A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Multi-Fiber Photometry to Record Neural Activity in Freely-Moving Animals
In This Article
Summary
This protocol details how to implement and perform multi-fiber photometry recordings, how to correct for calcium-independent artifacts, and important considerations for dual-color photometry imaging.
Abstract
Recording the activity of a group of neurons in a freely-moving animal is a challenging undertaking. Moreover, as the brain is dissected into smaller and smaller functional subgroups, it becomes paramount to record from projections and/or genetically-defined subpopulations of neurons. Fiber photometry is an accessible and powerful approach that can overcome these challenges. By combining optical and genetic methodologies, neural activity can be measured in deep brain structures by expressing genetically-encoded calcium indicators, which translate neural activity into an optical signal that can be easily measured. The current protocol details the components of a multi-fiber photometry system, how to access deep brain structures to deliver and collect light, a method to account for motion artifacts, and how to process and analyze fluorescent signals. The protocol details experimental considerations when performing single and dual color imaging, from either single or multiple implanted optic fibers.
Introduction
The ability to correlate neural responses with specific aspects of an animal’s behavior is critical to understand the role a particular group of neurons plays in directing or responding to an action or stimulus. Given the complexity of animal behavior, with the myriad of internal states and external stimuli that can affect even the simplest of actions, recording a signal with single-trial resolution equips researchers with the necessary tools to overcome these limitations.
Fiber photometry has become the technique of choice for many researchers in the field of systems neuroscience because of its relative simplicity compared to other in vivo recording techniques, its high signal-to-noise ratio, and the ability to record in a variety of behavioral paradigms1,2,3,4,5,6,7,8. Unlike traditional electrophysiological methods, photometry is the optical approach most commonly used in conjunction with genetically-encoded calcium indicators (GECIs, the GCaMP series)9. GECIs change their ability to fluoresce based on whether or not they are bound to calcium. Because the internal concentration of calcium in neurons is very tightly regulated and voltage-gated calcium channels open when a neuron fires an action potential, transient increases in internal calcium concentration, which result in transient increases in the ability of a GECI to fluorescence, can be a good proxy for neuronal firing9.
With fiber photometry, excitation light is directed down a thin, multimode optic fiber into the brain, and an emission signal is collected back up through the same fiber. Because these optic fibers are lightweight and bendable, an animal can move largely unhindered, making this technique compatible with a wide array of behavioral tests and conditions. Some conditions, such as rapid movements or bending of the fiber-optic patch cord beyond the radius at which it can maintain total internal reflection, can introduce signal artifacts. To disambiguate signal from noise, we can exploit a property of GCaMP known as the “isosbestic point.” Briefly, with GCaMP, as the wavelength of the excitation light is shifted to the left, its emission in the calcium-bound state decreases and the emission in the calcium-unbound state marginally increases. The point at which the relative intensity of these two emissions are equal is termed the isosbestic point. When GCaMP is excited at this point, its emission is unaffected by changes in internal calcium concentrations, and variance in the signal is most often due to attenuation of the signal from overbending of the fiber-optic patch cord or movement of the neural tissue relative to the implanted fiber.
Single unit electrophysiology is still the gold standard for freely-moving in vivo recordings due to its single-cell and single-spike level resolution. However, it can be difficult to pinpoint the molecular identity of the cells being recorded, and the post-hoc analysis can be quite laborious. While fiber photometry does not have single-cell resolution, it does allow researchers to ask questions impossible to address with traditional techniques. Combining viral strategies with transgenic animals, the expression of GECIs can be directed to genetically-defined neuronal types to record population- or projection-defined neural activity, which can be performed by monitoring calcium signal directly at axon terminals10,11. Moreover, by implanting multiple fiber-optic cannulas, it is possible to simultaneously monitor neural activity from several brain regions and pathways in the same animal12,13.
In this manuscript, we describe a technique for single and multi-fiber photometry, how to correct for calcium-independent artifacts, and detail how to perform mono- and dual-color recordings. We also provide examples of the types of questions it enables one to ask and their increasing levels of complexity (see Figure 1). The fiber photometry setup for multi-fiber recordings detailed in this protocol can be built using a list of materials found at https://sites.google.com/view/multifp/hardware (Figure 2).
It is essential that the system be equipped for both 410 nm and 470 nm excitation wavelengths for calcium-independent and calcium-dependent fluorescence emission from GCaMP6 or its variants. For custom-built setups or if there is no available software to run the system, the free, open source program Bonsai (http://www.open-ephys.org/bonsai/) can be used. Alternatively, fiber photometry can be run through MATLAB (e.g., https://github.com/deisseroth-lab/multifiber)12 or other programming language14. The software and hardware of the system should allow manipulation of both the 410 nm and 470 nm LEDs and the camera, extraction of images (Figure 2), and calculation of the mean fluorescent intensity in the regions of interest (ROIs) drawn around the fibers on the images. The output should be a table of mean intensity values recorded with the 470 nm and 410 nm LEDs from each fiber in the patch cord. When performing multi-fiber experiments, 400 µm bundled fibers may limit the movement of mice. In such cases, we recommend using 200 µm patch cords, which provide more flexibility. It may also be possible to use smaller dummy cables during training of mice.
It is crucial to be able to extract time points for events of interest during fiber photometry acquisition. If the system does not readily provide a built-in system to integrate TTLs for specific events, an alternative strategy is to assign a time stamp to individual time points recorded to align with specific times and events during the experiment. Time stamping can be done using the computer clock.
Access restricted. Please log in or start a trial to view this content.
Protocol
All experiments were done in accordance with the Institutional Animal Care and Use Committees of the University of California, San Diego, and the Canadian Guide for the Care and Use of Laboratory Animals and were approved by the Université Laval Animal Protection Committee.
1. Alignment of the optical path between the CMOS (complementary metal oxide semiconductor) camera and the individual or branching patch cord
- Loosen all screws on the 5-axis translator (11, Figure 2B).
- Screw in the patch cord (12, Figure 2B) to the adaptor [SMA (sub-miniature A) or FC (fiber optic connector)] that is affixed to the 5-axis translator.
- Turn on the 470 nm excitation light (1, Figure 2B) at low power (100 µW), and place the tip of the patchcord pointing to an autofluorescent plastic slide. This does not have any bearing on future recordings but is solely for visualizing the alignment process.
- Record from the CMOS camera (13, Figure 2B) in live mode. Increase the gain or adjust the lookup table (LUT) until the image is not entirely black. The point is to be able to see an image at the focal point of the objective (10, Figure 2B).
- Advance the 5-axis translator towards the objective, ensuring that the 470 nm light is centered on the fiber at the SMA or FC end of the patch cord, until an image can be resolved on the camera.
- Adjust the X and Y axes until the image is centered and well-resolved.
- Visualize the light emitted from the ferrule-end of the patch cord. It should appear as an isotropic circle. If a branching patch cord is used, the amount of light emitted at the ferrule-ends of each patch cord should be similar. If the circle is not isotropic or the emitted light is unequal, adjust the 5-axis translator in the X-Y axis.
2. Setup of ROIs around fibers for measurement of mean fluorescent intensity
- Turn on all the excitation lights to better visualize the fibers. Adjust the camera gain such that no pixels are saturated and a clear image of the fibers are present.
- Live record or take a preliminary image.
- Draw ROIs around the fibers and keep them for the measurement of the mean intensity values during recordings (Figure 2A).
- For multiple fiber recordings, test for independence in signals.
- Live record from all fibers.
- Point one fiber towards a light source and tap with a finger. Very large fluctuations should occur solely in that channel (acceptable leakage 1:1000).
- If the signals are not independent, redraw more conservative ROIs and repeat the independence test.
- To label and keep track of which ROI corresponds to which fiber, colored tape or nail polish can be applied to the end of the fibers. Take a picture prior to the start of any experiment as a secondary reminder.
3. Setup of recording arena
- Hang the patch cord above the arena using stands, clamps, or holders.
- Make sure that the animal can freely move throughout the entire arena, uninhibited by the length of the fiber.
- Whether an operant box or open field is used, ensure that the patch cord will be able to reach the animal with minimal bending. If this requires a nose poke, ensure that there is enough room overhead to prevent bending of the fiber. Avoid any excessive bending or twisting of the patch cord.
4. In vivo recordings
NOTE: The procedure of optic fiber cannula implantation for fiber photometry experiments is identical to the procedure for optogenetics as described in Sparta et al15. We recommend using dental cement (see Table of Materials), which provides robust anchoring of the headcap to the skull bone. Dental cement will be particularly useful in cases where anchoring screws cannot be used.
- Visually inspect the distal end of the fibers of the patch cord by eye and with a minifiber microscope. If the surface of the fibers is scratched, repolish the fibers using fiber polishing/lapping film with fine grit (1 µm and 0.3 µm).
- Clean the distal ends of the patch cord with 70% ethanol and a cotton tip applicator.
- Clean the fiber-optic cannulas using 70% ethanol and a cotton tip applicator.
- Connect the ferrule end of the patch cord to the implanted fiber using a ceramic split-sleeve covered with a black shrink tube. During the connection, make sure that the sleeve is tight, otherwise use a new sleeve.
NOTE: There will be a large amount of signal loss if there is any space between the patch cord ferrule and the implant, and the recordings will not work. - Allow the animal to recover for a few minutes prior to the start of behavioral testing.
- Start recording the optical signal and run the experiment.
- While recording, keep a careful eye on the live-trace to ensure quality recordings. The signal is expected to rapidly decrease as a function of time in the first 2 min of recording. This effect is caused by heat-mediated LED decay, whereby the increase in heat increases the resistance of the optical element.
- If a jump in the signal that exceeds the on/off kinetics of GCaMP occurs, this is often an indication that the sleeve is not tight enough and the space between the patch cord and the implant is changing. In this case, stop the experiment and reconnect the animal using a new sleeve.
5. Fiber photometry data analysis
NOTE: This is a method for data analysis that works well for most recordings. However, alternative approaches can be implemented. Example code for data analysis can be found here: https://github.com/katemartian/Photometry_data_processing.
- Extract mean fluorescence intensity values recorded from 470 nm (Int470) and 410 nm (Int410) LEDs, corresponding to each individual fiber.
- Smooth each signal using a moving mean algorithm (Figure 3A).
- Perform baseline correction of each signal (Figure 3A and 3B) using the adaptive iteratively reweighted Penalized Least Squares (airPLS) algorithm (https://github.com/zmzhang/airPLS) to remove the slope and low frequency fluctuations in signals.
- Standardize each signal using the mean value and standard deviation (Figure 3C):
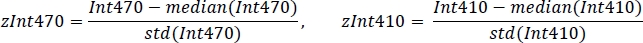
- Using non-negative robust linear regression, fit standardized zInt410 to zInt470 signals (Figure 3D) to the regression function:
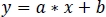
- Use the parameters of the linear regression (a, b) to find new values of zInt410 fitted to zInt470 (fitInt410, Figure 3D,E):
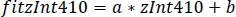
- Use the parameters of the linear regression (a, b) to find new values of zInt410 fitted to zInt470 (fitInt410, Figure 3D,E):
- Calculate the normalized dF/F (z dF/F) (Figure 3F):
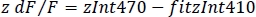
6. Simultaneous dual-color recordings
- Add to the photometry system a 560 nm LED to excite the red fluorescent calcium sensor and appropriate dichroic mirrors and filters (see Kim et al., 2016 for detailed description)12.
- Add an image splitter between the objective and the CMOS camera to separate the green and red emission wavelengths (see Figure 5). The image splitter will form two mirrored images on the camera sensor, corresponding to the red and green signals (e.g., a patch cord with 3 branches will create an image with 6 fibers).
- Draw ROIs around all fibers in both colors as detailed above. Make sure to clearly identify each ROI with the corresponding fiber and channel (green and red) (Figure 4A).
- Trigger simultaneous excitation with 470 nm and 560 nm LEDs and alternate them with 410 nm LED (Figure 5A).
7. Dual color data analysis
- Follow the steps in Section 5 to find fitInt410 for the Int470 signal and calculate z dF/F.
- Because the isosbestic point for red-shifted GECIs is generally unknown, the signal recorded with 410 nm LED in the green channel can be used for movement correction across both channels. Follow the steps in Section 5 to find fitInt410 for the Int560 signal and calculate z dF/F.
Access restricted. Please log in or start a trial to view this content.
Results
Neural correlates of behavioral responses can vary depending on a variety of factors. In this example, we used in vivo fiber photometry to measure the activity of axon terminals from the lateral hypothalamic area (LHA) that terminate in the lateral habenula (LHb). Wild type mice were injected with an adeno-associated virus (AAV) encoding GCaMP6s (AAV-hSyn-GCaMP6s) in the LHA and an optic fiber was implanted with the tip immediately above the LHb (Figure 4A). GCaMP6s expressi...
Access restricted. Please log in or start a trial to view this content.
Discussion
Fiber photometry is an accessible approach that allows researchers to record bulk-calcium dynamics from defined neuronal populations in freely-moving animals. This method can be combined with a wide range of behavioral tests, including “movement heavy” tasks such as forced swim tests2, fear-conditioning18, social interactions1,4, and others7,8
Access restricted. Please log in or start a trial to view this content.
Disclosures
Sage Aronson is the CEO and founder of Neurophotometrics Ltd., which sells multi-fiber photometry systems.
Acknowledgements
This work was supported by a grant from the Natural Sciences and Engineering Research Council of Canada (NSERC: RGPIN-2017-06131) to C.P. C. P. is a FRSQ Chercheur-Boursier. We also thank the Plateforme d’Outils Moléculaires (https://www.neurophotonics.ca/fr/pom) for the production of the viral vectors used in this study.
Access restricted. Please log in or start a trial to view this content.
Materials
| Name | Company | Catalog Number | Comments |
| 1/4"-20 Stainless Steel Cap Screw, 1" Long | Thorlabs | SH25S100 | |
| 1/4"-20 Stainless Steel Cap Screw, 1/2" Long | Thorlabs | SH25S050 | |
| 1/4"-20 Stainless Steel Cap Screw, 3/8" Long | Thorlabs | SH25S038 | |
| 1000 µm, 0.50 NA, SMA-SMA Fiber Patch Cable | Thorlabs | M59L01 | |
| 12.7 mm Optical Post | Thorlabs | TR30/M | |
| 12.7 mm Pedestal Post Holder | Thorlabs | PH20EM | |
| 15 V, 2.4 A Power Supply Unit with 3.5 mm Jack Connector for T-Cube | Thorlabs | KPS101 | |
| 20x objective | Thorlabs | RMS20X | #10 in Figure 2, #11 in Figure 5 |
| 30 mm Cage Cube with Dichroic Filter Mount | Thorlabs | CM1-DCH/M | #8-9 in Figure 2, #8-10 in Figure 5 |
| 405 nm LED | Doric Lenses | CLED_405 | #2 in Figure 2 |
| 410 nm bandpass filter | Thorlabs | FB410-10 | #5 in Figure 2; #7 in Figure 5 |
| 465 nm. LED | Doric Lenses | CLED_465 | #1 in Figure 2 |
| 470 nm bandpass filter | Thorlabs | FB470-10 | #4 in Figure 2; #6 in Figure 5 |
| 560 nm bandpass filter | Semrock | FF01-560/14-25 | #5 in Figure 5 |
| 560 nm LED | Doric Lenses | CLED_560 | #1 in Figure 3 |
| 5-axis kinematic Mount | Thorlabs | K5X1 | #11 in Figure 2, #12 in Figure 5 |
| Achromatic Doublet | Thorlabs | AC254-035-A-ML | #7 in Figure 2 |
| Adaptor for 405 collimator | Thorlabs | AD11F | #3 in Figure 2; #4 in Figure 5 |
| Adaptor for ajustable collimator | Thorlabs | AD127-F | #3 in Figure 2; #4 in Figure 5 |
| Aluminum Breadboard | Thorlabs | MB1824 | |
| Clamping Fork | Thorlabs | CF125 | |
| Cube connector | Thorlabs | CM1-CC | |
| Dual 493/574 dichroic | Semrock | FF493/574-Di01-25x36 | #10 in Figure 5 |
| Emission filter for GCaMP6 | Semrock | FF01-535/22-25 | #6 in Figure 2 |
| Enclosure with Black Hardboard Panels | Thorlabs | XE25C9 | |
| Externally SM1-Threaded End Cap for Machining | Thorlabs | SM1CP2M | |
| Fast-change SM1 Lens Tube Filter Holder | Thorlabs | SM1QP | #4-6 in Figure 2, #5-7 in Figure 5 |
| Fixed Collimator for 405 nm light | Thorlabs | F671SMA-405 | #3 in Figure 2; #4 in Figure 5 |
| Fixed collimator for 470 and 560 nm light | Thorlabs | F240SMA-532 | #3 in Figure 2; #4 in Figure 5 |
| Green emission filter | Semrock | FF01-520/35-25 | In light beam splitter |
| High-Resolution USB 3.0 CMOS Camera | Thorlabs | DCC3260M | #13 in Figure 2, #15 in Figure 5 |
| Light beam splitter | Neurophotometrics | SPLIT | #14 in Figure 5 |
| Longpass Dichroic Mirror, 425 nm Cutoff | Thorlabs | DMLP425R | #8 in Figure 2, #9 in Figure 5 |
| Longpass Dichroic Mirror, 495 nm Cutoff | Semrock | FF495-Di03 | #9 in Figure 2, #8 in Figure 5 |
| Metabond dental cement | C&B | ||
| M8 - M8 cable | Doric Lenses | Cable_M8-M8 | |
| Optic fiber cannulas | Doric Lenses | Need to specify that these will be used to photometry experiments requiring low autofluorescence | |
| Optic fiber Patchcords | Doric Lenses | Need to specify that these will be used to photometry experiments requiring low autofluorescence | |
| Red emission filter | Semrock | FF01-600/37-25 | In light beam splitter |
| T7 LabJack | LabJack | ||
| T-cube LED Driver | Thorlabs | LEDD1B | |
| USB 3.0 I/O Cable, Hirose 25, for DCC3240 | Thorlabs | CAB-DCU-T3 |
References
- Gunaydin, L. A., et al. Natural Neural Projection Dynamics Underlying Social Behavior. Cell. 157 (7), 1535-1551 (2014).
- Proulx, C. D., et al. A neural pathway controlling motivation to exert effort. Proceedings of the National Academy of Sciences of the United States of America. 115 (22), 5792-5797 (2018).
- Muir, J., et al. In Vivo Fiber Photometry Reveals Signature of Future Stress Susceptibility in Nucleus Accumbens. Neuropsychopharmacology. 43 (2), 255-263 (2017).
- Wang, D., et al. Learning shapes the aversion and reward responses of lateral habenula neurons. eLife. 6, (2017).
- de Jong, J. W., et al. A Neural Circuit Mechanism for Encoding Aversive Stimuli in the Mesolimbic Dopamine System. Neuron. 101 (1), 133-151 (2018).
- Lerner, T. N., et al. Intact-Brain Analyses Reveal Distinct Information Carried by SNc Dopamine Subcircuits. Cell. 162 (3), 635-647 (2015).
- Calipari, E. S., et al. In vivo imaging identifies temporal signature of D1 and D2 medium spiny neurons in cocaine reward. Proceedings of the National Academy of Sciences of the United States of America. 113 (10), 2726-2731 (2016).
- González, A. J., et al. Inhibitory Interplay between Orexin Neurons and Eating. Current Biology. 26 (18), 2486-2491 (2016).
- Chen, T. -W., et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature. 499 (7458), 295-300 (2013).
- Barker, D. J., et al. Lateral Preoptic Control of the Lateral Habenula through Convergent Glutamate and GABA Transmission. Cell Reports. 21 (7), 1757-1769 (2017).
- Siciliano, C. A., Tye, K. M. Leveraging calcium imaging to illuminate circuit dysfunction in addiction. Alcohol. 74, 47-63 (2018).
- Kim, C. K., et al. Simultaneous fast measurement of circuit dynamics at multiple sites across the mammalian brain. Nature Methods. 13 (4), 325-328 (2016).
- Sych, Y., Chernysheva, M., Sumanovski, L. T., Helmchen, F. High-density multi-fiber photometry for studying large-scale brain circuit dynamics. Nature Methods. 16 (6), 553-560 (2019).
- Akam, T., Walton, M. E. pyPhotometry: Open source Python based hardware and software for fiber photometry data acquisition. Scientific Reports. 9 (1), 3521(2019).
- Sparta, D. R., et al. Construction of implantable optical fibers for long-term optogenetic manipulation of neural circuits. Nature Protocol. 7 (1), 12-23 (2011).
- Stamatakis, A. M., et al. Lateral Hypothalamic Area Glutamatergic Neurons and Their Projections to the Lateral Habenula Regulate Feeding and Reward. The Journal of Neuroscience. 36 (2), 302-311 (2016).
- Tervo, G. D., et al. A Designer AAV Variant Permits Efficient Retrograde Access to Projection Neurons. Neuron. 92 (2), 372-382 (2016).
- Yu, K., da Silva, P., Albeanu, D. F., Li, B. Central Amygdala Somatostatin Neurons Gate Passive and Active Defensive Behaviors. The Journal of Neuroscience. 36 (24), 6488-6496 (2016).
- Falkner, A. L., Grosenick, L., Davidson, T. J., Deisseroth, K., Lin, D. Hypothalamic control of male aggression-seeking behavior. Nature Neuroscience. 19 (4), 596-604 (2016).
- Ren, J., et al. Anatomically Defined and Functionally Distinct Dorsal Raphe Serotonin Sub-systems. Cell. 175 (2), 472-487 (2018).
- Barnett, L. M., Hughes, T. E., Drobizhev, M. Deciphering the molecular mechanism responsible for GCaMP6m’s Ca2+-dependent change in fluorescence. PLOS ONE. 12 (2), 0170934(2017).
- Sun, F., et al. A Genetically Encoded Fluorescent Sensor Enables Rapid and Specific Detection of Dopamine in Flies, Fish, and Mice. Cell. 174 (2), 481-496 (2018).
- Patriarchi, T., et al. Ultrafast neuronal imaging of dopamine dynamics with designed genetically encoded sensors. Science. 360 (6396), (2018).
- Feng, J., et al. A Genetically Encoded Fluorescent Sensor for Rapid and Specific In Detection of Norepinephrine. Neuron. 102 (4), 745-761 (2019).
- Akerboom, J., et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Frontiers in Molecular Neuroscience. 6, 1-29 (2013).
- Dana, H., et al. Sensitive red protein calcium indicators for imaging neural activity. eLife. 5, (2016).
- Wang, H., Jing, M., Li, Y. Lighting up the brain: genetically encoded fluorescent sensors for imaging neurotransmitters and neuromodulators. Current Opinion in Neurobiology. 50, 171-178 (2018).
- Lu, L., et al. Wireless optoelectronic photometers for monitoring neuronal dynamics in the deep brain. Proceedings of the National Academy of Sciences. 115 (7), 1374-1383 (2018).
- Jennings, J. H., et al. Visualizing Hypothalamic Network Dynamics for Appetitive and Consummatory Behaviors. Cell. 160 (3), 516-527 (2014).
Access restricted. Please log in or start a trial to view this content.
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved