A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
A Set of Screening Techniques for a Quick Overview of the Neutrophil Function
In This Article
Summary
This protocol features a set of neutrophil functional assays to be used as a screening method to cover functions from different signaling pathways. The protocol includes an initial and simple evaluation of cell viability, purity, reactive oxygen species production, real-time migration, phagocytosis, and a preliminary suggestion of neutrophil extracellular traps.
Abstract
Neutrophils are known as one of the first lines of defense in the innate immune response and can perform many particular cellular functions, such as chemotaxis, reverse migration, phagocytosis, degranulation of cytotoxic enzymes and metabolites, and release of DNA as neutrophil extracellular traps (NETs). Neutrophils not only have tightly regulated signaling themselves, but also participate in the regulation of other components of the immune system. As fresh neutrophils are terminally differentiated, short-lived, and highly variable among individuals, it is important to make the most of the collected samples. Researchers often need to perform screening assays to assess an overview of the many neutrophil functions that may be affected by specific conditions under evaluation. A set of tests following a single isolation process of normal density neutrophils was developed to address this need, seeking a balance between speed, comprehensiveness, cost, and accuracy. The results can be used to reason and guide in-depth follow-up studies. This procedure can be carried out in an average time of 4 h and includes the evaluation of cell viability, reactive oxygen species (ROS) production, real-time migration, and phagocytosis of yeast on glass slides, leaving enough cells for more detailed approaches like omics studies. Moreover, the procedure includes a way to easily observe a preliminary suggestion of NETs after fast panoptic staining observed by light microscopy, with a lack of specific markers, albeit enough to indicate if further efforts in that way would be worthwhile. The diversity of functions tested combines common points among tests, reducing the analysis time and expenses. The procedure was named NeutroFun Screen, and although having limitations, it balances the aforementioned factors. Furthermore, the aim of this work is not a definite test set, but rather a guideline that can easily be adjusted to each lab's resources and demands.
Introduction
Neutrophils are the most abundant innate immune cells in human blood and are known to play a major part in infection and inflammation, being the first responders to arrive at the site of tissue damage1. In recent years, there has been a growing recognition of the crucial role that neutrophils play in a variety of diseases and in supporting homeostasis2. Neutrophils not only have tightly regulated signaling themselves, but also participate in the regulation of other components of the immune system3,4,5. Therefore, investigating neutrophils and their many unusual cellular functions, such as chemotaxis, reverse migration6, phagocytosis7, respiratory burst8, and the release of neutrophil extracellular traps (NETs)7, is imperative in numerous research contexts where it is necessary to assess the potential neutrophil functional, morphological, or molecular changes triggered by specific conditions under analysis.
Freshly isolated neutrophils are terminally differentiated, short-lived, highly dynamic, and easily activated9. However, an efficient storage method that does not affect the neutrophil responses has not yet been achieved, making it challenging to perform multiple assays that must be uninterrupted. Furthermore, previously described functional analyses10,11, based on assays that require cytometry and/or fluorescent staining, may not be a viable choice when a broad and initial evaluation of the neutrophil is needed.
To address these issues, this protocol describes a set of tests that can be carried out following a single isolation process, including the evaluation of cell viability, reactive oxygen species (ROS) production, real-time migration, and phagocytosis of Saccharomyces cerevisiae, whose results can be used to reason in-depth follow-up studies. This procedure, named NeutroFun Screen, was designed to encompass the leading effector activities, except degranulation, and can be completed in an average time of 4 h, including 1 h of activation. Additionally, the remaining cells can be used for more detailed approaches like omics studies. The advantage of this method lies in its balance between speed, comprehensiveness, cost, and accuracy.
Furthermore, there is a way to easily observe a preliminary suggestion of NETs, without specific markers, but enough to indicate if further efforts in that direction would be worthwhile. The diversity of functions tested aims to combine common points among tests, reducing the analysis time and expenses. The main goal of this method is to provide a balanced, functional analysis regarding speed, comprehensiveness, cost, and accuracy that allows for an overview of the neutrophil's response, making it a useful initial step in investigating the effects of novel stimuli on normal density neutrophils.
Protocol
All experiments strictly followed the ethical guidelines set by the institutional review board at the University of Brasilia (process 13364819.0.0000.5558), and samples were identified by codes to ensure donor anonymity. The cells were obtained from normal healthy male donors aged 18-35 years, who signed the informed consent and met the following eligibility criteria: non-smokers/vapers, no chronic health conditions, and no history of inflammatory conditions in the last 14 days.
1. Blood collection
- Aseptically place 0.3 mL of 5,000 IU/mL heparin (see Table of Materials) in a sterile 20 mL syringe to heparinize it.
- Apply a venous tourniquet about 4 in above the puncture site and identify the median cubital or cephalic vein for venipuncture.
NOTE: Ensure that the total tourniquet time does not exceed 1 min. - Clean the puncture site with 70% alcohol and perform the venipuncture.
- Gently invert the syringe three or four times after collecting the blood to mix the blood and heparin properly.
2. Neutrophil isolation
NOTE: Polymorphonuclear leukocytes (PMNs) are isolated through density gradient centrifugation followed by hypotonic lysis of the remaining red blood cells (RBC), as previously described11 with some changes. This method is not mandatory to perform the screening assays, and can be replaced as long as the chosen method results in a viability of >97%, priming or activation of <3% of the PMNs, and yields enough cells for all assays, replicates, and conditions. Performing these steps under aseptic conditions and using endotoxin-free solutions are mandatory to avoid cell activation.
- Make 12 mL dilutions of 60% and 70% separation media (commercially available; see Table of Materials) in 50 mL conical tubes.
- Prepare the gradient from bottom to top by adding 4 mL at a time of the 60% dilution over the 70% dilution, using a 5 mL pipette. Do this gently to prevent mixing the interface.
- Carefully layer 12 mL of heparinized blood on top of the density gradient. Centrifuge at 200 x g for 15 min at room temperature.
NOTE: From this step onward, until PMN activation, all the reagents and tubes used must be kept in a cooler filled with ice. - Discard the plasma/mononuclear cell layer, then gently transfer the layer above the erythrocyte pellet into two 15 mL conical tubes with approximately 7.5 mL in each. Make up the tube volume with Hank's balanced salt solution (HBSS; see Table of Materials).
- Centrifuge at 300 x g for 5 min at 19 °C.
- Wash the cell pellet with HBSS.
- Discard the supernatant by pouring the tube and gently resuspend the pellet in 7 mL of HBSS.
- Centrifuge at 300 x g for 5 min at 19 °C to remove all the separation media.
- Perform hypotonic lysis of the remaining RBCs.
- Discard the supernatant and combine the pellets in a single tube.
- Resuspend the RBC/PMN pellet in 3 mL of sterile H2O and add 3 mL of HBSS (2x) within 25 s to restore the osmolarity. Then, centrifuge at 300 x g for 5 min at 19 °C.
- Repeat steps 2.8.1 and 2.8.2 for a white, erythrocyte-free pellet.
NOTE: The supernatant must be removed as soon as possible to minimize the contact of neutrophils with RBC breakdown products. Alternatively, the second hypotonic lysis can be replaced by gently resuspending the residual RBC layer and removing all the supernatant, as the remaining RBCs will sediment above the PMN pellet.
- Discard the supernatant by pouring the tube, gently resuspend the PMNs in the remaining buffer, and transfer them to an ice-cold microtube.
NOTE: Ensure to annotate the volume while transferring the resuspended cells with a micropipette. - Transfer 3 x 1 µL of the cell suspension to a clean glass slide (three wells of 1 µL each) and stain with fast panoptic (see Table of Materials) for morphology and purity evaluation12.
- To stain with fast panoptic, immerse the slide five times in panoptic fixative n° 1, six times in eosin n° 2, and twice in hematoxylin n° 3, with each immersion lasting 1 s.
- Gently wash the glass slide with distilled water.
- Allow to drain and air-dry.
- Observe under a microscope and count 300 random cells in each well, thus differentiating the neutrophils from other granulocytes.
- Transfer 1 µL of the cell suspension to 49 µL of 0.2% trypan blue dye13 and count the cells using a Neubauer chamber, distinguishing between dead and viable cells.
- Adjust the cell concentration to 6,667 cells/µL using a solution of 50% autologous plasma and 50% HBSS supplemented with calcium and magnesium. Divide the 6,667 cells/µL suspension evenly among the microtubes corresponding to the conditions to be tested, including the negative control.
NOTE: Any cell concentration similar to the circulating neutrophils in the model organism can be used, but it is important to use the same cell concentration in all experiments for reproducibility.
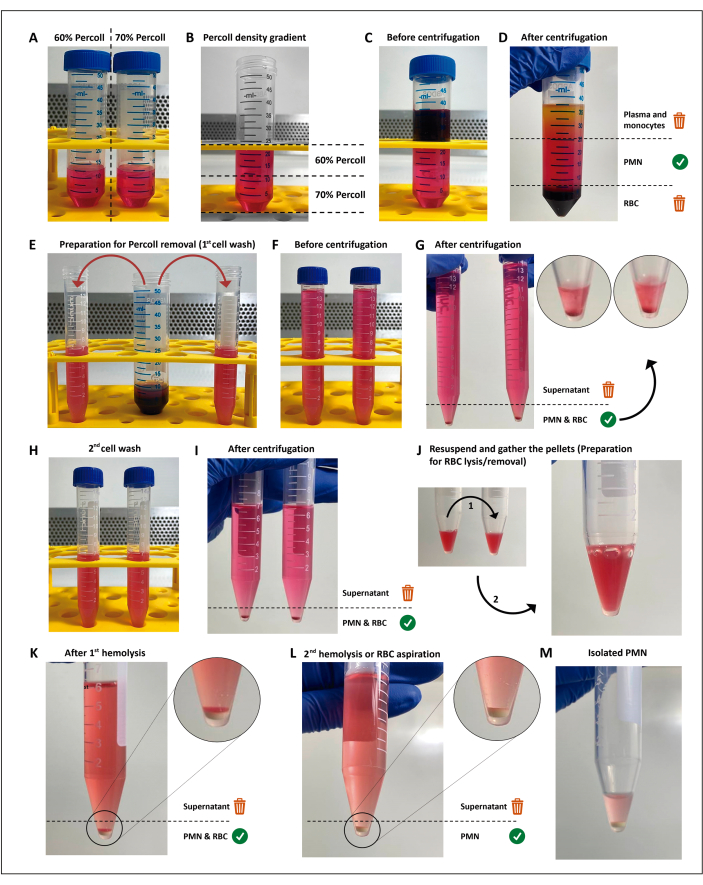
Figure 1: The neutrophil isolation protocol. Two concentrations of the separation media (percoll) (A) are stacked (B), then the blood is layered on top of the separation gradient (C). After centrifugation, the PMN is in the central layer (D), which is divided into two 15 mL tubes (E). The cell suspension is washed twice in HBSS and centrifuged (G-I) to remove the media, then the cells are resuspended, and residual RBCs are submitted to two rounds of hypotonic lysis (J-M). Please click here to view a larger version of this figure.
3. Preparation for neutrophil activation
- In 1.5 mL microtubes, prepare an activation system for each condition so that the final cell concentration is 6,600 cells/µL. For instance, to test the effects of 100 nM fMLP (N-formyl-methionyl-leucyl-phenylalanine; see Table of Materials), add 5 µL of 10 µM fMLP to 495 µL of the 6,667 cell/µL suspension. For the negative (unstimulated) control, add HBSS containing Ca2+ and Mg2+.
NOTE: To demonstrate this methodology, the following final concentrations of the stimuli were used: 100 nM fMLP, 16 µM of fallaxin, a naturally occurring antimicrobial peptide14, and 100 nM PMA (phorbol 12-myristate 13-acetate) (see Table of Materials). - Incubate at 37 °C without rotation.
NOTE: All aliquots for functional assays are taken from this cell suspension, hereafter referred to as the activation system.
4. Nitrotetrazolium blue chloride (NBT) assay for evaluating ROS production
- Preparation of NBT working solution: For each experimental condition, prepare an NBT (see Table of Materials) working solution of 6 mM using the following steps:
- Dissolve 0.0005 g of NBT in 10 µL of dimethyl sulfoxide (DMSO) and vortex for at least 15 min.
- Add 90 µL of HBSS Ca2+Mg2+ and vortex up to 2 min.
NOTE: All steps involving NBT must be performed in the dark.
- Perform NBT slide test.
- After 20 min of cell activation, gently mix the cell suspension and carefully transfer 2 µL of PMN to a clean glass slide. Incubate for 20 min in a humidified chamber at 37 °C.
NOTE: Do not spread the cell suspension too much over the slide; otherwise it may dry out before incubation. - Add 1 µL of the NBT working solution over the cells and further incubate for 20 min protected from light.
- Dry the slide with hot air and fix it with a drop of methanol in each well for 1 min. Stain with 0.03% safranin (see Table of Materials) for 1 min.
- Gently wash the glass slide with distilled water.
- Allow the slide to air-dry and observe under a microscope.
- Count 100 random cells in each well, differentiating neutrophils with and without formazan deposits.
- After 20 min of cell activation, gently mix the cell suspension and carefully transfer 2 µL of PMN to a clean glass slide. Incubate for 20 min in a humidified chamber at 37 °C.
- Perform NBT spectrophotometry assay.
- After 40 min of cell activation, gently mix the cell suspension and transfer 90 µL of PMNs from the activation system to a clean microtube. Then, carefully add 20 µL of the 6 mM NBT solution. Incubate in the dark for 20 min at 37 °C.
- Add 100 µL of 10% Sodium dodecyl sulfate (SDS; see Table of Materials) and vortex.
- Sonicate using a tip-sonicator at an amplitude of 60%, five cycles of 15 s each with 15 s intervals. Centrifuge at 12,000 x g for 5 min.
- Transfer 60 µL of the supernatant to a clear-bottom 96-well plate and measure the absorbance of the formazan product at 570 nm.
5. Phagocytosis assay
- Prepare a 33,000 yeasts/µL suspension for each condition, as described below:
- Add approximately 0.75 mg of dry yeast (Saccharomyces cerevisiae; see Table of Materials) to 200 µL of HBSS Ca2+Mg2+ and incubate in a thermomixer at 100 °C with 500 rpm for at least 15 min.
- Homogenize the mixture by vortexing and transfer 5 µL of yeast suspension to 45 µL of 0.2% trypan blue dye. Count the yeasts using a Neubauer chamber.
- Adjust the concentration of the initial suspension to 33,000 yeast cells/µL using HBSS Ca2+Mg2+. Keep the suspension on ice until use.
- After 20 min of cell activation, gently mix the cell suspension and transfer 5 µL of the activation system to 5 µL of the 33,000 yeast/µL Saccharomyces cerevisiae suspension in a new sterile microtube.
NOTE: The neutrophil to yeast ratio is 1:5 (PMN:yeast). - Immediately transfer 6 µL of the PMN/yeast suspension into three wells of a clean glass slide (2 µL each) and incubate the slide in a humidified chamber for 40 min.
NOTE: Do not spread the cell suspension too much over the slide; otherwise, it may dry out before incubation. - Dry the slide under hot air and stain with fast panoptic, as described in step 2.9 above.
NOTE: The third step of the fast panoptic staining is critical to the microscopy analysis of the slide. Staining the slide for ≥3 s at this step can render it unfit for analysis, since it will be difficult to differentiate yeast from neutrophil nuclear lobes. - Observe the slides under the microscope, counting 100 random neutrophils of each well and discriminating between PMNs positive and negative for phagocytosis.
NOTE: At least one yeast particle within or in direct contact with the PMN cellular membrane indicates a PMN positive for phagocytosis. If interested in the yeast/neutrophil ratio, count the number of yeast particles engulfed as well.
6. Real-time PMN chemotaxis assay
NOTE: The migration assay is performed similarly to the protocol described previoulsy15, with the following adaptations:
- Prepare the chemotactic gradient by adding 160 µL of chemoattractant (e.g., fMLP, IL-8, C5, or LTB4; see Table of Materials) to the lower chamber of an impedance-based real time cell analyzer (RTCA) plate. For negative controls and blanks, add 160 µL of HBSS Ca2+Mg2+.
- Attach the upper chamber and add 25 µL of HBSS Ca2+Mg2+. Incubate at room temperature for at least 1 h to form the chemotactic gradient.
- After 60 min of cell activation, gently mix the cell suspension and place 60 µL of cell suspension in the upper chamber. Add 60 µL of HBSS Ca2+Mg2+ to the blank.
- Place the RTCA plate and program the RTCA software to measure the cell index (CI) every 60 s for 2 h.
NOTE: The RTCA plates can be washed for reuse as previously described16. In summary, wash the RTCA chambers and electrodes with phosphate-buferred saline (PBS) three times, then with type I ultrapure water twice. Incubate the lower and upper chamber with 0.25% trypsin 0.53 mM ethylenediaminetetraacetic acid (EDTA) for 40 min. Wash with ultrapure water three times.
7. NET suggestive assay
- After 10 min of cell activation, gently mix the cell suspension and transfer 4 µL of the PMNs from each activation system under evaluation, divided into two wells of a clean glass slide. Incubate in a humidified chamber at 37 °C for 30 min.
- Add 1 µL of DNAse I to one of the wells and incubate for 20 min at 37 °C (in a wet chamber).
- Dry the slide and stain with fast panoptic, as previously described in step 2.9 of the "neutrophil isolation" section.
- Evaluate the slides under a microscope.
NOTE: Look for any indication of NET release, characterized by the presence of web-like structures. Once identified, confirm if the DNAse I treatment was capable of removing such structures. This assay is suggestive of NET formation, as additional tests are required to confirm their presence.
Results
The density-based isolation method used in this study (Figure 1) met the criteria for the proposed experiments. Neutrophil parameters obtained from this method included viability ≥98%, purity ≥94%, and cell yield ≥1.5 x 107, with no activation detectable by the screening tests. Two relevant steps in the isolation of PMNs are anticoagulation and RBC removal. Keeping the anticoagulated blood tube or syringe at a gentle rocking before layering over the density ...
Discussion
Neutrophils are highly dynamic and responsive cells that are short-lived and cannot yet be cryopreserved19, making investigations into their biology challenging. Therefore, it is essential to follow careful steps to obtain viable, enriched, and resting neutrophils11,20. This study employed a density-based isolation technique that emphasizes gentle and minimal manipulation, as well as the use of low temperatures until the activation step. A...
Disclosures
The authors declare no conflict of interest.
Acknowledgements
The authors acknowledge the following funding agencies: FAPDF, CNPq, CAPES, UnB, FINEP, and FINATEC.
Materials
| Name | Company | Catalog Number | Comments |
| CIM-Plate 16 | Agilent | 5665825001 | |
| CLARIOstar Plate Reader | BMG LABTECH | US Patent Number 9,733,124 Product details: MARS Data Analysis Software | |
| Dimethyl sulfoxide | Dinâmica | 1582 | |
| DNAse I | Sigma - Aldrich | DN 25 | |
| Ethylenediaminetetraacetic acid disodium salt dihydrate | Sigma - Aldrich | E5134 | |
| Fast panoptic stain | Laborclin | 620529 | |
| Glass slide | Exacta | 7102 | |
| Hank’s Balanced Salt Solution with calcium, with magnesium, without phenol red. | Sigma - Aldrich | 55037C | |
| Hank’s Balanced Salt Solution without calcium chloride, magnesium sulfate and sodium bicarbonate. | Sigma - Aldrich | H4641 | |
| Heparin | Blau | 7896014655229 | |
| Laminar flow cabinet | Veco | VLFS-12 | |
| Microscope | Zeiss | 415501-0101-002 | Product details: Primostar 1 |
| Mixing Block | BIOER | MB-102 | |
| Neubauer improved bright-lined | New Optik | 1110000 | |
| N-formyl-methionyl-leucyl-phenylalanine | Sigma - Aldrich | F3506 | |
| Nitroblue tetrazolium | Neon | CAS 298-83-9 | |
| Percoll | Cytiva | 17089101 | separation media |
| Phorbol 12-myristate 13-acetate | Sigma - Aldrich | P8139 | |
| Phosphate buffered saline tablet | Sigma - Aldrich | P4417 | |
| ROTOFIX 32 A | Hettich | 1206 | |
| Saccharomyces cerevisiae | Fleischmann | ||
| Safranin | Sigma - Aldrich | 50240 | |
| Sodium dodecyl sulfate | Cytiva | 17-1313-01 | |
| Sonicator | Qsonica | Q125 | |
| Trypan blue solution | Vetec | C.I. 23850 | |
| Vortex Genie 2 | Scientific Industries, Inc. | 0K-0500-902 | |
| xCELLigence Real-Time Cell Analysis (RTCA) DP (dual purpose) | Agilent | 380601050 | Product details: RTCA system composed of detection hardware, cell plates and software |
References
- Nauseef, W. M., Borregaard, N. Neutrophils at work. Nature Immunology. 15 (7), 602-611 (2014).
- Groeneweg, L., Hidalgo, A. Emerging roles of infiltrating granulocytes and monocytes in homeostasis. Cellular and Molecular Life Sciences. 77 (19), 3823-3830 (2020).
- Rosales, C., Lowell, C. A., Schnoor, M., Uribe-Querol, E. Neutrophils: their role in innate and adaptive immunity 2017. Journal of Immunology Research. 2017, 9748345 (2017).
- Castro, M., et al. Proteome analysis of resting human neutrophils. Protein & Peptide Letters. 13 (5), 481-487 (2006).
- Li, Y., et al. The regulatory roles of neutrophils in adaptive immunity. Cell Communication and Signaling. 17, 147 (2019).
- de Oliveira, S., Rosowski, E. E., Huttenlocher, A. Neutrophil migration in infection and wound repair: going forward in reverse. Nature Reviews Immunology. 16 (6), 378-391 (2016).
- Burn, G. L., Foti, A., Marsman, G., Patel, D. F., Zychlinsky, A. The neutrophil. Immunity. 54 (7), 1377-1391 (2021).
- El-Benna, J., et al. Priming of the neutrophil respiratory burst: role in host defense and inflammation. Immunological Reviews. 273 (1), 180-193 (2016).
- Castro, M. S., Cilli, E. M., Fontes, W. Combinatorial synthesis and directed evolution applied to the production of alpha-helix forming antimicrobial peptides analogues. Current Protein & Peptide Science. 7 (6), 473-478 (2006).
- Mihaila, A. C., et al. Transcriptional profiling and functional analysis of N1/N2 neutrophils reveal an immunomodulatory effect of S100A9-blockade on the pro-inflammatory N1 subpopulation. Frontiers in Immunology. 12, 708770 (2021).
- Kuhns, D. B., Priel, D. A. L., Chu, J., Zarember, K. A. Isolation and functional analysis of human neutrophils. Current Protocols in Immunology. 111 (1), 7-23 (2015).
- Paulíková, E., Kociková, A., Sabol, M. Modification of a panoptic method of staining isolated cells. Bratislavske Lekarske Listy. 94 (12), 638-640 (1993).
- Strober, W. Trypan blue exclusion test of cell viability. Current Protocols in Immunology. 111 (1), 1-3 (2015).
- Libério, M. S., et al. Anti-proliferative and cytotoxic activity of pentadactylin isolated from Leptodactylus labyrinthicus on melanoma cells. Amino Acids. 40 (1), 51-59 (2011).
- Cano, P. M., Vargas, A., Lavoie, J. P. A real-time assay for neutrophil chemotaxis. BioTechniques. 60 (5), 245-251 (2016).
- Stefanowicz-Hajduk, J., Adamska, A., Bartoszewski, R., Ochocka, J. R. Reuse of E-plate cell sensor arrays in the xCELLigence Real-Time Cell Analyzer. BioTechniques. 61 (3), 117-122 (2016).
- Björkstén, B., Nyström, K., Lindqvist, B. The nitroblue tetrazolium (NBT) test in endemic benign (epidemic) nephropathy. Acta Medica Scandinavica. 199 (1-6), 147-150 (1976).
- Aquino, E., et al. Proteomic analysis of neutrophil priming by PAF. Protein & Peptide Letters. 23 (2), 142-151 (2016).
- Blanter, M., Gouwy, M., Struyf, S. Studying neutrophil function in vitro: cell models and environmental factors. Journal of Inflammation Research. 14, 141-162 (2021).
- Hsu, A. Y., Peng, Z., Luo, H., Loison, F. Isolation of human neutrophils from whole blood and buffy coats. Journal of Visualized Experiments. (175), e62837 (2021).
- Moghadam, Z. M., Henneke, P., Kolter, J. From flies to men: ROS and the NADPH oxidase in phagocytes. Frontiers in Cell and Developmental Biology. 9, 628991 (2021).
- Pattan, S. S., Bhat, K. G., Pattar, G. D., Kuntagi, M. Comparison of three different techniques for isolation of neutrophils from blood and their utility in performing nitroblue tetrazolium test. International Journal of Basic and Applied Physiology. 8 (1), 41 (2019).
- Gooty, J. R., Shashirekha, A., Guntakala, V. R., Palaparthi, R. Estimation of phagocytic activity of polymorphonuclear leukocytes in chronic and aggressive periodontitis patients with nitroblue tetrazolium test. Journal of Indian Society of Periodontology. 23 (4), 316 (2019).
- Langer, S., et al. Clinical and laboratory profiles of 17 cases of chronic granulomatous disease in north India. Indian Journal of Hematology and Blood Transfusion. 37 (1), 45-51 (2021).
- Oualha, R., et al. Infection of human neutrophils with Leishmania infantum or Leishmania major strains triggers activation and differential cytokines release. Frontiers in Cellular and Infection Microbiology. 9, 153 (2019).
- Zilinskas, J., Zekonis, J., Zekonis, G., Valantiejiene, A., Periokaite, R. The reduction of nitroblue tetrazolium by total blood in periodontitis patients and the aged. Stomatologijal. 9 (4), 105-108 (2007).
- Benov, L. Improved formazan dissolution for bacterial MTT assay. Microbiology Spectrum. 9 (3), e01637 (2021).
- Chen, Y., Junger, W. G. Measurement of oxidative burst in neutrophils. Methods in Molecular Biology. 844, 115-124 (2012).
- Richardson, M. P., Ayliffe, M. J., Helbert, M., Davies, E. G. A simple flow cytometry assay using dihydrorhodamine for the measurement of the neutrophil respiratory burst in whole blood: comparison with the quantitative nitrobluetetrazolium test. Journal of Immunological Methods. 219 (1-2), 187-193 (1998).
- Jancinová, V., et al. The combined luminol/isoluminol chemiluminescence method for differentiating between extracellular and intracellular oxidant production by neutrophils. Redox Report. 11 (3), 110-116 (2006).
- Nosál, R., et al. Pharmacological intervention with oxidative burst in human neutrophils. Interdisciplinary Toxicology. 10 (2), 56-60 (2017).
- Mol, S., et al. Efficient neutrophil activation requires two simultaneous activating stimuli. International Journal of Molecular Sciences. 22 (18), 10106 (2021).
- Schneider, L., et al. Flow cytometry evaluation of CD14/CD16 monocyte subpopulations in systemic sclerosis patients: a cross sectional controlled study. Advances in Rheumatology. 61 (1), 27 (2021).
- Akin, E., Pelen, N. N., Tiryaki, I. U., Yalcin, F. Parameter identification for gompertz and logistic dynamic equations. PLoS One. 15 (4), e0230582 (2020).
- Guy, J. B., et al. Evaluation of the cell invasion and migration process: A comparison of the video microscope-based scratch wound assay and the boyden chamber assay. Journal of Visualized Experiments. (129), e56337 (2017).
- Brinkmann, V., et al. Neutrophil extracellular traps kill bacteria. Science. 303 (5663), 1532-1535 (2004).
- de Bont, C. M., Koopman, W. J. H., Boelens, W. C., Pruijn, G. J. M. Stimulus-dependent chromatin dynamics, citrullination, calcium signalling and ROS production during NET formation. Biochimica et Biophysica Acta. Molecular Cell Research. 1865, 1621-1629 (2018).
- Masuda, S., et al. Measurement of NET formation in vitro and in vivo by flow cytometry. Cytometry Part A. 91 (8), 822-829 (2017).
- Zharkova, O., et al. A flow cytometry-based assay for high-throughput detection and quantification of neutrophil extracellular traps in mixed cell populations. Cytometry Part A. 95 (3), 268-278 (2019).
- Hosseinnejad, A., et al. DNase I functional microgels for neutrophil extracellular trap disruption. Biomaterials Science. 10 (1), 85-99 (2022).
- Chrysanthopoulou, A., et al. Neutrophil extracellular traps promote differentiation and function of fibroblasts. The Journal of Pathology. 233 (3), 294-307 (2014).
- Tong, M., Abrahams, V. M. Visualization and quantification of neutrophil extracellular traps. Methods in Molecular Biology. 2255, 87-95 (2021).
- Santana, C. J. C., et al. Biological properties of a novel multifunctional host defense peptide from the skin secretion of the chaco tree frog, boana raniceps. Biomolecules. 10 (5), 790 (2020).
- Murphy, M. P., et al. Guidelines for measuring reactive oxygen species and oxidative damage in cells and in vivo. Nature Metabolism. 4 (6), 651-662 (2022).
- Boero, E., et al. Use of flow cytometry to evaluate phagocytosis of staphylococcus aureus by human neutrophils. Frontiers in Immunology. 12, 635825 (2021).
- Karsten, C. B., et al. A versatile high-throughput assay to characterize antibody-mediated neutrophil phagocytosis. Journal of Immunological Methods. 471, 46-56 (2019).
- Smirnov, A., Solga, M. D., Lannigan, J., Criss, A. K. Using imaging flow cytometry to quantify neutrophil phagocytosis. Methods in Molecular Biology. 2087, 127-140 (2020).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved