi, F. L., Bhattacharya, S., Huang, L. Protamine sulfate enhances lipid-mediated gene transfer. Gene Therapy. 4 (9), 961-968 (1997).
- Kirkling, M. E., et al. Notch signaling facilitates in vitro generation of cross-presenting classical dendritic cells. Cell Reports. 23 (12), 3658-3672 (2018).
- Pearce, E. J., Everts, B. Dendritic cell metabolism. Nature Reviews. Immunology. 15 (1), 18-29 (2015).
- Wculek, S. K., Khouili, S. C., Priego, E., Heras-Murillo, I., Sancho, D. Metabolic Control of Dendritic Cell Functions: Digesting Information. Frontiers in Immunology. 10 (775), (2019).
- Saxton, R. A., Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell. 168 (6), 960-976 (2017).
Przedruki i uprawnienia
Zapytaj o uprawnienia na użycie tekstu lub obrazów z tego artykułu JoVE
Zapytaj o uprawnieniaThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. Wszelkie prawa zastrzeżone