このコンテンツを視聴するには、JoVE 購読が必要です。 サインイン又は無料トライアルを申し込む。
Method Article
ショウジョウバエ幼虫の脳および細胞株からのPoly A尾長の測定
要約
プロトコルは他の種からのティッシュかセルのタイプに容易に合わせることができる ショウジョウバエ の神経系からの興味の遺伝子の多(A)長さを量化するための有効で、信頼できる方法を記述する。
要約
ポリアデニル化は、mRNA分子の3'末端にポリ(A)テールを付加する重要な転写後修飾です。ポリ(A)テールの長さは、細胞プロセスによって厳密に制御されています。mRNAポリアデニル化の調節不全は、遺伝子発現の異常や、がん、神経疾患、発達異常などのさまざまな疾患と関連しています。したがって、ポリアデニル化の動態を理解することは、mRNAプロセシングと転写後遺伝子制御の複雑さを解明するために不可欠です。
この論文では、 ショウジョウバエ の幼虫の脳と ショウジョウバエ のシュナイダーS2細胞から単離されたRNAサンプルのポリ(A)テールの長さを測定する方法を紹介します。私たちは、酵母ポリ(A)ポリメラーゼを用いてmRNAの3'末端にG/I残基を酵素的に付加するグアノシン/イノシン(G/I)テーリングアプローチを採用しました。この修飾により、RNAの3'末端が酵素分解から保護されます。次に、保護された全長のポリ(A)テールを、ユニバーサルアンチセンスプライマーを使用して逆転写します。続いて、逆転写に用いるユニバーサル配列オリゴとともに、目的遺伝子を標的とする遺伝子特異的オリゴを用いてPCR増幅を行う。
これにより、目的の遺伝子のポリ(A)テールを包含するPCR産物が生成されます。ポリアデニル化は均一な修飾ではなく、さまざまな長さのテールが生じるため、PCR産物はさまざまなサイズを示し、アガロースゲルに塗抹標本パターンをもたらします。最後に、PCR産物を高分解能キャピラリーゲル電気泳動にかけ、ポリ(A)PCR産物と遺伝子特異的PCR産物のサイズを用いて定量します。この手法は、ポリ(A)テールの長さを分析するための簡単で信頼性の高いツールを提供し、mRNA制御を支配する複雑なメカニズムについてより深い洞察を得ることができます。
概要
ほとんどの真核生物のmRNAは、標準的なポリ(A)ポリメラーゼによる非テンプレート化アデノシンの付加により、核内の3'末端で転写後ポリアデニル化されます。無傷のポリ(A)テールは、mRNAの核外輸送に不可欠であり1、ポリ(A)結合タンパク質との相互作用を促進して翻訳効率を高め2、分解に対する耐性を付与する3ため、mRNAのライフサイクル全体を通じて極めて重要です。場合によっては、ポリ(A)テールは、非標準的ポリ(A)ポリメラーゼによって促進され、細胞質内で伸長することもあります4。細胞質では、ポリ(A)テールの長さが動的に変化し、mRNA分子の寿命に影響を与えます。多数のポリメラーゼおよびデデニラーゼは尾の長さ5,6,7を調節するために知られている。例えば、ポリ(A)テールの短縮は並進抑制と相関し、ポリ(A)テールの延長は並進抑制を増強する8,9。
蓄積されたゲノム研究は、真核生物生物学のさまざまな側面にわたるポリ(A)尾の長さの基本的な重要性を実証しました。これには、生殖細胞の発生、初期胚発生、学習と記憶のためのニューロンシナプス可塑性、および炎症反応における役割が含まれます10。ポリ(A)テールの長さを測定するために開発された方法は数多くあります。例えば、RNase H/oligo(dT)アッセイでは、オリゴ(dT)の存在下または非存在下でのRNase Hを利用して、ポリ(A)テール長11,12を研究します。ポリ(A)テールを研究する他の方法には、cDNA末端の迅速増幅などの3'末端のPCR増幅、ポリ(A)テスト(RACE-PAT)12,13、リガーゼ媒介ポリ(A)テスト(LM-PAT)14などがあります。PATアッセイのさらなる修飾には、ePAT15およびsPAT16が含まれる。酵素的Gテーリング17,18または3'末端のG/Iテーリングは、PATアッセイの他のバリエーションです。これらの技術のさらなる修正には、高分解能ポリ(A)テスト(Hire-PAT)と呼ばれる高分解能分析のためのキャピラリーゲル電気泳動とともに蛍光標識されたプライマーの使用が含まれます19。これらのPCR駆動アッセイにより、ポリ(A)長の定量を迅速かつ高感度に行うことができます。
次世代シーケンシングの開発に伴い、PAL-seq20 やTAIL-seq21などのハイスループットシーケンシング法により、トランスクリプトームワイドスケールでのポリアデニル化解析が可能になりました。しかしながら、これらの方法は、36〜51ヌクレオチドの短い配列リードしか提供しない。そのため、FLAM-Seq22 は、完全長mRNAのグローバルテール長プロファイリング用に開発され、ロングリードを提供します。ナノポアテクノロジー23 は、ポリ(A)テール長の推定のために、PCRに依存しない直接RNA、または直接cDNAシーケンシングを提供します。ただし、これらのハイスループット方法には制限がないわけではありません。大量の出発物質を必要とし、高価で時間がかかります。さらに、希少な転写産物の解析は、ハイスループットの方法では非常に困難な場合があり、ロースループットのPCRベースの分析法は、パイロット実験や他の方法の検証のために、少数の転写産物を解析する必要がある場合に利点を提供します。
我々は最近、ショウジョウバエにおいてDscam1 mRNAが短いポリ(A)テールを含むことを実証し、G/Iテーリング法を用いてDscam1 3'UTR上の細胞質ポリ(A)結合タンパク質の非標準的結合を必要とすることを明らかにした24。ここでは、ショウジョウバエの神経系およびショウジョウバエS2細胞由来のmRNAの組織調製とポリ(A)長の定量化のための合理化された手順を提供します。
Access restricted. Please log in or start a trial to view this content.
プロトコル
1. ショウジョウバエ幼虫の飼育と選抜
- ハエ株(w1118、野生型)を標準的なハエの餌培地で25°Cの加湿インキュベーターで維持/培養します。
- 産卵後72時間で放浪する3齢 幼虫を10匹選別する。
- 幼虫を35mmの空のシャーレに入れ、鉗子を使用して水道水を含む新しいシャーレに幼虫を移して静かに洗浄します。これを2回行って、残っている食べ物を取り除きます。
2. ショウジョウバエ幼虫からの脳分離(図1)
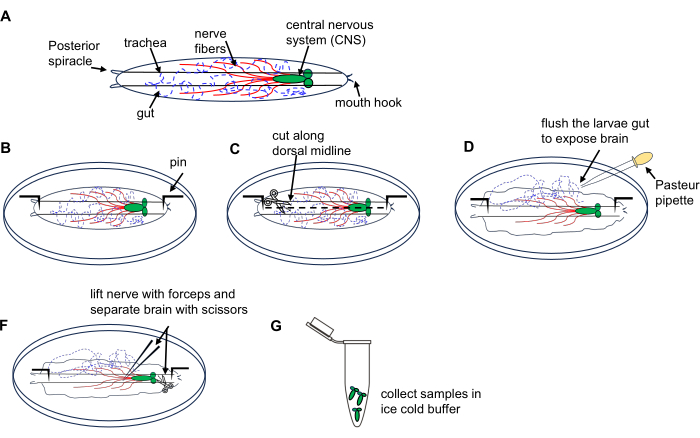
図1:ショウジョウバエ幼虫の脳の解剖(第3期)の徘徊期。 (A)ショウジョウバエ幼虫の模式図。(B-G)幼虫の解剖。この図の拡大版をご覧になるには、ここをクリックしてください。
- 氷冷PBSを含む解剖皿に幼虫10匹を置きます。
- 幼虫の背側を上にして(その長さに沿って走る気管管で識別されます)、両端を皿の底に固定し、後端の体壁に小さな切開を加えます。
- マイクロダイセクションハサミを使用して、背側の正中線に沿って前端に向かって体壁を切断します。
- パスツールピペット3xとPBSを皿に入れて幼虫の内部を短時間洗い流し、脳を露出させます。
- 鉗子を使用して脳を見つけて持ち上げ、マイクロダイセクションハサミを使用して慎重に分離します。
- 解剖した脳を、氷上で冷やしたPBSで満たされた1.5 mLの微量遠心チューブに移します。幼虫の脳みそを全て集める。セクション4で説明したように、RNAマイクロプレップキットを使用してRNA抽出を進めます。
注:組織の損傷やRNAの分解を防ぐために、15分以内に10匹の幼虫を解剖してください。
3. ショウジョウバエS2シュナイダー細胞
- ショウ ジョウバエ シュナイダー培地でショウジョウ バエ S2細胞を増殖させ、加湿インキュベーター内で25°Cで10%ウシ胎児血清(FBS)を8×106 〜10×10 6 細胞/mLの密度で、生存率90%以上で増殖させます。
- 50 mLの滅菌コニカルチューブ中で、25°Cに予熱した10% FBSを添加したシュナイダーショウジョウバエ培地で細胞を2.5×106細胞/mLに希釈します。
- 8 mLの細胞懸濁液(20 × 106細胞) を100 mmの培養プレートに移し、4 mLの培地を加えて12 mLにします(1日目)。
- 培養した細胞を加湿したインキュベーターで25°Cでインキュベートします。
注:細胞は12〜16時間後(2日目)にプレートに緩く付着します。 - 適切なDNAプラスミドを細胞にトランスフェクションします24。
- 加湿したインキュベーターで48時間インキュベートします。
- インキュベーション後、穏やかにピペッティングして5 mLの氷冷PBSを添加して細胞を回収します(4日目)。
- 細胞を15 mLチューブに移します。
- 1,000 × g 、4°Cで5分間遠心分離して細胞をペレット化します。
- 穏やかにピペッティングして細胞を氷冷PBSで2回すすぎ、1,000 × g で4°Cで5分間遠心分離して細胞を回収します。
- RNAミニプレップキットを用いてRNA抽出を行います。
注意: 滅菌層流フード内で次の手順を実行します。
4. ショウジョウバエ幼虫の脳とS2細胞からのトータルRNA抽出
- 幼虫の脳: 短時間の遠心分離(5,000 × gで8秒の短時間スピン)によりPBSを除去します。
- 600 μLのRNA溶解バッファーを添加し、プラスチック製の乳棒で10倍に均質化します。実体顕微鏡でチューブを目視検査し、完全に溶解することを確認します。
- 1,000 × g 、4°Cで5分間遠心分離し、組織の破片を除去します。透明化した上清をヌクレアーゼフリーの微量遠心チューブに移します。
- メーカーの指示に従って、RNAマイクロプレップキットを使用してRNAを単離します。
注:RNAマイクロプレップキットの使用は、サンプル中に存在するRNAの量が少ないため、幼虫の脳サンプルに不可欠です。 - S2 セル: PBSを除去し、メーカーの指示に従ってRNAを単離します。
- 分光光度法とアガロースゲル電気泳動により、RNAの収量と品質を測定します。
- 抽出したRNAの光学濃度をそれぞれA260 nm およびA280 nmで測定することにより、抽出されたRNAの純度と量を決定します。ダウンストリームアプリケーションでは、A260 nm/A280 nm 比が ≥2.0 で、RNA 濃度が >350 ng/μL であることを確認してください。
注:ショウ ジョウバエ の幼虫10匹の脳からの典型的なRNA収量は、5μL中~500-800 ng/μLまたは2.5-4 μgです。S2細胞の場合、収量は2~3μg/μL(15μL中15-30μg)です。単離されたRNAは、長期保存のために-80°Cで保存できます。
- 抽出したRNAの光学濃度をそれぞれA260 nm およびA280 nmで測定することにより、抽出されたRNAの純度と量を決定します。ダウンストリームアプリケーションでは、A260 nm/A280 nm 比が ≥2.0 で、RNA 濃度が >350 ng/μL であることを確認してください。
5. RNAゲルの調製と電気泳動
- 1.5% 変性 RNA ゲル (100 mL)
注:ホルムアルデヒドは、皮膚との接触や蒸気の吸入によって有毒です。化学ドラフトで取り扱ってください。- アガロース錠剤3錠(1.5 g)を82 mLのMOPS緩衝液(補足ファイル1)に、錠剤が完全に分解して微粒子を形成するまで溶解します。
- 溶液が透明になり、すべての粒子が完全に溶解するまで、アガロースラリーを電子レンジで加熱します。
- 溶液を~60°Cに冷却します。
- 18 mLの37%ホルムアルデヒドを加え、穏やかに渦巻いて混合します。溶液をキャスティングトレイに注ぎ、ドラフトで固めます。
- RNAサンプル調製と電気泳動
- RNA サンプルを 200 ng(5 μL 中)に希釈し、5 μL の 2x RNA ローディング色素を加えます。
- サンプルを70°Cのドライバスで5分間加熱します。
- 最初のレーンに 2 μL の RNA ラダーをロードし、隣接するレーンに 10 μL のサンプルをロードします。
- MOPSバッファー中、100V、60分間、5 x 6 cmのゲルで電気泳動を行います。
注:電気泳動条件は、アンプリコンのサイズに応じて調整してください。 - UVトランスイルミネーターでゲルを可視化します。
注:シングルバンド~600ヌクレオチドサイズの存在は、インタクトなRNA調製を示します( 図2Aを参照)。
6.ポリ(A)テール長測定
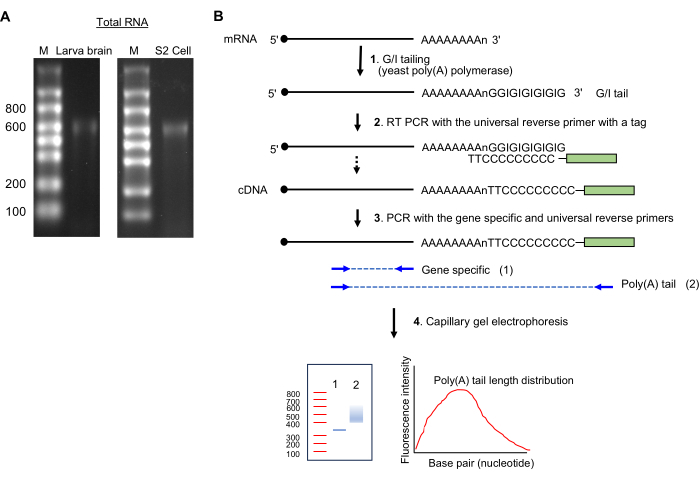
図2:RNAサンプル調製とポリ(A)テールアッセイ。 (A)RNAゲル画像は、ショウ ジョウバエ 幼虫の脳(左)とS2細胞(右)の1.5%ホルムアルデヒドアガロースゲル上の全RNAを示しています。一本鎖RNAラダーのサイズは、レーンMのヌクレオチドで示されています。 ~600 nt の主要な RNA バンディングは rRNA に由来します。(B) ポリ(A)テールアッセイの概略図。 略語:G / I =グアノシン/イノシン。 この図の拡大版をご覧になるには、ここをクリックしてください。
- GIテーリング(図2B)
- 試薬を氷上に保持し、最大14 μLのトータルRNAサンプル(1 μg)、4 μLの5x Tailバッファーミックス、および2 μLの10x Tail酵素ミックスの混合物(20 μL)を調製します。
- サーモサイクラーで37°Cで60分間インキュベートします。
- 1.5 μLのテールストップ溶液を加えます。氷の上に2分間置いてください。
注:逆転写に進むか、逆転写の準備が整うまでGIテールRNAサンプルを-80°Cで保存します。
- 逆転写とPCR増幅
- 混合物を調製し、 補足ファイル1に記載されている条件下でインキュベートすることにより、cDNAを合成します。
- cDNAサンプルを希釈し、 補足ファイル1に示すようにPCRを実行してDNAを増幅します。
7. アガロースゲル電気泳動によるPCR産物分析
- ステップ6.2.2のPCR産物の少量(2-5μL)を2.5%アガロースゲル上で、100Vで45分間電気泳動して分析し、品質管理を行います。
- ゲル抽出したPCRバンドをシーケンシングすることにより、遺伝子特異的反応およびテール特異的反応に対するPCRの特異性を検証します。
8. キャピラリー電気泳動
- 高感度DNAキットを備えたバイオアナライザーを使用して、遺伝子特異的およびポリ(A)特異的PCRからのPCR産物1 μL(0.5-5 ng/μL)に対して高分解能ゲル電気泳動を行います。実行が成功したことを示す、十分に分離されたピークを探します。
9.データ分析:ポリ(A)テール長測定(図3)
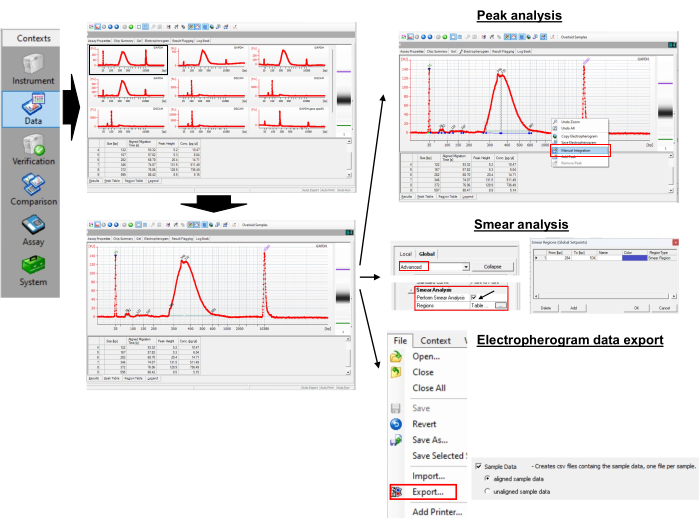
図3:ポリ(A)テール長とピーク値の測定。 この図の拡大版をご覧になるには、ここをクリックしてください。
- データへのアクセス
- データにアクセスするには、ソフトウェアでxadファイルを開きます。
- ツリービューパネルでサンプル名またはラダーを選択します。
注:これにより、選択したサンプルの結果がエレクトロフェログラムまたはゲル状の画像として表示されます。下部マーカー 35 塩基対 (bp) および上部マーカー 10,380 bp は、ラダーデータ(50-7,000 bp)をサンプルウェルからのデータに合わせるために使用される内部標準です。 - エレクトロフェログラムやゲル状の画像をズームインおよびズームアウトして、詳細を表示します。
- ピークおよびスミア分析の実行
- ピークサイズを取得するには、選択したサンプルのエレクトロフェログラムを開きます。
- エレクトロフェログラムを右クリックして手動 積分 を選択し、水平線をドラッグしてピークを手動で選択します。
- ピーク表でピーク値を観察します。ピーク高さが最大のピークを特定します。これは、個々のサンプルのポリ(A)テール長のピークです。示されている例では、346 bp です。
- トップメニューの[ セットポイントの表示/非表示 ]アイコンを有効にし、右側に新しいパネルがポップアップするのを待ちます。
- [詳細設定]を選択し、下にスクロールして[にじみ分析の実行]を見つけて、チェックボックスをオンにします。これにより、エレクトロフェログラムタブにRegionテーブルが追加されます。
- サンプルからエレクトロフェログラムを選択し、From [bp]とTo [bp]メニューを示すRegionテーブルに移動します。開始と終了 [bp] を設定するには、エレクトロフェログラムを右クリックして [領域] を選択し、[領域の追加] を追加します。
- [Region] テーブルの任意のセルを右クリックし、[Modify Regions] を選択すると、カスタム領域を設定できる小さな新しいウィンドウがポップアップ表示されます。
注:たとえば、GAPDHには300 bpから550 bpの領域を使用しました。遺伝子特異的 GAPDH PCR では、265 bp でピークが得られました。ユニバーサルプライマー(表1)は、アニーリングによりポリ(A)PCRの長さをG/IテールRNAに35 bp延長します。したがって、GAPDH RNA上の最初のアデニンヌクレオチドは300 bp(265 + 35)から始まります。ポリ(A)テールの最大長を任意に250(300 + 250 = 550)に制限しました。領域テーブルから、プログラムは領域内の平均サイズを 387 bp として返します。 - 式(1)を使用して、目的のmRNAのポリ(A)テール長を計算します。
ポリ(A)テールの長さ = (A - B - 35) (1)
ここで、Aはエレクトロフェログラム由来のポリ(A)特異的PCR産物の平均bp(すなわち、GAPDHについては387bp)、Bはエレクトロフェログラム由来の遺伝子特異的PCR産物のピークbp(すなわち、GAPDHについては265bp)、および「35」はユニバーサルリバースプライマータグの長さである。
注:上記の計算から、GAPDHの平均ポリ(A)テール長は387 - 265 - 35 = 87 bpです。
10. ポリ(A)テール長分布の可視化
- [ファイル] |エクスポート |サンプルデータ: サンプルデータを取得します。
注: エクスポートされた csv ファイルでは、X 軸に bp ではなく実行時間が表示されます。 - サンプルのエレクトロフェログラムに移動し、[エレクトロフェログラム]タブの[サイズを表示]をオンにすると、領域テーブルの領域テーブルがbpから実行時間に自動的に変換されます。この例では、70.39 秒から 86.28 秒は 300 bp から 550 bp に相当します。
- csv ファイルを開き、実行時間の 70.39 秒 から 86.28 秒 までの値を選択してグラフを生成します。bpサイズをグラフのX軸で可視化するには、bpサイズのエレクトロフェログラムを画像ファイルとしてエクスポートし、スプレッドシートで生成されたグラフに重ねて表示します。これは、ポリ(A)テール分布のbpサイズと適切に一致します。
Access restricted. Please log in or start a trial to view this content.
結果
ここでは、ショウジョウバエ幼虫の脳から採取したDscam1とGAPDHのポリ(A)尾長を解析しました(図4)。単離されたRNAは、品質管理のためにアガロースゲル上で可視化されました。約 600 ヌクレオチドサイズの 1 つの RNA バンドは、インタクトな RNA 調製を示します(図 2A)。RNAは、Agilent 2100バイオアナライザーを使用して、G/Iテーリング...
Access restricted. Please log in or start a trial to view this content.
ディスカッション
このプロトコルでは、私達はショウジョウバエS2セルからのサンプル準備と同様、第3のinstarの段階をさまようことからのショウジョウバエの幼虫の頭脳を解剖する技術を記述する。mRNAは不安定な性質を持っているため、サンプル採取には特別な注意が必要です。幼虫の脳解剖の場合、隔離中に脳が損傷してはならず、長期間溶液中に保管しないでください。解剖のラウ?...
Access restricted. Please log in or start a trial to view this content.
開示事項
著者には開示すべき利益相反はありません。
謝辞
本研究は、国立神経疾患・脳卒中研究所(National Institute of Neurological Disorders and Stroke Grant R01NS116463 to J.K.)と、国立衛生研究所(National Institutes of Health Grant P20GM103650)の支援を受けたネバダ大学リノ校の細胞・分子イメージングコア施設の支援を受けて、本研究で報告された研究に使用されました。
Access restricted. Please log in or start a trial to view this content.
資料
| Name | Company | Catalog Number | Comments |
| 3-(N-morpholino) propanesulfonic acid (MOPS) | Research Product Internation (RPI) | M92020 | |
| Agilent High Sensitivity DNA Kit | Agilent Technologies | 5067-4626 | |
| Agilent software 2100 expert free download demo | Agilent Technologies | https://www.agilent.com/en/product/automated-electrophoresis/bioanalyzer-systems/bioanalyzer-software/2100-expert-software-228259 | |
| Apex 100 bp-Low DNA Ladder | Genesee Scientific | 19-109 | |
| Bioanalyzer | Agilent 2100 Bioanalyzer G2938C | ||
| Diethyl pyrocarbonate (DEPC) | Research Product Internation (RPI) | D43060 | |
| DNA dye (Gel Loading Dye, Purple (6x) | New England biolabs | B7024S | |
| Drosophila S2 cell line | Drosophila Genomics Resource Center stock #181 | ||
| Drosophila Schneider’s Medium | Thermo Fisher Scientific | 21720024 | |
| Ehidium bromide | Genesee scientific | 20-276 | |
| Fetal bovine serum (FBS) | Sigma-Aldrich | F4135 | |
| Forceps Dumont 5 | Fine Science tools | 11254-20 | |
| Nuclease free water | Thermo Fisher Scientific | AM9932 | |
| PBS 10x | Research Product Internation (RPI) | P32200 | |
| Poly(A) Tail-Length Assay Kit | Thermo Fisher Scientific | 764551KT | |
| RiboRuler Low Range RNA Ladder | Thermo Fisher Scientific | SM1833 | |
| RNA Gel Loading Dye (2x) | Thermo Fisher Scientific | R0641 | |
| RNA microprep kit | Zymoresearch | R1050 | |
| RNA miniprep kit | Zymoresearch | R1055 | |
| Scissors-Vannas Spring Scissors - 2.5 mm Cutting Edge | Fine Science tools | 15000-08 | |
| TopVision Agarose Tablets | Thermo Fisher Scientific | R2802 | |
| Tris-Acetate-EDTA (TAE) | Thermo Fisher Scientific | B49 |
参考文献
- Stewart, M. Polyadenylation and nuclear export of mRNAs. Journal of Biological Chemistry. 294 (9), 2977-2987 (2019).
- Machida, K., et al. Dynamic interaction of poly(A)-binding protein with the ribosome. Scientific Reports. 8 (1), 17435(2018).
- Eisen, T. J., et al. The dynamics of cytoplasmic mRNA metabolism. Molecular Cell. 77 (4), 786-799 (2020).
- Liudkovska, V., Dziembowski, A. Functions and mechanisms of RNA tailing by metazoan terminal nucleotidyltransferases. Wiley Interdisciplinary Reviews RNA. 12 (2), e1622(2021).
- Goldstrohm, A. C., Wickens, M. Multifunctional deadenylase complexes diversify mRNA control. Nature Reviews Molecular Cell Biology. 9 (4), 337-344 (2008).
- Schmidt, M. J., Norbury, C. J. Polyadenylation and beyond: emerging roles for noncanonical poly(A) polymerases. Wiley interdisciplinary reviews RNA. 1 (1), 142-151 (2010).
- Laishram, R. S. Poly(A) polymerase (PAP) diversity in gene expression - Star-PAP vs canonical PAP. FEBS Letters. 588 (14), 2185-2197 (2014).
- Salles, F. J., Lieberfarb, M. E., Wreden, C., Gergen, J. P., Strickland, S. Coordinate initiation of Drosophila development by regulated polyadenylation of maternal messenger RNAs. Science. 266 (5193), 1996-1999 (1994).
- Wreden, C., Verrotti, A. C., Schisa, J. A., Lieberfarb, M. E., Strickland, S. Nanos and pumilio establish embryonic polarity in Drosophila by promoting posterior deadenylation of hunchback mRNA. Development. 124 (15), 3015-3023 (1997).
- Passmore, L. A., Coller, J. Roles of mRNA poly(A) tails in regulation of eukaryotic gene expression. Nature Reviews Molecular Cell Biology. 23 (2), 93-106 (2021).
- Murray, E. L., Schoenberg, D. R. Assays for determining poly(a) tail length and the polarity of mRNA decay in mammalian cells. Methods in Enzymology. 448, 483-504 (2008).
- Salles, F. J., Strickland, S. Analysis of poly(a) tail lengths by PCR: The PAT assay. Methods in Molecular Biology. 118, 441-448 (1999).
- Salles, F. J., Darrow, A. L., O'Connell, M. L., Strickland, S. Isolation of novel murine maternal mRNAs regulated by cytoplasmic polyadenylation. Genes and Development. 6 (7), 1202-1212 (1992).
- Salles, F. J., Strickland, S. Rapid and sensitive analysis of mRNA polyadenylation states by PCR. Genome Research. 4 (6), 317-321 (1995).
- Janicke, A., Vancuylenberg, J., Boag, P. R., Traven, A., Beilharz, T. H. ePAT: A simple method to tag adenylated RNA to measure poly(a)-tail length and other 3' RACE applications. RNA. 18 (6), 1289-1295 (2012).
- Minasaki, R., Rudel, D., Eckmann, C. R. Increased sensitivity and accuracy of a single-stranded DNA splint-mediated ligation assay (sPAT) reveals poly(a) tail length dynamics of developmentally regulated mRNAs. RNA Biology. 11 (2), 111-123 (2014).
- Martin, G., Keller, W. Tailing and 3'-end labeling of RNA with yeast poly(A) polymerase and various nucleotides. RNA. 4 (2), 226-230 (1998).
- Kusov, Y. Y., Shatirishvili, G., Dzagurov, G., Verena, G. M. A new G-tailing method for the determination of the poly(a) tail length applied to hepatitis a virus RNA. Nucleic Acids Research. 29 (12), 57(2001).
- Bazzini, A. A., Lee, M. T., Giraldez, A. J. Ribosome profiling shows that miR-430 reduces translation before causing mRNA decay in zebrafish. Science. 336 (6078), 233-237 (2012).
- Subtelny, A. O., Eichhorn, S. W., Chen, G. R., Sive, H., Bartel, D. P. Poly(a)-tail profiling reveals an embryonic switch in translational control. Nature. 508 (1), 66-71 (2014).
- Chang, H., Lim, J., Ha, M., Kim, V. N. TAIL-seq: Genome-wide determination of poly(a) tail length and 3' end modifications. Molecular Cell. 53 (6), 1044-1052 (2014).
- Legnini, I., Alles, J., Karaiskos, N., Ayoub, S., Rajewsky, N. FLAM-seq: Full-length mRNA sequencing reveals principles of poly(A) tail length control. Nature Methods. 16 (9), 879-886 (2019).
- Garalde, D. R., et al. Highly parallel direct RNA sequencing on an array of nanopores. Nature Methods. 15 (3), 201-206 (2018).
- Singh, M., Ye, B., Kim, J. H. Dual leucine zipper kinase regulates Dscam expression through a noncanonical function of the cytoplasmic poly(A)-binding protein. Journal of Neuroscience. 42 (31), 6007-6019 (2022).
- Macharia, R. W., Ombura, F. L., Aroko, E. O. Insects' RNA profiling reveals absence of "hidden break" in 28S ribosomal RNA molecule of onion thrips, Thrips tabaci. Journal of Nucleic Acids. 2015, 965294(2015).
- Miura, P., Sanfilippo, P., Shenker, S., Lai, E. C. Alternative polyadenylation in the nervous system: to what lengths will 3' UTR extensions take us. Bioessays. 36 (8), 766-777 (2014).
- Sement, F. M., et al. et al Uridylation prevents 3' trimming of oligoadenylated mRNAs. Nucleic Acids Research. 41 (14), 7115-7127 (2013).
Access restricted. Please log in or start a trial to view this content.
転載および許可
このJoVE論文のテキスト又は図を再利用するための許可を申請します
許可を申請さらに記事を探す
This article has been published
Video Coming Soon
Copyright © 2023 MyJoVE Corporation. All rights reserved