A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
An Iodide-Yellow Fluorescent Protein-Gap Junction-Intercellular Communication Assay
In This Article
Summary
Here, we present a protocol for a novel gap junction intercellular communication assay designed for the high-throughput screening of gap junction-modulating chemicals for drug discovery and toxicological assessment.
Abstract
Gap junctions (GJs) are cell membrane channels that allow diffusion of molecules smaller than 1 kDa between adjacent cells. As they have physiological and pathological roles, there is need of high-throughput screening (HTS) assays to identify GJ modulators in drug discovery and toxicology assays. A novel iodide-yellow fluorescent protein-gap junction-intercellular communication (I-YFP-GJIC) assay fulfills this need. It is a cell-based assay including acceptor and donor cells that are engineered to stably express a yellow fluorescent protein (YFP) variant, whose fluorescence is sensitively quenched by iodide, or SLC26A4, an iodide transporter, respectively. When iodide is added to a mixed culture of the two cell types, they enter the donor cells via the SLC26A4 transporter and diffuse to the adjacent acceptor cells via GJs where they quench the YFP fluorescence. YFP fluorescence is measured well by well in a kinetic mode. The YFP quenching rate reflects GJ activity. The assay is reliable and rapid enough to be used for HTS. The protocol for the I-YFP-GJIC assay using the LN215 cells, human glioma cells, is described.
Introduction
Gap junctions (GJs) act as intercellular channels to allow the diffusion of small molecules of <1 kDa such as nutrients, metabolites, and signaling molecules between adjacent cells. The junctional elements include a hemichannel or connexon in each cell, and each connexon constitutes six connexins (Cxs)1. GJs and Cxs have been used in toxicology assays of carcinogens such as polycyclic aromatic hydrocarbons (PAH), which are GJ inhibitors2,3,4. Disrupted GJIC has been associated with nongenotoxic carcinogenesis5,6. As a potential therapeutic target, GJ involvement has been reported in particular subtypes of seizures7,8, protection from cardiac and brain ischemia/reperfusion injury9, migraine with aura10, drug-induced liver injury6,11, and wound healing12. High-throughput screening (HTS) assays are required to identify GJ-modulating chemicals or antibodies for drug discovery, for toxicology assays, and to identify novel cellular regulators of GJ activity. HTS assays can also be used to investigate structure-activity relationships of GJ modulators2,13,14,15.
Some GJIC assays include dye transfer or dual patch clamp techniques. Lucifer yellow CH (LY) and calcein acetoxymethyl ester (calcein-AM) have been used in dye-transfer assays. Cells are not permeable to LY, which is introduced by microinjection, scrape loading, or electroporation. Once inside the cell, LY spreads into neighboring cells via GJs and GJ activity is assayed by the extent of the LY migration16. Calcein-AM assays usually involve gap-fluorescence recovery after photobleaching17,18. Calcein-AM is a cell-permeant dye that is converted intracellularly into impermeable calcein by an intrinsic esterase. The assay requires a confocal microscope to observe the transfer of calcein-AM into a cell from those surrounding it following laser photobleaching. If functional GJs are present, calcein-AM in adjacent cells enters the photobleached cells and the fluorescence is recovered. GJ activity is assayed by the extent of fluorescence recovery of the photobleached cells. Dye-transfer assays are laborious and time consuming or have low sensitivities. Dual patch clamping is an electrophysiological method that measures junctional conductance. It is relatively sensitive, with a direct dependence of conductance on the number of open GJs19; however, it is technically demanding, time consuming, and expensive20. The I-YFP-GJIC assay was developed for use in HTS.
