A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Recording Network Activity in Spinal Nociceptive Circuits Using Microelectrode Arrays
In This Article
Summary
The combined use of microelectrode array technology and 4-aminopyridine-induced chemical stimulation for investigating network-level nociceptive activity in the spinal cord dorsal horn is outlined.
Abstract
The roles and connectivity of specific types of neurons within the spinal cord dorsal horn (DH) are being delineated at a rapid rate to provide an increasingly detailed view of the circuits underpinning spinal pain processing. However, the effects of these connections for broader network activity in the DH remain less well understood because most studies focus on the activity of single neurons and small microcircuits. Alternatively, the use of microelectrode arrays (MEAs), which can monitor electrical activity across many cells, provides high spatial and temporal resolution of neural activity. Here, the use of MEAs with mouse spinal cord slices to study DH activity induced by chemically stimulating DH circuits with 4-aminopyridine (4-AP) is described. The resulting rhythmic activity is restricted to the superficial DH, stable over time, blocked by tetrodotoxin, and can be investigated in different slice orientations. Together, this preparation provides a platform to investigate DH circuit activity in tissue from naïve animals, animal models of chronic pain, and mice with genetically altered nociceptive function. Furthermore, MEA recordings in 4-AP-stimulated spinal cord slices can be used as a rapid screening tool to assess the capacity of novel antinociceptive compounds to disrupt activity in the spinal cord DH.
Introduction
The roles of specific types of inhibitory and excitatory interneurons within the spinal cord DH are being uncovered at a rapid rate1,2,3,4. Together, interneurons make up over 95% of the neurons in the DH and are involved in sensory processing, including nociception. Furthermore, these interneuron circuits are important for determining whether peripheral signals ascend the neuroaxis to reach the brain and contribute to the perception of pain5,6,7. To date, most studies have investigated the role of DH neurons at either the single-cell or whole-organism level of analysis using combinations of in vitro intracellular electrophysiology, neuroanatomical labeling, and in vivo behavioral analysis1,3,8,9,10,11,12,13,14. These approaches have significantly advanced the understanding of the role of specific neuron populations in pain processing. However, a gap remains in understanding how specific cell types and small macro-circuits influence large populations of neurons at a microcircuit level to subsequently shape the output of the DH, behavioral responses, and the pain experience.
One technology that can investigate macro-circuit or multicellular-level function is the microelectrode array (MEA)15,16. MEAs have been used to investigate nervous system function for several decades17,18. In the brain, they have facilitated the study of neuronal development, synaptic plasticity, pharmacological screening, and toxicity testing17,18. They can be used for both in vitro and in vivo applications, depending on the type of MEA. Furthermore, the development of MEAs has evolved rapidly, with different electrode numbers and configurations now available19. A key advantage of MEAs is their capacity to simultaneously assess electrical activity in many neurons with high spatial and temporal accuracy via multiple electrodes15,16. This provides a broader readout of how neurons interact in circuits and networks, under control conditions and in the presence of locally applied compounds.
One challenge of in vitro DH preparations is that ongoing activity levels are typically low. Here, this challenge is addressed in spinal cord DH circuits using the voltage-gated K+ channel blocker, 4-aminopryidine (4-AP), to chemically stimulate DH circuits. This drug has previously been used to establish rhythmic synchronous electrical activity in the DH of acute spinal cord slices and under acute in vivo conditions20,21,22,23,24. These experiments have used single-cell patch and extracellular recording or calcium imaging to characterize 4-AP-induced activity20,21,22,23,24,25. Together, this work has demonstrated the requirement of excitatory and inhibitory synaptic transmission and electrical synapses for rhythmic 4-AP-induced activity. Thus, the 4-AP response has been viewed as an approach that unmasks native polysynaptic DH circuits with biological relevance rather than as a drug-induced epiphenomenon. Furthermore, 4-AP-induced activity exhibits a similar response profile to analgesic and antiepileptic drugs as neuropathic pain conditions and has been used to propose novel spinally-based analgesic drug targets such as connexins20,21,22.
Here, a preparation that combines MEAs and chemical activation of the spinal DH with 4-AP to study this nociceptive circuitry at the macro-circuit, or network level of analysis, is described. This approach provides a stable and reproducible platform for investigating nociceptive circuits under naive and neuropathic 'pain-like' conditions. This preparation is also readily applicable to test the circuit-level action of known analgesics and to screen novel analgesics in the hyperactive spinal cord.
Protocol
Studies were carried out on male and female c57Bl/6 mice aged 3-12 months. All experimental procedures were performed in accordance with the University of Newcastle's Animal Care and Ethics Committee (protocols A-2013-312, and A-2020-002).
1. In vitro electrophysiology
- Preparation of solutions for spinal cord slice preparation and recording
- Artificial cerebrospinal fluid
NOTE: Artificial cerebrospinal fluid (aCSF) is used in an interface incubation chamber, where slices are stored until recording commences and during experiments as both perfusate and diluent for drugs. See Table 1 for the detailed composition.
- Artificial cerebrospinal fluid
| Chemical | aCSF (mM) | aCSF (g/100 mL) | Sucrose-substituted aCSF (mM) | Sucrose-substituted aCSF (g/100 mL) | High-potassium aCSF (mM) | High-potassium aCSF (g/100 mL) |
| Sodium chloride (NaCl) | 118 | 0.690 | - | - | 118 | 0.690 |
| Sodium hydrogen carbonate (NaHCO3) | 25 | 0.210 | 25 | 0.210 | 25 | 0.210 |
| Glucose | 10 | 0.180 | 10 | 0.180 | 10 | 0.180 |
| Potasium chloride (KCl) | 2.5 | 0.019 | 2.5 | 0.019 | 4.5 | 0.034 |
| Sodium dihydrogen phosphate (NaH2PO4) | 1 | 0.012 | 1 | 0.012 | 1 | 0.012 |
| Magnesium cloride (MgCl2) | 1 | 0.01 | 1 | 0.01 | 1 | 0.01 |
| Calcium chloride (CaCl2) | 2.5 | 0.028 | 2.5 | 0.028 | 2.5 | 0.028 |
| Sucrose | - | - | 250 | 8.558 | - | - |
Table 1: Artificial Cerebrospinal Fluid compositions. Abbreviation: aCSF = artificial cerebrospinal fluid.
- Prepare aCSF containing (in mM) 118 NaCl, 25 NaHCO3, 10 glucose, 2.5 KCl, 1 NaH2PO4, 1 MgCl2, and 2.5 CaCl2 by adding the required quantities of the above, excluding CaCl2, to 2 L of distilled water.
- Bubble the above solution with carbogen (95% O2, 5% CO2) for 5 min and add CaCl2.
NOTE: This step prevents CaCl2 precipitation, i.e., the solution should not turn cloudy. For drug application during experiments, dilute the drug stock solutions in aCSF to desired final concentrations.
- Sucrose-substituted artificial cerebrospinal fluid
NOTE: Sucrose-substituted aCSF is used during dissection and spinal cord slicing. As indicated by the name, sucrose is substituted for NaCl to reduce neuronal excitation during these procedures while maintaining osmolarity. See Table 1 for the detailed composition.- Prepare sucrose-substituted aCSF containing (in mM) 250 sucrose, 25 NaHCO3, 10 glucose, 2.5 KCl, 1 NaH2PO4, 1 MgCl2, and 2.5 CaCl2 by adding the required quantities of all of the above, excluding CaCl2, to 300 mL of distilled water.
- Bubble the solution with carbogen for 5 min and then add CaCl2.
- Store the solution in a -80 °C freezer for approximately 40 min or until the solution forms a slurry. Avoid freezing solid and use while in slurry consistency.
- Microelectrode array preparation
NOTE: The contact surface of the MEA requires a pretreatment to make it hydrophilic.- Before the experiment, fill the MEA well with either fetal bovine serum (FBS) or horse serum (HS) for 30 min.
- Remove the FBS or HS and thoroughly rinse MEA with approximately five washes of distilled water until the distilled water is no longer foamy. Fill the well with aCSF, ready for use.
- Acute spinal cord slice preparation
NOTE: The mouse spinal cord slice preparation is as previously described by Smith et al.2. Ideally, removal of the lumbosacral enlargement should take no more than 8-10 min (steps 1.3.2-1.3.11 below).- Deeply anesthetize the mouse with 100 mg/kg ketamine (i.p.) and then decapitate it using large surgical scissors.
- Remove the skin over the abdominal region by making a small cut in the skin at the level of the hips. Pull the skin on either side of the cut rostrally until all the skin is removed, i.e., from the top of the rib cage to the top of the pelvis (both ventrally and dorsally).
- Place the body on ice and use a ventral approach to expose the vertebral column by removing all the viscera and cutting through the ribs lateral to the sternum.
- Remove the ventral rib cage, both scapulae (cut off at approximately T2), and the lower limbs and pelvis (cut off at approximately the top of the sacrum).
- Transfer the vertebral column and rib preparation to a dissecting bath containing ice-cold sucrose aCSF. Pin all four corners of the preparation (ventral surface upwards) by placing pins through the lower back muscles and the attached upper ribs.
- Remove all muscle and connective tissue overlying the ventral surface of the vertebrae with rongeurs and identify the vertebral region over the lumbosacral enlargement, which lies approximately beneath the T12 to L2 vertebral bodies.
- Remove a vertebral body that is caudal to the lumbosacral enlargement region to provide access to the spinal cord as it sits in the vertebral canal.
- Using curved spring scissors, cut through the vertebral pedicles bilaterally while lifting and pulling the vertebral body rostrally to separate the ventral and dorsal aspects of the vertebrae and expose the spinal cord.
- Once the vertebral bodies are removed to reveal the lumbosacral enlargement, carefully clear the remaining roots that anchor the spinal cord with spring scissors until the cord floats free.
- Isolate the spinal cord with rostral and caudal cuts well above and below the lumbosacral enlargement, allowing the target region of the cord to 'float free.'
NOTE: The preferred slice orientation will determine how the cord is subsequently mounted for sectioning (Figure 1). - For transverse slices, lift the lumbosacral segment by an attached root and place it on a pre-cut polystyrene (Styrofoam) block (1 cm x 1 cm x 1 cm) with a shallow channel cut in the center. Use cyanoacrylate adhesive (see the Table of Materials) to attach the block and cord to the sectioning platform and place it in the cutting bath containing ice-cold sucrose aCSF (slurry).
NOTE: The shallow channel helps secure and orient the spinal cord, with the dorsal side exposed and the thoracic end of the cord at the bottom of the block. - For sagittal slices, lay a thin line of cyanoacrylate adhesive on the sectioning platform, lift the lumbosacral enlargement by an attached root, and place the cord along the line of glue, ensuring one lateral surface is in the adhesive and the other faces upwards. Place it in the cutting bath containing ice-cold sucrose aCSF (slurry).
- For horizontal slices, put a thin line of cyanoacrylate adhesive on the sectioning platform. Lift the lumbosacral enlargement by an attached root, and place the lumbosacral enlargement along the line of adhesive, ensuring the ventral surface is in the adhesive and the dorsal surface faces upwards. Use attached roots to position the cord. Place it in the cutting bath containing ice-cold sucrose aCSF (slurry).
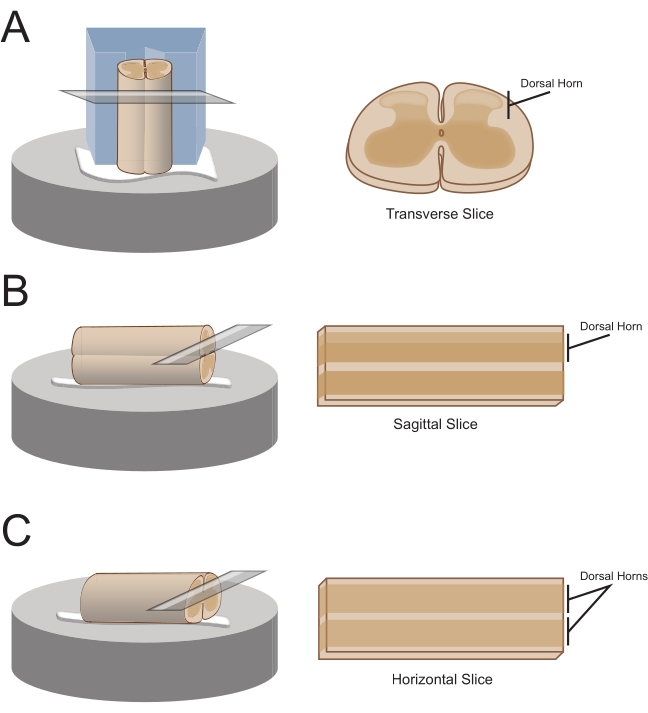
Figure 1: Spinal cord slice orientations, mounting and cutting methods. (A) Transverse slices require a Styrofoam cutting block with a supporting groove cut into it. The spinal cord is rested against the block in the support groove, the dorsal side of the cord facing away from the block. The block and cord are glued onto a cutting stage with cyanoacrylate adhesive. (B) Sagittal slices are prepared by placing a thin line of cyanoacrylate adhesive on the cutting stage and then positioning the spinal cord on its side on the glue. (C) Horizontal slices are prepared by placing a thin line of cyanoacrylate adhesive on the cutting stage and then positioning the spinal cord ventral side down on the glue. Please click here to view a larger version of this figure.
- Obtain 300 µm thick slices (L1-L5, same thickness regardless of orientation) using a vibrating microtome with the following settings: speed 0.06 mm/s, amplitude 2.50 mm, and calibrated to within ±0.02 height amplitude deviation.
- Transfer the slices to an air interface incubation chamber containing oxygenated aCSF.
- Before recording, allow the slices to equilibrate for 1 h at room temperature (20-24 °C).
- Microelectrode array recordings
NOTE: The following steps detail how to use record data from MEA-based experiments on spinal cord slices. Several MEA designs can be used depending on the experiment. Design details for MEAs used in these experiments are shown in Table 2 and Figure 2. Detailed design information has been published by Egert et al.26 and Thiebaud et al.27 for planar and 3-dimensional (3D) MEAs, respectively. Both MEA types are composed of 60 titanium nitride electrodes, with a silicon nitride insulating layer and titanium nitride tracks and contact pads.- Experimental setup
- Turn on the computer and interface board, and start the recording software.
- Load the pre-assembled recording template (Figure 3A). Name the files for the day in the recorder tab.
- Continuously bubble aCSF with carbogen (5% CO2, 95% O2) for the duration of the experiment.
- Turn the perfusion system on, which is controlled by a peristaltic pump. Place the inlet line into aCSF and the inlet end in a waste beaker. Prime the perfusion lines with aCSF.
- Prepare 4-AP and any other drug solutions by diluting stocks in 50 mL of aCSF to the required final concentration (e.g., 200 µM for 4-AP).
- Place the drug solutions in drug pots and bubble them with carbogen.
- 4-AP activity
- Following incubation, transfer a single slice from the incubator using a large-tip Pasteur pipette filled with aCSF.
- Place the slice in the MEA well and add additional aCSF.
- Position the slice over the 60-electrode recording array using a fine short hair paintbrush. Avoid contacting the electrodes with the paintbrush or dragging the tissue across the electrodes, especially if using 3D arrays.
NOTE: Depending on the MEA layout, this can be done with or without the assistance of a microscope for accurate positioning. - After positioning the slice, place a weighted net over the tissue to hold it in place and promote good contact with MEA electrodes.
NOTE: The slice may need repositioning following net placement. - Place the MEA in the recording headstage (Figure 2A,B).
- Check the position of the tissue over the electrodes using an inverted microscope (2x magnification) to confirm that as many electrodes as possible are under the superficial DH (SDH). Ensure that at least 2-6 electrodes do not contact the slice as these electrodes are important for subtracting noise and recording artefacts during analysis (Figure 2E).
- Turn on the camera, connect it to the device, and take a reference image of the slice relative to the MEA for use during analysis.
- Press Start DAQ in the recording software, and confirm that all electrodes are receiving a clear signal.
NOTE: If the signal is noisy, unclip the headstage, and clean both the MEA contact pads and gold spring contacts with 70% ethanol (use a laboratory wipe to ensure that the pads and contacts are dry after cleaning). If the signal is still noisy, turn off the malfunctioning electrodes in the recording software or note down for exclusion later during analysis. - Attach the perfusion inlet and outlet lines to the MEA-well (previously filled with aCSF) and turn the perfusion system on. Check the flow rate, ideally 4-6 bath volumes per minute, and ensure that the outflow is sufficient to prevent overflow of the superfusate.
- Allow the tissue to equilibrate for 5 min and then record 5 min of raw, unfiltered baseline data.
- Move the perfusion inlet line from aCSF to a 4-AP solution and wait for 12 min for the 4-AP-induced rhythmic activity to reach steady state (2 min for drugs to reach the bath and 10 min for the activity to peak and then plateau).
- Record 5 min of 4-AP-induced activity. Be prepared for subsequent recordings to test the drugs or to check the stability of 4-AP.
- Experimental setup
| Microelectrode Array Layouts | ||||
| Microelectrode Array Model | 60MEA 200/30iR-Ti | 60-3DMEA 100/12/40iR-Ti | 60-3DMEA 200/12/50iR-Ti | 60MEA 500/30iR-Ti |
| Planar or 3-Dimensional (3D) | Planar | 3D | 3D | Planar |
| Electrode Grid | 8 x 8 | 8 x 8 | 8 x 8 | 6 x 10 |
| Electrode Spacing | 200 µm | 100 µm | 200 µm | 500 µm |
| Electrode Diameter | 30 µm | 12 µm | 12 µm | 30 µm |
| Electrode Height (3D) | N/A | 40 µm | 50 µm | N/A |
| Experiments | Transverse slice | Transverse slice | Sagittal + Horizontal | Sagittal + Horizontal |
Table 2: Microelectrode array layouts.
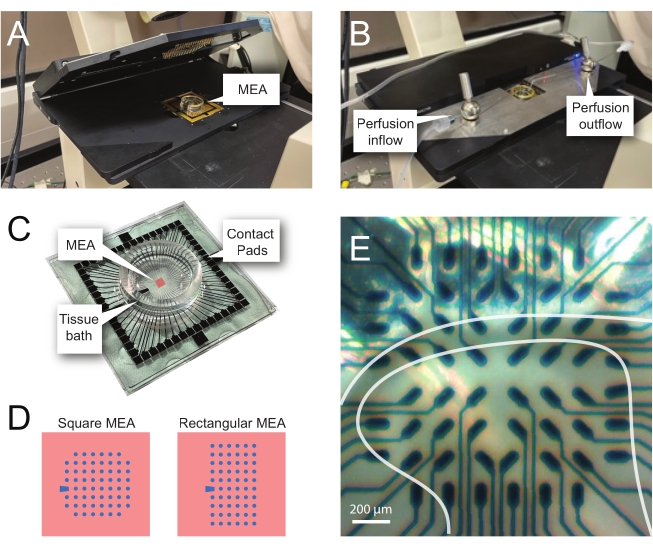
Figure 2: Tissue positioning on the microelectrode array. (A) Image shows an open MEA headstage with an MEA placed in position. (B) Same as A with MEA headstage closed for recordings and tissue perfusion system in place. (C) Image shows an MEA as supplied by the manufacturer. Contact pads, which interface with the gold springs of the headstage, and the MEA tissue bath that holds the tissue bathing solution and tissue slice are shown. The area highlighted by the red square in the center is the location of the electrode array. (D) Schematics show the two MEA electrode configurations used in this study, with further details presented in Table 2. The reference electrode is denoted by the blue trapezoid. The left MEA electrode layout shows a 60-electrode square configuration, used most in the presented work-models 60MEA200/30iR-Ti with 30 μm diameter electrodes spaced 200 μm apart, or 200 μm spaced and 100 μm spaced 3-dimensional MEAs (60MEA200/12/50iR-Ti and 60MEA100/12/40iR-Ti) with electrodes 12 μm in diameter and either 50 μm or 40 μm high, respectively. The left MEA electrode layout shows a 6 x 10 electrode rectangular layout-60MEA500/30iR-Ti. (E) High-magnification image of a 60MEA100/12/40iR-Ti square MEA with transverse spinal cord slice positioned for recording. The slice sits on electrode rows 3-8. The top row of electrodes, which do not contact any tissue, serve as reference electrodes. The SDH area appears as a semitransparent band. In this case, the SDH overlies electrodes in rows 4, 5, and 6 and columns 2, 3, 4, 5, and 7 of the MEA. Scale bar = 200 µm. Abbreviations: MEA = microelectrode array; SDH = superficial dorsal horn. Please click here to view a larger version of this figure.
- Changing slices
- Following each recording session, rinse the lines with aCSF.
- Remove the MEA from the headstage.
- Remove the net and the tissue from the MEA well, rinse them well with aCSF, and repeat the above steps with a new slice.
2. Data processing and analysis
NOTE: The following steps detail how to use the analysis software for MEA experiments on spinal cord slices. One of the 60 electrodes serves as an internal reference (marked by a trapezoid in Figure 2 C,D), while between four and twenty-five of the remaining 59 are positioned under the SDH in an adult mouse spinal cord slice. Subsequent analysis detects extracellular action potential (EAP) and local field potential (LFP) waveforms (see Figure 3B for examples) from the raw signal in this region.
- Raw data processing
- Open the analysis software and load the pre-made analysis layout (Figure 3B).
- Open the file of interest and deselect the reference electrode (electrode 15 in 8 x 8 MEA- or electrode E1 in 6 x 10 MEA-configuration) and any electrodes deemed to be excessively noisy.
- Set the time window for analysis (0:00 → 5:00 min).
- Move to the Cross-channel filter tab. Select Complex reference and select the Reference Electrodes based on the image taken and notes made during the experiment (i.e., those electrodes not under tissue). To apply and check this, press Explore before continuing.
- Move to the EAP filter tab and apply a 2nd order high pass Butterworth filter (200 Hz cut off) to remove LFP activity.
- Move to the LFP filter tab and apply a 2nd order band pass Butterworth filter (delta frequencies of 0.5-4 Hz) to remove EAP activity.
- Move to the EAP detector tab and select Auto threshold. Tick Rising and Falling edge boxes and set the Dead time to 0.5 ms.
- Set Positive and Negative thresholds based on the data. Inspect the data by returning to the Raw data analyzer screen, moving the time marker, and then returning to the EAP detector tab and pressing Explore. Repeat until satisfied that the set detection threshold is capturing EAPs without capturing noise/non-physiological activity. Use the reference electrodes to identify noise/non-physiological activity.
NOTE: It is necessary to ensure a minimal number of EAPs are detected in reference electrodes where physiological activity will not be occurring. However, rather slight deviations in baseline might be falsely detected as EAPs. This is while still aiming to maximize the number of real events detected in the active electrodes. - Move to the LFP detector tab, select the Manual threshold, tick Rising and Falling edge boxes, and set the Dead time to 3 ms.
- Repeat step 2.1.8 for one electrode by selecting an electrode with LFP activity. Once satisfied, select Apply to all as thresholds will only be applied to a single electrode when undertaking manual thresholding.
- While examining LFP data in the Detector tab, note the maximum number of threshold crossings for the one LFP waveform and maximum time separation of threshold crossings for the one LFP waveform for use in later analysis.
- Press Start analysis.
- When the analysis is complete, move to the EAP analyzer tab and export the data. Do the same on the LFP analyzer tab.
- Repeat this process for all other files from the same slice.
- Following data export, convert the files to xlsx format so they can be read by the programming script used. Name the files according to the following convention for the provided script to read them: experiment name (e.g., sample data) - slice number (e.g., S1) - recording number (e.g., R1) - activity type (e.g., spikes or SPs, corresponding to EAPs or LFPs, respectively).
NOTE: The EAP analysis described here treats spiking from individual channels as a single population, even though this activity would commonly arise from multiple neurons in close proximity to the recording electrode. If the number of neurons contributing to EAPs in a channel is desired, multispike sorting techniques described elsewhere can be applied to distinguish distinct populations of spikes based on waveform characteristics28.
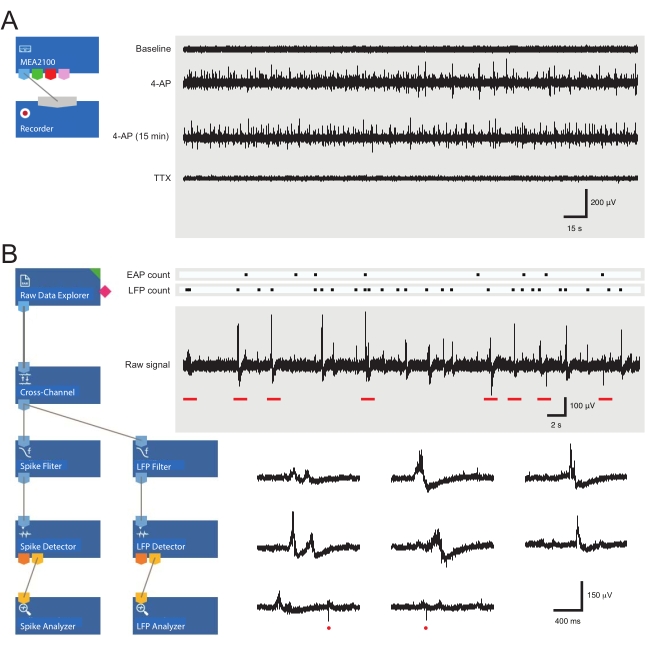
Figure 3: Data recording and analysis tool layouts and example microelectrode array recordings showing extracellular action potential and local field potential waveforms. (A) Schematic shows preconfigured recording templateused for the acquisition of MEA data. Linking the MEA2100 and the recording (headstage/amplifier) tool enables the data to be named and saved. Four example traces of raw data (right, 5-min epochs) were collected by one MEA channel showing activity at baseline, 12 min after 4-AP application, a further 15 min after established 4-AP activity, and following bath application of TTX (1 µM). Note, the addition of 4-AP (second trace) produces a clear increase in background noise and EAP/LFP activity. Importantly, the activity remains relatively stable for at least 15 min after 4-AP-induced activity is established (third trace). Addition of TTX (1 µM) abolishes all activity (bottom trace). (B) Schematic (left) shows analyzer software configuration for data analysis. The raw data explorer tool is used to import recordings collected by recording software. These data are then run through a cross-channel filter tool that subtracts the selected reference electrode(s) signal(s) from other electrodes to remove background noise. Data pass through the EAP filter and the LFP filter tools to optimize signal-to-noise relationships for each waveform. Following this step, the EAP path data enter the EAP detector tool, where thresholds are set. EAPs are detected and then sent to the EAP analyzer tool where the latencies of each event are recorded and exported as a txt. file. An identical workflow occurs for LFP data using a corresponding LFP toolkit. Right traces show data from a single MEA channel containing various extracellular waveforms. Location of EAP and LFP signals are highlighted in the above 'count rasters.' Lower traces are epochs from upper recording (denoted by red bars) showing waveforms on an expanded timescale, including various LFP signals (note the variety of appearances) and individual extracellular EAPs (red circles). Note, LFP/EAP waveform and polarity vary relative to the number of neurons producing these signals, their proximity to the recording electrode, and their location in relation to the nearby electrode(s). Abbreviations: MEA = microelectrode array; EAP = extracellular action potential; LFP = local field potential; 4-AP = 4-aminopyridine; TTX = tetrodotoxin. Please click here to view a larger version of this figure.
- Synchronicity analysis
NOTE: Synchronicity, or the number of 'coincident' events between two electrodes, was determined using the coincidence criterion within the A-SPIKE-synchronization method outlined by Satuvuori et al.29. The script used here only compares electrodes adjacent to one another for efficiency (i.e., horizontal, vertical, and diagonal neighbors); however, the script could be rewritten to compare all electrodes if required.- Perform data analysis using a custom programming script, which extracts latency timestamps for each electrode from the .xlsx files.
NOTE: This can be done manually. - In step 2.1.11, record the maximum number of threshold crossings and maximum time separation of threshold crossings for the one LFP waveform. Modify the script for inputting these LFP-defining parameters for each slice before running the script.
NOTE: Thresholding previously performed in analysis software clearly captures EAPs as a single event. However, LFPs are composed of a variable number of peaks depending on the shape of the waveform and the subsequent number of threshold crossings by the one event. - Modify the script to input the electrodes of interest before analysis.
- To determine synchronicity (defined in the script by modifiable time frames for synchronous activity to occur within), separate and analyze the extracted latencies to detect coincident events.
NOTE: The script allows the maximum time between coincident events to be set. These are set at 20 ms for EAPs and 200 ms for LFPs. - Run the script to extract latency timestamps.
NOTE: The .xlsx output file contains the interpretations of latency data, which are EAP and LFP counts, frequencies, and coincident event counts for individual electrodes and whole slices. These data are used to assess the frequency, EAP/LFP counts, number of active electrodes, number of coincident events, number of linked electrodes, and the average strength of these linkages.
- Perform data analysis using a custom programming script, which extracts latency timestamps for each electrode from the .xlsx files.
Results
Model of network activity in the spinal cord dorsal horn
Application of 4-AP reliably induces synchronous rhythmic activity in the spinal cord DH. Such activity presents as increased EAPs and LFPs. The later signal is a low-frequency waveform, which has previously been described in MEA recordings30. Changes in EAP and/or LFP activity following drug application reflect altered neural activity. Examples of EAPs and LFPs are shown in Figure 3B and ...
Discussion
Despite the importance of the spinal DH in nociceptive signaling, processing, and the resulting behavioral and emotional responses that characterize pain, the circuits within this region remain poorly understood. A key challenge in investigating this issue has been the diversity of neuron populations that comprise these circuits6,31,32. Recent advances in transgenic technologies, led by optogenetics and chemogenetics, are beginn...
Disclosures
The authors have no conflicts of interest to declare.
Acknowledgements
This work was funded by the National Health and Medical Research Council (NHMRC) of Australia (grants 631000, 1043933, 1144638, and 1184974 to B.A.G. and R.J.C.) and the Hunter Medical Research Institute (grant to B.A.G. and R.J.C.).
Materials
| Name | Company | Catalog Number | Comments |
| 4-aminopyridine | Sigma-Aldrich | 275875-5G | |
| 100% ethanol | Thermo Fisher | AJA214-2.5LPL | |
| CaCl2 1M | Banksia Scientific | 0430/1L | |
| Carbonox (Carbogen - 95% O2, 5% CO2) | Coregas | 219122 | |
| Curved long handle spring scissors | Fine Science Tools | 15015-11 | |
| Custom made air interface incubation chamber | |||
| Foetal bovine serum | Thermo Fisher | 10091130 | |
| Forceps Dumont #5 | Fine Science Tools | 11251-30 | |
| Glucose | Thermo Fisher | AJA783-500G | |
| Horse serum | Thermo Fisher | 16050130 | |
| Inverted microscope | Zeiss | Axiovert10 | |
| KCl | Thermo Fisher | AJA383-500G | |
| Ketamine | Ceva | KETALAB04 | |
| Large surgical scissors | Fine Science Tools | 14007-14 | |
| Loctite 454 Instant Adhesive | Bolts and Industrial Supplies | L4543G | |
| MATLAB | MathWorks | R2018b | |
| MEAs, 3-Dimensional | Multichannel Systems | 60-3DMEA100/12/40iR-Ti, 60-3DMEA200/12/50iR-Ti | 60 titanium nitride (TiN) electrodes with 1 internal reference electrode, organised in an 8x8 square grid. Electrodes are 12 µm in diameter, 40 µm (100/12/40) or 50 µm (200/12/50) high and equidistantly spaced 100 µm (100/12/40) or 200 µm (200/12/50) apart. |
| MEA headstage | Multichannel Systems | MEA2100-HS60 | |
| MEA interface board | Multichannel Systems | MCS-IFB 3.0 Multiboot | |
| MEA net | Multichannel Systems | ALA HSG-MEA-5BD | |
| MEA perfusion system | Multichannel Systems | PPS2 | |
| MEAs, Planar | Multichannel Systems | 60MEA200/30iR-Ti, 60MEA500/30iR-Ti | 60 titanium nitride (TiN) electrodes with 1 internal reference electrode, organised in either a 8x8 square grid (200/30) or a 6x10 rectangular grid (500/30). Electrodes are 30 µm in diameter and equidistantly spaced 200 µm (200/30) or 500 µm (500/30) apart. |
| MgCl2 | Thermo Fisher | AJA296-500G | |
| Microscope camera | Motic | Moticam X Wi-Fi | |
| Multi Channel Analyser software | Multichannel Systems | V 2.17.4 | |
| Multi Channel Experimenter software | Multichannel Systems | V 2.17.4 | |
| NaCl | Thermo Fisher | AJA465-500G | |
| NaHCO3 | Thermo Fisher | AJA475-500G | |
| NaH2PO4 | Thermo Fisher | ACR207805000 | |
| Rongeurs | Fine Science Tools | 16021-14 | |
| Small spring scissors | Fine Science Tools | 91500-09 | |
| Small surgical scissors | Fine Science Tools | 14060-09 | |
| Sucrose | Thermo Fisher | AJA530-500G | |
| Superglue | cyanoacrylate adhesive | ||
| Tetrodotoxin | Abcam | AB120055 | |
| Vibration isolation table | Newport | VH3048W-OPT | |
| Vibrating microtome | Leica | VT1200 S |
References
- Smith, K. M., et al. Calretinin positive neurons form an excitatory amplifier network in the spinal cord dorsal horn. eLife. 8, 49190 (2019).
- Smith, K. M., et al. Functional heterogeneity of calretinin-expressing neurons in the mouse superficial dorsal horn: implications for spinal pain processing. The Journal of physiology. 593 (19), 4319-4339 (2015).
- Boyle, K. A., et al. Defining a spinal microcircuit that gates myelinated afferent input: Implications for tactile allodynia. Cell Reports. 28 (2), 526-540 (2019).
- Browne, T. J., et al. Transgenic cross-referencing of inhibitory and excitatory interneuron populations to dissect neuronal heterogeneity in the dorsal horn. Frontiers in Molecular Neuroscience. 13, 32 (2020).
- Graham, B. A., Hughes, D. I. Rewards, perils and pitfalls of untangling spinal pain circuits. Current Opinion in Physiology. 11, 35-41 (2019).
- Todd, A. J. Neuronal circuitry for pain processing in the dorsal horn. Nature Reviews Neuroscience. 11 (12), 823-836 (2010).
- Hughes, D. I., Todd, A. J. Central nervous system targets: inhibitory interneurons in the spinal cord. Neurotherapeutics. 17 (3), 874-885 (2020).
- Duan, B., et al. Identification of spinal circuits transmitting and gating mechanical pain. Cell. 159 (6), 1417-1432 (2014).
- Hachisuka, J., Chiang, M. C., Ross, S. E. Itch and neuropathis itch. Pain. 159 (3), 603 (2018).
- Foster, E., et al. Targeted ablation, silencing, and activation establish glycinergic dorsal horn neurons as key components of a spinal gate for pain and itch. Neuron. 85 (6), 1289-1304 (2015).
- Bourane, S., et al. Identification of a spinal circuit for light touch and fine motor control. Cell. 160 (3), 503-515 (2015).
- Cheng, L., et al. Identification of spinal circuits involved in touch-evoked dynamic mechanical pain. Nature neuroscience. 20 (6), 804-814 (2017).
- Peirs, C., et al. Mechanical allodynia circuitry in the dorsal horn is defined by the nature of the injury. Neuron. 109 (1), 73-90 (2021).
- Huang, J., et al. Circuit dissection of the role of somatostatin in itch and pain. Nature Neuroscience. 21 (5), 707-716 (2018).
- Obien, M. E. J., Deligkaris, K., Bullmann, T., Bakkum, D. J., Frey, U. Revealing neuronal function through microelectrode array recordings. Frontiers in Neuroscience. 8, 423 (2015).
- Nam, Y., Wheeler, B. C. In vitro microelectrode array technology and neural recordings. Critical Reviews in Biomedical Engineering. 39 (1), 45-61 (2011).
- Johnstone, A. F., et al. Microelectrode arrays: a physiologically based neurotoxicity testing platform for the 21st century. Neurotoxicology. 31 (4), 331-350 (2010).
- Stett, A., et al. Biological application of microelectrode arrays in drug discovery and basic research. Analytical and Bioanalytical Chemistry. 377 (3), 486-495 (2003).
- Xu, L., et al. Trends and recent development of the microelectrode arrays (MEAs). Biosensors and Bioelectronics. 175 (1), 112854 (2020).
- Chapman, R. J., Cilia La Corte, P. F., Asghar, A. U. R., King, A. E. Network-based activity induced by 4-aminopyridine in rat dorsal horn in vitro is mediated by both chemical and electrical synapses. The Journal of Physiology. 587, 2499-2510 (2009).
- Ruscheweyh, R., Sandkühler, J. Epileptiform activity in rat spinal dorsal horn in vitro has common features with neuropathic pain. Pain. 105 (1-2), 327-338 (2003).
- Kay, C. W., Ursu, D., Sher, E., King, A. E. The role of Cx36 and Cx43 in 4-aminopyridine-induced rhythmic activity in the spinal nociceptive dorsal horn: an electrophysiological study in vitro. Physiological Reports. 4 (14), 12852 (2016).
- Jankowska, E., Lundberg, A., Rudomin, P., Sykova, E. Effects of 4-aminopyridine on synaptic transmission in the cat spinal cord. Brain Research. 240 (1), 117-129 (1982).
- Semba, K., Geller, H. M., Egger, M. D. 4-Aminopyridine induces expansion of cutaneous receptive fields of dorsal horn cells. Brain Research. 343 (2), 398-402 (1985).
- Ruscheweyh, R., Sandkühler, J. Long-range oscillatory Ca2+ waves in rat spinal dorsal horn. European Journal of Neuroscience. 22 (8), 1967-1976 (2005).
- Egert, U., et al. A novel organotypic long-term culture of the rat hippocampus on substrate-integrated multielectrode arrays. Brain Research Protocols. 2 (4), 229-242 (1998).
- Thiebaud, P., De Rooij, N., Koudelka-Hep, M., Stoppini, L. Microelectrode arrays for electrophysiological monitoring of hippocampal organotypic slice cultures. IEEE Transactions on Biomedical Engineering. 44 (11), 1159-1163 (1997).
- Rey, H. G., Pedreira, C., Quiroga, R. Q. Past, present and future of spike sorting techniques. Brain Research Bulletin. 119, 106-117 (2015).
- Satuvuori, E., et al. Measures of spike train synchrony for data with multiple time scales. Journal of Neuroscience Methods. 287, 25-38 (2017).
- Mendis, G. D. C., Morrisroe, E., Reid, C. A., Halgamuge, S. K., Petrou, S. Use of local field potentials of dissociated cultures grown on multi-electrode arrays for pharmacological assays. 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society. , 952-956 (2016).
- Hughes, D. I., et al. Morphological, neurochemical and electrophysiological features of parvalbumin-expressing cells: a likely source of axo-axonic inputs in the mouse spinal dorsal horn. The Journal of Physiology. 590 (16), 3927-3951 (2012).
- Peirs, C., Seal, R. P. Neural circuits for pain: recent advances and current views. Science. 354 (6312), 578-584 (2016).
- Li, J., Baccei, M. L. Developmental regulation of membrane excitability in rat spinal lamina I projection neurons. Journal of Neurophysiology. 107 (10), 2604-2614 (2012).
- Li, J., Baccei, M. L. Pacemaker neurons within newborn spinal pain circuits. Journal of Neuroscience. 31 (24), 9010-9022 (2011).
- Sandkühler, J., Eblen-Zajjur, A. Identification and characterization of rhythmic nociceptive and non-nociceptive spinal dorsal horn neurons in the rat. Neuroscience. 61 (4), 991-1006 (1994).
- Lucas-Romero, J., Rivera-Arconada, I., Roza, C., Lopez-Garcia, J. A. Origin and classification of spontaneous discharges in mouse superficial dorsal horn neurons. Scientific Reports. 8 (1), 9735-9735 (2018).
- Antonio, L., et al. L. al. In vitro seizure like events and changes in ionic concentration. Journal of Neuroscience Methods. 260, 33-44 (2016).
- Avoli, M., Jefferys, J. G. Models of drug-induced epileptiform synchronization in vitro. Journal of Neuroscience Methods. 260, 26-32 (2016).
- Taccola, G., Nistri, A. Low micromolar concentrations of 4-aminopyridine facilitate fictive locomotion expressed by the rat spinal cord in vitro. Neuroscience. 126 (2), 511-520 (2004).
- Mitra, P., Brownstone, R. M. An in vitro spinal cord slice preparation for recording from lumbar motoneurons of the adult mouse. Journal of Neurophysiology. 107 (2), 728-741 (2012).
- Egert, U., Heck, D., Aertsen, A. Two-dimensional monitoring of spiking networks in acute brain slices. Experimental Brain Research. 142 (2), 268-274 (2002).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved