A subscription to JoVE is required to view this content. Sign in or start your free trial.
Method Article
Two-photon Imaging of Microglial Processes' Attraction Toward ATP or Serotonin in Acute Brain Slices
* These authors contributed equally
In This Article
Summary
Microglia, the resident immune cells of the brain, respond quickly with morphological changes to modifications of their environment. This protocol describes how to use two-photon microscopy to study the attraction of microglial processes toward serotonin or ATP in acute brain slices of mice.
Abstract
Microglial cells are resident innate immune cells of the brain that constantly scan their environment with their long processes and, upon disruption of homeostasis, undergo rapid morphological changes. For example, a laser lesion induces in a few minutes an oriented growth of microglial processes, also called "directional motility", toward the site of injury. A similar effect can be obtained by delivering locally ATP or serotonin (5-hydroxytryptamine [5-HT]). In this article, we describe a protocol to induce a directional growth of microglial processes toward a local application of ATP or 5-HT in acute brain slices of young and adult mice and to image this attraction over time by multiphoton microscopy. A simple method of quantification with free and open-source image analysis software is proposed. A challenge that still characterizes acute brain slices is the limited time, decreasing with age, during which the cells remain in a physiological state. This protocol, thus, highlights some technical improvements (medium, air-liquid interface chamber, imaging chamber with a double perfusion) aimed at optimizing the viability of microglial cells over several hours, especially in slices from adult mice.
Introduction
Microglial cells are the brain's resident macrophages and play a role in both physiological and pathological conditions1,2. They have a highly branched morphology and are constantly extending and retracting their processes3,4. This "scanning" behavior is believed to be related and necessary to the survey of their surroundings. The morphological plasticity of microglia is expressed in three modes. First, some compounds rapidly modulate microglial morphology: the addition of ATP5,6 or NMDA5,7 in the medium bathing acute brain slices increases the complexity of microglial ramifications, whereas norepinephrine decreases it6. These effects either are directly mediated by microglial receptors (for ATP and norepinephrine) or require an ATP release from neurons (for NMDA). Second, the growth and retraction speed of microglial processes, called motility or "surveillance", can be affected by extracellular factors8, homeostasis disruptions9,10, or mutations9,10,11. Third, in addition to these isotropic changes of morphology and motility, microglia have the capacity to extend their processes directionally toward a pipette delivering ATP3,5,12,13,14, in culture, in acute brain slices or in vivo, or delivering 5-HT in acute brain slices15. Such oriented growth of microglial processes, also called directional motility, was first described as a response to a local laser lesion3,4. Thus, physiologically, it may be related to the response to injury or required for targeting microglial processes toward synapses or brain regions requiring pruning during development15,16, or in physiological17,18,19 or pathological situations9,18,19,20 in adulthood. The three types of morphological changes rely on different intracellular mechanisms11,13,20, and one given compound does not necessarily modulate all of them (e.g., NMDA, which acts indirectly on microglia, has an effect on morphology but does not induce directional motility5,7). Therefore, when aiming to characterize the effect of a compound, a mutation or a pathology on microglia, it is important to characterize the three components of their morphological plasticity. Here, we describe a method to study the directional growth of microglial processes toward a local source of compound, which is, here, ATP or 5-HT.
There are several models to study microglia processes' attraction: primary cultures in 3D environment6,18,19, acute brain slices6,13,15, and in vivo imaging3,13. The in vivo approach is the best to preserve the physiological state of microglia. However, intravital imaging of deep regions requires complex surgical procedures and, therefore, it is often limited to superficial cortical layers. The use of microglia primary culture is the easiest technique to test a large number of conditions with a limited number of animals. Nevertheless, it is impossible to obtain the same cell morphology as in vivo, and cells lose their physiological interactions with neurons and astrocytes. Acute brain slices represent a compromise between these two approaches. This model allows researchers to study brain structures which are otherwise difficult to reach and to image with high resolution in vivo, and to investigate slices from neonatal stages, whereas transcranial microscopy is mostly performed at adulthood. Finally, it makes it possible to observe in real-time the effects of local drug application, and to repeat experiments while using a limited number of animals. Nonetheless, an issue with acute brain slices is the limited time (a few hours) during which the cells remain alive, notably for slices from mice older than two weeks, and the potential change of microglia morphology over time21,22.
Here, we describe a protocol to prepare acute brain slices of young and adult Cx3cr1GFP/+ mice up to two months old, with the preservation of microglia morphology and motility for several hours. We, then, describe how to use these slices to study the attraction of microglial processes toward compounds like ATP or 5-HT.
Access restricted. Please log in or start a trial to view this content.
Protocol
All experiments were approved by the local ethical committee (Darwin Committee, agreements #1170 and #10921).
1. Preparation of Glass Micropipettes for the Local Application of Compounds
- Prepare pipettes from borosilicate thin-wall glass capillaries with an electrode puller. Adjust the parameters to obtain pipettes with a 4 - 5 µm diameter at their extremity. Figure 2D shows one pipette in brightfield at low magnification.
2. Solutions
- Ensure that only glassware that has been cleaned by an autoclave cycle, followed by rinsing 2x - 3x with ultrapure water, will be used. Never use glassware that has been in contact with paraformaldehyde.
- Prepare a 2 mol·L-1 CaCl2 stock solution by dissolving 14.7 mg of CaCl2·2H2O in 50 mL of water of high purity (ultrapure water, resistance 18.2 MΩ; the traces of metal in distilled water or tap water can lead to suboptimal slice quality due to pro-oxidative effects).
- Store this stock solution at room temperature for a maximum of one month.
- On the day of the experiment, prepare 1 L of choline-aCSF (artificial cerebrospinal fluid) solution, whose composition is 110 mmol·L-1 choline Cl, 25 mmol·L-1 glucose, 25 mmol·L-1 NaHCO3, 7 mmol·L-1 MgCl2, 11.6 mmol·L-1 ascorbic acid, 3.1 mmol·L-1 sodium pyruvate, 2.5 mmol·L-1 KCl, 1.25 mmol·L-1 NaH2PO4, and 0.5 mmol·L-1 CaCl2, 0.5.
- To prepare this solution, add, in the following order, to a 1 L graduated flask: 0.186 g of KCl, 0.195 g of NaH2PO4, 2.04 g of acid ascorbic, 2.1 g of NaHCO3, and 4.5 g of glucose.
- Fill about half of the final volume with ultrapure water and stir until complete dissolution.
- Add 0.34 g of sodium pyruvate and 15.36 g of choline Cl.
NOTE: It is convenient to first dissolve the choline Cl with 5 to 10 mL of the solution prepared in step 2.3.2 before adding it to the whole solution. - Add 7 mL of 1 mol·L-1 MgCl2 and 250 µL of 2 mol·L-1 CaCl2 (prepared in step 2.2) to the solution.
- Fill the graduated flask up to 1 L with ultrapure water.
- With a vapor pressure osmometer, check that the osmolarity is between 300 and 310 mΩ. If not, adjust it with glucose.
- Check the pH after carbogenation (i.e., bubbling with “carbogen”, a mix of 95% O2/5% CO2) and adjust it, if necessary, to 7.3 - 7.4 with 10 M NaOH.
- Transfer the solution to a glass bottle for storage. Keep the bottle in the fridge until use (STEP 3.1).
NOTE: It is recommended to make a fresh solution on the day of the experiment. However, if necessary, choline-aCSF can be stored up to two days at 4 °C.
- On the day of the experiment, prepare 1 L of an aCSF solution, whose composition is 124 mmol·L-1 NaCl, 26.2 mmol·L-1 NaHCO3, 25 mmol·L-1 glucose, 2.5 mmol·L-1 KCl, 2 mmol·L-1 CaCl2, 1 mmol·L-1 MgCl2, and 1.25 mmol·L-1 NaH2PO4.
- To prepare this solution, add, in the following order, to a graduated flask: 0.150 g of NaH2PO4, 0.186 g of KCl, 2.2 g of NaHCO3, 4.5 g of glucose, and 7.3 g of NaCl. Bring the solution to a volume of 1 L with ultrapure water and stir it vigorously on a stir plate.
- Add 1 mL of 1 mol·L-1 MgCl2 and 1 mL of 2 mol·L-1 CaCl2 to the solution and transfer the aCSF solution to a glass bottle for storage.
- Check whether the osmolarity is 300 - 310 mΩ·L-1 and, if not, adjust it with glucose.
- Check the pH after carbogenation (i.e., bubbling with “carbogen”) and adjust it, if necessary, to 7.3 - 7.4 with 10 M NaOH.
- Transfer the solution to a glass bottle for storage. Keep the bottle in the fridge until use (step 3.1).
NOTE: It is recommended to make a fresh solution on the day of the experiment. However, an alternative is to prepare a 10x stock solution containing NaCl, NaHCO3, KCl, and NaH2PO4 at 10x the final concentration, which can be stored for no more than one week at 4 °C. Make the final aCSF on the day of the experiment by diluting the 10x stock solution with ultrapure water and adding the glucose, CaCl2, and MgCl2.
- Prepare the drug solutions on the day of the experiment. Use the aCSF solution to bring them to the final concentrations which are, here, 500 µmol·L-1 for ATP and 5 µmol·L-1 for 5-HT.
NOTE: For ATP, a stock solution can be prepared (e.g., 50 mM ATP in water), stored in aliquoted form at -20 °C, and diluted with aCSF to the final concentration on the day of the experiment. In contrast, the 5-HT (serotonin-HCl) solution must be prepared from powder on the day of the experiment, at 1 mg·mL-1 in water, kept at 4 °C to avoid 5-HT oxidation, and diluted in aCSF at the time of the experiment.
3. Preparation of Acute Brain Slices
- Preparation of the dissection area
- Prepare 70 mL of ice-cold choline-aCSF in an 80 mL beaker placed on ice, to be used for cardiac perfusion, rapid cooling down of the brain, and slicing. Prepare 150 mL of choline-aCSF in a 200 mL crystallizing dish, placed in a heated water bath maintained at 32 °C. Place a nylon mesh strainer in the crystallizing dish to retain the slices. This will be used to let the slices recover for 10 min just after slicing.
- At least 30 min before starting the dissection (section 3.2), start bubbling these two solutions (70 mL of choline-aCSF on ice and 150 mL of choline-aCSF at 32° C) with carbogen. Maintain constant carbogenation during the entire procedure.
- Prepare the interface chamber device (Figure 1C), which will be used to keep slices until their use.
- In a sealed food box (10 x 10 cm or 10 cm in diameter, 8 cm in height), installed on a magnetic stirrer, place a 200 mL crystallizing dish with a bar magnet.
- Add 200 mL of aCSF in this crystallizing dish and place the 3D-printed interface slice holder on top of it (the interface slice holder is composed of two perfectly fitting parts, with a polyamide mesh stretched between them, Figure 1A,B).
- Remove excess volume from the crystallizing dish to keep only a thin film of solution covering the mesh of the interface slice holder. This will later create a fine rim of solution surrounding the slices (but without covering them).
- Put a few millimeters of aCSF at the bottom of the food box and start bubbling it with carbogen (at first use, make a small hole in the sealed food box wall to make sure the tubing can enter the box).
- Close the sealed box while maintaining constant carbogenation. This will create a humidified 95% O2/5% CO2 rich environment in which the slices will be transferred after their recovery in choline-aCSF and maintained before they are imaged. This device is hereafter referred to as the “interface chamber” (Figure 1C).
- Brain dissection and slicing
- Anesthetize the mouse with an intraperitoneal injection of 50 mg·mL-1 pentobarbital (0.15 mL/20 g of mouse body weight), immobilize it, expose the heart, and perform a cardiac perfusion with 10 mL of ice-cold, carbogenated, choline-aCSF (see step 3.1.1), with a peristaltic pump. Observe the pallor of the liver as an indicator of a good perfusion. The perfusion lasts less than 5 min.
- Decapitate the mouse and cut the skin to expose the skull. With big scissors, apply two transversal cuts from the large foramen and one long sagittal cut and, using fine forceps, remove the skull plates.
- Quickly and gently extract the brain (in less than 1 min) and place it for 1 min in the 80 mL beaker containing the remaining (~60 mL) ice-cold choline-aCSF (still under constant carbogenation), in order to cool it down.
- Transfer the brain onto a filter paper previously wet with aCSF.
- Cut out the brain according to the brain region of interest and preferred angle of slicing. For example, to image the thalamus or the hippocampus on coronal slices, cut out with a scalpel blade the cerebellum and, then, about 2 mm from the rostral and caudal extremities of the brain.
NOTE: It is important to remove brain parts that are too rostral or too caudal because the smaller the region to trim before reaching the area of interest, the faster the slicing. A total time for slicing (step 3.2.7) of less than 20 min is recommended. - For coronal slices, position and glue (with cyanoacrylate glue) the caudal face of the brain onto a 10 cm Petri dish, glued on the cutting block of a vibrating slicer and position it in the reservoir chamber of the vibrating slicer, which is positioned in a larger chamber filled with ice. Then, fill the Petri dish with all the remaining ice-cold choline-aCSF.
- While keeping constant the 95% O2/5% CO2 bubbling of the ice-cold choline-aCSF, cut 300 µm-thick coronal slices (speed: 0.08 mm·s-1, blade vibration: 60 Hz, vibration amplitude: 1 mm).
- Collect the brain slices with a wide-mouth (4 mm in diameter) disposable transfer pipette, one by one after every single pass of the blade, to avoid the accumulation of toxic components released by the periphery of the slices. Take care to avoid air bubbles during the transfer and place each slice in the choline-aCSF at 32 °C for about 10 min for recovery.
- With the transfer pipette, place the slices onto pieces of lens-cleaning paper topped with a drop of choline-aCSF. Aspirate the excess of choline-aCSF and, with the spatula, place the slices, laid on the lens-cleaning tissue, on the mesh of the interface chamber containing carbogenated aCSF at room temperature (see 3.1.3.5). Let the slice recover in this environment for at least 30 min.
NOTE: After this, the slices are ready and can be used for microglia imaging for up to 6 h after the brain extraction from young (less than one-month-old) mice and up to 4 h after the brain extraction from two-months-old adults.
4. Two-photon Microscopy
- Parameters setting
- Switch on the multiphoton system (hybrid detectors, laser, scanner, electro-optic modulator, microscope).
- Tune the laser at 920 nm, check that the laser is mode-locked, and set the power at 5% - 15% and the gain at 10%. This corresponds to a power of 3 - 5 mW under the objective. Ensure that the nondescanned detectors are engaged and the appropriate emission and excitation filters installed.
- Set parameters of the imaging software to the following values: for the frame size, 1024 x 1024 pixels corresponding to an area of 295.07 x 295.07 µm; for the zoom, 2. If the signal is very noisy, apply a line average of 2. For the pixel dynamics, set the imaging software at 12 bits or more.
NOTE: Images with a higher bit value allow researchers to distinguish smaller differences in fluorescence intensity than images with a lower bit value: a change of one gray value in an 8-bit image would correspond to a change of 16 gray values in a 12-bit and of 256 gray values in a 16-bit image. Therefore, higher-bit images are more appropriate for quantitative analysis, but as their size increases with bit depth, storage capacity, and computing power can become limiting. - Select the scan mode XYZT with a Z-interval range at 2 µm and a T-interval of 2 min.
NOTE: The x,y and z resolution are determined by the Nyquist sampling theorem. A Z-step size around 0.8 would be optimal to resolve microglia processes (with a diameter of <1 µm), but the optical resolution of multiphoton microscopy is limiting (at 920 nm with a 0.95 NA objective, the axial resolution is around 1 µm). On top of that physical barrier, in a live-imaging experiment, the sensitivity or signal-to-noise ratio, the resolution, the speed, and the total observation time matter. Taking into account all these parameters, a z-step of 2 µm (as in numerous studies3,11,14), an image size of 1024 x 1024 pixels, and a high-speed acquisition using a resonant scanner coupled to HyD detectors (it takes around 15 s to acquire 50 z-plans) were selected here. The frequency of acquisitions is one XYZT series every 2 min and the total duration is 30 min. If the set-up is not fast or sensitive enough, it is possible to reduce the lateral resolution (down to 512 x 512) or the number of z-slices (by imaging exclusively in the z-depth which exhibits the strongest fluorescence [i.e., not the deepest z-slices where fluorescence is faint]), or to decrease the speed of the scanner. The axial resolution can also be decreased by increasing the z-step up to 3 µm, but as this may impact the quantification, all experiments to be compared should be performed with the same z-step.
NOTE: It is possible to perform similar experiments on slices from CX3CR1creER-YFP mice18, a mouse line used to induce genetic deletion in microglia only, and in which microglia constitutively express yellow fluorescent protein (YFP). However, the expression level of YFP is very low compared to green fluorescent protein (GFP) in CX3CR1GFP/+ mice; thus, imaging is possible but challenging and requires the optimization of the acquisition parameters. It is recommended to adjust them as follows. - Tune the laser at 970 nm (which is better adapted to YFP excitation than 920 nm), the power at 50%, and the gain at 50%, which corresponds to a laser power under the objective of 5 - 6 mW.
- Set a line average of 4 (or more) to improve the signal-to-noise ratio.
- Positioning of the slice and of the glass micropipette, and the local application of the compound
- Connect the peristaltic pump to the recording chamber, 30 min before starting the recording. After cleaning the whole perfusion system with 50 mL of ultrapure water, start the perfusion of the recording chamber with aCSF (50 mL) contained in a glass beaker under constant carbogenation. Throughout the experiment, keep the circulating aCSF to 32 °C with an inline microheater or a Peltier heater.
NOTE: A specific perfusion chamber with top and bottom perfusion is designed to optimize the oxygenation on both sides of the slice. The perfusion chamber is composed of two perfectly fitting parts, with a polyamide mesh stretched between them (Figure 2A,B). Compared with other types of chambers, where the slice is directly laying on a glass coverslip, this chamber reduces neuronal death in the bottom part of the slice, improves viability, and reduces the slice movements induced by its swelling. - With a wide-mouth disposable transfer pipette, transfer the brain slice to be imaged to the aCSF beaker to remove the lens paper, let it sink (as a proof that no air bubble is attached), and transfer it to the recording (perfusion) chamber.
- Position a slice holder (a hairpin made of platinum with the two branches joined by parallel nylon threads) on the slice to minimize slice movement due to the perfusion flow.
- Use the bright-field illumination to target the brain region of interest (exposure time: 50 to 80 ms) using a low magnification objective (5X or 10X). Switch to the higher magnification (25x with a 0.35x lens) water immersion objective and adjust the position.
NOTE: Avoid to image fields close to the slice holder’s nylon threads as they can block the light and locally deform the slice. Make sure that the area of interest is flat. If necessary, remove the slice holder in order to reposition the slice and/or the slice holder. - Use the fluorescence illumination to locate fluorescent microglial cells to be imaged in the field (exposure time: 250 - 500 ms).
NOTE: This step allows researchers to check the presence of cells in the region of interest and their fluorescence intensity, and to control for the amount of cellular debris. - Backfill the pipette with 10 µL of aCSF with ATP, 5-HT, or the drug of interest at its final concentration. Point the tip downward and gently shake the drug-filled pipette to remove any air bubbles trapped in the tip.
NOTE: If the solution to be injected tends to form bubbles, consider using borosilicate pipettes with an internal filament. Leakage of ATP out of the pipette can attract microglial processes even before the injection (if this occurs, it will be visible at the analysis step). Although this should be moderate with the ATP concentration used (500 µmol·L-1), if it is an issue, consider prefilling the micropipette with 2 mL of aCSF prior to adding the ATP (or other compound) solution at step 4.2.6. - Mount the filled pipette in a pipette holder, connected with transparent tubing to a 5 mL syringe, with a plunger positioned at the 5 mL position. The pipette holder itself is mounted onto a three-axis micromanipulator.
- Under bright-field illumination, use the micromanipulator to position the pipette in the center of the field. For a reproducible and optimal centering, display and use the rulers on the image.
- Lower the pipette gently toward the slice, controlling and adjusting the objective at the same time, until the pipette tip lightly touches the surface of the slice. Stopping the descent of the pipette as soon as it is visible that the slice has been touched allows the pipette tip to penetrate 80 - 100 µm of the surface of the slice (see Figure 3B).
- Tune the laser (see the parameters above) and switch the microscope to the multiphoton mode. Make sure that the chamber is screened from any light source (e.g., a computer screen). Switch on the nondescanned detectors and set the gain. Use a lookup table (LUT) with a color-coded upper limit to avoid saturating the pixels in the image.
- Determine the thickness of the slice to be imaged (i.e., the upper and lower z-positions where fluorescence is detectable [usually between 220 and 290 µm in total]).
NOTE: At the surface of the slice, there is an increased density of processes and possibly of microglia, often with an unusual morphology, in comparison with the inside of the slice. This accumulation will be more striking with time (i.e., more visible in the last than in the first brain slice to be imaged). Therefore, the z-planes in the first ~30 µm should not be used for the analysis and can even be skipped for the acquisition. - Start recording for a total duration of 30 min (or more if desired) and after a 5 min baseline, locally apply the compound to be tested (without interrupting the imaging). To do this, slowly press the plunger of the syringe connected to the micropipette, from the 5 mL to the 1 mL position (in about 5 s). Resistance when pressing the plunger must be felt immediately. If not, the tip might be broken.
NOTE: For a trained experimenter, the injections with this method are reproducible, but alternatively to the manual manipulation of a syringe, the pipette could be linked to an automated pressure ejection system to allow a better control of the volume delivered. The injection creates a physical distortion of the slice at the site of the injection. This distortion is visible a posteriori in the first two or three images after the injection but should not be visible on the fourth image, (i.e., 8 min after the injection). If it persists, consider changing the parameters for the pipette preparation. - At the end of the acquisition (30 min), discard the micropipette and remove the slice. If desired, fix the slice for further immunolabeling. For example, the SNAPSHOT method is optimized for the fixation and staining of thick slices23.
- Prior to starting to image a new slice, make the 2D movie (section 5.1) in order to check that microglia have a normal morphology and are moving and, thus, that the slices are healthy.
- Connect the peristaltic pump to the recording chamber, 30 min before starting the recording. After cleaning the whole perfusion system with 50 mL of ultrapure water, start the perfusion of the recording chamber with aCSF (50 mL) contained in a glass beaker under constant carbogenation. Throughout the experiment, keep the circulating aCSF to 32 °C with an inline microheater or a Peltier heater.
5. Analysis of the Attraction of Microglial Processes
- 2D projection and drift correction
- Open the file (.LIF) with Fiji24.
- If necessary, make a substack (Image/Stacks/Tools/Make Substack) with only the z-planes of interest. For example, exclude the z-planes corresponding to the surface of the slice if they have been acquired but are not to be used for the analysis (see the NOTE after step 4.2.11) and the deepest z-planes with no fluorescence. The final stack generally contains 90 - 110 z-slices (180 - 220 µm).
- Launch the Z project function (Image/Stacks/Z Project") and select the Max Intensity projection type to make the projections of the z-stack acquired at each time point.
- Launch the MultiStackReg plugin (Plugin/Registration/MultiStackReg), selecting Action 1: Align and Transformation: Rigid Body to correct slight drifts that may have occurred during the acquisition. Save this 2D movie as a new file (.tiff).
- Data processing
- Open this new file with Icy25.
- Draw a circular R1 region of interest (ROI) of 35 µm in diameter, centered on the injection site (identified notably by the shadow of the pipette and the distortion created at the time of injection).
- Use the plugin ROI intensity evolution and measure the mean intensity over time in R1.
- Save the results to an .XLS file.
- Quantification and representation of the results
- To quantify the microglial response over time, determine at each time point
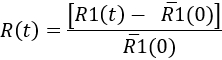
Here,`R1(0) is the mean of the R1(t) values before the injection. Then, the results can be represented as a kinetic of the microglial response, or at a specific time point (see Figure 7).
- To quantify the microglial response over time, determine at each time point
Access restricted. Please log in or start a trial to view this content.
Results
This protocol describes a method to induce, observe, and quantify the oriented growth of microglial processes toward a locally applied compound, for example, ATP or 5-HT, in acute brain slices from young or adult (at least up to two-month-old) mice. Among the factors that contribute to maintaining brain slices from adult animals in a healthy state for several hours is the use of two tools designed to optimize cell survival at two steps of the protocol. First, the interface slice holder in...
Access restricted. Please log in or start a trial to view this content.
Discussion
By maintaining, unlike in dissociated or organotypic slice culture, a structural integrity with limited network adjustments, acute brain slices allow researchers to study microglia in their physiological environment. However, one of the major limitations is the fact that the slicing procedure creates injuries that can rapidly compromise the viability of neurons, particularly in the adult brain. As microglia are particularly reactive to cell damage, it is important to limit neuronal cell death as much as possible to prese...
Access restricted. Please log in or start a trial to view this content.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank the Cell and Tissue Imaging Facility of the Institut du Fer à Moulin, where all image acquisition and analysis have been performed. This work has been supported in part by the Centre National de la Recherche Scientifique, the Institut National de la Santé et de la Recherche Médicale, the Sorbonne Université Sciences, and by grants from Sorbonne Universités-Pierre et Marie Curie University (Emergence-UPMC program 2011/2014), the Fondation pour la Recherche sur le Cerveau, the Fondation de France, the Fondation pour la Recherche Médicale "Equipe FRM DEQ2014039529", the French Ministry of Research (Agence Nationale pour la Recherche ANR-17-CE16-0008 and the Investissements d'Avenir programme "Bio-Psy Labex" ANR-11-IDEX-0004-02) and a Collaborative Research in Computational Neuroscience program, National Science Foundation/French National Agency for Research (number : 1515686). All the authors are affiliated to research groups which are members of the Paris School of Neuroscience (ENP) and of the Bio-Psy Labex. F.E. is a Ph.D. student affiliated with Sorbonne Université, Collège Doctoral, F-75005 Paris, France, and is funded by the Bio-Psy Labex. V.M. is a post-doctoral fellow funded by the Collaborative Research in Computational Neuroscience program, National Science Foundation/French National Agency for Research (number: 1515686). The authors thank Marta Kolodziejczak who participated in the initiation of the project.
Access restricted. Please log in or start a trial to view this content.
Materials
| Name | Company | Catalog Number | Comments |
| for pipettes preparation | |||
| Clark Borosilicate Thin Wall Capillaries | Harvard Apparatus | 30-0065 | Borosilicate Thin Wall without Filament, 1.5 mm OD, 1.17 mm ID, 75 mm L , Pkg. of 225 |
| DMZ Universal Puller | Zeitz Instrumente | ||
| Name | Company | Catalog Number | Comments |
| for solutions | |||
| Calcium Chloride dihydrate (CaCl2) | Sigma | C5080 | |
| Choline Chloride | Sigma | C7527 | |
| D-(+)-Glucose | Sigma | G8270 | |
| L-Ascorbic acid | Sigma | A5960 | |
| Magnesium Chloride solution 1 M (MgCl2) | Sigma | 63020 | |
| Potassium chloride SigmaUltra >99.0% (KCl) | Sigma | P9333 | |
| Sodium bicarbonate (NaHCO3) | Sigma | S5761 | |
| Sodium Chloride (NaCl) | Sigma | S5886 | |
| Sodium phosphate monobasic | Sigma | S5011 | |
| Sodium pyruvate | Sigma | P2256 | |
| Ultrapure water | MilliQ | for all the solutions | |
| Name | Company | Catalog Number | Comments |
| for slice preparation | |||
| 2x 200 mL crystalizing dishes | |||
| 80 mL Pyrex beaker | |||
| Antlia-3C Digital Peristaltic pump | DD Biolab | 178961 | For mice perfusion and 2-photon chamber perfusion (aCSF) |
| Carbogen 5% CO2/95% O2 | Air Liquide France Industrie | ||
| Dolethal | Vetoquinol | Dolethal 50 mg/mL | |
| Filter papers (Whatman) | Sigma | WHA1001042 | Whatman qualitative filter paper, Grade 1 (Pore size: 11µM) |
| Fine Scissors - Sharp | Fine Science Tools | 14060-60 | |
| Food box 10 cm diameter, 8 cm Height | |||
| glue (ethyl cyanoacrylate) | Loctite | super glue 3 power flex | |
| Hippocampal Tool (spatula) | Fine Science Tools | 10099-15 | The largest extremity has to be angled at 90° |
| Ice | |||
| Iris Forceps (curved) | Moria | MC31 | |
| Lens cleaning tissue | THOR LABS | ||
| Nylon mesh strainer | diameter 7 cm | ||
| Razor blades | Electron Microscopy Sciences | 72000 | For the slicer |
| scalpel blade | |||
| Slice interface holder | home-made, the file for 3D printing is provided in Supplemental Material | ||
| Surgical Scissors - Sharp | Fine Science Tools | 14002-14 | |
| Vibrating slicer | Thermo Scientific | 720-2709 | Model: HM 650V (Vibrating blade microtome) |
| Water bath | Set at 32 °C (first recovery step) | ||
| Name | Company | Catalog Number | Comments |
| for slice imaging | |||
| × 25 0.95 NA water-immersion objective | Leica Microsystems (Germany) | HCX Irapo | |
| 2-photon MP5 upright microscope with resonant scanners (8 kHz) and two HyD Hybrid detectors | Leica Microsystems (Germany) | ||
| Antlia-3C Digital Peristaltic pump | DD Biolab | 178961 | For 2-photon chamber perfusion with aCSF |
| Carbogen 5% CO2/95% O2 | Air Liquide France Industrie | I1501L50R2A001 | |
| Chameleon Ultra2 Ti:sapphire laser | Coherent (Germany) | ||
| disposable transfer pipettes , wide mouth | ThermoFischer scientific | for example : 232-11 | 5.8 mL with fin tip, but we cut it (approx 7 cm) to have a 4 mm diameter mouth |
| emission filter SP680 | Leica Microsystems (Germany) | ||
| fluorescent cube containing a 525/50 emission filter and a 560 dichroic filter (for fluorescence collection) | Leica Microsystems (Germany) | ||
| glass beaker with 50 mL of ACSF to maintain constant perfusion of the slice | |||
| Heating system | Warner Instrument Corporation | Automatic Heater Controller TC-324B | to maintain perfusion solution at 32 °C |
| perfusion chamber | home-made, the file for 3D printing is provided in Supplemental Material | ||
| slice holder ("harp") | home made : hairpin made of platinum with the two branches joined by parallel nylon threads | ||
| Name | Company | Catalog Number | Comments |
| for slice stimulation | |||
| Adenosine 5′-triphosphate disodium salt hydrate (ATP) | Sigma | A-26209 | to be prepared ex-temporaneously : 1 mg/mL (3 mM) stock solution prepared the day of the experiment, kept at 4 °C (a few hours) and diluted just before use |
| Fluorescein (optional) | Sigma | F-6377 | use at 1 µM final |
| Micromanipulator | Luigs and Neumann | SM7 | connected to the micropipette holde |
| Micropipette holder | same as for eletrophysiology | ||
| Serotonin hydrochloride | Sigma | H-9523 | aliquots of 50mM stock solution in H20 kept at -20 °C. 500 µM solution prepared the day of the experiment. |
| Syringe 5 mL (without needle) | Terumo medical products | SS+05S1 | |
| Transparent tubing | Fischer Scientific | 11750105 | Saint Gobain Performance Plastics™ Tygon™ E-3603 Non-DEHP Tubing |
| Name | Company | Catalog Number | Comments |
| for image analysis | |||
| Fiji | https://fiji.sc | Schindelin, J. et al Nat. Methods (2012) doi 10.1038 | |
| Icy | Institut Pasteur | http://icy.bioimageanalysis.org | de Chaumont, F. et al. Nat. Methods (2012) |
| Name | Company | Catalog Number | Comments |
| mice | |||
| CX3CR1-GFP mice | Jung et al, 2000 | male or females, P3 to 2 months-old ; we have backcrossed these mice on 129sv background. | |
| CX3CR1creER-YFP mice | Parkhurst et al 2013 | male or females, P3 to 2 months-old ; we have backcrossed these mice on 129sv background. |
References
- Salter, M. W., Stevens, B. Microglia emerge as central players in brain disease. Nature Publishing Group. 23 (9), 1018-1027 (2017).
- Tay, T. L., Savage, J., Hui, C. W., Bisht, K., Tremblay, M. -È Microglia across the lifespan: from origin to function in brain development, plasticity and cognition. The Journal of Physiology. , (2016).
- Davalos, D., et al. ATP mediates rapid microglial response to local brain injury in vivo. Nature Neuroscience. 8 (6), 752-758 (2005).
- Nimmerjahn, A. Resting Microglial Cells Are Highly Dynamic Surveillants of Brain Parenchyma in Vivo. Science. 308 (5726), 1314-1318 (2005).
- Dissing-Olesen, L., et al. Activation of neuronal NMDA receptors triggers transient ATP-mediated microglial process outgrowth. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience. 34 (32), 10511-10527 (2014).
- Gyoneva, S., Traynelis, S. F. Norepinephrine modulates the motility of resting and activated microglia via different adrenergic receptors. Journal of Biological Chemistry. 288 (21), 15291-15302 (2013).
- Eyo, U. B., et al. Neuronal hyperactivity recruits microglial processes via neuronal NMDA receptors and microglial P2Y12 receptors after status epilepticus. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience. 34 (32), 10528-10540 (2014).
- Hristovska, I., Pascual, O. Deciphering Resting Microglial Morphology and Process Motility from a Synaptic Prospect. Frontiers in Integrative Neuroscience. 9, 1231(2016).
- Avignone, E., Lepleux, M., Angibaud, J., Nägerl, U. V. Altered morphological dynamics of activated microglia after induction of status epilepticus. Journal of Neuroinflammation. 12, 202(2015).
- Abiega, O., et al. Neuronal Hyperactivity Disturbs ATP Microgradients, Impairs Microglial Motility, and Reduces Phagocytic Receptor Expression Triggering Apoptosis/Microglial Phagocytosis Uncoupling. PLoS Biology. 14 (5), e1002466(2016).
- Madry, C., et al. Microglial Ramification, Surveillance, and Interleukin-1β Release Are Regulated by the Two-Pore Domain K+Channel THIK-1. Neuron. 97 (2), 299-312 (2018).
- Honda, S., et al. Extracellular ATP or ADP induce chemotaxis of cultured microglia through Gi/o-coupled P2Y receptors. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience. 21 (6), 1975-1982 (2001).
- Haynes, S. E., et al. The P2Y12 receptor regulates microglial activation by extracellular nucleotides. Nature Neuroscience. 9 (12), 1512-1519 (2006).
- Wu, L. -J., Vadakkan, K. I., Zhuo, M. ATP-induced chemotaxis of microglial processes requires P2Y receptor-activated initiation of outward potassium currents. Glia. 55 (8), 810-821 (2007).
- Kolodziejczak, M., et al. Serotonin Modulates Developmental Microglia via 5-HT 2BReceptors: Potential Implication during Synaptic Refinement of Retinogeniculate Projections. ACS Chemical Neuroscience. 6 (7), 1219-1230 (2015).
- Schafer, D. P., et al. Microglia Sculpt Postnatal Neural Circuits in an Activity and Complement-Dependent Manner. Neuron. 74 (4), 691-705 (2012).
- Pfeiffer, T., Avignone, E., Nägerl, U. V. Induction of hippocampal long-term potentiation increases the morphological dynamics of microglial processes and prolongs their contacts with dendritic spines. Scientific Reports. 6, 32422(2016).
- Parkhurst, C. N., et al. Microglia Promote Learning-Dependent Synapse Formation through Brain-Derived Neurotrophic Factor. Cell. 155 (7), 1596-1609 (2013).
- Wu, Y., Dissing-Olesen, L., Macvicar, B. A., Stevens, B. Microglia: Dynamic Mediators of Synapse Development and Plasticity. Trends in Immunology. 36 (10), 605-613 (2015).
- Ohsawa, K., et al. P2Y12 receptor-mediated integrin-beta1 activation regulates microglial process extension induced by ATP. Glia. 58 (7), 790-801 (2010).
- Kurpius, D., Wilson, N., Fuller, L., Hoffman, A., Dailey, M. E. Early activation, motility, and homing of neonatal microglia to injured neurons does not require protein synthesis. Glia. 54 (1), 58-70 (2006).
- Stence, N., Waite, M., Dailey, M. E. Dynamics of microglial activation: a confocal time-lapse analysis in hippocampal slices. Glia. 33 (3), 256-266 (2001).
- Dissing-Olesen, L., Macvicar, B. A. Fixation and Immunolabeling of Brain Slices: SNAPSHOT Method. Current Protocols in Neuroscience. 71, 1.23.1-12 (2015).
- Schindelin, J., et al. Fiji: an open-source platform for biological-image analysis. Nature Methods. 9 (7), 676-682 (2012).
- de Chaumont, F., et al. Icy: an open bioimage informatics platform for extended reproducible research. Nature Methods. 9 (7), 690-696 (2012).
- Aitken, P. G., et al. Preparative methods for brain slices: a discussion. Journal of Neuroscience Methods. 59 (1), 139-149 (1995).
- Paris, I., et al. ProMoIJ: A new tool for automatic three-dimensional analysis of microglial process motility. Glia. 66 (4), 828-845 (2018).
- Pagani, F., et al. Defective microglial development in the hippocampus of Cx3cr1 deficient mice. Frontiers in Cellular Neuroscience. 9 (229), 111(2015).
- Ting, J. T., Daigle, T. L., Chen, Q., Feng, G. Acute brain slice methods for adult and aging animals: application of targeted patch clamp analysis and optogenetics. Methods in Molecular Biology. , Clifton, NJ. 221-242 (2014).
- Mainen, Z. F., et al. Two-photon imaging in living brain slices. Methods. 18 (2), San Diego, CA. 231-239 (1999).
- Tanaka, Y., Tanaka, Y., Furuta, T., Yanagawa, Y., Kaneko, T. The effects of cutting solutions on the viability of GABAergic interneurons in cerebral cortical slices of adult mice. Journal of Neuroscience Methods. 171 (1), 118-125 (2008).
- Gyoneva, S., et al. Systemic inflammation regulates microglial responses to tissue damage in vivo. Glia. 62 (8), 1345-1360 (2014).
- Heindl, S., et al. Automated Morphological Analysis of Microglia After Stroke. Frontiers in Cellular Neuroscience. 12, 106(2018).
- Dailey, M. E., Eyo, U., Fuller, L., Hass, J., Kurpius, D. Imaging microglia in brain slices and slice cultures. Cold Spring Harbor Protocols. 12 (12), 1142-1148 (2013).
Access restricted. Please log in or start a trial to view this content.
Reprints and Permissions
Request permission to reuse the text or figures of this JoVE article
Request PermissionThis article has been published
Video Coming Soon
Copyright © 2025 MyJoVE Corporation. All rights reserved