Figure 1 illustrates the components and steps of the I-YFP GJIC assay, which utilizes acceptor cells expressing an iodide-sensitive YFP variant bearing H148Q and I152L (YFPQL) and donor cells expressing an iodide transporter (SLC26A4)21. The two mutations carried by YFPQL allow quenching of fluorescence by iodide22. Iodides are added to co-cultured acceptor and donor cells; they do not enter the acceptor cells, but are taken up by the SLC26A4 transporters present on the donor cells. Iodides in the donor cells diffuse through functioning GJs into adjacent acceptor cells where they quench the YFPQL fluorescence. If GJs are closed or blocked by inhibitors, iodide cannot enter the acceptor cells to quench the fluorescence. The YFPQL quenching rate reflects GJ activity. The I-YFP GJIC assay procedure is neither complicated nor time consuming. It is compatible with HTS and can be used to test the effects of a large number of compounds on GJ activity in a relatively short period. It requires only acceptor and donor cells, and two balanced salt solutions. The protocol described below is based on LN215 cells whose major Cx is Cx4321. The LN215-YFPQL receptor and LN215-I− donor cells were generated by transduction with lentiviruses expressing YFPQL or SLC26A421,23.
Access restricted. Please log in or start a trial to view this content.
Protocol
1. Generation of lentiviruses expressing YFPQL and SLC26A4
- Grow HEK293T human embryonic kidney cells to 80% confluency on 100 mm culture plates. Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum, 100 U/mL penicillin, and 100 µg/mL streptomycin is the culture medium used throughout the protocol to maintain HEK293T and other cells mentioned below.
- Coat 6-well culture plates by adding 2 mL of 0.005% of sterile poly-L-lysine (PLL) solution to each well for 10 min. Aspirate the PLL solution and rinse the surface twice with 2 mL of sterile water.
- Wash the HEK293T cells with 10 mL of phosphate buffered saline (PBS) and treat the cell monolayers in each 100 mm dish with 2 mL of 0.25% trypsin-EDTA solution at 37 °C for 3 min. Add 5 mL of culture medium and resuspend the cells.
- Count the cells in a hemocytometer and adjust the density of the cell suspension to 250,000 cells/mL in culture medium and add 500,000 cells in 2 mL of culture medium to each lysine-coated well of 6-well plates. Incubate the cells in a humidified 5% CO2/95% air atmosphere at 37 °C for 24 h and then replace the culture medium with DMEM without penicillin or streptomycin.
- In a 1.5 mL tube, dilute 20 µL of transfection reagent with 500 µL of DMEM without serum or antibiotics. Mix gently by pipetting and let stand at room temperature for 5 min.
- Meanwhile, pipette 250 µL of DMEM into each of two 1.5 mL tubes and then add 1500 ng of pLVX-EIP-YFPQL or pLenti6P-SLC26A4, 1225 ng of psPAX2 and 375 ng of pMD2.G to each. The two lentiviral plasmids have been previously described21. Add 250 µL of diluted transfection reagent to each plasmid tube, mix gently and incubate for 20 min at room temperature.
- After 20 min, add 500 µL of transfection reagent and plasmid complexes in the 1.5 mL tubes dropwise to each culture plate well in step 1.4 and mix by rocking the plate back and forth. Incubate cells at 37 °C in a CO2 incubator for 12 h.
- Replace the medium with 2.5 mL of fresh medium and incubate for an additional 48 h. Then place the culture plate on ice for 5 min to keep the conditioned medium containing lentivirus chilled to maintain infectivity.
- Harvest the media containing lentiviruses and transfer to 15 mL conical tubes. Centrifuge at 3,000 x g at 4 °C for 3 min and then remove floating HEK293T cells from the supernatant by filtration at 0.4 µm.
- Store the media containing lentiviruses at 4 °C for use within 2 days. For later use, store 200 µL aliquots at −80 °C.
2. Generation of LN215-YFPQL and LN215-I− cells by lentiviral transduction
- Grow LN215 cells in 100 mm culture plates to 80% confluency in DMEM supplemented with 10% fetal bovine serum (FBS) 100 U/mL penicillin, and 100 µg/mL streptomycin as described above.
NOTE: If the I-YFP-GJIC assay is conducted using a different cell line, use the appropriate culture medium. LN215-YFPQL, and LN215-I- cells can be provided by the University-Industry foundation, Yonsei University. Please contact the corresponding author. - One day before transduction, wash the cells twice with 10 mL of PBS, treat with 2 mL of 0.25% trypsin-EDTA at 37 °C for 3 min. Resuspend the cells in 5 mL of culture medium with a 10 mL serologic pipette and adjust the density to 50,000 cells/mL. Add 20,000 cells in 400 µL of media to each well of a 24-well culture plate for treatment as no virus control, YFPQL, and SLC26A4 cells.
- After 24 h of incubation at 37 °C, transduce two wells by replacing the culture medium with 400 µL of a 1:1 mixture of pLVX-EIP-YFPQL or pLenti6P-SLC26A4 lentivirus and fresh culture medium supplemented with polybrene at a final concentration of 4 µg/mL. For no virus controls, replace with culture medium.
- Incubate the cells at 37 °C for 15 h, aspirate the medium containing lentiviruses, add fresh culture medium, and incubate the cells for an additional 72 h.
CAUTION: To prevent contamination of lentivirus between wells, use new tips or pipets for each well when you aspirate culture medium containing lentivirus or dispense fresh growth medium. - Wash the cells in each well twice with 0.5 mL of PBS, treat with 300 µL of trypsin-EDTA for 3 min. Resuspend the cells in 2 mL of culture medium and plate in six-well culture plates with 2 µg/mL puromycin.
- Culture the cells in media containing 2 µg/mL puromycin until all cells in the control well are dead (round-shaped or floating when observed in microscope), which usually takes a week. Refresh the culture media containing puromycin every other day during the selection period. If LN215-YFPQL or LN215-I- cultures become confluent before selection has completed, transfer the cells to 100 mm plate and continue selection as in step 2.5.
3. Preparation of solutions required for the assay
- Prepare 500 mL of C-solution (10 mM HEPES, 140 mM NaCl, 10 mM glucose, 5 mM KCl, 1 mM MgCl2, and 1 mM CaCl2) and 500 mL of I-solution (10 mM HEPES, 140 mM NaI, 10 mM glucose, 5 mM KCl, and 1 mM CaCl2).
- Adjust the pH of both solutions to 7.4 with 1 N NaOH, sterilize the solutions by filtrations at 0.4 µm for storage. Store at 4 °C for up to 1 month. Check the pH before using.
4. Plating the LN215-YFPQL and LN215-I− cells
- Culture LN215-YFPQL and LN215-I− cells in 100 mm plates separately in culture medium to reach the populations required for assay. LN215-YFPQL and LN215-I− cells in 40% and 80% confluency in 100 mm plates, respectively, are sufficient for a 96-well plate assay.
- One day before conducting the I-YFP GJIC assay, wash each 100 mm culture plate with 10 mL of PBS. Treat each plate with 2 mL of 0.25% trypsin-EDTA solution and incubate at 37 °C for 5 min. Resuspend the cells in each plate in 4 mL of culture medium and transfer to 15 mL conical tubes.
- Pellet the cells by centrifugation at 1,000 x g for 3 min. Discard the supernatant and resuspend each cell pellet with 5 mL of culture medium. Break up any cell clumps into single cells by pipetting up and down about 20 times with a 10 mL serological pipette.
- Count the cells in a hemocytometer, and dilute the cells in the culture medium to make cell suspensions of LN215-YFPQL at 80,000 cells/mL and LN215-I− at 160,000 cells/mL.
- Mix 7 mL of LN215-YFPQL and 7 mL of LN215-I− cell suspensions in a reservoir. Add 100 µL of the mixture to each well of a 96-well cell culture plate using a multichannel pipette.
NOTE: To add 100 µL of the mixture in each well of 96-well cell plate, about 10 mL of mixed cell suspension is needed. It is recommended to make more cell suspension than needed. - Incubate the cells in humidified 5% CO2/95% air at 37 °C for 24 h. The LN215-YFPQL and LN215-I−cell culture should be 100% confluent when the assay is conducted.
5. Conducting the I-YFP GJIC assay
NOTE: Use a fluorescence microscope with 20x magnification, and a GFP filter to check the 96-well plates to be sure there are no clumps of LN215-YFPQL or LN215-SLC26A4 cells and that the cell cultures are fully confluent and well distributed before conducting the assay.
- At least 30 min before doing the assay, turn on a microplate and set to 37 °C.
- Wash the tubing of an automated injector with 3 mL of 70% ethanol, 3 mL of distilled water, and then 3 mL of I-solution at a flow rate of 300 µL/s.
- Warm the C- and I-solutions to 37 °C in the water bath.
NOTE: As 100 µL of each solution is needed for each well of the 96-well plate, about 10 mL of each solution is needed for each assay. An additional 10 mL of the I-solution is needed for priming each plate (total of 20 mL) and an additional 25 mL of the C-solution is needed for washing each plate (total of 35 mL). - Aspirate the growth medium or invert the plate to empty it; tap out residual medium.
NOTE: Residual fetal bovine serum in the growth media causes background fluorescence and a decline in assay quality. - Add 200 µL of C-solution to each well from a reservoir using a multichannel pipette. Aspirate the C-solution or invert the plate to empty and tap out residual solution.
- Add 50 µL µL C-solution, 1 µL of 2.5 mM chemical stock (see Table 1) or dimethylsulfoxide (DMSO) as a vehicle, and then 50 µL µL C-solution to each well with a multichannel pipette.
NOTE: Most reagents in a chemical library are dissolved in DMSO and up to 1% (v/v) is allowed in most cell-based assays24. As DMSO has a higher density than water, reagents dissolved in DMSO tend to go down to the bottom when added to the culture plate wells, which disturbs the concentrations of the assay solutions. This can be circumvented by adding 50 µL of C-solution, reagents in DMSO, and 50 µL C of -solution in order. The last 50 µL of C-solution is for mixing. - Incubate the cells at 37 °C in air, not in 5% CO2. The incubation time can be modified, but 10 min is usually sufficient for ion-channel modulators to act.
- During incubation, set the microplate reader program to inject 100 µL of I-solution to each well at 1 s and to measure the fluorescence for 10 s at 0.4 s intervals. Set the reader to read fluorescence from the bottom. The recommended injection speed is 135 µL/s. Set the excitation wavelength to 485 nm and read the emission at 520 nm.
NOTE: Detailed settings for the microplate reader programs are as follows.- Click the Manage protocols button.
- Click the New Button. Select the Fluorescence Intensity in the measurement method section, and Well Mode in the reading mode section. Next, click the OK button. New tab will appear.
- In the Basic Parameters menu, set the excitation wavelength to 485 nm and the emission at 520 nm. Select the Bottom Optic to read fluorescence from the bottom. Set measurement start time to be "0 s", number of intervals to be "25", number of flashes per well, and interval to be "20", and interval time to be "0.4 s".
- In the Layout menu, draw region of the plate to be read.
- In the Concentrations/Volumes/Shacking menu, set the microplate reader to inject 100 µL of I-solution to each well with 135 µL/s injection speed.
NOTE: Avoid faster injection speeds because they can result in detachment of cells. - In the Injection time menu, set the injection start time to be 1 s. Then, click the Start Measurement button.
- After incubation, place the 96-well plates in the microplate reader and start the measurement by clicking Start Measurement button again.
6. Calculation of GJIC activity
- Calculate the percentages of YFPQL quenching and GJIC activity as23
YFPQL quenching (%) =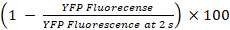
GJIC activity (%) =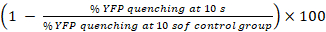
NOTE: In principle, GJIC activity should be calculated from the difference of the percentages of YFP fluorescence in wells with acceptor cells and donor cells and the corresponding acceptor-cell only wells. However, as LN215-YFPQL cells show negligible YFP quenching by iodide after 10 s, we do not take the background YFP quenching into account when conducting HTS using LN215-YFPQL and LN215-I- cells.
Access restricted. Please log in or start a trial to view this content.
Results
Twenty-nine 96-well culture plates were used to screen 2,320 chemicals to identify novel GJIC modulators by I-YFP GJIC assay using the LN215-YFPQL and LN215-I− cells. The results obtained with a representative plate are shown in Figure 2. The percentage of YFP fluorescence in each well is shown as a line graph in Figure 2A and the percentage of GJIC activity in each well is shown in th...
Access restricted. Please log in or start a trial to view this content.
Discussion
The I-YFP-GJIC assay can be used for HTS because it is robust, rapid, and inexpensive. An HTS assay is considered robust if the Z'-factor is above 0.525. See Zhang et al. for a description of the statistical analysis used to assessing the suitability of HTS assays25. When LN215 cells were used, the Z'-factor was >0.5 without any assay optimization. If other cell types are used in the assay and its Z'-factor is <0.5, the robustness can be improved by exte...
Access restricted. Please log in or start a trial to view this content.
Disclosures
The authors have no conflicts of interest to disclose.
Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2011-0023701, 2016R1D1A1A02937397, and 2018R1A6A1A03023718).
Access restricted. Please log in or start a trial to view this content.
Materials
| Name | Company | Catalog Number | Comments |
| 96-well plate | SPL | 30096 | |
| Calcium chloride (CaCl2) | Sigma | C5670 | I-solution |
| D-(+)-Glucose | Sigma | G7021 | C-solution, I-solution |
| Dimethyl sulfoxide (DMSO) | sigma | 276855 | |
| HEPES | Sigma | RES6003H-B7 | C-solution, I-solution |
| Lipofectamine 2000 | Invitrogen | 11668-027 | transfection reagent |
| Magnesium chloride hexahydrate (MgCl2 6H2O) | Sigma | M2393 | C-solution |
| Microplate reader | BMG LabTech | POLARstar Omega 415-1618 | |
| pMD2.G | Addgene | #12259 | |
| Polybrene | sigma | H9268 | |
| Poly-L-lysine solution | sigma | P4707 | |
| Potassium chloride (KCl) | Sigma | P5405 | C-solution, I-solution |
| psPAX2 | Addgene | #12260 | |
| Puromycin Dihydrochloride | sigma | P8833 | |
| Sodium chloride (NaCl) | Sigma | S5886 | C-solution, I-solution |
| Sodium hyroxide (NaOH) | Sigma | S2770 | |
| Sodium Iodide (NaI) | Sigma | 383112 | I-solution |
References
- Goodenough, D. a, Goliger, J. a, Paul, D. L. Connexins, connexons, and Intercellular Communication. Annual Review of Biochemistry. 65, 475-502 (1996).
- Upham, B. L., Weis, L. M., Trosko, J. E. Modulated Gap Junctional Intercellular Communication as a Biomarker of PAH Epigenetic Toxicity: Structure-Function Relationship. Environmental Health Perspectives. 106, 975(1998).
- U, J. E. T., Chang, C., Upham, B., Wilson, M. Epigenetic toxicology as toxicant-induced changes in intracellular signalling leading to altered gap junctional intercellular communication. Toxicology Letters. , 71-78 (1998).
- Yamasaki, H. Role of disrupted gap junctional intercellular communications in detection and characterization of carcinogens. Mutation Research - Reviews in Genetic Toxicology. , (1996).
- Yamasaki, H., et al. Gap junctional intercellular communication and cell proliferation during rat liver carcinogenesis. Environmental Health Perspectives. 101, suppl 5 191-197 (1993).
- Vinken, M., et al. Gap junctional intercellular communication as a target for liver toxicity and carcinogenicity. Critical Reviews in Biochemistry and Molecular Biology. 44, 201-222 (2009).
- Fonseca, C. G., Green, C. R., Nicholson, L. F. B. Upregulation in astrocytic connexin 43 gap junction levels may exacerbate generalized seizures in mesial temporal lobe epilepsy. Brain Research. 929 (1), 105-116 (2002).
- Garbelli, R., et al. Expression of connexin 43 in the human epileptic and drug-resistant cerebral cortex. Neurology. 76 (10), 895-902 (2011).
- Schulz, R., et al. Connexin 43 is an emerging therapeutic target in ischemia/reperfusion injury, cardioprotection and neuroprotection. Pharmacology and Therapeutics. 153, 90-106 (2015).
- Sarrouilhe, D., Dejean, C., Mesnil, M. Involvement of gap junction channels in the pathophysiology of migraine with aura. Frontiers in Physiology. , (2014).
- Patel, S. J., et al. Gap junction inhibition prevents drug-induced liver toxicity and fulminant hepatic failure. Nature Biotechnology. 30 (2), 179-183 (2012).
- Kandyba, E. E., Hodgins, M. B., Martin, P. E. A murine living skin equivalent amenable to live-cell imaging: analysis of the roles of connexins in the epidermis. Journal of Investigative Dermatology. 128 (4), 1039-1049 (2008).
- Upham, B. L., et al. Differential roles of 2 , 6 , and 8 carbon ceramides on the modulation of gap junctional communication and apoptosis during carcinogenesis. Cancer Letters. 191, 27-34 (2003).
- Upham, B. L., et al. Structure-activity-dependent regulation of cell communication by perfluorinated fatty acids using in vivo and in vitro model systems. Environmental Health Perspectives. 117 (4), 545-551 (2009).
- Weis, L. M., et al. Bay or Baylike Regions of Polycyclic Aromatic Hydrocarbons Were Potent Inhibitors of Gap Intercellular Communication. Environmental Health Perspectives. 106 (1), 17-22 (1998).
- el-Fouly, M. H., Trosko, J. E., Chang, C. C. Scrape-Loading and Dye Transfer: A Rapid and Simple Technique to Study Gap Junctional Intercellular Communication. Experimental Cell Research. 168 (2), 422-430 (1987).
- Wade, M. H., Trosko, J. E., Schindler, M. Photobleaching Assay of Gap Junction- Mediated Communication Between Human Cells. Advancement of Science. 232 (4749), 525-528 (2010).
- Abbaci, M., et al. In vitro characterization of gap junctional intercellular communication by gap-FRAP technique. Proceedings of SPIE. , 585909(2005).
- Neyton, J., Trautmann, A. Single-channel currents of an intercellular junction. Nature. 317 (6035), 331-335 (1985).
- Wilders, R., Jongsma, H. J. Limitations of the dual voltage clamp method in assaying conductance and kinetics of gap junction channels. Biophysical Journal. 63 (4), 942-953 (1992).
- Lee, J. Y., Choi, E. J., Lee, J. A new high-throughput screening-compatible gap junctional intercellular communication assay. BMC Biotechnology. 15 (1), 1-9 (2015).
- Galietta, L. J. V., Haggie, P. M., Verkman, A. S. Green fluorescent protein-based halide indicators with improved chloride and iodide affinities. FEBS Letters. 499 (3), 220-224 (2001).
- Lee, J. Y., Yoon, S. M., Choi, E. J., Lee, J. Terbinafine inhibits gap junctional intercellular communication. Toxicology and Applied Pharmacology. 307, 102-107 (2016).
- Hughes, J., Rees, S., Kalindjian, S., Philpott, K. Principles of early drug discovery. British Journal of Pharmacology. 162 (6), 1239-1249 (2011).
- Zhang, J. H., Chung, T. D. Y., Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. Journal of Biomolecular Screening. 4 (2), 67-73 (1999).
- Upham, B. L. Role of Integrative Signaling Through Gap Junctions in Toxicology. Current Protocols in Toxicology. , (2011).
- Schalper, K. A., et al. Modulation of gap junction channels and hemichannels by growth factors. Molecular BioSystems. 8 (3), 685-698 (2012).
- Kulkarni, G. V., Mcculloch, C. A. G. Serum deprivation induces apoptotic cell death in a subset of Balb / c 3T3 fibroblasts. Journal of Cell Science. 1179, 1169-1179 (1994).
- Harris, A. L., Locke, D. Permeability of Connexin Channels. Connexins. , 165-206 (2009).
- Choi, E. J., Yeo, J. H., Yoon, S. M., Lee, J. Gambogic Acid and Its Analogs Inhibit Gap. Junctional Intercellular Communication. 9, 1-10 (2018).
Access restricted. Please log in or start a trial to view this content.
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved