A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Assessing Protein Interactions in Live-Cells with FRET-Sensitized Emission
In This Article
Erratum Notice
Summary
Förster Resonance Energy Transfer (FRET) between two fluorophore molecules can be used for studying protein interactions in the living cell. Here, a protocol is provided as to how to measure FRET in live cells by detecting sensitized emission of the acceptor and quenching of the donor molecule using confocal laser scanning microscopy.
Abstract
Förster Resonance Energy Transfer (FRET) is the radiationless transfer of energy from an excited donor to an acceptor molecule and depends upon the distance and orientation of the molecules as well as the extent of overlap between the donor emission and acceptor absorption spectra. FRET permits to study the interaction of proteins in the living cell over time and in different subcellular compartments. Different intensity-based algorithms to measure FRET using microscopy have been described in the literature. Here, a protocol and an algorithm are provided to quantify FRET efficiency based on measuring both the sensitized emission of the acceptor and quenching of the donor molecule. The quantification of ratiometric FRET in the living cell not only requires the determination of the crosstalk (spectral spill-over, or bleed-through) of the fluorescent proteins but also the detection efficiency of the microscopic setup. The protocol provided here details how to assess these critical parameters.
Introduction
Microscopy-based analysis of Förster Resonance Energy Transfer (FRET) permits assessment of interactions between proteins in live cells. It provides spatial and temporal information, including information on where in the cell and in which subcellular compartment the interaction takes place and if this interaction changes over time.
Theodor Förster laid the theoretical foundation of FRET in 19481. FRET is a radiationless transfer of energy from an excited donor to an acceptor molecule and depends upon the distance of the molecules and the relative orientation of their transition dipoles as well as the overlap between the donor emission and acceptor absorption spectra. The rate of energy transfer is inversely proportional to the sixth power of the donor-acceptor distance. Thus, FRET can be used to measure molecular proximity in the range of 1-10 nm.
FRET competes with other de-excitation processes of the donor molecule and results in the so-called donor-quenching and sensitized emission of the acceptor. Donor-quenching is a reduction of the number of emitted donor photons, while sensitized emission is an increase in emitted acceptor photons. Many microscopic FRET analyses use fluorescence intensity measurements, including acceptor photobleaching2, donor photobleaching2, or FRET-sensitized photobleaching of the acceptor3.
Here, a step-by-step experimental protocol and mathematical algorithm are presented to quantify FRET using donor quenching and acceptor sensitized emission4,5, a method often referred to as ratiometric FRET. Many protocols on how to approximate sensitized emission have been published, few have quantified the absolute FRET efficiency6,7,8,9. The quantification of FRET efficiencies in the living cell requires determining (i) the crosstalk (spectral spill-over, or bleed-through) of the fluorescent proteins and, also (ii) the detection efficiency of the microscopic setup. While crosstalk can be assessed by imaging cells expressing only one of the fluorophores, the assessment of the relative detection efficiency of the donor and acceptor fluorescence is more complicated. It requires the knowledge of at least the ratio of the number of donor and acceptor molecules giving rise to the measured signals. The number of fluorophores expressed in live cells varies, however, from cell to cell and is unknown. The so-called α factor characterizes the relative signal strengths from a single excited donor and acceptor molecule. Knowledge of the factor is a prerequisite for quantitative ratiometric FRET measurements in samples with variable acceptor-to-donor molecule ratios as encountered during live-cell imaging with fluorescent proteins. Using a 1-to-1 donor-acceptor fusion protein as a calibration probe permits the determination of the α factor and also serves as a positive control. This genetically coupled probe is expressed by cells in unknown total amounts but in a fixed and known relative amount of one-to-one. The following protocol lays out how to construct the 1-to-1 probe and how to use it for quantification of FRET efficiency. A spreadsheet that includes all formulae can be found in the supplement and can be used by the readers to enter their own measurements in the respective columns as outlined below.
While the protocol uses the GFP-Cherry donor/ acceptor pair, the presented approach can be performed with any other FRET pair. The Supplementary File 1 provides details on cyan-yellow pairs.
Access restricted. Please log in or start a trial to view this content.
Protocol
1. Plasmid construction
- For generating the eGFP-mCherry1 fusion probe, use an N1 mammalian cell expression vector (see Table of Materials) with mCherry110 inserted using the restriction sites AgeI and BsrGI.
- Use the following oligonucleotides to amplify eGFP11 without a stop codon as SalI-BamHI fragment: N-terminal primer 5'-AAT TAA CAG TCG ACG ATG GTG AGC AAG GGC GAG G 3' and C-terminal primer 5'-AAT ATA TGG ATC CCG CTT GTA CAG CTC GTC CAT GC 3'.
- Insert this SalI-BamHI fragment into the multiple cloning site of the N1 vector to introduce RNPPV linker (five amino-acid) linker between the green and red fluorescent protein.
NOTE: This linker yields a mean FRET efficiency for the GFP-Cherry donor-acceptor pair of about 0.25 -0.3 (Figure 1A). The choice of stiff12 and helical13 linkers of varying lengths to scale measured FRET efficiencies has been discussed elsewhere but is not required for our purpose of the fusion protein. Going forward for simplicity we will call the fluorescent proteins 'GFP' and 'Cherry'.
2. Cell culture and transfection
- Use any cell line, e.g., NRK cells, for FRET experiments in media, e.g., Dulbecco's modified Eagle's media (DMEM), without phenol red, to reduce background fluorescence. For the same reason, the usage of phenol red free trypsin is advised.
- Once cells are 80% confluent, detach cells with 1 mL of 0.05% trypsin-EDTA, count the number of cells in suspension using a Neubauer chamber and seed about 10,000 cells per well of an 8-well chambered cover glass; alternatively, from a confluent cell culture grown in T25 flask, use 1 drop of cell suspension from a 2 mL pipette or 3 drops from a 5-mL cell suspension from a confluent culture grown in a T 12.5-cell culture flask.
- Grow cells in 8-well chambers (0.8 cm2/ well) with #1.0 cover glass for fluorescence live-cell microscopy at standard cell culture conditions (37°C and 5% CO2).
- 24 h after plating transfect the cells using an appropriate commercially available transfection media (see Table of Materials), with GFP, Cherry, GFP/ Cherry mix (1:1 mix, i.e., 0.8 μg and 0.8 μg GFP and Cherry plasmid DNA), and the GFP-Cherry chimera.
- For transfection, use 5 μL of the transfection reagent in 45 μL of DMEM and 1.6 μg of plasmid DNA. Stir by gently flicking the microcentrifuge tube.
- After 15 min incubation of the mix at room temperature, add 1-2 μL of the transfection reagent mixture to each well of the 8-well chamber slide. Return the chambered cover glass to the incubator.
- Let 20 h after transfection elapse before live-cell imaging, to allow for proper fluorescent protein expression, folding and maturation, especially of the red fluorophore.
3. FRET Imaging
- Image transfected cells in a humidified and heated environmental chamber at 37 °C. To buffer the cell media at physiological pH, use CO2 gas set to 5% flow, or add 20 mM HEPES to render the cell media CO2-independent.
- Use a confocal laser scanning microscope. Set the excitation and emission as follows to optimize the signal and minimize cross-talk.
- Use the 488-nm line of the argon ion laser to excite GFP and the 561-nm diode pumped solid state laser (or 543-nm Helium Neon laser, depending upon available laser lines) to excite Cherry.
- Set the following in the software of a commercial confocal microscope. Set the Dichroic mirror to 488/ 561 by button click using the pull-down menu. Collect fluorescence using 488-nm laser light for excitation in channel 1 through an emission band of 505 - 530 nm (or 505 - 550 nm) and in channel 2 with a long pass filter >585 nm and use the 561-nm laser light for excitation in channel 3 with a long pass filter > 585 nm (type in wavelengths). Band pass filters e.g., 590 - 650 nm or similar can also be used which have the advantage of excluding Raman-scattering.
- Excite with the two lasers sequentially and set the imaging mode to Switch after each line so that the excitation of the 512 x 512 pixels image alternates after each line (and not after each frame which would abrogate the ability to detect FRET due to diffusion of the labeled proteins while recording the images with different excitations; button click).
- Set up a mini-time series of three images by button clicks to detect if significant photobleaching occurs, and potentially reduce the laser power. Photobleaching of less than 1% is optimal. High laser intensity can also lead to absorption saturation reducing the apparent FRET efficiency14. Laser power, up to 10-20 μW measured at the objective lens are safe to use.
- First, image cells expressing the GFP-Cherry fusion construct. Set the parameters that define the time-integrated laser intensity per pixel in a confocal image, i.e., the pixel dwell time in microseconds, the acousto-optical tunable filter (AOTF) transmission in percent, and the zoom.
- Image cells using a 63x oil objective and Zoom set to 3x. This provides sufficient magnification and resolution to image cells in its entirety. Aim for a pixel size of 70-80 nm.
- Set Pixel dwell time to 2-4 μs and AOTF transmission for the 488-nm and 561-nm laser such that images have a good signal-to-noise ratio without bleaching and no pixels showing fluorescence intensity saturation. It is advantageous to adjust the laser power of 488 and 561 such that signal levels in channel 1 and channel 3 are similar.
- Set the Photomultiplier (averaging mode) gain to 600-800.
- Image with these settings 15-20 cells expressing the GFP-Cherry fusion protein. 15-20 cells provide good statistics while keeping the total time of a FRET measurement session limited to a few hours to help ensure stability of the microscopic set-up.
- Image with the same settings cells expressing GFP, Cherry, GFP and Cherry and non-transfected cells. Search for expressing cells in the green channel or red channel, respectively.
- Then, image 15-20 cells co-expressing proteins of interest coupled to GFP and Cherry, respectively. Searching for expressing cells, avoid long exposure of the cells in order to not bleach the fluorescent proteins. Cherry has a lower photostability than GFP, and bleaching Cherry, the acceptor compromises FRET analysis.
NOTE: Absorption and emission spectra of GFP and Cherry is shown in Supplementary Figure 1. After measuring for 5-6 h, it is advisable to repeat imaging a few cells expressing the GFP-Cherry chimera at the end of the imaging session to document that the set-up remained stable and detected FRET efficiencies of the GFP-Cherry fusion protein did not significantly change during the course of an imaging session.
4. Image analysis for detecting absolute FRET efficiencies using donor quenching and sensitized emission
NOTE: Here, a practical step-by-step guide as to how to determine FRET efficiency with the use of the attached spreadsheet (Supplementary File 2) is provided. Theory and derivation of the presented equations can be found in detail in previous publications4,15,16,17. With the described settings, the following fluorescence intensities are collected.
- Measure the donor signal I1 in channel 1, the donor channel, with 488-nm excitation and an emission band of 505-530 nm.
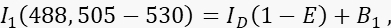
where ID is the unquenched donor signal in channel 1 that would be measured in the absence of an acceptor, is the mean FRET efficiency, and B1 the average background signal in channel 1. - Measure the acceptor signal I3 in channel 3, the acceptor channel, with 561-nm excitation and emission at >585 nm.
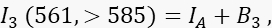
where IA is the acceptor signal and B3 the background in channel 3. - Measure the FRET signal in channel 2, the transfer channel, with 488-nm excitation and emission at >585 nm.
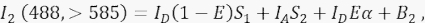
Where, the signal in channel 2 is a sum of four different components: (i) ID(1 - E)S1 is the spectral spill over from the quenched donor signal into the >585 detection channel (with the cross-talk factor S1), (ii) IAS2 is the acceptor signal from the direct excitation by 488-nm light (with the cross-talk factor S2), (iii) IDEα is the sensitized emission of the acceptor by FRET from the excited donor molecule (α will be detailed further in 4.8. - 4.10.), and (iv) B2 is the background signal. - Measure average background intensities in channels 1, 2, 3 in non-transfected or mock-transfected cells; either is fine with negligible difference. For all cell measurements, use the free-hand tool to delineate regions of interests and avoid perinuclear vesicles with increased autofluorescence. It is important to avoid significant autofluorescence from these perinuclear vesicles.
- Enter the measurements into the columns X, Y, and Z of the provided spreadsheet. Average background intensities in the 3 channels are entered into A2, B2, and C2 of the excel spreadsheet (Supplementary File 2).
- Measure average intensities in channels 1, 2, 3 of cells expressing GFP or Cherry alone, and enter the measurements into columns C, D, E, and N, O, P. Respective background intensities are subtracted (in F, G, H, and Q, R, S).
- In order to calculate E, the FRET efficiency, determine the cross-talk factors S1 and S2. The spectral cross-talk factor S1 is calculated from cells expressing only GFP
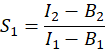
in column I. Enter the mean value for S1 into cell D2 on the excel spreadsheet. - Calculate the spectral cross-talk factor S2 from cells expressing only Cherry
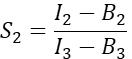
in column T. Enter the mean for S2 into cell E2 on the excel spreadsheet. - Ensure that the α factor relates the signal from any given number of excited GFP molecules in channel 1 to the signal of an equal number of excited Cherry molecules in channel 2, and is defined by
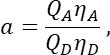
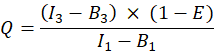
where QA and QD are the fluorescence quantum yields of Cherry and GFP; ηA and ηD the detection efficiencies of acceptor and donor fluorescence in channels 2 and 1, respectively.
NOTE: The α factor could be determined from two samples expressing known absolute amounts of GFP and Cherry. It is, however, impossible to know the exact amount of GFP and Cherry expressed in a cell. Therefore, we calculated the factor by using cells that express the GFP-Cherry fusion protein. Here, while the absolute amount is still unknown, the ratio of donor and acceptor molecules is known to be one. - Measure average intensities in channels 1, 2, 3 of cells expressing the GFP-Cherry fusion protein, and enter the measurements into the columns AE, AF, AG. Background intensities are subtracted (in AH, AI, AJ).
- Calculate the α factor (AJ column) from the fluorescence intensities in channel 1, 2 and 3 of the GFP-Cherry fusion protein as follows:
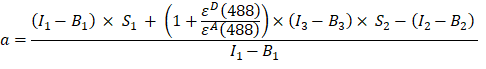
- Background-corrected intensities measured in channel 1 (I1 - B1), 2 (I2 - B2) and 3 (I3 - B3), respectively, are measured using the GFP-Cherry chimera. The spectral cross talk factor S1 was determined using cells expressing GFP only (see 4.7.). εD and εA are the extinction coefficients of GFP, the donor, and Cherry, the acceptor, at 488 nm, and can be determined from the literature (εGFP = 53,000 M-1cm-1)18 and the absorption curve of Cherry (εCherry ≈ 5560 M-1cm1). The ratio
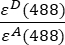 has been entered in cell G2 of the excel spreadsheet. Enter the mean value for the α factor into J2.
has been entered in cell G2 of the excel spreadsheet. Enter the mean value for the α factor into J2. - Use the determined α factor for the calculation of the FRET efficiency, E, as follows (column AK):
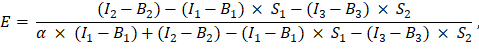
- Alternatively, determine FRET efficiency, E, for the negative controls, i.e., the co-expression of GFP and Cherry and the expression of GFP alone by adding the measurements of channels 1, 2, and 3 in the excel sheet in column AD, AE, and AF under the GFP-Cherry fusion protein measurements. Determine FRET efficiencies between the GFP and Cherry labeled proteins of interest in the same way.
- Determine the unquenched donor intensity, ID as (I1 - B1)/(1 - E), and the acceptor intensity as IA = I3 - B3; these values are proportional to the expression levels of the tagged proteins.
- Determine the corrected acceptor-to-donor intensity ratio (Q) of the GFP-Cherry fusion protein as follows (column AL):
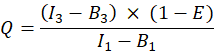
- For other co-transfected cells, calculate the acceptor-to-donor molecular ratio NA/ND as follows:
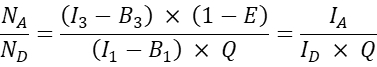
NOTE: The rationale to determine the NA/ND ratio and plot the mean cellular FRET efficiency E versus NA/ND, is that one donor molecule can transfer energy to multiple acceptors while an acceptor molecule can only receive energy from one donor at a given time. Even if only one acceptor can interact with a donor because of the stoichiometry of the interaction, an increase of acceptor concentration is expected to increase the fraction of donors in complex with the acceptor because of the law of mass action. Thus, for a fixed (or narrow range of) donor expression, the FRET efficiency should rise with increasing NA/ND. When plotting the FRET efficiency E versus NA/ND for the co-expression of GFP and Cherry, i.e. the negative probe, however, an increase in NA/ND should not result in an increase in FRET efficiency (at least at sufficiently low acceptor concentrations where random FRET due to the vicinity of acceptor dyes to donor dyes does not occur).
Access restricted. Please log in or start a trial to view this content.
Results
Figure 1 shows the images obtained in the donor channel, channel 1 (488, 505-530 nm), the transfer channel, channel 2 (488, >585 nm), and the acceptor channel, channel 3 (561, >585 nm), respectively. Representative images of cells expressing GFP only, Cherry only, co-expressing GFP and Cherry, and expressing the GFP-Cherry fusion protein. The mean cellular FRET efficiencies calculated in NRK cells expressing GFP-Cherry fusion protein (positive co...
Access restricted. Please log in or start a trial to view this content.
Discussion
The presented protocol details the use of the genetically coupled one-to-one fluorescent protein calibration probe for quantifying FRET using the detection of sensitized emission of the acceptor and quenching of the donor molecule by confocal microscopy. This method can be applied to assess protein interactions in the physiological context of the living cell in different subcellular compartments. Spatial resolution can be further improved by applying the presented algorithm to calculate FRET efficiencies in each pixel of...
Access restricted. Please log in or start a trial to view this content.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We would like to thank the Neuroscience Imaging Service at Stanford University School of Medicine for providing equipment and space for this project. This research was supported by intramural funding of the Stanford Cancer Institute and the Gynecologic Oncology Division Stanford as well as GINOP-2.3.2-15-2016-00026, GINOP-2.3.3-15-2016-00030, NN129371, ANN135107 from the National Research, Development and Innovation Office, Hungary.
Access restricted. Please log in or start a trial to view this content.
Materials
| Name | Company | Catalog Number | Comments |
| 0.5% Trypsin-EDTA without phenol red (10x) | Thermo Fisher Scientific | 15400054 | |
| Clontech mCherry N1 vector | Addgene | 3553 | |
| DMEM without phenol red | Thermo Fisher Scientific | 11054020 | |
| Fugene 6 | Promega | E2691 | |
| HEPES | Thermo Fisher Scientific | 15630080 | |
| LabTek 8-well chambers #1.0 | Thermo Fisher Scientific | 12565470 | |
| L-Glutamine (200 mM) | Thermo Fisher Scientific | 25030081 |
References
- Förster, T. Zwischenmolekulare Energiewanderung und Fluoreszenz. Annal der Physik. 437, 55-75 (1948).
- Jovin, T. M., Arndt-Jovin, D. J. Luminescence digital imaging microscopy. Annual Review of Biophysics and Biophysical Chemistry. 18, 271-308 (1989).
- Mekler, V. M. A photochemical technique to enhance sensitivity of detection of fluorescence resonance energy transfer. Photochemistry and Photobiology. 59, 615-620 (1994).
- Vamosi, G., et al. Conformation of the c-Fos/c-Jun complex in vivo: A combined FRET, FCCS, and MD-modeling study. Biophysical Journal. 94 (7), 2859-2868 (2008).
- Renz, M., Daniels, B. R., Vamosi, G., Arias, I. M., Lippincott-Schwartz, J. Plasticity of the asialoglycoprotein receptor deciphered by ensemble FRET imaging and single-molecule counting PALM imaging. Proceedings of the National Academy of Science U. S. A. 109 (44), 2989-2997 (2012).
- van Rheenen, J., Langeslag, M., Jalink, K. Correcting confocal acquisition to optimize imaging of fluorescence resonance energy transfer by sensitized emission. Biophysical Journal. 86 (4), 2517-2529 (2004).
- Muller, S. M., Galliardt, H., Schneider, J., Barisas, B. G., Seidel, T. Quantification of Forster resonance energy transfer by monitoring sensitized emission in living plant cells. Frontiers in Plant Sciences. 4, 413(2013).
- Gates, E. M., LaCroix, A. S., Rothenberg, K. E., Hoffman, B. D. Improving quality, reproducibility, and usability of FRET-based tension sensors. Cytometry Part A. 95 (2), 201-213 (2019).
- Menaesse, A., et al. Simplified instrument calibration for wide-field fluorescence resonance energy transfer (FRET) measured by the sensitized emission method. Cytometry Part A. , 24194(2020).
- Shaner, N. C., et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nature Biotechnology. 22 (12), 1567-1572 (2004).
- Cormack, B. P., Valdivia, R. H., Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene. 173, 1 Spec No 33-38 (1996).
- Nagy, P., et al. Novel calibration method for flow cytometric fluorescence resonance energy transfer measurements between visible fluorescent proteins. Cytometry Part A. 67 (2), 86-96 (2005).
- Arai, R., Ueda, H., Kitayama, A., Kamiya, N., Nagamune, T. Design of the linkers which effectively separate domains of a bifunctional fusion protein. Protein Engineering. 14 (8), 529-532 (2001).
- Szendi-Szatmari, T., Szabo, A., Szollosi, J., Nagy, P. Reducing the detrimental effects of saturation phenomena in FRET microscopy. Analytical Chemistry. 91 (9), 6378-6382 (2019).
- Tron, L., et al. Flow cytometric measurement of fluorescence resonance energy transfer on cell surfaces. Quantitative evaluation of the transfer efficiency on a cell-by-cell basis. Biophysical Journal. 45 (5), 939-946 (1984).
- Sebestyen, Z., et al. Long wavelength fluorophores and cell-by-cell correction for autofluorescence significantly improves the accuracy of flow cytometric energy transfer measurements on a dual-laser benchtop flow cytometer. Cytometry. 48 (3), 124-135 (2002).
- Szaloki, N., et al. High throughput FRET analysis of protein-protein interactions by slide-based imaging laser scanning cytometry. Cytometry Part A. 83 (9), 818-829 (2013).
- McRae, S. R., Brown, C. L., Bushell, G. R. Rapid purification of EGFP, EYFP, and ECFP with high yield and purity. Protein Expression and Purification. 41 (1), 121-127 (2005).
- Kremers, G. J., Goedhart, J., van Munster, E. B., Gadella, T. W. Cyan and yellow super fluorescent proteins with improved brightness, protein folding, and FRET Forster radius. Biochemistry. 45 (21), 6570-6580 (2006).
- Bajar, B. T., Wang, E. S., Zhang, S., Lin, M. Z., Chu, J. A Guide to Fluorescent Protein FRET Pairs. Sensors (Basel). 16 (9), (2016).
- Scott, B. L., Hoppe, A. D. Optimizing fluorescent protein trios for 3-Way FRET imaging of protein interactions in living cells. Science Reports. 5, 10270(2015).
- Chen, Y. C., Spring, B. Q., Clegg, R. M. Fluorescence lifetime imaging comes of age how to do it and how to interpret it. Methods Molecular Biology. 875, 1-22 (2012).
- King, C., Sarabipour, S., Byrne, P., Leahy, D. J., Hristova, K. The FRET signatures of noninteracting proteins in membranes: Simulations and experiments. Biophysical Journal. 106 (6), 1309-1317 (2014).
Access restricted. Please log in or start a trial to view this content.
Erratum
Formal Correction: Erratum: Assessing Protein Interactions in Live-Cells with FRET-Sensitized Emission
Posted by JoVE Editors on 3/14/2023. Citeable Link.
An erratum was issued for: Assessing Protein Interactions in Live-Cells with FRET-Sensitized Emission. The Authors section was updated from:
György Vámosi1
Sarah Miller2
Molika Sinha2
Gabor Mocsár1
Malte Renz2
1Department of Biophysics and Cell Biology, Faculty of Medicine, University of Debrecen
2Gynecologic Oncology Division, Stanford University School of Medicine
to:
György Vámosi1
Sarah Miller2
Molika Sinha2
Maria Kristha Fernandez2
Gabor Mocsár1
Malte Renz2
1Department of Biophysics and Cell Biology, Faculty of Medicine, University of Debrecen
2Gynecologic Oncology Division, Stanford University School of Medicine
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved