A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Superior Auto-Identification of Trypanosome Parasites by Using a Hybrid Deep-Learning Model
In This Article
Summary
Worldwide medical blood parasites were automatically screened using simple steps on a low-code AI platform. The prospective diagnosis of blood films was improved by using an object detection and classification method in a hybrid deep learning model. The collaboration of active monitoring and well-trained models helps to identify hotspots of trypanosome transmission.
Abstract
Trypanosomiasis is a significant public health problem in several regions across the world, including South Asia and Southeast Asia. The identification of hotspot areas under active surveillance is a fundamental procedure for controlling disease transmission. Microscopic examination is a commonly used diagnostic method. It is, nevertheless, primarily reliant on skilled and experienced personnel. To address this issue, an artificial intelligence (AI) program was introduced that makes use of a hybrid deep learning technique of object identification and object classification neural network backbones on the in-house low-code AI platform (CiRA CORE). The program can identify and classify the protozoan trypanosome species, namely Trypanosoma cruzi, T. brucei, and T. evansi, from oil-immersion microscopic images. The AI program utilizes pattern recognition to observe and analyze multiple protozoa within a single blood sample and highlights the nucleus and kinetoplast of each parasite as specific characteristic features using an attention map.
To assess the AI program's performance, two unique modules are created that provide a variety of statistical measures such as accuracy, recall, specificity, precision, F1 score, misclassification rate, receiver operating characteristics (ROC) curves, and precision versus recall (PR) curves. The assessment findings show that the AI algorithm is effective at identifying and categorizing parasites. By delivering a speedy, automated, and accurate screening tool, this technology has the potential to transform disease surveillance and control. It could also assist local officials in making more informed decisions on disease transmission-blocking strategies.
Introduction
Trypanosomiasis is a significant challenge to global health issues due to a variety of zoonotic species causing human disease with a wide range of geographical distribution outside the African and American continents, such as South and Southeast Asia1,2,3. Human African trypanosomiasis (HAT) or sleeping sickness, is caused by Trypanosoma brucei gambiense and T. b. rhodesiense which produce the chronic and acute forms, respectively, representing the major spread in Africa. The causative parasite belongs to the Salivaria group due to the transmission by infected saliva of Tsetse flies4. Whereas, the well-known American trypanosomiasis (Chagas's disease) caused by T. cruzi has been a public health concern for non-endemic countries; including Canada, the USA, Europe, Australia, and Japan, because of the frequent migration of individuals from endemic areas5. The trypanosome infection belongs to the Stercoraria group because it is transmitted by the infected feces of reduviid bugs. The trypanosomiases and trypanosomoses (Surra disease) caused by the T. evansi infection are endemic in Africa, South America, Western and Eastern Asia, and South and Southeast Asian countries3,6. Although human trypanosomiasis caused by the trypanosome has been reported3,4,7,8,9,10,11,12, the route of transmission of the parasite infection is debated: either the mechanical or infected blood through hematophagous insects such as tsetse flies and tabanids or horse flies6,7,8,9,10,12,13,14. No case report has been found in Thailand, however, a high prevalence of the T. evansi infection in dog15, racing horses, and water buffalo in the eastern region has been published16, suggesting an acquired transmission between domestic animals would have occurred. Several atypical human infections caused by animal trypanosomes (T. vivax, T. b. brucei, T. congolense, T. lewisi, and T. evansi) were reported, which are not the classical forms of human trypanosomes17. Awareness about atypical human infections might be underestimated, highlighting the need for improved diagnostic tests and field investigations for detection and confirmation of these atypical cases, and allowing for proper control and treatment of animal pathogenic diseases that affect global livestock, food security18, and human healthcare. This led to the development of a potential strategy integrated with an existing common method (microscopic examination) to rapidly screen blood samples in remote areas during active surveillance, enabling the identification of the hotspot zones for restricting and controlling the disease.
Having a sporadic incidence of the Surra disease in a wide range of domestic animals such as dromedaries, cattle, equines, and dogs that evoke a euryxenous T. evansi may be zoonotic to humans1,4,13,14. Human infection seems impossible because a trypanolytic factor in human serum, expressed from a sra-like gene, is capable of preventing human T. brucei and T. congolense12,19. Furthermore, as the first case report from India demonstrates, the illness has no association with immunocompromised HIV patients4. As described above, the possible human infection may be related to a high-density lipoprotein deficiency with abnormal function of the trypanosome lytic factor, which is a rare autosomal recessive genetic disorder, namely Tangier disease4. In 2016, a Vietnamese patient was discovered to possess two wild-type APOL1 alleles and a serum APOL1 concentration within the normal range. However, the theory of APOL-1 deficiency is no longer considered valid12. Therefore, one possible mechanism of trypanosome infection is direct contact of a wound with infected animal blood during occupational animal farming4,12. Microscopic examination reveals that T. evansi morphology is a monomorphic form of the trypomastigote including a predominant long slender, flagellated, and dividing trypanosome which is similar to their relative species of T. brucei1,12,13. The nucleus is in the central position with a visible small kinetoplast in the posterior position. A previous study indicated that the parasite can exist in two comparable forms, known as the classical and truncated forms. However, it remains necessary to confirm their respective pathogenic effects on hosts20. The course of symptoms varies ranging from intermittent fever associated with chills and sweating. Suramin, fortunately, is a successful first-line therapy for early-stage human African trypanosomiasis with no invasion of the central nervous system (CNS), healing patients in India and Vietnam4,12,21.
Except for clinical sign examination, several diagnostic methods for T. evansi parasites exist, including parasitological microscopic observation4,9,12, serological4,8,9,10,12, and molecular biological tests4,12. Thin-blood films stained with Giemsa are often used to visualize the parasite present under microscopic examination, which is routinely and commonly used22. However, the procedure appears to be feasible; nonetheless, it is time-consuming and labor-intensive, has inter-rater assessment variability, is sensitive to only an acute phase, and requires a personal trainee23. Both molecular biology and serological testing also needed highly skilled personnel to perform multiple processes of sample preparation, including extracting and purifying the samples before testing them with expensive apparatus, which is difficult to standardize, risk of contamination with extra-parasitic materials, and discrepancies in results24. Based on the rationale described above, rapid and early screening technology is needed to support the field surveillance study and ensure that the survey result is reported in a timely manner to identify the hotspot zone for further control of the disease transmission1,8. Computerized-based devices (CAD) have been proposed as an innovative technology for medical fields, including histopathological and cytopathological tasks25. The CAD mentioned above was performed at high speed and computed using pattern recognition, namely, artificial intelligence (AI). The AI method is accomplished using convolutional neural network algorithms that can be used to deal with a large number of dataset samples, especially, a supervised learning approach that trains a well-trained model upon data consumption.
In general, AI is the ability of computers to solve tasks that require expert intelligence, such as data labeling. Machine learning (ML), a subfield of AI, is represented as a computer system with two different processes comprised of feature extraction and pattern recognition. Deep learning (DL), or advanced ML algorithms, refers to the development of computerized programs and devices comparing human-like performance with levels of accuracy greater and equal to that accomplished by human professionals26. Currently, the role of DL in medical and veterinary fields is promisingly expanding and revolutionizing communicable disease prevention with the aim of recent prevention and guiding it to individual health staff22,27. The potential DL application is limitless with quality labels and a large number of augmented datasets, freeing specialists to manage the project task. Specifically, an advance in the digital image along with computer-assisted analysis, improved the automatic diagnostic and screening in five categories of pathology reported; including static, dynamic, robotic, whole slide imaging, and hybrid methods28. It is necessary to consider that the integration of DL algorithm approaches and digital image data could encourage local staff to utilize the technology in their daily practices.
Previously, the increase in prediction accuracy of using a hybrid model had been proven27. To identify the trypanosome parasite in microscopic images, this research presents two hybrid models, incorporating the YOLOv4-tiny (object detection) and Densenet201 (object classification) algorithms. Among several detection models, YOLOv4-tiny with a CSPDarknet53 backbone showed high performance as a prediction result in terms of localization and classification29. Since the real-time detector has modified the optimal balance among the input network resolution, the amount of the convolutional layer, the total parameter, and the number of layer outputs, it has improved prioritizing fast operating speeds and optimizing for parallel computations when compared to previous versions. Dense Convolutional Network (DenseNet) is another popular model that achieves state-of-the-art results across competitive datasets. DenseNet201 yielded a similar validation error comparable to that of ResNet101; however, DenseNet201 has fewer than 20 million parameters, which is less than ResNet101's more than 40 million parameters30. Therefore, the DenseNet model could improve prediction accuracy with an increasing number of parameters with no sign of overfitting. Here, an artificial intelligence (AI) program utilizes a hybrid deep learning algorithm with deep detection- and classification neural network backbones on the in-house CiRA CORE platform. The developed program can identify and classify the protozoan trypanosome species, namely Trypanosoma cruzi, T. brucei, and T. evansi, from oil-immersion microscopic images. This technology has the potential to revolutionize disease surveillance and control by providing a rapid, automated, and accurate screening method. It could aid local staff in making more informed decisions on transmission-blocking strategies for parasitic protozoan disease.
Protocol
Archived blood films and project design were approved by the Institutional Biosafety Committee, the Institutional Animal Care and Use Committee of the Faculty of Veterinary Science, Chulalongkorn University (IBC No. 2031033 and IACUC No. 1931027), and Human Research Ethics Committee of King Mongkut's Institute of Technology Ladkrabang (EC-KMITL_66_014).
1. Preparation of raw images
- The image dataset preparation
- Obtain at least 13 positive slides with blood-parasite infections, including T. brucei, T. cruzi, and T. evansi, confirmed by parasitologist experts. Separate the 13 slides for training (10 slides) and testing (three slides).
- Acquire images of the Giemsa stained-thin blood films described above under an oil-immersion field of a light microscope with a digital camera. Obtain images containing multiple objects of the trypomastigotes of all three parasite species under microscopic examination; look for a slender shape, long tails, an undulating membrane, and a kinetoplast at the anterior end.
NOTE: Creating both thick and thin smears would enhance the detection of acute phase trypanosomiasis31. The blood collection by finger-prick is recommended by WHO32. Nevertheless, thin films are more effective in identifying Trypanosoma cruzi and other species, as these organisms tend to become distorted in thick films33. In light of this, we utilized thin blood film images to maintain the appropriate morphology of the parasites for this study. - Store all images in a parasite-specific folder with the following specifications: 1,600 x 1,200 pixels, 24-bit depth, and JPG file format. Split the images into the training and test sets at a ~6:1 ratio.
NOTE: See https://gitlab.com/parasite3/superior-auto-identification-of-medically-important-trypanosome-parasites-by-using-a-hybrid-deep-learning-model/-/blob/main/JOVEimage.zip; 650 images were split to train (560 images) and test (90 images) model. - Define the region of interest as a rectangular label for two classes: trypanosomes and non-trypanosomes. Use the auto-cropping module to crop all detected images by using the well-trained object detection model. The auto-cropping module is the module developed in the in-house CiRA CORE program (see Table of Materials). Collect a single object per image for training the object classification.
NOTE: For this paper, 1,017 images were split for training (892 images) and testing (126 images). The model training was performed with four labeled classes, including leukocyte, T. brucei, T. cruzi, and T. evansi.
2. Training process with in-house CiRA CORE platform
- Starting a new project
- Open the CiRA CORE application from the computer desktop (see Table of Materials) and create a new project by double-clicking on the program's icon.
- Choose the operation icon on the left vertical toolbar to select the required tools.
- Object detection model training
- Select the training-DL model function for data labeling and training by using the drag-and-drop method. Go to the General toolbar | CiRA AI | Drag DeepTrain | Drop DeepTrain on the screen (right-hand side).
NOTE: For additional options, right-click on the selected tool and perform the appropriate functions: Copy, Cut, or Delete. - Import the images using DeepTrain tool's settings. Click on the Load images button and navigate to the image directory. Label the objects by holding the left-click and naming the selected object. Adjust the rectangle line thickness and font size by clicking on the Display Setting button and Save GT as a .gt file in the same directory.
NOTE: Save as needed to avoid any undesired conditions such as power shortage, automatic program closures, and hanging within the labeling process. - Prior to model training, expand the data to gather sufficient information using the four augmentation techniques: Rotation, Contrast, Noise, and Blur. Click the Gen Setting button to access this feature.
- Initiate model training by clicking the Training button in the DeepTrain tool. The training part has two sub-functions: Generate Training Files and Train. Under the Generate Training Files function, select the desired models, batch size, and subdivisions. Click the Generate button to generate data and save it in the directory. In the Train function, choose the following options: i) use another generated training location for conditions and backup, ii) use prebuilt weights for continued training, or iii) override parameters for current training design. This will design the model configuration and training conditions.
NOTE: The generation process time depends on the image file size, augmentation usage, and available memory space. - Once all necessary configurations are complete, begin the model training by clicking on the Train button. Allow the program to continuously execute, evaluating the training loss and adjusting the weight of the dataset during the training process. If the model achieves optimal loss, save the trained weight file in the specified directory by clicking on the Export button.
- Select the training-DL model function for data labeling and training by using the drag-and-drop method. Go to the General toolbar | CiRA AI | Drag DeepTrain | Drop DeepTrain on the screen (right-hand side).
3. Object detection model evaluation
- Select the object detection model evaluation function for model evaluation using the drag-and-drop method. Go to the Plugin toolbar | Evaluate | Drag EvalDetect | Drop EvalDetect on the screen (right-hand side).
- Click on Setting and wait for three functions: Detection, Evaluate, and Plot. Initiate model evaluation by importing the trained weight file from the directory (step 2.2.5) by clicking on Load Config.
- Under the Detection function, select non-maximum suppression (NMS) value to enhance accuracy by eliminating redundant false positive (FP) detections. NMS removes duplicate model-generated detections for improved reliability.
- Proceed with the following steps under the Evaluation function:
- Import test images from the image file directory by clicking on Browse. Import the GT file from the directory where it was saved in step 2.2.2 by clicking on Load GT.
- Choose the Intersection over Union (IoU) value to assess accuracy on the specific image test dataset.
- Click the Evaluation button to assess the detection model in the specified directory. Once the evaluation is completed, the results will be automatically saved as a CSV file in the same directory, sorted by class name. This CSV file will provide essential parameters such as True Positive (TP), False Positive (FP), False Negative (FN), Recall, and Precision for each class.
- To plot the Precision-Recall (PR) curve, follow these steps under the Plot function: Import the CSV files from the previous section (step 3.4) directory by clicking on Browse. Choose classes from the list and click the Plot button to display the editable PR curve image.
- Finally, to save an image with the AUC values of the PR curve in the required image format at the specified directory, click on the Save button of the image.
4. Image cropping for a single object per image
- Prior to cropping the images, complete the following steps:
- Import the images from the image file directory by accessing the settings of the Image Slide tool.
- Import the trained weight file (saved in step 2.2.8) by accessing the settings of the Deep Detect tool. Click on the Config button | + button, select the backend (CUDA or CPU), provide a name, click OK, choose the weight file directory, and click Choose. Within the Deep Detect tool, select the detection parameters (threshold and non-maxima suppression (nms)); drawing parameters; tracking parameters; and region of interest (ROI) parameters.
- Select the directory where the cropped images will be saved by accessing the settings of the Deep Crop tool. Click Browse | choose the directory to save the cropped images | click Choose | select the image format (jpg or png) | enable the Auto Save option.
- Crop images to obtain a single object per image for image classification and segmentation. To carry out this process, utilize four tools and establish connections between them: go to the General toolbar | General | Button Run. Next, navigate to General toolbar | CiRA AI | DeepDetect; then, go to General toolbar | CiRA AI | DeepCrop. Finally, go to Image toolbar | Acquisition | ImageSlide.
- Once all the necessary settings are in place, initiate the image cropping process by clicking on the Button Run tool.
- Obtain a new image training dataset consisting of single-object images with a size of 608 x 608.
5. Image classification as model training
- Use drag-and-drop to select the image classification model training function for data training. Go to the Image toolbar | DeepClassif | Drag ClassifTrain | Drop ClassifTrain on the screen.
- Import images for model training using the ClassifTrain tool's settings. Click on the Open folder button and navigate to the desired image directory. Before training, expand the data by clicking on the Augmentation button for more information using techniques such as Rotation, Contrast, Flipping (horizontal and/or vertical), Noise, and Blur.
- To commence model training, click on the GenTrain button of the ClassifTrain tool. Under the GenTrain function, select the models, batch size, and subdivisions. Assign a directory to save the generated file. Click the Generate button to proceed with data for training. In the Train function, tick the appropriate options: Continue training with default weight or custom weight.
NOTE: The generation process may take time depending on factors such as image file size, augmentation usage, class balancing, and available memory space. - Once all preparations are complete, initiate the model training by clicking the Start button. Allow the program to execute continuously, evaluating the training loss and adjusting the weight of the dataset during the training process. If the model achieves the desired level of loss, save the trained weight file to the specified directory by clicking on the Export button.
6. Classification model evaluation
- Select the image classification model evaluation function for model evaluation using the drag-and-drop method. Go to the Plugin toolbar | Evaluate | Drag EvaluateClassif | Drop EvaluateClassif on the screen (the right-hand side).
- Click on Setting to access additional functions within the EvaluateClassif tool, namely Evaluate and PlotROC.
- To initiate model evaluation, click on the Evaluate button in the EvaluateClassif tool. Follow these steps under the Evaluate function.
- Import the test images from the image file directory by clicking on the Load folder image. Import the trained weight file from the directory (saved in step 5.4) by clicking on Load Config. Click the Start button to evaluate the classification model.
- Once the evaluation is complete, save the evaluated file as CSV in the specified directory by clicking on the Export to CSV button. For evaluation of data at every threshold, save the CSV file with class names in the specified directory by clicking on Start all threshold. The saved CSV file includes parameters such as Recall (True Positive Rate), False Positive Rate, and Precision for each class.
- To plot the Receiver Operating Characteristics (ROC) curve, click on the PlotROC button within the EvaluateClassif tool. Follow these steps under the PlotROC function.
- Import CSV files from the directory obtained earlier by clicking on Browse. Inspect the imported class list and select each class label to plot the ROC curve.
- Click the Plot button to visualize the ROC curve as an image. Make the desired edits to adjust image properties, including font size, font colors, rounding the decimal, line styles, and line colors.
- Finally, save an image of the ROC curve with the AUC values in the required image format at the specified directory by clicking on the Save button.
7. Testing the process with the CiRA CORE application
- Object detection as model testing
- To perform model testing, utilize four tools and establish connections between them. Go to the General toolbar | General | Button Run. Then, General toolbar | General | Debug. After that, click on General toolbar | CiRA AI | DeepDetect, and finally Image toolbar | Acquisition | ImageSlide.
- Before testing the images, follow these steps:
- Import the test images from the image file directory by clicking on the Setting option in the Image Slide tool.
- Import the saved trained weight file from step 2.2.8 by clicking on the Setting option in the DeepDetect tool. Click on the Config button, then the + button, select the backend (CUDA or CPU), provide a name, click OK, choose the weight file directory, and click Choose. Under the DeepDetect tool, select the detection parameters (Threshold and nms), drawing parameters, tracking parameters, and ROI parameters.
- View the test image results by clicking on the image function in the Debug tool.
- Finally, check the predicted results for each image by clicking on the Run button on the Button Run tool.
- Image classification as model testing
- To perform model testing, utilize four tools and establish connections between them. Go to the General toolbar | General | Button Run; then, General toolbar | Debug. After that, navigate to Image toolbar | Acquisition | ImageSlide, and finally, Image toolbar | DeepClassif | DeepClassif.
- Before testing the images, follow these steps:
- Import the test images from the image file directory by clicking on the Setting option in the Image Slide tool.
- Import the saved trained weight file from section 5.5 by clicking on the Setting option in the DeepClassif tool. Click on the Config button | + button | select the backend (CUDA or CPU) | provide a name | click OK | choose the weight file directory | click Choose. Under the DeepClassif tool, select the classification parameters (Threshold and number of top-class predictions), Guide map parameters (threshold, alpha, beta, and color map), and various parameters in the color map.
- View the test image results by clicking on the image function in the Debug tool.
- Finally, check the predicted results for each image by clicking on the Run button on the Button Run tool.
8. Hybrid (detection and classification) as model testing
- To perform this model testing, utilize four tools and establish connections between them. Go to the General toolbar | General | ButtonRun. Then, General toolbar | General | Debug. After that, Image toolbar | Acquisition | ImageSlide, and finally, Image toolbar | DeepComposite | DeepD->C.
- Before testing the images, follow these steps: Import test images from the image file directory by clicking on the Setting option in the Image Slide tool. Import the two saved trained weight files from section 2.1.5 and section 4.4 by clicking on the Setting option in the DeepD->C tool:
- For the Detect function, click on the Config button |+ button, select the backend (CUDA or CPU) | provide a name | click OK | choose the weight file directory | click Choose. Under the Detect function, select the detection parameters (Threshold and nms), drawing parameters, tracking parameters, and ROI parameters.
- For the Classif function, click on the Config button |+ button, select the backend (CUDA or CPU) | provide a name | click OK | choose the weight file directory | click Choose. Under the Classif function, select the classification parameters (Threshold and number of top-class predictions) and Guide map parameters (threshold, alpha, beta, and color map).
- View the test image results by clicking on the image function in the Debug tool. Finally, check the predicted results for each image by clicking on the Run button on the Button Run tool.
9. Five-fold cross-validation
NOTE: To validate the performance of the proposed model more effectively, K-fold cross-validation is used.
- Divide the dataset into five sections, corresponding to the five folds of cross-validation. During each iteration of model training and testing, use one section as the validation set for testing and the remaining four sections for training. Repeat this process five times, with each fold being used as the validation set once.
- For Folds 1 through 5:
- Repeat section 5 to train the model using the training data from the four folds.
- Repeat section 7.2 to test the model using the remaining fold as the test set.
10. Model evaluation
- Confusion matrix
- Based on the test results, the four conditions will happen as follows:
- True Positive (TP): When the input image is true, and the prediction is also true.
- False Positive (FP): When the input image is false, but the prediction is true.
- False Negative (FN): When the input image is true, but the prediction is false.
- True Negative (TN): When the input image is false, and the prediction is also false.
- Using these four conditions, evaluate the performances with the confusion matrix.
- Based on the test results, the four conditions will happen as follows:
- Performance evaluations
- The most commonly used classification performance metrics are accuracy, precision, recall, specificity, and F1-score values. Calculate all evaluation metrics in equations (1-6) used to evaluate model performance from values from the confusion matrix.
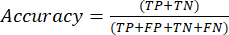 (1)
(1)
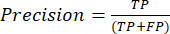 (2)
(2)
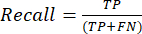 (3)
(3)
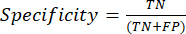 (4)
(4)
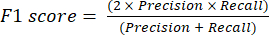 (5)
(5)
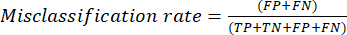 (6)
(6)
- The most commonly used classification performance metrics are accuracy, precision, recall, specificity, and F1-score values. Calculate all evaluation metrics in equations (1-6) used to evaluate model performance from values from the confusion matrix.
- ROC curve
NOTE: The ROC curve is a performance measure for classification problems with different threshold settings. The area under the ROC curve (AUC) represents the degree or measure of separability, while the ROC is a probability curve.- The ROC curve is a two-dimensional graph with the true positive rate (TPR) and false positive rate (FPR) values plotted on the Y and X axes, respectively. Construct the ROC curves using the TPR and TFR values obtained from the confusion matrix. The TPR value is the same as the sensitivity; calculate the FPR value using the equation (7).
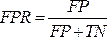 (7)
(7) - After obtaining the TPR and FPR values, plot the ROC curve using the Jupyter Notebook open-source web tool in a Python environment. The AUC is an effective way to assess the performance of the proposed model in ROC curve analysis.
- The ROC curve is a two-dimensional graph with the true positive rate (TPR) and false positive rate (FPR) values plotted on the Y and X axes, respectively. Construct the ROC curves using the TPR and TFR values obtained from the confusion matrix. The TPR value is the same as the sensitivity; calculate the FPR value using the equation (7).
- PR curve
- Use the PR curve to evaluate models by measuring the area under the PR curve. Construct the PR curve by plotting the models' precision and recall using the model's confidence threshold functions. Because the PR curve is also a two-dimensional graph, plot Recall on the x-axis and Precision on the y-axis.
- Plot the PR curve, like the ROC curve, using the open-source Jupyter Notebook web tool in a Python environment. The area under the Precision-Recall curve (AUC) score is also helpful in multilabel classification.
Results
In this study, hybrid deep learning algorithms were proposed to help automatically predict the positivity of a blood sample with a trypanosome parasite infection. Archived, Giemsa-stained blood films were sorted to localize and classify the parasitized versus non-parasitic by using the object detection algorithm based on a darknet backbone neural network. Within any rectangular box prediction result obtained by the previous model, the best-selected classification model was developed to classify all three species of medic...
Discussion
Microscopic observation for Trypanosoma protozoa infection is early and commonly used, especially during surveillance in remote areas where there is a lack of skilled technicians and labor-intensive and time-consuming processes that are all obstacles to reporting the health organization timely. Although molecular biology techniques such as immunology and polymerase chain reaction (PCR) have been approved as high-sensitivity methods to support the effectiveness of lab findings, expensive chemicals, apparatus, and professi...
Disclosures
All authors have no financial disclosures and no conflicts of interest.
Acknowledgements
This work (Research grant for New Scholar, Grant No. RGNS 65 - 212) was financially supported by the Office of the Permanent Secretary, Ministry of Higher Education, Science, Research and Innovation (OPS MHESI), Thailand Science Research and Innovation (TSRI) and King Mongkut's Institute of Technology Ladkrabang. We are grateful to the National Research Council of Thailand (NRCT) [NRCT5-RSA63001-10] for funding the research project. M.K. was funded by Thailand Science Research and Innovation Fund Chulalongkorn University. We also thank the College of Advanced Manufacturing Innovation, King Mongkut's Institute of Technology, Ladkrabang who have provided the deep learning platform and software to support the research project.
Materials
| Name | Company | Catalog Number | Comments |
| Darknet19, Darknet53 and Densenet201 | Gao Huang, Z. L., Laurens van der Maaten. Densely Connected Convolutional Networks. arXiv:1608.06993 [cs.CV]. (2016) | https://github.com/liuzhuang13/DenseNet | Deep convolutional neural network model that can function to classification Generic name: YOLO model/ detection model? |
| Olympus CX31 Model CX31RRBSFA | Olympus, Tokyo, Japan | SN 4G42178 | A light microscope |
| Olympus DP21-SAL U-TV0.5XC-3 | Olympus, Tokyo, Japan | SN 3D03838 | A digital camera Generic name: Classification models/ densely CNNs |
| Window 10 | Microsoft | Window 10 | Operation system in computers |
| YOLO v4-tiny | Naing, K. M. et al. Automatic recognition of parasitic products in stool examination using object detection approach. PeerJ Comput Sci. 8 e1065, (2022). | https://git.cira-lab.com/users/sign_in | Deep convolutional neural network model that can function to both localization and also classification |
| https://git.cira-lab.com/users/sign_in |
References
- Kasozi, K. I., et al. Epidemiology of trypanosomiasis in wildlife-implications for humans at the wildlife interface in Africa. Frontiers in Veterinary Science. 8, 621699 (2021).
- Ola-Fadunsin, S. D., Gimba, F. I., Abdullah, D. A., Abdullah, F. J. F., Sani, R. A. Molecular prevalence and epidemiology of Trypanosoma evansi among cattle in peninsular Malaysia. Acta Parasitologica. 65 (1), 165-173 (2020).
- Aregawi, W. G., Agga, G. E., Abdi, R. D., Buscher, P. Systematic review and meta-analysis on the global distribution, host range, and prevalence of Trypanosoma evansi. Parasites & Vectors. 12 (1), 67 (2019).
- Joshi, P. P., et al. Human trypanosomiasis caused by Trypanosoma evansi in India: the first case report. The Am Journal of Tropical Medicine and Hygiene. 73 (3), 491-495 (2005).
- Lidani, K. C. F., et al. Chagas disease: from discovery to a worldwide health problem. Frontiers in Public Health. 7, 166 (2019).
- Sazmand, A., Desquesnes, M., Otranto, D. Trypanosoma evansi. Trends in Parasitology. 38 (6), 489-490 (2022).
- Powar, R. M., et al. A rare case of human trypanosomiasis caused by Trypanosoma evansi.Indian. Journal of Medical Microbiology. 24 (1), 72-74 (2006).
- Shegokar, V. R., et al. Short report: Human trypanosomiasis caused by Trypanosoma evansi in a village in India: preliminary serologic survey of the local population. American Journal of Tropical Medicine and Hygiene. 75 (5), 869-870 (2006).
- Haridy, F. M., El-Metwally, M. T., Khalil, H. H., Morsy, T. A. Trypanosoma evansi in dromedary camel: with a case report of zoonosis in greater Cairo, Egypt. Journal of the Egyptian Society of Parasitology. 41 (1), 65-76 (2011).
- Dey, S. K. CATT/T.evansi antibody levels in patients suffering from pyrexia of unknown origin in a tertiary care hospital in Kolkata. Research Journal of Pharmaceutical, Biological and Chemical Sciences. 5, 334-338 (2014).
- Dakshinkar, N. P., et al. Aberrant trypanosomias in human. Royal Veterinary Journal of India. 3 (1), 6-7 (2007).
- Vn Vinh Chau, N., et al. A clinical and epidemiological investigation of the first reported human infection with the zoonotic parasite Trypanosoma evansi in Southeast Asia. Clinical Infectious Diseases. 62 (8), 1002-1008 (2016).
- Misra, K. K., Roy, S., Choudhary, A. Biology of Trypanosoma (Trypanozoon) evansi in experimental heterologous mammalian hosts. Journal of Parasitic Diseases. 40 (3), 1047-1061 (2016).
- Nakayima, J., et al. Molecular epidemiological studies on animal trypanosomiases in Ghana. Parasites & Vectors. 5, 217 (2012).
- Riana, E., et al. The occurrence of Trypanosoma in bats from Western Thailand. The 20th Chulalongkorn University Veterinary Conference CUVC 2021: Research in practice. 51, (2021).
- Camoin, M., et al. The Indirect ELISA Trypanosoma evansi in equids: optimisation and application to a serological survey including racing horses, in Thailand. BioMed Research International. 2019, 2964639 (2019).
- Truc, P., et al. Atypical human infections by animal trypanosomes. PLoS Neglected Tropical Diseases. 7 (9), 2256 (2013).
- Desquesnes, M., et al. Diagnosis of animal trypanosomoses: proper use of current tools and future prospects. Parasites & Vectors. 15 (1), 235 (2022).
- Da Silva, A. S., et al. Trypanocidal activity of human plasma on Trypanosoma evansi in mice. Revista Brasileira de Parasitologia Veterinaria. 21 (1), 55-59 (2012).
- Desquesnes, M., et al. Trypanosoma evansi and surra: a review and perspectives on transmission, epidemiology and control, impact, and zoonotic aspects. BioMed Research International. 2013, 321237 (2013).
- World Health Organization. A new form of human trypanosomiasis in India. Description of the first human case in the world caused by Trypanosoma evansi. Weekly Epidemiological Record. 80 (7), 62-63 (2005).
- Naing, K. M., et al. Automatic recognition of parasitic products in stool examination using object detection approach. PeerJ Computer Science. 8, 1065 (2022).
- Wongsrichanalai, C., Barcus, M. J., Muth, S., Sutamihardja, A., Wernsdorfer, W. H. A review of malaria diagnostic tools: microscopy and rapid diagnostic test (RDT). American Journal of Tropical Medicine and Hygiene. 77, 119-127 (2007).
- Rostami, A., Karanis, P., Fallahi, S. Advances in serological, imaging techniques and molecular diagnosis of Toxoplasma gondii infection. Infection. 46 (3), 303-315 (2018).
- Ahmad, Z., Rahim, S., Zubair, M., Abdul-Ghafar, J. Artificial intelligence (AI) in medicine, current applications and future role with special emphasis on its potential and promise in pathology: present and future impact, obstacles including costs and acceptance among pathologists, practical and philosophical considerations. A comprehensive review. Diagnostic Pathology. 16 (1), 24 (2021).
- Sarker, I. H. Deep learning: a comprehensive overview on techniques, taxonomy, applications and research directions. SN Computer Science. 2 (6), 420 (2021).
- Kittichai, V., et al. Classification for avian malaria parasite Plasmodium gallinaceum blood stages by using deep convolutional neural networks. Scientific Reports. 11 (1), 16919 (2021).
- Baskota, S. U., Wiley, C., Pantanowitz, L. The next generation robotic microscopy for intraoperative teleneuropathology consultation. Journal of Pathology Informatics. 11, 13 (2020).
- Bochkovskiy, A., Wang, C. -. Y., Liao, H. -. Y. M. YOLOv4: optimal speed and accuracy of object detection. arXiv. , 10934 (2004).
- Huang, G., Liu, Z., vander Maaten, L., Weinberger, K. Q. Densely connected convolutional networks. arXiv. , 06993 (2018).
- . CDC-DPDx. Diagnostic procedures - Blood specimens Available from: https://www.cdc.gov/dpdx/diagosticprocedures/blood/specimenproc.html#print (2020)
- Control and surveillance of African trypanosomiasis: report of a WHO expert committee. WHO Technical Report Series 881 Available from: https://iris.who.int/bitstream/handle/10665/42087/WHO_TRS_881.pdf?sequence=1 (1998)
- Leber, A. L. Detection of blood parasites. Clinical Microbiology Procedures Handbook. , (2022).
- Huang, L. -. P., Hong, M. -. H., Luo, C. -. H., Mahajan, S., Chen, L. -. J. A vector mosquitoes classification system based on edge computing and deep learning. Proceedings-2018 Conmference on Technologies and Applications of Artifical Intelligence. , 24-27 (2018).
- Cihan, P., Gökçe, E., Kalipsiz, O. A review of machine learning applications in veterinary field. Kafkas Universitesi Veteriner Fakultesi Dergisi. 23 (4), 673-680 (2017).
- Berrar, D. Cross-validation. Encyclopedia of Bioinformatics and Computational Biology. 1, 542-545 (2019).
- Gaithuma, A. K., et al. A single test approach for accurate and sensitive detection and taxonomic characterization of Trypanosomes by comprehensive analysis of internal transcribed spacer 1 amplicons. PLoS Neglected Tropical Diseases. 13 (2), 0006842 (2019).
- Vijayalakshmi, A., Rajesh Kanna, B. Deep learning approach to detect malaria from microscopic images. Multimedia Tools and Applications. 79 (21-22), 15297-15317 (2019).
- Morais, M. C. C., et al. Automatic detection of the parasite Trypanosoma cruzi in blood smears using a machine learning approach applied to mobile phone images. PeerJ. 10, 13470 (2022).
- Uc-Cetina, V., Brito-Loeza, C., Ruiz-Pina, H. Chagas parasite detection in blood images using AdaBoost. Computational and Mathematical Methods in Medicine. 2015, 139681 (2015).
- Zhang, C., et al. Deep learning for microscopic examination of protozoan parasites. Computational and Structural Biotechnology Journal. 20, 1036-1043 (2022).
- Sarataphan, N., et al. Diagnosis of a Trypanosoma lewisi-like (Herpetosoma) infection in a sick infant from Thailand. Journal of Medical Microbiology. 56, 1118-1121 (2007).
- Desquesnes, M., et al. A review on the diagnosis of animal trypanosomoses. Parasites & Vectors. 15 (1), 64 (2022).
- Fuhad, K. M. F., et al. Deep learning based automatic malaria parasite detection from blood smear and its smartphone based application. Diagnostics (Basel). 10 (5), 329 (2020).
- Christian Matek, S. S., Spiekermann, K., Marr, C. Human-level recognition of blast cells in acute myeloid leukaemia with convolutional neural networks. Nature Machine Intelligence. 1, 538-544 (2019).
- Hamdan, S., Ayyash, M., Almajali, S. Edge-computing architectures for internet of things applications: a survey. Sensors (Basel). 20 (22), 6441 (2020).
- Visser, T., et al. A comparative evaluation of mobile medical APPS (MMAS) for reading and interpreting malaria rapid diagnostic tests. Malaria Journal. 20 (1), 39 (2021).
- Giorgi, E., Macharia, P. M., Woodmansey, J., Snow, R. W., Rowlingson, B. Maplaria: a user friendly web-application for spatio-temporal malaria prevalence mapping. Malaria Journal. 20 (1), 471 (2021).
- Rajaraman, S., Jaeger, S., Antani, S. K. Performance evaluation of deep neural ensembles toward malaria parasite detection in thin-blood smear images. PeerJ. 7, 6977 (2019).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved