A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Ion Mobility-Mass Spectrometry Techniques for Determining the Structure and Mechanisms of Metal Ion Recognition and Redox Activity of Metal Binding Oligopeptides
In This Article
Summary
Ion mobility-mass spectrometry and molecular modeling techniques can characterize the selective metal chelating performance of designed metal-binding peptides and the copper-binding peptide methanobactin. Developing new classes of metal chelating peptides will help lead to therapeutics for diseases associated with metal ion misbalance.
Abstract
Electrospray ionization (ESI) can transfer an aqueous-phase peptide or peptide complex to the gas-phase while conserving its mass, overall charge, metal-binding interactions, and conformational shape. Coupling ESI with ion mobility-mass spectrometry (IM-MS) provides an instrumental technique that allows for simultaneous measurement of a peptide’s mass-to-charge (m/z) and collision cross section (CCS) that relate to its stoichiometry, protonation state, and conformational shape. The overall charge of a peptide complex is controlled by the protonation of 1) the peptide’s acidic and basic sites and 2) the oxidation state of the metal ion(s). Therefore, the overall charge state of a complex is a function of the pH of the solution that affects the peptides metal ion binding affinity. For ESI-IM-MS analyses, peptide and metal ions solutions are prepared from aqueous-only solutions, with the pH adjusted with dilute aqueous acetic acid or ammonium hydroxide. This allows for pH dependence and metal ion selectivity to be determined for a specific peptide. Furthermore, the m/z and CCS of a peptide complex can be used with B3LYP/LanL2DZ molecular modeling to discern binding sites of the metal ion coordination and tertiary structure of the complex. The results show how ESI-IM-MS can characterize the selective chelating performance of a set of alternative methanobactin peptides and compare them to the copper-binding peptide methanobactin.
Introduction
Copper and zinc ions are essential for living organisms and crucial to processes including oxidative protection, tissue growth, respiration, cholesterol, glucose metabolism, and genome reading1. To enable these functions, groups such as the thiolate of Cys, imidazole of His2,3, (more rarely) thioether of methionine, and carboxylate of Glu and Asp selectively incorporate metals as cofactors into the active sites of metalloenzymes. The similarity of these coordination groups raises an intriguing question regarding how the His and Cys ligands selectively incorporate either Cu(I/II) or Zn(II) to ensure correct functioning.
Selective binding is often accomplished by acquisition and trafficking peptides, which control Zn(II) or Cu(I/II) ion concentrations4. Cu(I/II) is highly reactive and causes oxidative damage or adventitious binding to enzymes, so its free concentration is tightly regulated by copper chaperones and copper-regulating proteins that transport it safely to various locations in the cell and tightly control its homeostasis5,6. Disruption of copper metabolism or homeostasis is directly implicated in Menkes and Wilson’s disease7 as well as cancers7 and neural disorders, such as prion8 and Alzheimer’s disease9.
Wilson’s disease is associated with increased copper levels in the eyes, liver and sections of the brain, where the redox reactions of Cu(I/II) produces reactive oxygen species, causing hepatolenticular and neurological degeneration. Existing chelation therapies are the small thiol amino acid penicillamine and triethylenetetramine. Alternatively, the methanotrophic copper-acquisition peptides methanobactin (mb)10,11 exhibit therapeutic potential because of their high binding affinity for Cu(I)12. When the methanobactin (mb-OB3b) from Methylosinus trichosporium OB3b was studied in an animal model of Wilson’s disease, copper was efficiently removed from the liver and excreted through the bile13. In vitro experiments confirmed that mb-OB3b could chelate the copper from the copper metallothionein contained in the liver cytosol13. Laser ablation inductively coupled plasma mass spectrometry imaging techniques have investigated the spatial distribution of copper in Wilson’s disease liver samples14,15,16 and shown that mb-OB3b removes the copper with short treatment periods of only 8 days17.
The mb-OB3b will also bind with other metal ions, including Ag(I), Au(III), Pb(II), Mn(II), Co(II), Fe(II), Ni(II), and Zn(II)18,19. Competition for the physiological Cu(I) binding site is exhibited by Ag(I) because it can displace Cu(I) from the mb-OB3b complex, with both Ag(I) and Ni(II) also showing irreversible binding to Mb which cannot be displaced by Cu(I)19. Recently, a series of alternative methanobactin (amb) oligopeptides with the 2His-2Cys binding motif have been studied20,21, and their Zn(II) and Cu(I/II) binding properties characterized. Their primary amino acid sequences are similar, and they all contain the 2His-2Cys motif, Pro and an acetylated N-terminus. They mainly differ from mb-OB3b because the 2His-2Cys motif replaces the two enethiol oxazolone binding sites of mb-OB3b.
Electrospray ionization coupled with ion mobility-mass spectrometry (ESI-IM-MS) provides for a powerful instrumental technique for determining the metal-binding properties of peptides because it measures their mass-to-charge (m/z) and collision cross section (CCS) while conserving their mass, charge, and conformational shape from the solution-phase. The m/z and CCS relate to the peptides stoichiometry, protonation state, and conformational shape. Stoichiometry is determined because the identity and number of each element present in the species is explicitly identified. The overall charge of the peptide complex relates to the protonation state of the acidic and basic sites and the oxidation state of the metal ion(s). The CCS gives information of the conformational shape of the peptide complex because it measures the rotational averaged size which relates to the tertiary structure of the complex. The overall charge state of the complex is also a function of pH and affects the peptide’s metal ion binding affinity because the deprotonated basic or acidic sites such as the carboxyl, His, Cys and Tyr are also the potential binding sites for the metal ion. For the analyses, the peptide and metal ion are prepared in aqueous solutions with the pH adjusted by dilute aqueous acetic acid or ammonium hydroxide. This allows for the pH dependence and metal ion selectivity to be determined for the peptide. Furthermore, the m/z and CCS determined by ESI-IM-MS can be used with B3LYP/LanL2DZ molecular modeling to discover the type of metal ion coordination and tertiary structure of the complex. The results shown in this article reveal how ESI-IM-MS can characterize the selective chelating performance of a set of amb peptides and compare them to the copper-binding peptide mb-OB3b.
Protocol
1. Preparation of reagents
- Culture Methylosinus trichosporium OB3b, isolate the Cu(I)-free mb-OB3b18,22,23, freeze-dry the sample and store at -80 °C until use.
- Synthesize the amb peptides (>98% purity for amb1, amb2, amb4; >70% purity for amb7), freeze-dry the samples, and store them at -80 °C until use.
- Purchase >98% purity manganese(II) chloride, cobalt(II) chloride, nickle(II) chloride, copper(II) chloride, copper(II) nitrate, silver(I) nitrate, zinc(II) chloride, iron(III) chloride and lead(II) chloride.
- Purchase the poly-DL-alanine polymers used as calibrants for measuring the collision cross sections of the amb species and HPLC grade or higher ammonium hydroxide, glacial acetic acid, and acetonitrile.
2. Preparation of stock solution
- Peptide stock solution
- Weigh accurately, using at least three significant figures, the mass of 10.0–20.0 mg of the mb-OB3b or amb in a 1.7 mL plastic vial.
NOTE: The weighed mass should yield either 12.5 mM or 1.25 mM, depending on the solubility of the peptide, when 1.00 mL of deionized (DI) water is added. - Using a pipet, add 1.00 mL of deionized water (>17.8 MΩ cm) to the weighed peptide sample to yield either the 12.5 mM or 1.25 mM solution. Place cap securely and mix thoroughly with at least 20 inversions.
- Using a micropipet dispense 50.0 μL aliquots from the peptide sample into individually labelled 1.5 mL vials and store them at -80 °C until use.
- Weigh accurately, using at least three significant figures, the mass of 10.0–20.0 mg of the mb-OB3b or amb in a 1.7 mL plastic vial.
- Metal ion stock solutions
- Weigh accurately, using at least three significant figures, the mass of 10.0–30.0 mg of the metal chloride or silver nitrate in a 1.7 mL vial.
NOTE: The weighed mass should yield 125 mM when 1.00 mL of DI water is added. - Add the 1.00 mL of DI water to the weighed metal sample in the 1.7 mL vial to yield the 125 mM solution. Place cap securely and mix thoroughly with at least 20 inversions.
- Weigh accurately, using at least three significant figures, the mass of 10.0–30.0 mg of the metal chloride or silver nitrate in a 1.7 mL vial.
- Ammonium hydroxide stock solutions: prepare a 1.0 M acetic acid solution by diluting 57 μL of the 99.5% acetic acid solution with DI water to a final volume of 1.00 mL. Prepare a 1.0 M ammonium hydroxide solution by diluting 90 μL of the 21% ammonium hydroxide solution with DI water to a final volume of 1.00 mL. Make two successive dilutions of each solution by taking 100 μL of the 1.0 M solutions to prepare 0.10 M and 0.010 M acetic acid and ammonium hydroxide solutions.
- Poly-DL-alanine stock solution: prepare the poly-DL-alanine (PA) by weighing 1.0 mg of PA and dissolving in 1.0 mL of DI water to give 1,000 ppm. Mix thoroughly. Using a micropipet, dispense 50.0 μL aliquots, and place each into a 1.7 mL vial and store at -80 °C.
3. Electrospray-ion mobility-mass spectrometry analysis
- Clean the ESI entrance tubing and needle capillary thoroughly with about 500 μL of 0.1 M glacial acetic acid, 0.1 M ammonium hydroxide, and finally DI water.
- Thaw a 50.0 μL aliquot of the 1,000 ppm PA stock solution and dilute it with 450 μL of DI water to give a 100 ppm PA. Pipet 100.0 μL of this solution and dilute it to 1.00 mL with 500 μL of DI water and 500 μL of acetonitrile to give 10 ppm PA solution.
- Collect the negative and positive ion IM-MS spectra of the 10 ppm PA solution for 10 min each using native ESI-IM-MS conditions as described in the discussion section.
- Thaw a 50.0 μL aliquot of the 12.5 mM or 1.25 mM amb stock solution and make successive dilutions with DI water to give a final concentration of 0.125 mM amb. Mix thoroughly each dilution.
- Pipet 100.0 μL of the 125 mM metal ion stock solution, place in a 1.7 mL vial and dilute to 1.00 mL with DI water to give 12.5 mM metal ion. Repeat with two more successive dilutions to give a final 0.125 mM metal ion concentration. Mix thoroughly each dilution.
- Pipet 200.0 μL of the 0.125 mM amb into a 1.7 mL vial, dilute with 500 μL of DI water, and mix the solution thoroughly.
- Adjust the pH of the sample to 3.0 by adding 50 μL of 1.0 M acetic acid solution.
- Add 200.0 μL of the 0.125 mM metal ion to the pH-adjusted sample. Add DI water to yield a final volume of 1.00 mL of the sample, mix thoroughly, and allow the sample to equilibrate for 10 min at RT.
- Using a blunt nose syringe take 500 μL of the sample and collect the negative and positive ion ES-IM-MS spectra for 5 min each. Use the remaining 500 μL of the sample to record its final pH using a calibrated micro pH electrode.
- Repeat steps 3.6–3.9, while modifying step 3.7 to adjust the pH to 4.0, 5.0, 6.0, 7.0, 8.0, 9.0, or 10.0 by adding new volumes of the 0.010 M, 0.10 M, or 1.0 M acetic acid or ammonium hydroxide solutions.
- Collect the negative and positive ion ESI-IM-MS spectra of the 10 ppm PA solution for 10 min each.
4. Preparation of the metal ion titration of amb samples
- Follow the steps described in steps 3.1–3.5.
- Pipet 200.0 μL of the 0.125 mM amb into a 1.7 mL vial, dilute with 500.0 μL of DI water and mix the solution thoroughly.
- Adjust the pH of the sample to pH = 9.0 by adding 80 μL of the 0.010 M ammonium hydroxide solution.
- Add 28 μL of the 0.125 mM metal ion solution to give 0.14 molar equivalents of the metal ion, add DI water to make the final volume of the sample 1.00 mL, mix thoroughly, and allow the sample to equilibrate for 10 min at RT.
- Using a blunt nose syringe take 500 μL of the sample and collect the negative and positive ion ESI-IM-MS spectra for 5 min each. Use the remaining 500 μL of the sample to record its final pH using a calibrated micro pH electrode.
- Repeat steps 4.2–4.5, while modifying step 4.3 to add an appropriate volume of the 0.125 mM metal ion solution to give either 0.28, 0.42, 0.56, 0.70, 0.84, 0.98, 1.12, 1.26, or 1.40 molar equivalents.
- Collect the negative and positive ion IM-MS spectra of the 10 ppm PA solution for 10 min each.
5. Analysis of ESI-IM-MS pH titration data
- From the IM-MS spectra identify which charged species of ambs are present by matching them to their theoretical m/z isotope patterns.
- Open MassLynx and click on Chromatogram to open the Chromatogram window.
- Go to the File menu and Open to locate and open the IM-MS data file.
- Extract the IM-MS spectrum by right-clicking and dragging across the chromatogram and releasing. The spectrum window will open showing the IM-MS spectrum.
- In the spectrum window, click on Tools and Isotope model. In the isotope modeling window, enter the molecular formula of the amb species, check the Show charged ion box, and enter the charge state. Click OK.
- Repeat to identify all the species in the IM-MS spectrum and record their m/z isotope range.
- For each amb species, separate any coincidental m/z species and extract their arrival time distributions (ATD) using their m/z isotope patterns to identify them.
- In MassLynx click on DriftScope to open the program. In DriftScope click on File and Open to locate and open the IM-MS data file.
- Use the mouse and left-click to zoom in on m/z isotope pattern of the amb species.
- Use the Selection tool and left mouse button to select the isotope pattern. Click the Accept current selection button.
- To separate any coincidental m/z species use the Selection tool and left mouse button to select the ATD time-aligned with isotope pattern of the amb species. Click the Accept current selection button.
- To export the ATD, go to File | Export to MassLynx, then select Retain Drift Time and save file in appropriate folder.
- Determine the centroid of the ATD and integrate the area under the ATD curve as a measure of the species population.
- In the Chromatogram window of MassLynx open the saved exported file. Click on Process | Integrate from the menu. Check the ApexTrack Peak Integration box and click OK.
- Record the centroid ATD (tA) and the integrated area as shown on the Chromatogram window. Repeat for all saved amb and PA IM-MS data files.
- Use the integrated ATD for all extracted amb species of either the positive or negative ions at each titration point to normalize to a relative percentage scale.
- Enter the identities of the amb species and their integrated ATD at each pH into a spreadsheet.
- For each pH, use the sum of the integrated ATDs to normalize the individual amb’s ATD to a percentage scale.
- Plot the percent intensities of each amb species vs. pH in a graph to show how the population of each species varies as a function of pH.
6. Collision cross-sections
- Using a spreadsheet, convert the CCSs (Ω) of PA negative25,26 and positive27 ions measured in He buffer gas28 to corrected CCS (Ωc) using Equation 1 below, where: z = ion charge; ec = electron charge (1.602×10-19 C); mN2 = mass of N2 gas (Da); and mion = ion mass.29
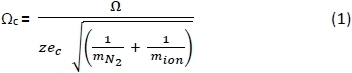
- Convert the average arrival times (tA) of the PA calibrants and amb species into drift times (tD) using Equation 2 below, where: c = the enhanced duty cycle delay coefficient (1.41), and m/z is the mass-to-charge of the peptide ion.

- Plot the PA calibrants’ tD vs. their Ωc. Then, using a least-squares regression fit of Equation 3 shown below, determine the A' and B values, where: A' is the correction for the temperature, pressure, and electric field parameters; and B compensates for the nonlinear effect of the IM device.

- Using these A' and B values and the centroid tD value from the ATD of the ambs determine their Ωc using Equation 3 and their Ω using Equation 1. This method provides CCSs for the peptide species with estimated absolute errors of about 2%25,26,27.
7. Computational methods
- Use the B3LYP/LanL2DZ level of theory, comprising of the Becke 3-parameter hybrid functionals30 and the Dunning basis set31 and electron core potentials32,33,34 to locate geometry-optimized conformers for all possible types of coordinations of the observed m/z amb species35.
NOTE: For details of how to build and submit calculations refer to the GaussView usage in Supplementary File. - Compare the predicted free energy of each of the conformers and calculate their theoretical CCSs using the ion-scaled Lennard-Jones (LJ) method from the Sigma program36.
- From the lowest free energy conformers determine which conformer exhibits the LJ CCS which agrees with the IM-MS measured CCS to identify the tertiary structure and type of coordination for the conformers observed in the experiment.
Results
Metal binding of amb1
The IM-MS study20 of amb1 (Figure 1A) showed that both copper and zinc ions bound to amb1 in a pH-dependent manner (Figure 2). However, copper and zinc bound to amb1 through different reaction mechanisms at different coordination sites. For example, adding Cu(II) to amb1 resulted in oxidation of amb1 (amb
Discussion
Critical steps: conserving solution-phase behaviors for examination via ESI-IM-MS
Native ESI instrumental settings must be used that conserve the peptides stoichiometry, charge state, and conformational structure. For native conditions, the conditions in the ESI source such as the cone voltages, temperatures, and gas flows have to be optimized. Also, the pressures and voltages in the source, trap, ion mobility, and transfer traveling waves (especially the DC trap bias that controls injection voltag...
Disclosures
The authors have nothing to disclose.
Acknowledgements
This material is based upon work supported by the National Science Foundation under 1764436, NSF instrument support (MRI-0821247), Welch Foundation (T-0014), and computing resources from the Department of Energy (TX-W-20090427-0004-50) and L3 Communications. We thank the Bower’s group of University of California - Santa Barbara for sharing the Sigma program and Ayobami Ilesanmi for demonstrating the technique in the video.
Materials
| Name | Company | Catalog Number | Comments |
| acetonitrile HPLC-grade | Fisher Scientific (www.Fishersci.com) | A998SK-4 | |
| ammonium hydroxide (trace metal grade) | Fisher Scientific (www.Fishersci.com) | A512-P500 | |
| cobalt(II) chloride hexahydrate 99.99% | Sigma-Aldrich (www.sigmaaldrich.com) | 255599-5G | |
| copper(II) chloride 99.999% | Sigma-Aldrich (www.sigmaaldrich.com) | 203149-10G | |
| copper(II) nitrate hydrate 99.99% | Sigma-Aldrich (www.sigmaaldrich.com) | 229636-5G | |
| designed amb1,2,3,4,5,6,7 peptides | Neo BioLab (neobiolab.com) | designed peptides were synthized by order | |
| designed amb5B,C,D,E,F peptides | PepmicCo (www.pepmic.com) | designed peptides were synthized by order | |
| Driftscope 2.1 software program | Waters (www.waters.com) | software analysis program | |
| Freeze-dried, purified, Cu(I)-free mb-OB3b | cultured and isolated in the lab of Dr. DongWon Choi (Biology Department, Texas A&M-Commerce) | ||
| glacial acetic acid (Optima grade) | Fisher Scientific (www.Fishersci.com) | A465-250 | |
| Iron(III) Chloride Anhydrous 98%+ | Alfa Aesar (www.alfa.com) | 12357-09 | |
| lead(II) nitrate ACS grade | Avantor (www.avantormaterials.com) | 128545-50G | |
| manganese(II) chloride tetrahydrate 99.99% | Sigma-Aldrich (www.sigmaaldrich.com) | 203734-5G | |
| MassLynx 4.1 | Waters (www.waters.com) | software analysis program | |
| nickel chloride hexahydrate 99.99% | Sigma-Aldrich (www.sigmaaldrich.com) | 203866-5G | |
| poly-DL-alanine | Sigma-Aldrich (www.sigmaaldrich.com) | P9003-25MG | |
| silver nitrate 99.9%+ | Alfa Aesar (www.alfa.com) | 11414-06 | |
| Waters Synapt G1 HDMS | Waters (www.waters.com) | quadrupole - ion mobility- time-of-flight mass spectrometer | |
| zinc chloride anhydrous | Alfa Aesar (www.alfa.com) | A16281 |
References
- Dudev, T., Lim, C. Competition among Metal Ions for Protein Binding Sites: Determinants of Metal Ion Selectivity in Proteins. Chemical Reviews. 114 (1), 538-556 (2014).
- Sovago, I., Kallay, C., Varnagy, K. Peptides as complexing agents: Factors influencing the structure and thermodynamic stability of peptide complexes. Coordination Chemistry Reviews. 256 (19-20), 2225-2233 (2012).
- Sóvágó, I., Várnagy, K., Lihi, N., Grenács, &. #. 1. 9. 3. ;. Coordinating properties of peptides containing histidyl residues. Coordination Chemistry Reviews. 327, 43-54 (2016).
- Rubino, J. T., Franz, K. J. Coordination chemistry of copper proteins: How nature handles a toxic cargo for essential function. Journal of Inorganic Biochemistry. 107 (1), 129-143 (2012).
- Robinson, N. J., Winge, D. R. Copper Metallochaperones . Annual Review of Biochemistry. 79, 537-562 (2010).
- Scheiber, I. F., Mercer, J. F. B., Dringen, R. Metabolism and functions of copper in brain. Progress in Neurobiology. 116, 33-57 (2014).
- Tisato, F., Marzano, C., Porchia, M., Pellei, M., Santini, C. Copper in Diseases and Treatments, and Copper-Based Anticancer Strategies. Medicinal Research Reviews. 30 (4), 708-749 (2010).
- Millhauser, G. L. Copper and the prion protein: Methods, structures, function, and disease. Annual Review of Physical Chemistry. 58, 299-320 (2007).
- Arena, G., Pappalardo, G., Sovago, I., Rizzarelli, E. Copper(II) interaction with amyloid-beta: Affinity and speciation. Coordination Chemistry Reviews. 256 (1-2), 3-12 (2012).
- Kim, H. J., et al. Methanobactin, a copper-acquisition compound from methane-oxidizing bacteria. Science. 305 (5690), 1612-1615 (2004).
- Di Spirito, A. A., et al. Methanobactin and the link between copper and bacterial methane oxidation. Microbiology Molecular Biology Reviews. 80 (2), 387-409 (2016).
- Kenney, G. E., Rosenzweig, A. C. Chemistry and biology of the copper chelator methanobactin. ACS Chemical Biology. 7 (2), 260-268 (2012).
- Summer, K. H., et al. The biogenic methanobactin is an effective chelator for copper in a rat model for Wilson disease. Journal of Trace Elements in Medicine and Biology. 25 (1), 36-41 (2011).
- Hachmoeller, O., et al. Investigating the influence of standard staining procedures on the copper distribution and concentration in Wilson's disease liver samples by laser ablation-inductively coupled plasma-mass spectrometry. Journal of Trace Elements in Medicine and Biology. 44, 71-75 (2017).
- Hachmoeller, O., et al. Spatial investigation of the elemental distribution in Wilson's disease liver after D-penicillamine treatment by LA-ICP-MS. Journal of Trace Elements in Medicine and Biology. 44, 26-31 (2017).
- Hachmoeller, O., et al. Element bioimaging of liver needle biopsy specimens from patients with Wilson's disease by laser ablation-inductively coupled plasma-mass spectrometry. Journal of Trace Elements in Medicine and Biology. 35, 97-102 (2016).
- Mueller, J. C., Lichtmannegger, J., Zischka, H., Sperling, M., Karst, U. High spatial resolution LA-ICP-MS demonstrates massive liver copper depletion in Wilson disease rats upon Methanobactin treatment. Journal of Trace Elements in Medicine and Biology. 49, 119-127 (2018).
- Choi, D. W., et al. Spectral and thermodynamic properties of Ag(I), Au(III), Cd(II), Co(II), Fe(III), Hg(II), Mn(II), Ni(II), Pb(II), U(IV), and Zn(II) binding by methanobactin from Methylosinus trichosporium OB3b. Journal of Inorganic Biochemistry. 100, 2150-2161 (2006).
- McCabe, J. W., Vangala, R., Angel, L. A. Binding Selectivity of Methanobactin from Methylosinus trichosporium OB3b for Copper(I), Silver(I), Zinc(II), Nickel(II), Cobalt(II), Manganese(II), Lead(II), and Iron(II). Journal of the American Society of Mass Spectrometry. 28, 2588-2601 (2017).
- Sesham, R., et al. The pH dependent Cu(II) and Zn(II) binding behavior of an analog methanobactin peptide. European Journal of Mass Spectrometry. 19 (6), 463-473 (2013).
- Wagoner, S. M., et al. The multiple conformational charge states of zinc(II) coordination by 2His-2Cys oligopeptide investigated by ion mobility - mass spectrometry, density functional theory and theoretical collision cross sections. Journal of Mass Spectrom. 51 (12), 1120-1129 (2016).
- Bandow, N. L., et al. Isolation of methanobactin from the spent media of methane-oxidizing bacteria. Methods in Enzymology. 495, 259-269 (2011).
- Choi, D. W., et al. Spectral and thermodynamic properties of methanobactin from γ-proteobacterial methane oxidizing bacteria: a case for copper competition on a molecular level. Journal of Inorganic Biochemistry. 104 (12), 1240-1247 (2010).
- Pringle, S. D., et al. An investigation of the mobility separation of some peptide and protein ions using a new hybrid quadrupole/travelling wave IMS/oa-ToF instrument. International Journal of Mass Spectrometry. 261 (1), 1-12 (2007).
- Forsythe, J. G., et al. Collision cross section calibrants for negative ion mode traveling wave ion mobility-mass spectrometry. Analyst. 14 (20), 6853-6861 (2015).
- Allen, S. J., Giles, K., Gilbert, T., Bush, M. F. Ion mobility mass spectrometry of peptide, protein, and protein complex ions using a radio-frequency confining drift cell. Analyst. 141 (3), 884-891 (2016).
- Bush, M. F., Campuzano, I. D. G., Robinson, C. V. Ion Mobility Mass Spectrometry of Peptide Ions: Effects of Drift Gas and Calibration Strategies. Analytical Chemistry. 84 (16), 7124-7130 (2012).
- Salbo, R., et al. Traveling-wave ion mobility mass spectrometry of protein complexes: accurate calibrated collision cross-sections of human insulin oligomers. Rapid Communications in Mass Spectrometry. 26 (10), 1181-1193 (2012).
- Smith, D. P., et al. Deciphering drift time measurements from travelling wave ion mobility spectrometry-mass spectrometry studies. European Journal of Mass Spectrometry. 15 (2), 113-130 (2009).
- Becke, A. D. Density-functional thermochemistry. III. The role of exact exchange. Journal of Chemical Physics. 98 (7), 5648-5652 (1993).
- Dunning, T. H., Hay, P. J. Gaussian basis sets for molecular calculations. Modern Theoretical Chemistry. 3, 1-27 (1977).
- Hay, P. J., Wadt, W. R. Ab initio effective core potentials for molecular calculations. Potentials for potassium to gold including the outermost core orbitals. Journal of Chemical Physics. 82 (1), 299-310 (1985).
- Hay, P. J., Wadt, W. R. Ab initio effective core potentials for molecular calculations. Potentials for the transition metal atoms scandium to mercury. Journal of Chemical Physics. 82 (1), 270-283 (1985).
- Wadt, W. R., Hay, P. J. Ab initio effective core potentials for molecular calculations. Potentials for main group elements sodium to bismuth. Journal of Chemical Physics. 82 (1), 284-298 (1985).
- . Gaussian 09, Revision C.01. Gaussian, Inc. , (2012).
- Wyttenbach, T., von Helden, G., Batka, J. J., Carlat, D., Bowers, M. T. Effect of the long-range potential on ion mobility measurements. Journal of the American Society of Mass Spectrometry. 8 (3), 275-282 (1997).
- Choi, D., et al. Redox activity and multiple copper(I) coordination of 2His-2Cys oligopeptide. Journal of Mass Spectrometry. 50 (2), 316-325 (2015).
- Rigo, A., et al. Interaction of copper with cysteine: stability of cuprous complexes and catalytic role of cupric ions in anaerobic thiol oxidation. Journal of Inorganic Biochemistry. 98 (9), 1495-1501 (2004).
- Vytla, Y., Angel, L. A. Applying Ion Mobility-Mass Spectrometry Techniques for Explicitly Identifying the Products of Cu(II) Reactions of 2His-2Cys Motif Peptides. Analytical Chemistry. 88 (22), 10925-10932 (2016).
- Choi, D., Sesham, R., Kim, Y., Angel, L. A. Analysis of methanobactin from Methylosinus trichosporium OB3b via ion mobility mass spectrometry. European Journal of Mass Spectrometry. 18 (6), 509-520 (2012).
- Martell, A. E., Motekaitis, R. J. NIST Standard Reference Database 46. Institute of Standards and Technology. , (2001).
- Pesch, M. L., Christl, I., Hoffmann, M., Kraemer, S. M., Kretzschmar, R. Copper complexation of methanobactin isolated from Methylosinus trichosporium OB3b: pH-dependent speciation and modeling. Journal of Inorganic Biochemistry. 116, 55-62 (2012).
- Amin, E. A., Truhlar, D. G. Zn Coordination Chemistry: Development of Benchmark Suites for Geometries, Dipole Moments, and Bond Dissociation Energies and Their Use To Test and Validate Density Functionals and Molecular Orbital Theory. Journal of Chemical Theory and Computation. 4 (1), 75-85 (2008).
- Sorkin, A., Truhlar, D. G., Amin, E. A. Energies, Geometries, and Charge Distributions of Zn Molecules, Clusters, and Biocenters from Coupled Cluster, Density Functional, and Neglect of Diatomic Differential Overlap Models. Journal of Chemical Theory and Computation. 5 (5), 1254-1265 (2009).
- Lillo, V., Galan-Mascaros, J. R. Transition metal complexes with oligopeptides: single crystals and crystal structures. Dalton Transactions. 43 (26), 9821-9833 (2014).
- Choutko, A., van Gunsteren, W. F. Conformational Preferences of a beta-Octapeptide as Function of Solvent and Force-Field Parameters. Helvetica Chimica Acta. 96 (2), 189-200 (2013).
- Angel, L. A. Study of metal ion labeling of the conformational and charge states of lysozyme by ion mobility mass spectrometry. European Journal of Mass Spectrometry. 17 (3), 207-215 (2011).
- Kelso, C., Rojas, J. D., Furlan, R. L. A., Padilla, G., Beck, J. L. Characterisation of anthracyclines from a cosmomycin D-producing species of Streptomyces by collisionally-activated dissociation and ion mobility mass spectrometry. European Journal of Mass Spectrometry. 15 (2), 73-81 (2009).
- El Ghazouani, A., et al. Copper-binding properties and structures of methanobactins from Methylosinus trichosporium OB3b. Inorganic Chemistry. 50 (4), 1378-1391 (2011).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved