A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Pseudotyped Viruses As a Molecular Tool to Monitor Humoral Immune Responses Against SARS-CoV-2 Via Neutralization Assay
* These authors contributed equally
In This Article
Summary
Pseudotyped viruses (PVs) are replication-defective virions that are used to study host-virus interactions under safer conditions than handling authentic viruses. Presented here is a detailed protocol that shows how SARS-CoV-2 PVs can be used to test the neutralizing ability of patients' serum after COVID-19 vaccination.
Abstract
Pseudotyped viruses (PVs) are molecular tools that can be used to study host-virus interactions and to test the neutralizing ability of serum samples, in addition to their better-known use in gene therapy for the delivery of a gene of interest. PVs are replication defective because the viral genome is divided into different plasmids that are not incorporated into the PVs. This safe and versatile system allows the use of PVs in biosafety level 2 laboratories. Here, we present a general methodology to produce lentiviral PVs based on three plasmids as mentioned here: (1) the backbone plasmid carrying the reporter gene needed to monitor the infection; (2) the packaging plasmid carrying the genes for all the structural proteins needed to generate the PVs; (3) the envelope surface glycoprotein expression plasmid that determines virus tropism and mediates viral entry into the host cell. In this work, SARS-CoV-2 Spike is the envelope glycoprotein used for the production of non-replicative SARS-CoV-2 pseudotyped lentiviruses.
Briefly, packaging cells (HEK293T) were co-transfected with the three different plasmids using standard methods. After 48 h, the supernatant containing the PVs was harvested, filtered, and stored at -80 °C. The infectivity of SARS-CoV-2 PVs was tested by studying the expression of the reporter gene (luciferase) in a target cell line 48 h after infection. The higher the value for relative luminescence units (RLUs), the higher the infection/transduction rate. Furthermore, the infectious PVs were added to the serially diluted serum samples to study the neutralization process of pseudoviruses' entry into target cells, measured as the reduction in RLU intensity: lower values corresponding to high neutralizing activity.
Introduction
Pseudotyped viruses (PVs) are molecular tools used in microbiology to study host-virus and pathogen-pathogen interactions1,2,3,4. PVs consist of an inner part, the viral core that protects the viral genome, and an outer part, the envelope glycoproteins on the surface of the virus that defines the tropism5. A pseudovirus is replication-incompetent in the target cell because it does not contain all the genetic information to generate new viral particles. This combination of peculiar features makes PVs a safe alternative to a wildtype virus. Wildtype viruses, on the other hand, are highly pathogenic and cannot be used in BSL 2 laboratories for analysis6.
The infectivity of PVs can be monitored by a reporter gene, usually coding for a fluorescent protein (GFP, RFP, YFP) or an enzyme that produces chemiluminescent products (luciferase). This is contained in one of the plasmids used for PV production and incorporated in the genome of the pseudovirus7.
Several types of PV cores currently exist, including lentiviral-derived particles based on the HIV-1 genome. The great advantage of HIV-1-based PVs over other platforms is their intrinsic integration process in the target cell genome8. Although HIV-1 is a highly contagious virus and is the causative agent of AIDS, these lentiviral vectors are safe to use because of the extensive optimization steps over the years. Optimal safety conditions were achieved with the introduction of 2nd-generation lentiviral vectors, in which viral genes were depleted without influencing transduction capabilities9. The 3rd and 4th generations contributed to the increased safety of lentiviral vector handling with the further splitting of the viral genome into separate plasmids10, 11. The latest generations of PVs are generally employed to produce lentiviral vectors for gene therapy.
PVs can be used to study interactions between viruses and host cells, during both the production and the infection phases. PVs are especially employed in pseudovirus neutralization assays (PVNA). PVNAs are widely validated to assess the neutralization potential of serum or plasma by targeting the viral glycoprotein on the PV's envelope12,13. Neutralization activity, expressed as the inhibitory concentration 50 (IC50), is defined as the dilution of serum/plasma that blocks 50% of viral particle entry14. In this protocol, we described the set-up of a PVNA to test the antibody activity against Severe Acute Respiratory Syndrome - Coronavirus 2 (SARS-CoV-2) in sera collected before and after receiving a booster vaccine dose.
Protocol
The present protocol has been approved by and follows the guidelines of the Ethical Committee of the University of Verona (approval protocol number 1538). Informed written consent was obtained from the human subjects participating in the study. Whole blood samples were collected from healthcare worker (HCW) volunteers who were in the process of receiving anti-SARS-CoV-2 vaccines. These samples were collected in plastic tubes containing anticoagulants for the subsequent isolation of serum15.
All the following processes must be performed in a Class-2 biological hood, working under sterile conditions. Virus handling must be performed with care, and all waste products must be neutralized in a diluted bleach solution. An overview of the protocol is displayed in Figure 1.
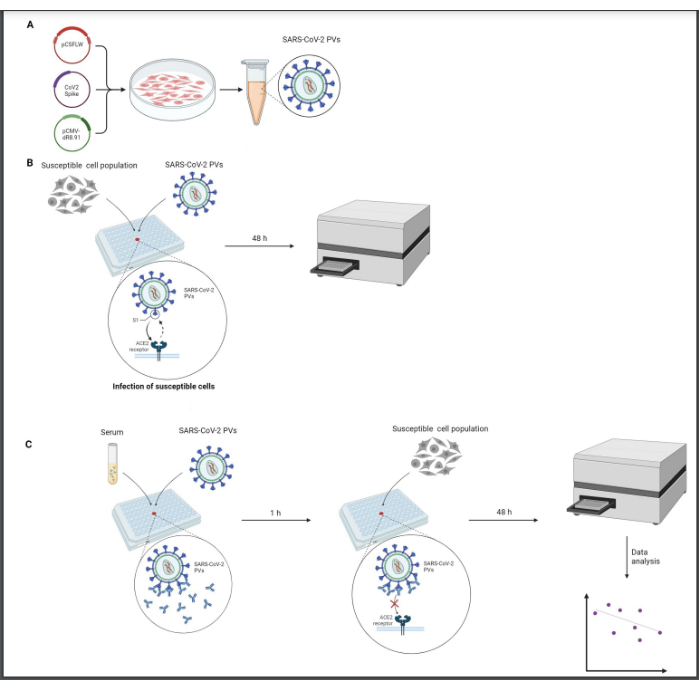
Figure 1: Graphical representation of a neutralization assay. (A) PV production, (B) PV titration, and (C) neutralization assay. All the procedures are performed in a class-2 biological hood under sterile conditions. Titration step (B) needs to be performed to standardize the infectivity levels of PVs before use in the neutralization assay (C). This figure was created with BioRender. Please click here to view a larger version of this figure.
1. SARS-CoV-2 PVs production and infectivity test
- Seed 5 x 105 HEK293T cells in complete Dulbecco's Modified Eagle Medium (DMEM, high-glucose, 10% foetal bovine serum (FBS), 1% L-glutamine, 1% penicillin/streptomycin) in a 6-well plate (6WP) to reach a suitable cell density compatible with the transfection reagent used. In the case of performing transfection with polyehtylenimine (PEI) (prepare the reagent following the manufacturer instructions), ensure that the cells reach 40-60% density on the day of transfection (step 1.3). Keep the cells in a humidified incubator at 37 °C and 5% CO2.
- Prior to transfection, replace the spent cell medium with fresh medium without antibiotics (DMEM, high-glucose, 10% FBS, 1% L-glutamine) to achieve higher transfection efficiency.
NOTE: The day after seeding, HEK293T cells are ready to be transfected. - Transfect adherent HEK293T cells with a suitable transfection reagent according to the manufacturer's instructions. If using PEI, prepare two mixes and follow the steps below.
- To prepare mix A, add 500 ng of pCMV-dR8.91 packaging plasmid16, 750 ng of pCSFLW reporter plasmid16, and 450 ng of SARS-CoV-2 Spike expressing plasmid in 100 µL of reduced serum medium.
- To prepare mix B, add 17.5 µL of PEI (concentration: 1 mg/mL) to 100 µL of the reduced serum medium.
- Allow both mixes to incubate at room temperature (RT) for 5 min. Next, mix the contents of both tubes together by adding the PEI mix B to DNA mix A.
- Incubate the tube for 20-30 min at RT. Flick the tube gently every 3-4 min to enhance the mixing. Finally, add the mixture to the HEK293T cells.
- 16-20 h after the transfection, replace the culture medium with fresh, complete DMEM. Incubate at 37 °C and 5% CO2, to allow for the production of PVs by transfected cells.
- 72 h after the transfection, harvest the supernatant containing PVs. Then centrifuge at 1600 x g for 7 min at room temperature to remove cell debris and dead cells and filter it through a 0.45 µm cellulose acetate filter.
- OPTIONAL STEP: To increase the final yield of PV titer, perform multiple transfections, pool the cell media containing PVs, and concentrate it using concentrating tubes.
- Proceed directly with the next steps ("PVs titration", section 2) or aliquot the PV-containing medium in suitable tubes to store at -80 °C until use. Prepare an additional aliquot (400-500 µL) to be used for titration.
NOTE: Making multiple aliquots will guarantee reproducibility between experiments by avoiding excessive thaw-freeze cycles.
2. PVs titration
- Use the fresh PV-containing medium for the next steps or thaw the testing aliquot (step 1.7) to perform the titration of the new viral stock. Freezing aliquots of the same PV stock will guarantee reproducibility.
- Add 50 µL of complete DMEM (or complete medium compatible with the target cell line in usage) in all the wells of a 96 well-plate (96WP) necessary to test in duplicate the PV stock, leaving row "A" empty. Add 100 µL of PVs stock to row "A". Based on the number of preparations to be tested, leave one column without the virus as a "cell only" control (Figure 2).
- Pipette 50 µL from row A to row B and repeat this process up to row G to obtain serial dilutions of the initial stock. Discard the excess volume from the last row.
- Detach cellsusing trypsin/ethylenediaminetetraacetic acid 1x (EDTA) in Dulbecco's phosphate buffer saline 1x (DPBS 1x), after removing the spent medium and washing cells with DPBS 1x twice. Prepare cells to a density of 4 x 105 cells/mL.
NOTE: In this protocol, PVs infection was tested on the susceptible cell line HEK293T/ACE2; such cells were derived from HEK293T, transduced using a lentiviral vector to express ACE2 receptor. - Add 50 µL of the cell suspension into each well to ensure a cell count of 2 x 104 cells per well.
- Incubate at 37 °C and 5% CO2, for 48 h.
- After the incubation, perform the Luciferase assay to obtain the reading as per the manufacturer's instructions. Add 100 µL of the luciferase reagent to the wells and incubate in the dark at RT for 2 min. Move the content of each well to a black 96 well plate (compatible with the available plate reader) and read the plates in a 96 well plate reader.
NOTE: The luminometer used for the luciferase readout will produce a spreadsheet file with the raw, unprocessed data that will be used for downstream analysis (in this case, an Excel file). The virus' infectivity will be expressed as relative luminescence units (RLU) (described in paragraph 4.1).
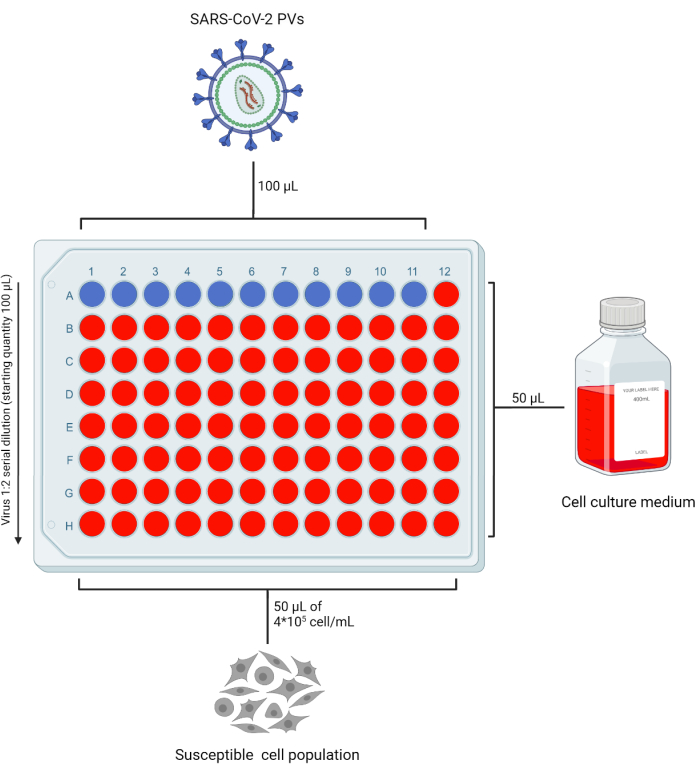
Figure 2: Representative layout of a 96 well plate for PVs titration. A fixed volume of PV-containing supernatant is added to row A, columns 1-11, and serially diluted. The last column is left as the "cell only" control. This figure was created with BioRender. Please click here to view a larger version of this figure.
3. Neutralization assay
- Thaw patients' sera on ice. Inactivate serum samples by incubating them at 56 °C for 30 min.
- In a 96 well plate, add 50 µL of the fresh, complete DMEM (or complete medium compatible with the target cell line used)in each of the following wells: from row B (columns 1-10) to row H (columns 1-10). Put 95 µL of the fresh, complete DMEM in row A (columns 1-10). Add 50 µL and 100 µL of complete DMEM into the wells of columns 11 and 12, respectively. These will be the infected (virus control, or VC) and uninfected (cell only, or CC) controls, respectively (Figure 3).
- Add 5 µL of heat-inactivated serum/plasma samples in row A (columns 1-10). Each sample will be in duplicate. With a multichannel pipette, mix the samples in the first row and move 50 µL of medium containing serum from row A to row B. Repeat this process up to the last row (Figure 3). Discard the remaining 50 µL.
- Thaw the necessary number of PVs' aliquots and dilute to ≥ 104 RLU/mL. Add 50 µL of the diluted PV-containing medium to each well (from column 1 to column 11) using a multichannel pipette to reach a 1:1 dilution of heat inactivated serum/plasma to virus. Incubate at 37 °C and 5% CO2, for 1 h to allow the antibodies in the serum samples to bind to the SARS-CoV-2 spike protein on the PVs.
- Prepare at least 5 mL suspension of susceptible cells (HEK293T/ACE2) at a cell density of 4 x 105 cells/mL. Add 50 µL of the cell suspension to each well and incubate at 37 °C and 5% CO2, for 48 h.
- After the incubation, perform the luciferase assay reading according to the manufacturer's instructions, as described in step 2.7.
NOTE: The luminometer used for luciferase readout will produce a spreadsheet file (in this case, .xlsx) with the raw, unprocessed data that will be used for downstream analysis (the Luciferase assay file).
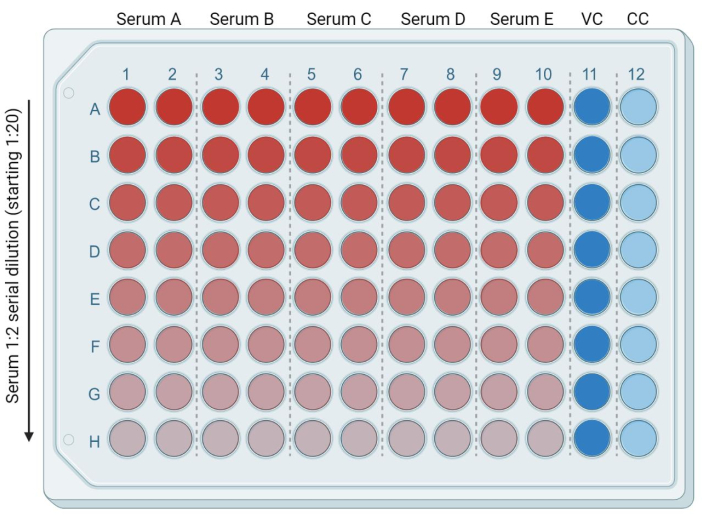
Figure 3: Plate representation based on serum dilution. Bright red corresponds to a higher quantity of serum, and bright blue lane (column 11) corresponds to infected cell control (VC, virus control). Light blue lane (column 12) corresponds to uninfected cells (CC, cell control). This figure was created with BioRender. Please click here to view a larger version of this figure.
4. Titration analysis
- On the Luciferase assay file, assign the names/titles to the corresponding samples.
- Multiply the RLU measure by the dilution factors (from the top to the bottom of the grid: 20x, 40x, 80x, 160x, 320x, 640x, 1,280x, 2,560x) to obtain RLU/mL. If different dilution factors are used, change the multiplication factors accordingly.
- Calculate the average RLU/mL for each PV preparation.
5. PVs neutralization assay analysis
- On the Luciferase assay spreadsheet file (in this case, .xlsx), assign the corresponding titles to the tested samples. Enter the dilution factor of the sample (40s, 80x, 160x, 320x, 640x, 1,280x, 2,560x, 5,120x). Calculate the Log10 of the dilution factors.
- Calculate the average RLU of uninfected and infected control (Figure 3, columns 11 and 12, respectively). These values will be useful for the normalization in step 5.5.
- Open a new document for data analysis. Select X/Y analysis, input X as Numbers and Y as Enter 2 replicate values in side-by-side sub-columns.
- Enter Log10 (dilution) values as X numbers. Enter the duplicate RLU of the samples.
- Go to Analyze > Normalize > Flag all the samples on the same sheet. Input the average VC and CC values in How is 0% defined?, and How is 100% defined?, respectively. Click OK.
- On the normalized data sheet, go to Analyze > XY analyses > Nonlinear analyses (curve fit). Flag all the samples and click OK. For the Dose-response - Inhibition, select log(inhibitor) vs normalized response - variable slope.
- Under Constrain, change HillSlope to Must be less than 0.
- Under Output, flag Create summary table and graph. Click on OK to obtain the final analyses. A working sheet with a template for the analysis is provided in Supplementary File 1.
Results
This protocol describes the production of SARS-CoV-2 PVs and a downstream application of these PVs to analyze the neutralization activity of serum/plasma of subjects receiving anti-COVID-19 vaccination17. Furthermore, this protocol can be applied to produce pseudotypes of each SARS-CoV-2 variant of concern (VOC) to test the evolution of the neutralizing response. Despite this protocol facilitating the study of humoral immune response after COVID-19 vaccination, it can be adapted to easily test the...
Discussion
Although using a wildtype virus simulates the actual infection, lentiviral PVs are a safer option to study the mechanisms associated with viral entry and infection without the strict safety requirements necessary to work with pathogenic viruses4,20,21. PVs are composed of a replication-defective viral core surrounded by the surface envelope glycoprotein of a pathogenic virus which is the objective of the study.
Disclosures
The authors declare to have no conflict of interest.
Acknowledgements
We acknowledge the contribution of the health-care workers volunteers. This project was supported by the Department of Excellence 2023/2027, MUR, Italy. AR and DZ were supported by PRIN2022 (EU fundings; NextGenerationEU)
Materials
| Name | Company | Catalog Number | Comments |
| 0.45 μm filter | SARSTEDT | 83 1826 | |
| 6-well plate | SARSTEDT | 83 3920 | |
| 96-well plate | SARSTEDT | 8,33,924 | |
| Amicon Ultra-15 Centrifugal Filter Units | Merck | 10403892 | |
| Black Opaque 96-well Microplate | Perkin Elmer | 60005270 | |
| Dulbecco's Modified Eagle Medium | SIGMA-ALDRICH | D6546 - 500ML | |
| Dulbecco's phosphate buffered saline (PBS 1x) | AUROGENE | AU-L0615-500 | |
| Foetal Bovine Serum | AUROGENE | AU-S1810-500 | |
| Graphpad Prism version 7 | graphpad dotmatics | NA | In the manuscript, we replace the commercial name with 'data analysis program' |
| HEK293T cells | ATCC | CRL-3216 | |
| HEK293T/ACE2 cells | ATCC | CRL-3216 | HEK293T has been transduced to overexpress ACE2 with a lentiviral vector. |
| L-glutamine | AUROGENE | AU-X0550-100 | |
| Luminometer - Victor3 | Perkin Elmer | HH35000500 | In the manuscript, we replace the commercial name with 'luminometer' |
| Opti-MEM | Thermo Fisher Scientific | 11058021 | In the manuscript, we replace the commercial name with 'reduced serum medium' |
| p8.91 packaging plasmid | Di Genova et al., 2021 | A kind gift from Prof. Nigel Temperton (ref 16.) | |
| pCSFLW reporter plasmid | Di Genova et al., 2021 | A kind gift from Prof. Nigel Temperton (ref 16.) | |
| Penicillin/streptomycin | AUROGENE | AU-L0022-100 | |
| Polyethylenimine, branched (PEI) (25 kDa) | SIGMA-ALDRICH | 408727 | |
| RRL.sin.cPPT.SFFV/Ace2.IRES-puro.WPRE (MT126) | Addgene | 145839 | This plasmid was used to generate HEK293Tcells/ACE2 |
| SARS-CoV-2 Spike expressing plasmid | Addgene | pGBW-m4137382 | |
| steadylite plus Reporter Gene Assay System | Perkin Elmer | 6066759 | In the manuscript, we replaced the commercial name with 'luciferase reading reagent' |
| Trypsin EDTA 1x | AUROGENE | AU-L0949-100 |
References
- Ozaki, D. A., et al. International technology transfer of a GCLP-compliant HIV-1 neutralizing antibody assay for human clinical trials. Plos One. 7 (1), e30963 (2012).
- Pouget, M., et al. Generation of liposomes to study the effect of Mycobacterium tuberculosis lipids on HIV-1 cis- and trans-infections. International Journal of Molecular Sciences. 22 (4), 1945 (2021).
- McKay, L. G. A., et al. The HCV envelope glycoprotein down-modulates NF-κB signalling and associates with stimulation of the host endoplasmic reticulum stress pathway. Frontiers in Immunology. 13, 831695 (2022).
- Xiang, Q., Li, L., Wu, J., Tian, M., Fu, Y. Application of pseudovirus system in the development of vaccine, antiviral-drugs, and neutralizing antibodies. Microbiological Research. 258, 126993 (2022).
- Li, Q., Liu, Q., Huang, W., Li, X., Wang, Y. Current status on the development of pseudoviruses for enveloped viruses. Reviews in Medical Virology. 28, e1963 (2018).
- D'Apice, L., et al. Comparative analysis of the neutralizing activity against SARS-CoV-2 Wuhan-Hu-1 strain and variants of concern: Performance evaluation of a pseudovirus-based neutralization assay. Frontiers in Immunology. 13, 981693 (2022).
- Falzarano, D., Groseth, A., Hoenen, T. Development and application of reporter-expressing mononegaviruses: current challenges and perspectives. Antiviral Research. 103, 78-87 (2014).
- Gutierrez-Guerrero, A., Cosset, F. -. L., Verhoeyen, E. Lentiviral vector pseudotypes: Precious tools to improve gene modification of hematopoietic cells for research and gene therapy. Viruses. 12, 1016 (2020).
- Zufferey, R., Nagy, D., Mandel, R. J., Naldini, L., Trono, D. Multiply attenuated lentiviral vector achieves efficient gene delivery in vivo. Nature Biotechnology. 15 (9), 871-875 (1997).
- Dull, T. A third-generation lentivirus vector with a conditional packaging system. Journal of Virology. 72 (11), 8463-8471 (1998).
- Berkhout, B. A Fourth generation lentiviral Vector: Simplifying genomic gymnastics. Molecular Therapy. 25 (8), 1741-1743 (2017).
- Wu, X. Development and evaluation of a pseudovirus-luciferase assay for rapid and quantitative detection of neutralizing antibodies against Enterovirus 71. Plos One. 8 (6), e64116 (2013).
- Ferrara, F., et al. Development of lentiviral vectors pseudotyped with Influenza B hemagglutinins: application in vaccine immunogenicity, mAb potency, and sero-surveillance studies. Frontiers in Immunology. 12, 661379 (2021).
- Hu, J., et al. Development of cell-based pseudovirus entry assay to identify potential viral entry inhibitors and neutralizing antibodies against SARS-CoV-2. Genes & Diseases. 7 (4), 551-557 (2020).
- Dalle Carbonare, L., et al. Serology study after BTN162b2 vaccination in participants previously infected with SARS-CoV-2 in two different waves versus naïve. Communications Medicine. 1 (1), 38 (2021).
- Di Genova, C., et al. Production, titration, neutralisation, storage and lyophilisation of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) lentiviral pseudotypes. Bio-protocol. 11 (21), e4236 (2021).
- Chmielewska, A. M., Czarnota, A., Bieńkowska-Szewczyk, K., Grzyb, K. Immune response against SARS-CoV-2 variants: The role of neutralization assays. NPJ Vaccines. 6 (1), 1-8 (2021).
- Chen, Q., et al. Development and optimization of a sensitive pseudovirus-based assay for HIV-1 neutralizing antibodies detection using A3R5 cells. Human Vaccines & Immunotherapeutics. 14 (1), 199-208 (2018).
- Gauger, P. C., Vincent, A. L. Serum virus neutralization assay for detection and quantitation of serum neutralizing antibodies to influenza A virus in swine. Methods in Molecular Biology (Clifton, N.J). 2123, 321-333 (2020).
- Miglietta, R., Pastori, C., Venuti, A., Ochsenbauer, C., Lopalco, L. Synergy in monoclonal antibody neutralization of HIV-1 pseudoviruses and infectious molecular clones. Journal of Translational Medicine. 12 (1), 346 (2014).
- Chen, M., Zhang, X. -. E. Construction and applications of SARS-CoV-2 pseudoviruses: A mini review. International Journal of Biological Sciences. 17 (6), 1574-1580 (2021).
- Zipeto, D., et al. Induction of human immunodeficiency virus neutralizing antibodies using fusion complexes. Microbes and Infection. 8 (6), 1424-1433 (2006).
- WHO Coronavirus (COVID-19) Dashboard. Available from: https://covid19.who.int (2022)
- Zhou, P. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 579 (7798), 270-273 (2020).
- Chen, X., Huang, H., Ju, J., Sun, R., Zhang, J. Impact of vaccination on the COVID-19 pandemic in U.S. states. Scientific Reports. 12 (1), 1554 (2022).
- Stefani, C., Fantoni, T., Bissoli, M., Thomas, J., Ruggiero, A. HIV and SARS-CoV-2 Co-Infection: From Population Study Evidence to In Vitro Studies. Life. 12 (12), 2089 (2022).
- Watson, O. J., et al. Global impact of the first year of COVID-19 vaccination: a mathematical modelling study. The Lancet Infectious Diseases. 22 (9), 1293-1302 (2022).
- Cantoni, D. Analysis of antibody neutralisation activity against SARS-CoV-2 variants and seasonal human coronaviruses NL63, HKU1, and 229E induced by three different COVID-19 vaccine olatforms. Vaccines. 11 (1), 58 (2023).
- Siracusano, G., et al. Different decay of antibody response and VOC sensitivity in naïve and previously infected subjects at 15 weeks following vaccination with BNT162b2. Journal of Translational Medicine. 20 (1), 22 (2022).
- Ruggiero, A. SARS-CoV-2 vaccination elicits unconventional IgM specific responses in naïve and previously COVID-19-infected individuals. eBioMedicine. 77, (2022).
- Piubelli, C. Subjects who developed SARS-CoV-2 specific IgM after vaccination show a longer humoral immunity and a lower frequency of infection. eBioMedicine. 89, 104471 (2023).
- Zhang, G. F. Infectivity of pseudotyped SARS-CoV-2 variants of concern in different human cell types and inhibitory effects of recombinant spike protein and entry-related cellular factors. Journal of Medical Virology. 95 (1), e28437 (2023).
- da Costa, K. A. S. Influenza A (N1-N9) and Influenza B (B/Victoria and B/Yamagata) neuraminidase pseudotypes as tools for pandemic preparedness and improved influenza vaccine design. Vaccines. 10 (9), 1520 (2022).
- Condor Capcha, J. M. Generation of SARS-CoV-2 spike pseudotyped virus for viral entry and neutralization assays: a 1-week protocol. Frontiers in Cardiovascular Medicine. 7, 618651 (2021).
- Diomede, L., et al. Doxycycline inhibition of a pseudotyped virus transduction does not translate to inhibition of SARS-CoV-2 infectivity. Viruses. 13 (9), 1745 (2021).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved