A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Mechano-Node-Pore Sensing: A Rapid, Label-Free Platform for Multi-Parameter Single-Cell Viscoelastic Measurements
* These authors contributed equally
In This Article
Summary
Presented here is a method to mechanically phenotype single cells using an electronics-based microfluidic platform called mechano-node-pore sensing (mechano-NPS). This platform maintains moderate throughput of 1-10 cells/s while measuring both the elastic and viscous biophysical properties of cells.
Abstract
Cellular mechanical properties are involved in a wide variety of biological processes and diseases, ranging from stem cell differentiation to cancer metastasis. Conventional methods for measuring these properties, such as atomic force microscopy (AFM) and micropipette aspiration (MA), capture rich information, reflecting a cell's full viscoelastic response; however, these methods are limited by very low throughput. High-throughput approaches, such as real-time deformability cytometry (RT-DC), can only measure limited mechanical information, as they are often restricted to single-parameter readouts that only reflect a cell's elastic properties. In contrast to these methods, mechano-node-pore sensing (mechano-NPS) is a flexible, label-free microfluidic platform that bridges the gap in achieving multi-parameter viscoelastic measurements of a cell with moderate throughput. A direct current (DC) measurement is used to monitor cells as they transit a microfluidic channel, tracking their size and velocity before, during, and after they are forced through a narrow constriction. This information (i.e., size and velocity) is used to quantify each cell's transverse deformation, resistance to deformation, and recovery from deformation. In general, this electronics-based microfluidic platform provides multiple viscoelastic cell properties, and thus a more complete picture of a cell's mechanical state. Because it requires minimal sample preparation, utilizes a straightforward electronic measurement (in contrast to a high-speed camera), and takes advantage of standard soft lithography fabrication, the implementation of this platform is simple, accessible, and adaptable to downstream analysis. This platform's flexibility, utility, and sensitivity have provided unique mechanical information on a diverse range of cells, with the potential for many more applications in basic science and clinical diagnostics.
Introduction
Single cells are dynamic, viscoelastic materials1. A multitude of internal and external processes, (e.g., onset of mitosis or remodeling of the extracellular matrix [ECM]), influence their structure and composition2,3,4, often resulting in distinct biophysical properties that complement their current state. In particular, mechanical properties have been shown to be important biomarkers of cellular development, physiology, and pathology, yielding valuable quantitative information that can supplement canonical molecular and genetic approaches5,6,7. For example, Li et al. recently described the mechanical differences between drug-resistant and drug-responsive acute promyelocytic leukemia cells, while also using RNA-seq to uncover differentially-expressed cytoskeleton-associated genes8. By understanding the complex interplay between single-cell mechanics and cellular function, mechanophenotyping has broader applications in transforming basic science and clinical diagnostics9.
The most widely adopted tool for measuring single-cell mechanics is atomic force microscopy (AFM). While AFM enables a high-resolution, localized measurement of cellular mechanical properties, it remains limited to a throughput of <0.01 cells/s10. Alternatively, optical stretchers, which use two divergent laser beams to trap and deform suspended single cells11, are limited to marginally higher throughputs of <1 cell/s12. Recent advances in microfluidic technologies have enabled a new generation of devices for rapid, single-cell, mechanical assessment12,13. These techniques employ narrow constriction channels14,15, shear flow16, or hydrodynamic stretching17 to deform cells quickly at throughputs of 10-1,000 cells/s18. While the measurement rate of these approaches is considerably faster than conventional techniques, they often trade high-throughput capabilities for limited mechanical readouts (Supplementary Table 1). All the aforementioned rapid microfluidic methods focus on basic, single-parameter metrics, such as transit time or deformability ratios, that only reflect a cell's elastic properties. However, given the intrinsic viscoelastic nature of single cells, a robust and thorough mechanical characterization of cells requires consideration of not only elastic components but also viscous responses.
Mechano-node-pore sensing (mechano-NPS)2,8 (Figure 1A) is a microfluidic platform that addresses existing limitations with single-cell mechanophenotyping. This method enables the measurement of multiple biophysical parameters simultaneously, including cell diameter, relative deformability, and recovery time from deformation, with a moderate throughput of 1-10 cells/s. This technique is based on node-pore sensing (NPS)19,20,21,22,23,24, which involves using a four-point probe measurement to measure the modulated current pulse produced by a cell transiting a microfluidic channel that has been segmented by wider regions, referred to as "nodes". The modulated current pulse is a result of the cell partially blocking the flow of current in the segments (i.e., "pores") and nodes, with more current blocked in the former than in the latter. In mechano-NPS, one segment, the "contraction channel", is narrower than a cell diameter; consequently, a cell must deform to transit the entire channel (Figure 1B). Cell diameter can be determined by the magnitude of the subpulse produced when the cell transits the node-pores prior to the contraction channel (Figures 1B,C). Here, |ΔInp|, the current drop when the cell is in the pore, is proportional to the volume ratio of the cell to the pore, Vcell/Vpore2,8,19. Cell stiffness can be determined by ΔTc, the duration of the dramatically larger subpulse produced when the cell transits the contraction channel (Figures 1B,C). A stiffer cell will take longer to transit the channel than a softer one2,8. Finally, cell "recovery", the cell's ability to return to its original size and shape post deformation, can be determined by the series of subpulses produced as the cell transits the node-pores after the contraction channel (Figures 1B,C). The recovery time, ΔTr, is the time it takes for the current subpulses to return to the magnitude of the previous subpulses, prior to the cell being squeezed. Overall, the modulated current pulses produced as a cell transits the microfluidic channel are recorded and analyzed to extract the relevant single-cell mechanical parameters (Figure 1D)2,8.
The reproducibility and ease of use of this electronics-based microfluidic platform have been previously demonstrated25. Additionally, the platform presents a low barrier to entry for single-cell mechanophenotyping. Standard soft lithography is employed to fabricate microfluidic devices. The measurement hardware consists of inexpensive components, including a simple printed circuit board (PCB), power supply, preamplifier, data acquisition board (DAQ), and computer. Finally, user-friendly code is available for data acquisition and analysis, enabling straightforward implementation. This mechanophenotyping technique can distinguish populations of non-malignant and malignant breast and lung epithelial cell lines, discriminate between sublineages in primary human mammary epithelial cells, and characterize the effects of cytoskeletal perturbations and other pharmacological agents2,8. Overall, this platform is an effective approach for the mechanophenotyping of single cells.
Protocol
1. Design device geometry
- Choose the width of the sizing and recovery segments so that it is wider than the diameter of the largest cells to be measured but also maintains a sufficient signal-to-noise ratio (SNR). See Supplementary Table 2 for examples of different sizing and recovery segment widths for various cell lines.
- Choose the contraction segment width to apply a 30%-40% strain to the average size of the cells that are to undergo mechanophenotyping. Strain is defined as
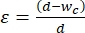 , where d is cell diameter and wc is the contraction channel width2,8. See Supplementary Table 2 for different contraction segment widths for various cell lines.
, where d is cell diameter and wc is the contraction channel width2,8. See Supplementary Table 2 for different contraction segment widths for various cell lines.
NOTE: If one wishes to compare cell types or conditions with substantially different diameters, separate device designs should be used with contraction segment widths specific to each cell type/condition. - Design a reference device for each unique device geometry. This is necessary for determining De, the effective diameter of the sizing pore segment of the microfluidic channel.
NOTE: The reference device uses the same geometry as the primary device. The only modification is that the contraction segment should be equal in width to the sizing pore segment to allow for calibration with polystyrene beads of a known size. Widening the contraction prevents the polystyrene beads from clogging the contraction channel during calibration. The calibration process is further described in steps 4.1 and 5.3.1. Calibration can also be achieved using a commercially available cell counter, in which case, no reference device is needed. This process is described in step 4.2. - Choose the channel height such that the largest cells of interest can fully elongate without restriction within the contraction segment2. Ensure that the channel height is larger than hmin
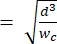 (this assumes the cell is spherical pre-deformation, and that isometric deformation occurs along the channel length and height during deformation).
(this assumes the cell is spherical pre-deformation, and that isometric deformation occurs along the channel length and height during deformation).
NOTE: Given the magnitude of a current subpulse,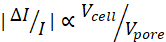 , the larger the hmin is, the lower the overall SNR will be.
, the larger the hmin is, the lower the overall SNR will be. - Design and create a photomask using computer-aided design software with the chosen channel widths. An example file is provided in Supplementary File 1. Scale the microfluidic mask design by 1.5% to account for polydimethylsiloxane (PDMS) shrinkage after peeling from the negative master.
NOTE: An array of devices can be included on a single mask as long as the overall array does not exceed the size of the wafer (Supplementary Figure 1A). - Design and create a photomask with electrodes that will be used to perform a four-point probe measurement of the microfluidic device current (Figure 1D). An example file is provided in Supplementary File 1.
NOTE: An array of electrodes can be included on a single mask as long as the array does not exceed the size of the glass slide (Supplementary Figure 1B).
2. Fabricate devices (Figure 2)
- Prepare electrode patterns on a glass substrate.
- Spin coat, pattern, and process a positive photoresist onto a plain glass slide according to the product data sheet. An example of this procedure is outlined in Supplementary File 2.
- Perform metal deposition, lift-off, and gold etching.
- Perform thin film deposition of 75 Å Ti, 250 Å Pt, and 250 Å Au onto the slide. An example of this procedure using electron-gun evaporation is outlined in Supplementary File 3.
- Immerse the slide in acetone for 15 min to perform a lift-off of excess metal.
- In a fume hood, use a disposable pipette to drop-cast gold etchant onto the region of electrodes that will be exposed to the microfluidic channel, as shown in Supplementary Figure 2. Be cautious to avoid dropping etchant elsewhere on the slide.
CAUTION: Gold etchant can cause skin and eye irritation. Do not breathe vapors, and do not ingest. Handle with care, wear appropriate personal protective equipment (PPE), and discard waste according to local disposal regulations. - Rinse the slide with deionized (DI) water and dry it with dry nitrogen (N2).
- If multiple electrodes are printed on the same glass slide, dice the slide into individual chips.
- Use a glass cutting tool to score the slide along the patterned electrode boundaries.
- Break the glass along the score to partition the slide into individual chips.
- Visually inspect the electrodes under a microscope. Ensure that individual electrodes are not electrically open or that electrodes are not shorted together.
- Fabricate a negative master mold for channels.
- Spin coat, pattern, and process an SU-8 epoxy resist onto a polished silicon wafer according to the product data sheet. An example of this procedure is outlined in Supplementary File 2.
- Measure feature heights using a profilometer and visually inspect the features under a microscope (Supplementary Figure 3). Ensure that the desired geometries are well-defined.
- Mold PDMS channels with soft lithography.
- Prepare PDMS by weighing an elastomer and a crosslinker at a 10:1 mass ratio in a disposable cup.
NOTE: For a wafer with a 3 in diameter, 30 g of PDMS is sufficient. - Mix the PDMS vigorously for 30 s with a disposable fork, until the PDMS is opaque with bubbles.
- De-gas the PDMS in a vacuum chamber for approximately 30-90 min, or until the PDMS is transparent with no visible bubbles.
- Place the wafer with the SU-8 master mold into a disposable Petri dish and pour PDMS over the center of the wafer.
- Place the Petri dish containing the PDMS and wafer in a vacuum chamber and de-gas for approximately 30 min, or until no bubbles remain in the PDMS.
- Bake the PDMS at 80 °C for 2 h in an oven or on a hot plate.
- With a sharp blade, cut and remove the PDMS from the SU-8 negative master.
- Dice the molded PDMS slab into individual molds using a sharp blade
- Core the inlet and outlet access holes using a disposable biopsy punch. For best results, use a new punch for each PDMS slab. A sharper punch produces smooth-edged holes, minimizing particulates that could obstruct the contraction channel.
NOTE: The diameter of the access holes should be slightly less than the outer diameter of the tubing. For example, if using polytetrafluoroethylene (PTFE) tubing with an outer diameter of 1/32 in, a 1.5 mm hole should be punched.
- Prepare PDMS by weighing an elastomer and a crosslinker at a 10:1 mass ratio in a disposable cup.
- Bond a glass/electrode substrate to the PDMS channels.
- Clean the electrode glass slides with methanol (≥99.8%). Dry with dry N2.
- Clean the PDMS device with scotch tape, followed by a rinse with isopropyl alcohol (IPA) and deionized water (DI; 18 MΩ/cm2). Dry with dry N2. Then, clean with scotch tape once more.
- Place the glass substrate with prefabricated electrodes and the prepared PDMS mold (feature side up) into a plasma cleaner.
- Expose both to oxygen plasma for 2 min (100-300 mTorr, 30 W).
- Align and place the PDMS mold with the feature side face-down onto the glass substrate with prefabricated electrodes.
NOTE: Bonding is instantaneous once the plasma-treated PDMS and glass come into contact; consequently, further alignment modifications will not be possible. To facilitate alignment, 20 µL of a 2:1 dilution of methanol in DI water may be pipetted onto the plasma-treated glass surface. The methanol solution acts as a physical barrier between the treated glass and PDMS, allowing for alignment adjustments. If using methanol, bake the aligned and mated device at 50 °C for 2 h to evaporate the solution and complete the bonding process. - Visually inspect the bonded device under a microscope. Ensure that the electrodes and channel geometries are properly aligned.
3. Measure cells (Figure 1D)
- Prepare the pressure source, PCB, benchtop hardware, and data acquisition software.
- Connect the microfluidic device to the PCB using the clamp. An example of the PCB is provided in Supplementary File 4 (GERBER files) and Supplementary File 5 (schematic, board, and PCB parts list files).
- Align the clamp's spring-loaded pins with the electrode contact pads on the microfluidic device and align the clamp's header pins with the holes on the PCB.
- Firmly insert the clamp's header pins into the PCB holes, making sure the spring-loaded pins stay aligned with the electrode contact pads.
- Set up and connect the electronic hardware.
- Connect two of the power supply's output ports to the PCB's supply voltage port with a double banana plug-to-Bayonet Neill-Concelman (BNC) female adapter and a BNC cable.
- Turn on the power supply. Set the output connected to the BNC's inner conductor to +15 V and set the other output to -15 V. Enable both outputs to power the circuit.
- Connect the third of the power supply's output ports to the input voltage port of the PCB with a BNC cable. Set the output to the desired applied voltage, but do not enable it until starting the experiment.
- Connect the PCB's output current port to the input of the current preamplifier with a BNC cable.
- Connect the output of the current preamplifier to one analog input on the BNC terminal block of the data acquisition system with a BNC cable. Optionally, connect an analog low-pass filter in line with the BNC cable to filter out high-frequency interference.
NOTE: To improve the SNR, the PCB and device may be housed within a thick metal enclosure. All BNC cables and fluidic tubing can be routed through holes drilled into the enclosure.
- Install and set up the required software on the personal computer (PC)
- Power on and connect the pressure controller to the PC. Install any required pressure controller software as per the manufacturer's instructions.
- Install MATLAB and the Data Acquisition Toolbox on the PC. Ensure the required drivers for the data acquisition system are installed so that the MATLAB Data Acquisition Toolbox interface can detect it.
- Download the included data acquisition script, "NPS.m", from https://github.com/sohnlab/node-pore-sensing-public.
- Open and configure the data acquisition script.
- Set the correct values to initialize the data acquisition session, which includes the Vendor ID, the DAQ's Device ID, and the analog input channel number (lines 34-36 in the included script).
NOTE: The Device ID can be found using the function "daq.getDevices" or "daqlist". - Set the desired sample rate for the acquisition (line 23 in the included script). For optimal results, it should be set to at least 10 kHz.
- Set the correct values to initialize the data acquisition session, which includes the Vendor ID, the DAQ's Device ID, and the analog input channel number (lines 34-36 in the included script).
- Connect the microfluidic device to the PCB using the clamp. An example of the PCB is provided in Supplementary File 4 (GERBER files) and Supplementary File 5 (schematic, board, and PCB parts list files).
- Prepare the cell suspension.
- Prepare a solution of 2% fetal bovine serum (FBS) in 1x phosphate buffered saline (PBS), and filter with a 0.22 µm filter.
- Culture and prepare the cells according to the appropriate cell-culture protocol of the cell line of choice. Suspend the cells in the prepared solution of 2% FBS in 1x PBS at a concentration of 1-5 x 105 cells/mL. Keep the cells on ice for the duration of the experiments.
- Measure the physical properties of the cells.
- Load the cell sample into the tubing and connect it to the device inlet.
- Cut 30 cm of PTFE tubing with a razor blade or sharp knife.
- Attach one end of the tubing to a luer lock syringe. Use the syringe to draw up the cell sample into the other end of the tubing.
- Carefully insert the tubing into the inlet of the device.
- Connect the opposite end of the tubing to the microfluidic pressure controller.
NOTE: A filter can be added between the microfluidic pressure controller and the tubing to prevent liquid backflow into the pressure controller.
- Run the experiment.
- Set the desired constant driving pressure on the pressure controller software and allow the sample to fill the device.
NOTE: The pressure is typically 2-21 kPa. The flow speed must be slow enough to allow for clearly defined pulses but fast enough to allow for adequate throughput.- If bubbles form in the microfluidic channels, use dead-end filling: plug the device outlet and apply a low pressure to the inlet to force air out through the gas-permeable PDMS. Leaving bubbles in the channel will lead to an unstable current baseline and prevent accurate measurements.
- If debris clogs the microfluidic channel, dislodge it by lightly pressing on the top of the PDMS device while applying the driving pressure, "pulsing" a higher pressure by toggling the pressure on and off, or removing the tubing and reinserting it. If the debris remains, it may be necessary to switch to a new device.
- Set the desired voltage by rotating the Voltage knob on the power supply and enable the voltage by pressing the On button.
NOTE: Voltage is typically 1-5 V. Choose the lowest voltage necessary for an adequate SNR. The same voltage should be used across all conditions to be compared. - Turn on the current preamplifier and set the sensitivity (A/V) as low as possible; alternatively, set the gain (V/A) as high as possible without overloading the preamplifier or exceeding the maximum analog input voltage of the DAQ. In this study, the sensitivity was set to 10-7 A/V.
NOTE: The proper sensitivity/gain value will depend on both the applied voltage as well as the baseline resistance of the microfluidic channel. - Press the green Run button in the MATLAB ribbon menu to begin the data acquisition script NPS.m and start sampling and saving the data.
- To end the experiment, press the Stop button in the lower left corner of the figure window to stop the data acquisition script. Disable the power supply output by pressing the On button. Set the pressure source to zero pressure in the pressure controller software.
- At this point, the experiment can be paused to do one or more of the following:
- Replace the current device with a new one.
- Reload the tubing with more cell samples.
NOTE: To avoid sample cross-contamination, use new devices to measure cells of different types or conditions. - Unclamp the device from the PCB and examine the channel's condition under a microscope. To restart the experiment using the same device, care must be taken not to introduce air bubbles. It may be necessary to apply gentle pressure to the syringe plunger to keep the cell sample at the very end of the tubing while inserting it into the device inlet.
- Set the desired constant driving pressure on the pressure controller software and allow the sample to fill the device.
- Load the cell sample into the tubing and connect it to the device inlet.
4. Calibrate the microfluidic device
- Option 1: Measure the polystyrene beads in reference devices.
- Choose a polystyrene bead size that is smaller than the sizing channel.
- Add 1.5% Tween and polystyrene beads to the filtered PBS and FBS solution used during the cell experiments, at a concentration of 1-3 x 105 beads/mL.
- Proceed with the experiment as outlined in section 3, using the reference device described in step 1.3, and apply the same voltage used during experimentation. Use the average magnitude of the current drop produced as beads transit the sizing pores and the known diameter of the beads to calculate De, as described in section 5.
- Option 2: Independently measure the cell size with a separate measurement device.
- Instead of following the protocol in step 4.1, use a commercially available cell size measurement instrument to measure the average size of cells in the sample. In this case, no reference device is needed. Use the average current drop produced as cells transit the sizing pore and the measured average cell diameter to calculate De as described in section 5.
5. Analyze data to extract cell phenotypes
NOTE: Data processing can be performed using the MATLAB command-line interface program file mNPS_procJOVE.m at https://github.com/sohnlab/NPS-analysis-JOVE. See Supplementary File 6 for more instructions.
- Preprocess the data (Figure 3A).
- Compute the measured electrical current by applying the gain value used in the current-to-voltage preamplifier to the raw data acquired by the DAQ.
- Remove high-frequency noise by applying a rectangular smoothing function and/or a low-pass filter to the raw current measurement. Then, resample the filtered data to a lower sample rate. Also, compute the corresponding timestamp data at this lower sample rate.
- Compute a fitted baseline current signal by applying a method such as asymmetric least-squares smoothing26.
- Compute the approximate first derivative (difference signal) of the preprocessed current data by taking the difference between subsequent data points.
- Identify cell events and extract subpulse data (Figure 3B).
- Search for candidate cell events by examining the preprocessed data. Reject cell events that overlap with other cell events (i.e., coincidence events) (Supplementary Figure 4), exhibit a poor baseline fit, or have an unexpected or erroneous pulse shape (e.g., where a clog may have been present in the channel).
- Extract subpulse data for each cell event.
- Each node-pore segment will appear as a corresponding subpulse within the overall signal pulse (Figures 1B, C). Identify the start of each subpulse by computing the timepoint when the difference signal reaches a local minimum value. Identify the end of each subpulse by computing the timepoint when the difference signal reaches a local maximum value.
- Determine the width of each subpulse as the elapsed time between the start and end time points. Determine the amplitude of each subpulse by computing the mean of the difference between the measured current and the baseline current for all data points between the start and end time points.
- Determine the cell mechanophenotype for each cell event based on subpulse data.
- Determine the cell diameter d based on the equation defined by Deblois and Bean24:
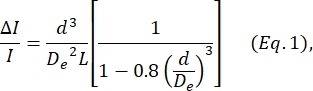
where ΔI/I is the mean ratio of subpulse amplitude to baseline current in the sizing subpulses, De is the effective diameter of the channel (measured in step 4), and L is the total length of the node-pore channel.- De is determined by calculating the average ΔI/I produced by a set of particles of a known diameter (either cells or beads, see step 4), using that known diameter as d, and solving Eq. 1 for De.
- Quantify the cell's resistance to deformation.
- Determine the fluid velocity Uflow by calculating the mean cell velocity in the sizing subpulses, using the known segment lengths and measured duration of each subpulse.
- Determine the whole-cell deformability index (wCDI), defined by Kim et al.2 as:
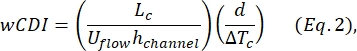
where Lc is the length of the contraction segment, hchannel is the channel height, and ΔTc is the duration of the contraction subpulse.
- Identify the cell's recovery time from deformation, defined as the first recovery subpulse with an amplitude within 8% of the mean amplitude from the sizing subpulse2.
- Calculate the cell's transverse deformation within the contraction segment.
- Calculate the effective diameter of the contraction segment (De,c) as defined by Kim et al.2:
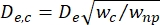 , where wc is the width of the contraction segment and wnp is the width of all other segments.
, where wc is the width of the contraction segment and wnp is the width of all other segments. - Calculate the equivalent spherical diameter dc of the cell within the contraction by again using the equation defined by Deblois and Bean24:
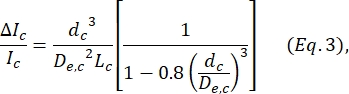
where ΔIc/Ic is the ratio of subpulse amplitude to baseline current in the contraction subpulse and Lc is the length of the contraction segment. - Calculate the cell's elongation length Ldeform as described by Kim et al.2:
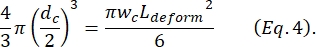
- Finally, compute the cell's transverse deformation δdeform, which is defined by Kim et al.2 to be
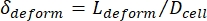 .
.
- Calculate the effective diameter of the contraction segment (De,c) as defined by Kim et al.2:
- Determine the cell diameter d based on the equation defined by Deblois and Bean24:
Results
The mechanophenotyping platform presented here is a simple and versatile approach for measuring the biophysical properties of single cells with moderate throughput. Cells are flowed through the microfluidic channel (Figure 1A) using constant pressure-driven flow. As the cells transit, the length of the microfluidic channel and the current pulses produced are recorded using the data acquisition hardware. The acquired signal (Figure 1B,C) is then ...
Discussion
Measuring the mechanical properties of single cells using this mechanophenotyping technique consists of three stages: device fabrication, data acquisition, and data analysis. Within each stage, there are notable aspects that may significantly impact the experimental results. During device fabrication, consistent channel geometries and device-to-device uniformity are essential for accurate and repeatable results. Specifically, the sidewalls of each device should be relatively smooth (Figure 4Ai
Disclosures
L. L. S holds US patent No. 11,383,241: "Mechano-node-pore sensing", J. Kim, S. Han, and L. L. Sohn, issued July 12, 2022.
Acknowledgements
This research was supported by grants from NIBIB 1R01EB024989-01 and NCI 1R01CA190843-01. A. L. and R. R. were supported by an H2H8 Association Graduate Research Fellowship. K. L. C. was supported by a National Science Foundation Graduate Research Fellowship and a Siebel Scholar Fellowship.
Materials
| Name | Company | Catalog Number | Comments |
| Acetone | J.T. Baker | 5356-05 | Purity (GC) ≥ 99.5% (https://us.vwr.com/store/product/6057739/acetone-99-5-vlsi-j-t-baker) |
| Aluminum Foil | n/a | n/a | |
| Analog Low-Pass Filter | ThorLabs | EF504 | ≤240 kHz Passband, Coaxial BNC Feedthrough (https://www.thorlabs.com/thorproduct.cfm?partnumber=EF504#ad-image-0) |
| Biopsy Punch | Integra Miltex | 33-31AA-P/25 | 1mm, Disposable, with Plunger (https://mms.mckesson.com/product/573313/Miltex-33-31AA-P25) |
| Blade | n/a | n/a | |
| BNC Cable | Pomona Electronics | 2249-C-12 | https://www.digikey.com/en/products/detail/pomona-electronics/2249-C-12/603323?utm_adgroup=Coaxial%20Cables%20%28RF%29&utm_source=google&utm_ medium=cpc&utm_campaign= Shopping_Product_Cable%20Assemblies_NEW&utm_term= &utm_content=Coaxial%20Cables%20%28RF%29&gclid=Cj0KCQjwlK-WBhDjARIsAO2sErQqnVJ pj5OXVObuTI8ZUf1ZeIn7zvzGnx mCWdePrG6SdEJMF3X6ubUaAs w-EALw_wcB |
| Cleanroom Polyester Swab | Thermo Fisher Scientific | 18383 | https://www.fishersci.com/shop/products/texwipe-cleantip-alpha-polyester-series-swabs-6/18383 |
| Current Preamplifier | DL Instruments | 1211 | https://www.brltest.com/index.php?main_page=product_info&products_ id=1419 |
| Custom PCB (w/ components) | n/a | n/a | see Supplemental files 4 and 5 |
| DAQ Terminal Block | National Instruments | BNC-2120 | https://www.ni.com/en-in/support/model.bnc-2120.html |
| DAQ to BNC-2110 cable | National Instruments | SHC68-68-EPM | https://www.ni.com/en-in/support/model.shc68-68-epm.html |
| Data Acquisition Board (DAQ) | National Instruments | PCI-6251 | https://www.ni.com/docs/en-US/bundle/pci-6251-feature/page/overview.html |
| Dessicator | Thermo Fisher Scientific | 5311-0250 | https://www.thermofisher.com/order/catalog/product/5311-0250 |
| Female BNC To Banana Plug Adapter | Pomona Electronics | 72909 | https://www.digikey.com/en/products/detail/pomona-electronics/72909/1196318 |
| Fetal Bovine Serum (FBS) | VWR | 89510-186 | https://us.vwr.com/store/product/18706419/avantor-seradigm-select-grade-usda-approved-origin-fetal-bovine-serum-fbs |
| Glass Cutter | Chemglass | CG-1179-21 | https://chemglass.com/plate-glass-cutters-diamond-tips |
| Gold Etchant TFA | Transene | NC0977944 | https://www.fishersci.com/shop/products/NC0977944/NC0977944 |
| Hot Plate | Thermo Fisher Scientific | SP131825 | |
| Isopropyl Alcohol | Spectrum Chemical | I1056-4LTPL | Purity (GC) ≥99.5% (https://www.spectrumchemical.com/isopropyl-alcohol-99-percent-fcc-i1056) |
| Metal Hardware Enclosure | Hammond Manufacturing | EJ12126 | https://www.digikey.com/en/products/detail/hammond-manufacturing/EJ12126/2423415 |
| Methanol | Sigma-Aldrich | 34860 | Purity (GC) ≥99.8% (https://www.sigmaaldrich.com/IN/en/substance/methanol320467561) |
| MF-321 Developer | Kayaku Advanced Materials | n/a | https://kayakuam.com/products/mf-321/ |
| MICROPOSIT S1813 Positive Photoresist | DuPont | n/a | https://kayakuam.com/products/microposit-s1800-g2-series-photoresists/ |
| Phosphate Buffered Saline (PBS) | Thermo Fisher Scientific | 10010049 | https://www.thermofisher.com/order/catalog/product/10010049?SID=srch-hj-10010049 |
| Photomask | Fineline Imaging | n/a | Photomask are custom ordered from our CAD designs (https://www.fineline-imaging.com/) |
| Plain Glass Microscope Slide | Fisher Scientific | 12-553-5B | Material: Soda Lime, L75 x W50 mm, Thickness: 0.90–1.10 mm |
| Plasma Cleaner | Harrick Plasma | PDC-001 | https://harrickplasma.com/plasma-cleaners/expanded-plasma-cleaner/ |
| Plastic Petri Dish | Thermo Fisher Scientific | FB0875712 | 100 mm (https://www.fishersci.com/shop/products/fisherbrand-petri-dishes-clear-lid-raised-ridge-100-x-15mm/FB0875712) |
| Pressure Controller | Fluigent | MFCS-EZ | https://www.fluigent.com/research/instruments/pressure-flow-controllers/mfcs-series/ |
| Pressure Controller Software | Fluigent | MAESFLO | |
| Programming & Computation Software | MATLAB | R2021b | for data acquisition and analysis (https://www.mathworks.com/products/matlab.html) |
| PTFE Tubing | Cole Parmer | 06417-31 | 0.032" ID x 0.056" (https://www.coleparmer.com/i/masterflex-transfer-tubing-microbore-ptfe-0-032-id-x-0-056-od-100-ft-roll/0641731) |
| Scepter 2.0 Handheld Automatic Cell Counter | Millapore Sigma | PHCC20060 | https://www.sigmaaldrich.com/IN/en/product/mm/phcc20060 |
| Silicon Wafer | Wafer World | 2885 | 76.2 mm, Single Side Polished (https://www.waferworld.com/product/2885) |
| Spin Coater | n/a | n/a | |
| SU-8 3025 Negative Photoresist | Kayaku Advanced Materials | n/a | https://kayakuam.com/products/su-8-2000/ |
| SU8 Developer | Kayaku Advanced Materials | n/a | https://kayakuam.com/products/su-8-developer/ |
| Sygard 184 Polydimethlysiloxane | Dow Chemical | 4019862 | https://www.ellsworth.com/products/by-market/consumer-products/encapsulants/silicone/dow-sylgard-184-silicone-encapsulant-clear-0.5-kg-kit/ |
| Tape | Scotch | 810-341296 | https://www.staples.com/Scotch-Magic-Tape-810-3-4-x-36-yds-1-Core/product_130567?cid=PS:GS:SBD:PLA:OS&gclid= Cj0KCQjwlK-WBhDjARIsAO 2sErRwzrrgjU0NjFkDkne1xm vT7ekS3tdzvAgiMDwPoxocgH VTQZi7vJgaAvQZEALw_wcB |
| Titanium, Platinum, Gold | n/a | n/a | |
| Triple Output Power Supply | Keysight | E36311A | https://www.newark.com/keysight-technologies/e36311a/dc-power-supply-3o-p-6v-5a-prog/dp/15AC9653 |
| UV Mask Aligner | Karl Suss America | MJB3 Mask Aligner |
References
- Pegoraro, A. F., Janmey, P., Weitz, D. A. Mechanical properties of the cytoskeleton and cells. Cold Spring Harbor Perspectives in Biology. 9 (11), 022038 (2017).
- Kim, J., et al. Characterizing cellular mechanical phenotypes with mechano-node-pore sensing. Microsystems & Nanoengineering. 4, 17091 (2018).
- Mierke, C. T. Bidirectional mechanical response between cells and their microenvironment. Frontiers in Physics. 9, 619 (2021).
- Kumar, S., Weaver, V. M. Mechanics, malignancy, and metastasis: The force journey of a tumor cell. Cancer and Metastasis Reviews. 28 (1), 113-127 (2009).
- Nia, H. T., Munn, L. L., Jain, R. K. Physical traits of cancer. Science. 370 (6516), (2020).
- Fletcher, D. A., Mullins, R. D. Cell mechanics and the cytoskeleton. Nature. 463 (7280), 485-492 (2010).
- Wirtz, D., Konstantopoulos, K., Searson, P. C. The physics of cancer: The role of physical interactions and mechanical forces in metastasis. Nature Reviews Cancer. 11 (7), 512-522 (2011).
- Li, B., et al. Mechanical phenotyping reveals unique biomechanical responses in retinoic acid-resistant acute promyelocytic leukemia. iScience. 25 (2), 103772 (2022).
- Kozminsky, M., Sohn, L. L. The promise of single-cell mechanophenotyping for clinical applications. Biomicrofluidics. 14 (3), 031301 (2020).
- Li, M., Dang, D., Liu, L., Xi, N., Wang, Y. Atomic force microscopy in characterizing cell mechanics for biomedical applications: A review. IEEE Transactions on Nanobioscience. 16 (6), 523-540 (2017).
- Wottawah, F., et al. Optical rheology of biological cells. Physical Review Letters. 94 (9), 1-4 (2005).
- Darling, E. M., Di Carlo, D. High-throughput assessment of cellular mechanical properties. Annual Review of Biomedical Engineering. 17 (1), 35-62 (2015).
- Carey, T. R., Cotner, K. L., Li, B., Sohn, L. L. Developments in label-free microfluidic methods for single-cell analysis and sorting. Wiley Interdisciplinary Reviews: Nanomedicine and Nanobiotechnology. 11 (1), 1529 (2019).
- Bagnall, J. S., et al. Deformability of tumor cells versus blood cells. Scientific Reports. 5, 18542 (2015).
- Byun, S., et al. Characterizing deformability and surface friction of cancer cells. Proceedings of the National Academy of Sciences. 110 (19), 7580-7585 (2013).
- Otto, O., et al. Real-time deformability cytometry: On-the-fly cell mechanical phenotyping. Nature Methods. 12 (3), 199-202 (2015).
- Gossett, D. R., et al. Hydrodynamic stretching of single cells for large population mechanical phenotyping. Proceedings of the National Academy of Sciences. 109 (20), 7630-7635 (2012).
- Guck, J., Chilvers, E. R. Mechanics meets medicine. Science Translational Medicine. 5 (212), 3-6 (2013).
- Balakrishnan, K. R., et al. Node-pore sensing: A robust, high-dynamic range method for detecting biological species. Lab on a Chip. 13 (7), 1302-1307 (2013).
- Carbonaro, A., Sohn, L. L. A resistive-pulse sensor chip for multianalyte immunoassays. Lab on a Chip. 5 (10), 1155-1160 (2005).
- Saleh, O. A., Sohn, L. L. Direct detection of antibody-antigen binding using an on-chip artificial pore. Proceedings of the National Academy of Sciences. 100 (3), 820-824 (2003).
- Saleh, O. A., Sohn, L. L. An artificial nanopore for molecular sensing. Nano Letters. 3 (1), 37-38 (2003).
- Saleh, O. A., Sohn, L. L. Quantitative sensing of nanoscale colloids using a microchip Coulter counter. Review of Scientific Instruments. 72 (12), 4449-4451 (2001).
- DeBlois, R. W., Bean, C. P. Counting and sizing of submicron particles by the resistive pulse technique. Review of Scientific Instruments. 41 (7), 909-916 (1970).
- Li, B., et al. Evaluating sources of technical variability in the mechano-node-pore sensing pipeline and their effect on the reproducibility of single-cell mechanical phenotyping. PLoS ONE. 16 (10), 0258982 (2021).
- Zhang, Z. M., Chen, S., Liang, Y. Z. Baseline correction using adaptive iteratively reweighted penalized least squares. Analyst. 135 (5), 1138-1146 (2010).
- Alibert, C., Goud, B., Manneville, J. B. Are cancer cells really softer than normal cells. Biology of the Cell. 109 (5), 167-189 (2017).
- Fujiwara, I., Zweifel, M. E., Courtemanche, N., Pollard, T. D. Latrunculin A accelerates actin filament depolymerization in addition to sequestering actin monomers. Current Biology. 28 (19), 3183-3192 (2018).
- Saleh, O. A. . A novel resistive pulse sensor for biological measurements. , (2003).
- Dokukin, M. E., Guz, N. V., Sokolov, I. Quantitative study of the elastic modulus of loosely attached cells in AFM indentation experiments. Biophysical Journal. 104 (10), 2123-2131 (2013).
- Li, Q., Lim, C. T., Goh, J. C. H., et al. Probing the elasticity of breast cancer cells using AFM. 13th International Conference on Biomedical Engineering. IFMBE Proceedings. 23, 2122-2125 (2009).
- Rother, J., et al. Atomic force microscopy-based microrheology reveals significant differences in the viscoelastic response between malign and benign cell lines. Open Biology. 4 (5), 140046 (2014).
- Li, Q., et al. AFM indentation study of breast cancer cells. Biochemical and Biophysical Research Communications. 374 (4), 609-613 (2008).
- Xu, C., et al. Elasticity measurement of breast cancer cells by atomic force microscopy. Proc. SPIE 9230. Twelfth International Conference on Photonics and Imaging in Biology and Medicine. (PIBM 2014). 92300, (2014).
- Alcaraz, J., et al. Microrheology of human lung epithelial cells measured by atomic force microscopy. Biophysical Journal. 84 (3), 2071-2079 (2003).
- Li, M., Dang, D., Liu, L., Xi, N., Wang, Y. Atomic force microscopy in characterizing cell mechanics for biomedical applications: A review. IEEE Transactions on Nanobioscience. 16 (6), 523-540 (2017).
- Urbanska, M., et al. A comparison of microfluidic methods for high-throughput cell deformability measurements. Nature Methods. 17, 587-593 (2020).
- Hill, R. T., Chilkoti, A. Surface Patterning. Biomaterials Science: An Introduction to Materials: Third Edition. , 276-301 (2013).
- Wang, Z., Volinsky, A. A., Gallant, N. D. Crosslinking effect on polydimethylsiloxane elastic modulus measured by custom-built compression instrument. Journal of Applied Polymer Science. 131 (22), 41050 (2014).
- Gibson, L. J. The hierarchical structure and mechanics of plant materials. Journal of the Royal Society Interface. 9 (76), 2749-2766 (2012).
- Stephens, A. D., Banigan, E. J., Adam, S. A., Goldman, R. D., Marko, J. F. Chromatin and lamin a determine two different mechanical response regimes of the cell nucleus. Molecular Biology of the Cell. 28 (14), 1984-1996 (2017).
- Rosenbluth, M. J., Lam, W. A., Fletcher, D. A. Force microscopy of nonadherent cells: A comparison of leukemia cell deformability. Biophysical Journal. 90 (8), 2994-3003 (2006).
- Evers, T. M. J., Holt, L. J., Alberti, S., Mashaghi, A. Reciprocal regulation of cellular mechanics and metabolism. Nature Metabolism. 3 (4), 456-468 (2021).
- Balakrishnan, K. R., et al. Node-pore sensing enables label-free surface-marker profiling of single cells. Analytical Chemistry. 87 (5), 2988-2995 (2015).
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionExplore More Articles
This article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved